Abstract
The genus Laurencia, a category of marine red algae, is well recognized for producing a large variety of natural products (NPs) that are both chemically intriguing and structurally distinct. The aim of this research was to identify NPs with potential anti-SARS-CoV-2 activity. The crystals of the proteins RdRp and nsp15 were obtained from the RCSB protein database. About 300 NPs were discovered using the PubChem, ChemSpider, and CMNPD databases. The program Autodock Vina was used to conduct the molecular docking procedure once the proteins and ligands were prepared. Before running MD simulations using the CABS-flex 2.0 website, binding affinity assessments and interactions between amino acids were carefully reviewed. Only nine NPs were shortlisted to be examined further. Bromophycolide R, S, and bromophycoic acid C show the tendency to inhibit RdRp by β-hairpin motif binding at the N-terminal known as Active site 2 (AS2), whereas the other four NPs, bromophycolide E, H, P, and thyrsenol A, may effectively inhibit RdRp through interactions via C-terminal, also known as the Active site 1 (AS1). For the enzyme nsp15, bromophycoic B, C, and floridoside showed plausible interactions. In conclusion, out of nine, seven candidates shortlisted for RdRp exhibited strong interactions with the key residues in the AS1 and AS2 regions. Bromophycoic acid C may work as a dual inhibitor due to its favorable interactions with the nsp15 protein and RdRp’s N-terminal, with affinities of −8.5 and −8.2 kcal/mol, respectively.
Keywords:
COVID-19; SARS-CoV-2; RdRp; Nsp12; Nsp15; endoribonuclease; Laurencia; natural products; anti-viral; red algae 1. Introduction
The life-threatening coronavirus infection in 2019 was anticipated to be caused by the positive-sense, a single-strand RNA virus referred to as the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). As of July 2023, there were over 768 million confirmed cases and about 7.0 million fatalities globally, according to the WHO dashboard https://covid19.who.int/ (accessed on 23 July 2023). Even if the severity of the illness is lessened by existing vaccinations and antiviral medical treatment, people can still contract the illness, especially if the virus comprises unique mutations [1,2]. Moreover, vaccine efficiency is still poor, and certain antiviral medications suggested for the treatment of SARS-CoV-2 have been reported to cause viral modifications [3], making it unlikely that this will be enough to halt the spread on its own. At this point, it is quite likely that SARS-CoV-2 will spread throughout the world’s population [4].
Viral RNA-dependent RNA polymerase (RdRp), also known as nonstructural protein 12 (Nsp12), is one of the many possible therapeutic targets and is regarded as a promising target [5]. For viral replication, RdRp is essential [6]. It has conserved structural and core sequence characteristics among RNA viruses, enabling a wide range of antiviral activity [7]. In SARS-CoV-2, the RdRp complex made up of three non-structural proteins 12, 7, and 8, is necessary for RNA replication. Whilst nsp7 and nsp8 are auxiliary stimulators of polymerase activity, nsp12 functions as RdRp and is an important component [8]. There is an excellent effort being made by scientists all around the world to help elucidate feasible alternatives for inhibiting SARS-CoV-2 RdRp. Numerous studies have been conducted to date [9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47]. One medication that targets RdRp is Remdesivir, which is the only one that the US Food and Drug Administration has licensed for the treatment of COVID-19. In several clinical investigations, it has been shown that Remdesivir greatly increases COVID-19 recovery [48,49,50]. Remdesivir was proven to be most efficacious when provided in early infection [51,52]. Nevertheless, investigations have shown that Remdesivir did not statistically significantly outperform the standard of treatment in terms of clinical outcomes [53,54,55], particularly in the case of patients who had severe symptoms and required oxygen support [56]. Remdesivir may produce better results when combined with additional medications or cutting-edge therapies such as monoclonal antibodies when used alone [57,58].
A recent research investigation, again looking at the use of corticosteroids, revealed the impact on viral replication and host inflammation. It was observed that the steroid drugs Ciclesonide and mometasone showed dosage dependent influence on viral proliferation and cytotoxicity [59]. According to recent findings, Ciclesonide either directly or indirectly interacts with nsp15 [60]. A nidoviral RNA uridylate-specific endoribonuclease is called nsp15 (NendoU). It possesses an EndoU family C-terminal catalytic domain [61]. The nsp15 was formerly believed to be involved directly in viral replication; however, it has now been discovered that this is not the case. Moreover, nsp15’s ability to modulate immune system response by meddling with innate immune response has just been de-scribed [62]. It was also mentioned that nsp15 may be involved in RNA disintegration, which assists in concealing it from the host defenses. This nsp15 is described as being crucial to coronavirus biology when taking into account its different activities.
More than any other form of life, viruses have remained impervious to treatments and prevention. Only a few medications are now available to treat viral illnesses effectively. There is an increasing amount of interest in using natural products as a source for novel medications [63]. Effective medications are desperately needed to combat the SARS-CoV-2 infection, including small-molecule inhibitors [64,65], bioactive natural products [66,67,68], and conventional medicine [69]. Marine red algae is seen as a vital source of novel therapeutic compounds [70,71]. Natural compounds from the red algal genus Laurencia may prove to be vital in the treatment of SARS-CoV-2 infection due to their antiviral potency [72,73].
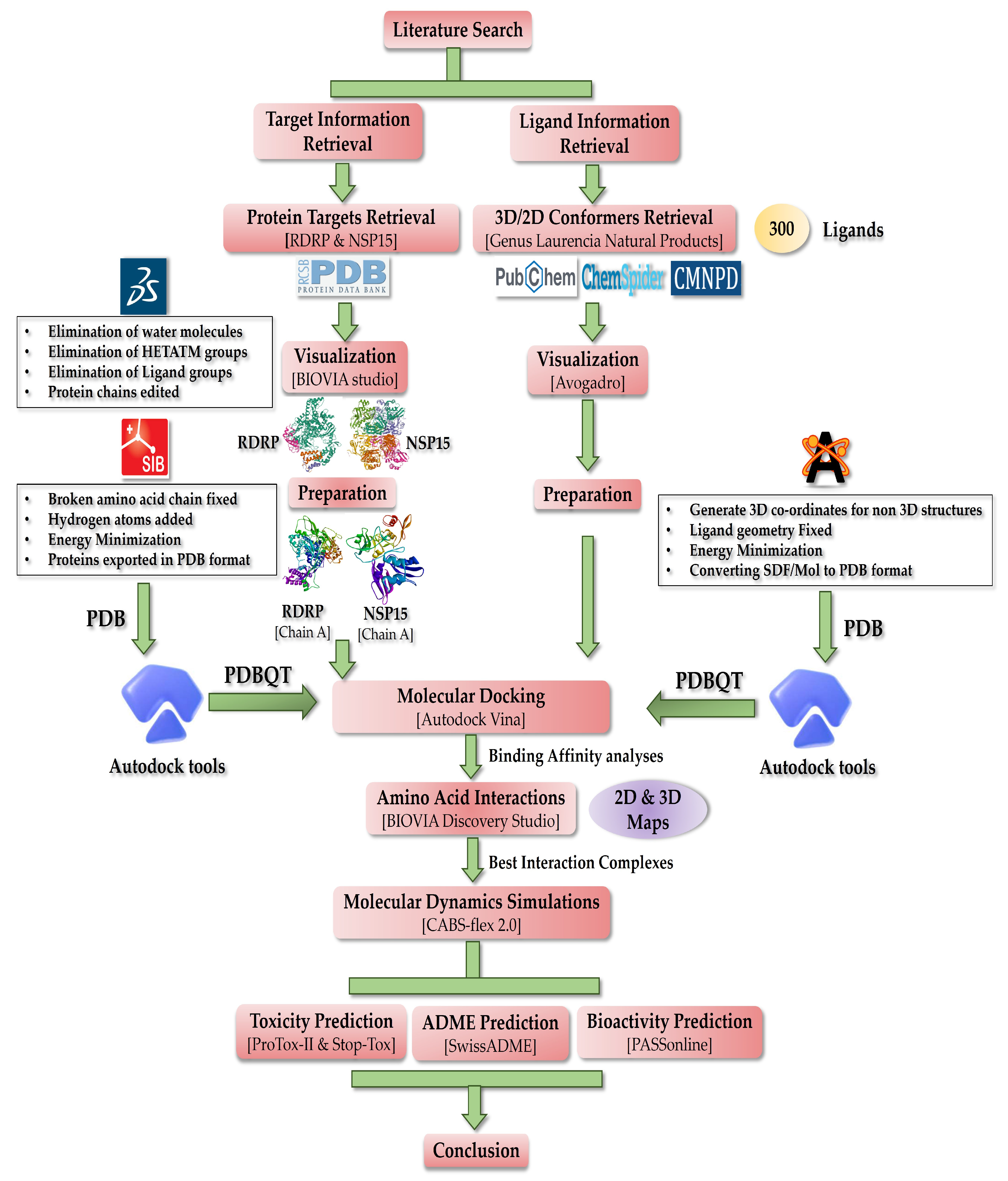
The purpose of this study is in silico evaluation of antiviral activity of 300 natural products from a marine red algal genus Laurencia (refer to Table S1). By focusing more on the amino acid interactions than just the binding affinities, this study was able to identify a small number of NPs from a total of 300 NPs [74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172]. This study is geared toward suggesting some low molecular weight NPs as potential SARS-CoV-2 RdRp (nsp12) and endoribonuclease (nsp15) inhibitors. For the RdRp enzyme, the two docking sites employed in the study are frequently referred to as active site 1 (AS1) and active site 2 (AS2); these active sites are hypothesized to be linked allosterically and coordinate the enzyme activity [173]. As there are not many studies emphasizing the relevance of docking the N-terminal (AS2), it was the major emphasis of this investigation. To ascertain if lead compounds are effective antiviral candidates, it is very crucial to take into account the activity of both AS1 and AS2 [173]. Thus, a two-phase docking strategy was used. First, the β-hairpin motif situated in the NiRAN region of the N-terminal was targeted, since it is crucial for stabilizing the complete protein and ensuring optimal functioning. Second, the top ligands shortlisted from the N-terminal docking were then further docked in the C-terminal portion of RdRp (AS1), consisting of the main catalytic core responsible for RNA replication. Finally, the toxicity and pharmacokinetic characteristics of the screened NPs identified for both target proteins were explored. A schematic depicting the workflow used for this computational study is shown in Figure 1.
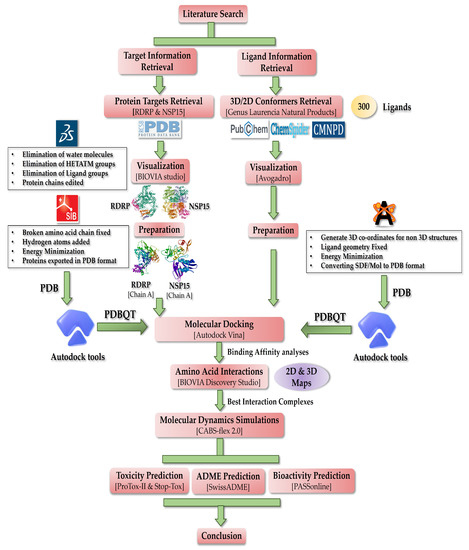
Figure 1.
Schematic of the workflow describing methodology applied in this work.
2. Materials and Methods
2.1. Retrieval of Proteins
Data on the common COVID-19 proteins were found in the literature. The RCSB PDB database was used to locate the crystal structures of SARS-CoV-2 RdRp and nsp15, which were produced using electron microscopy and X-ray diffraction techniques, respectively [174]. The RNA-dependent RNA polymerase, generally known as RdRp or nsp12, and its 3D structure (PDB ID: 7BV2) resolved at 2.50 Å, were used for this investigation [175]. The 3D structure of non-structural protein number 15, also known as nsp15, which was chosen (PDB ID 6VWW), resolved at 2.20 Å [61]. The targets were selected on the basis of resolution and PDB validation reports for optimum metrics to ensure a better docking process.
2.2. Retrieval of Ligands
A thorough literature search was conducted in order to learn more about the natural compounds produced by the Laurencia genus of the marine red algae. Despite the fact that there are hundreds of natural compounds, this study was restricted to about 300 naturally occurring products from this genus (refer to Table S1). To find 2D and/or 3D conformers of all 300 ligands, a thorough search was conducted. The PubChem database’s 3D conformers of ligands were initially obtained in SDF chemical format [176]. In the event that 3D structures were not available, 2D conformers were downloaded. The ligands that were absent from the PubChem database were obtained from the ChemSpider database and retrieved in the Mol chemical format using the same database [177]. The 2D Mol format structures of ligands that were not accessible in the PubChem or ChemSpider databases were obtained using the CMNPD database [178].
2.3. Visualization Tools
Using BIOVIA Discovery Studio Visualizer 4.5, the protein structures of RdRp and nsp15 were studied [179]. This gave us the necessary details on the quantity of amino acid chains, connected ligands, HETATM groups, etc. However, Avogadro software was first employed for visualization and afterwards for the preparation of ligands [180,181].
2.4. Pre-Docking Preparations
Using the data gathered throughout the visualization process, BIOVIA Discovery Studio Visualizer prepared both target proteins. The nsp15 protein only had chains A and B, but the RdRp protein contained three amino acid chains, such as A, B, and C. Both proteins demonstrated the presence of ligand groups, HETATM, and water molecules. However, RdRp was also found to have a nucleic acid group. In order to get rid of unnecessary molecules (H2O, HETATM, nucleic acid, and undesired amino acid chains), 3D structures were cleaned and trimmed. For both proteins to be utilized in this investigation, Chain A was chosen, but the ligand group was left intact. Hydrogen atoms were also added, energy minimization was performed, and Swiss-pdb viewer software [182] was utilized to perhaps restore the fragmented amino acid chain. After the energy minimization procedure, the ligand groups were eliminated; this step was important to prevent potential disruption of the active site. Additionally, these structures were stored in the Swiss-pdb viewer’s PDB format before being converted utilizing Autodock tools to the necessary PDBQT format [183].
The Avogadro program was used to construct the 2D and 3D conformers of ligands acquired in SDF and Mol chemical formats. The 2-dimensional structures were first opened in Avogadro to generate 3D coordinates, which turned them into 3-dimensional structures. All 2D structures were thus transformed to 3D conformers and stored. Once all 300 products had been converted to 3D, each one was individually put through a process of geometry optimization and energy minimization. All ligands that were processed were exported in PDB format. Finally, Gasteiger charges were added using the auto dock tools by converting ligands from PDB to PDBQT format.
2.5. Docking Protocol Validation
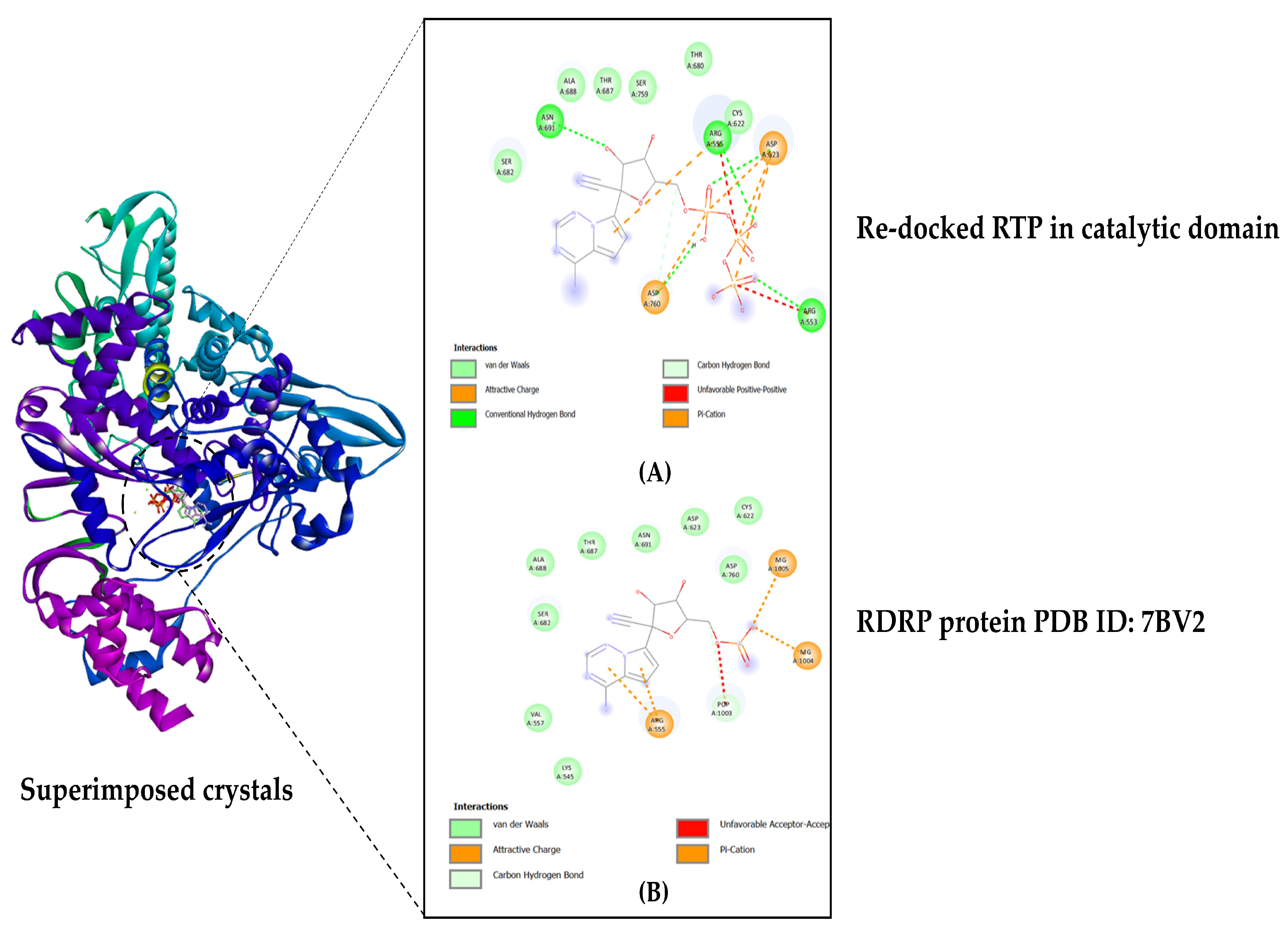
By re-docking the original ligand RTP (remdesivir triphosphate) in the catalytic domain on the C-terminal, the docking technique was validated. On the crystal structure of the protein that was re-docked with the native RTP, the original crystal structure (PDB ID: 7BV2) was superimposed. Analyses were performed on the map of amino acid interactions and docking score was noted. To evaluate the precision of the employed protocol, the RMSD value was determined (see Figure 2).
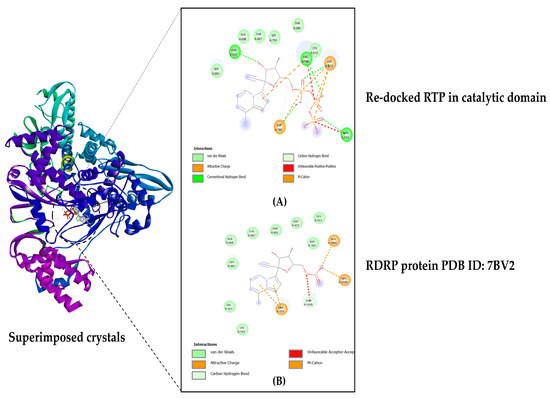
Figure 2.
Docking validation: (A) RTP re-docked in the active site pocket located in the C-terminal; (B) Original protein crystal from RCSB PDB, ID: 7BV2.
2.6. Molecular Docking
For the rigid protein flexible ligand molecular docking approach, Autodock Vina 1.2.0 was employed [184,185]. The Grid box was constructed and positioned suitably using knowledge of key amino acids and active sites for RdRp and nsp15. The RdRp protein file in PDBQT format was first opened in the program, and 300 ligands were individually docked (see Table 1). RdRp’s cavity volume was 1248 Å3, whereas the nsp15 protein’s active site volume was 83 Å3. For RdRp phase 1 docking, the grid box was constructed over the β-hairpin motif and positioned at X = 113 Å, Y = −87 Å, Z = 79 Å. In the phase 2 docking, the top NPs identified in phase 1 docking were docked in the grid box placed at the catalytic core region X = 99 Å, Y = 96 Å, Z = 104 Å. In the case of nsp15 protein, grid box was placed at X = −93 Å, Y = 19 Å, Z = −27 Å. The nsp15 protein was also docked with all 300 ligands (see Table 2). To confirm the correctness of the outcome, the full docking process was carried out twice. Additionally, all protein–ligand complexes in PDB format were acquired in order to use BIOVIA Discovery Studio Visualizer 4.5 to show amino acid interactions (see Table 3) in 2D and 3D maps (see Figures 3–6).
2.7. MD Simulation
Using the CABS-flex 2.0 website, MD simulations were performed for protein–ligand complexes exhibiting the desired amino acid interactions [186]. This service effectively assesses protein flexibility using quick simulations. PDB files for the best complexes were posted to the server. The server gave default settings for the simulation parameters, which included the number of cycles (50), cycles between trajectory frames (50), simulation temperature (1.4), and seed for the random number generator. At default settings, these MD simulation sessions are considered equivalent to 10 ns of computer performed simulations. Root Mean Square Fluctuation (RMSF) plots were produced by simulation sessions which were used further in this study (see Figures 7–9 and Tables S13–S15).
2.8. Toxicity Profile Assessment
ProTox II and StopTox web servers were employed to assess the potential toxicity of the top-ranking ligands [187,188]. ProTox-II reduces the requirement for animal testing by making quick predictions about a tiny molecule’s potential toxicity in a virtual environment. Important details regarding the ligands, including LD50 values (mg/kg body weight), toxicity class, and toxicity endpoints (hepatotoxicity, carcinogenicity, immunotoxicity, mutagenicity, and final cytotoxicity), were given by this website (see Table 4). Estimates of the likelihood that ligands may cause acute toxicity are quickly and accurately provided by the user-friendly program StopTox. Acute oral, dermal, and inhalation toxicity information, as well as information on the likelihood of skin and eye irritation and corrosion, skin sensitization, and other critical information, were all provided by this server (refer to Table 5).
2.9. Pharmacokinetics Analysis
The canonical SMILES (Table S16, ESI) on the Swiss-ADME service was used to investigate the drug-like qualities of natural compounds that had previously been selected as the best [189]. The five Lipinski parameters (Ro5) were observed and recorded as the first act of observation. These criteria included the molecular weight (≤500), consensus log p-value (≤5), and the number of hydrogen bond donors and acceptors (≤5). Two more criteria were included—as the topological polar surface area (≤140) and the number of rotatable bonds (≤10)—in order to more accurately assess the likelihood that the ligand will be orally active (refer to Table 6). Further, the water solubility, bioavailability, and GI absorption probability were noted (see Table 7). A simple mathematical equation that produced the absorption percentage was used to more precisely measure GI absorption [190,191]:
Additionally, the Swiss-ADME boiled egg graph was used to analyze the cell permeability of NPs (see Figure 10). Finally, utilizing the PASS online webserver [192], possible biological activity prediction was carried out (in Table 8).
3. Results and Discussion
3.1. Binding Affinity Studies
Due to the significant number of NPs used in this investigation, the data for the binding affinity for both RdRp and nsp15 were separated into three groups. First, the classes of ligands with low affinity were those having docking scores between −4.0 and −4.9 kcal/mol and −5.0 and −5.9 kcal/mol. Second, it was considered that the ligands having docking values between −6.0 and −7.9 kcal/mol were of intermediate affinity. These low and moderate ligands were further placed in subcategories, whereas high-affinity ligands were the final group.
3.1.1. Docking β-Hairpin Motif Region (RdRp): N-Terminal
The NiRAN region and the palm subdomain of RdRp are linked by a β-hairpin motif. This motif stabilizes the overall protein structure of RdRp, aiding in its optimum functioning. Important amino acids in this region are TYR: 32, LYS: 47, TYR: 122, TYR: 129, HIS: 133, ASN: 138, ASP: 140, THR: 141, SER: 709, and ASN: 781 [193,194,195,196]. Targeting this region might inhibit RdRp by disruption of conformational dynamics triggered by ligand binding. In simple terms, there might be structural re-arrangements that ultimately alter the protein function [197]. The residue TYR: 129 from AS2 (N-terminal) and SER: 709 from AS1 (C-terminal) are conserved among all coronaviruses. Targeting these residues will most likely compromise the inter-domain connections, which will eventually have a detrimental effect on the activity of RNA synthesis [173]. The results of the phase 1 docking process for the β-hairpin motif of RdRp were divided into the three major groups.
Ten NPs made up the first grouping in the low category because they had the lowest docking scores for the RdRp protein (see Table 1). The affinities of laurencenyne, halomon, and 9-octadecanoic acid were the lowest at −4.0 kcal/mol. 6, 8-cycloeudesmane had the greatest affinity score, which was closely followed by 1-methyl-2, 3 and 5-tribromoindole at −4.9 and −4.8 kcal/mol, respectively (refer to Table S2, ESI). The second subgroup had about 172 NPs, 19 of which had a low docking value of −5.0 kcal/mol. Nevertheless, only five of the NPs in this group showed the maximum docking scores in this class, which was −5.9 kcal/mol (refer to Table S3, ESI). These were (6R, 9R, 10S)-10-bromo-9-hydroxychamigra-2, 7(14)-diene, aldingenin C, axinysone B, aplysiolic acid, and compositacin C, J. There were two subcategories within the moderate class. As well, 61 NPs with lower-moderate scores were put in the first sub-category, 20 of which had affinities of −6.0 kcal/mol; nevertheless, five NPs, including 11,14-dihydroaplysia-5,11,14,15-tetrol, beta-sitosterol, callophycoic acid I, cholest-5-en-3-alpha-ol, and saringosterol, had the lowest score in the group at −6.9 kcal/mol (refer to Table S4, ESI). The second subcategory of upper-moderate ligands included 39 NPs, of which 6 NPs had a low docking score of −7.0 kcal/mol while 4 NPs, namely beta-cryptoxanthin, brassicasterol, bromophycoic acid E, and bromophycolide A, displayed −7.9 kcal/mol affinity (refer to Table S5, ESI). Ultimately, we identified 18 NPs in category 3 with the greatest affinities (see Table 1). Lithothamin A had the greatest docking score for RdRp of any ligand, with a value of −9.0 kcal/mol, followed closely by dehydrothyrsiferol, bromophycolide S, callicladol, and bromophycolide E, which had values of −8.9, −8.8, −8.7, and −8.5 kcal/mol, respectively (Table S6, ESI).

Table 1.
Docking output for the N-Terminal β-hairpin region of RdRp enzyme.
Table 1.
Docking output for the N-Terminal β-hairpin region of RdRp enzyme.
| RdRp Protein | ||||
|---|---|---|---|---|
| Low | Moderate | High | ||
| −4.0 to −4.9 kcal/mol | −5 to −5.9 kcal/mol | −6 to −6.9 kcal/mol | −7 to −7.9 kcal/mol | −8.0 kcal/mol and Above |
| 1-methyl-2,3,5-tribromoindole | (−)-3-(E)-bromomethylidene-10beta-bromo-beta-chamigrene | 1,2-Dehydro-3,4-epoxypalisadin B | 5-alpha-cholestane-3,6-dione | Bromophycoic acid B |
| 6,8-cycloeudesmane | (+)-3-(Z)-bromomethylidene-10beta-bromo-beta-chamigrene | 2-Hydroxyluzofuranone A, B | 6-hydroxycholest-4-en-3-one | Bromophycoic acid C |
| 9-octadecanoic acid | (5S)-5-Acetoxy-beta-bisabolene | 3 alpha-hydroperoxy-3-epiaplysin | 10-acetoxyangasiol | Bromophycolide E |
| 14-methylpentadecanoic acid | (6R,9R,10S)-10-bromo-9-hydroxychamigra-2,7(14)-diene | 3-Bromo-4,5-dihydroxybenzaldehyde | 10-epi-Dehydrothyrsiferol | Bromophycolide H |
| beta-Synderol | (10R)-10-Bromo-alpha-chamigrene | 3-Bromobarekoxide | 13-Hydroxyprethyrsenol A | Bromophycolide K |
| Halomon | 2-bromospironippol | 3-epi-Perforenone A | 15,16-Dehydrovenustatriol | Bromophycolide L |
| Luzonensin | (3Z)-laurenyne | 3R,4S-luzonolone | 15-DehydroxythyrsenolA | Bromophycolide P |
| Hordenine | 3,4-epoxypalisadin B | 3S, 4R-luzonolone | 16-Hydroxydehydrothyrsiferol | Bromophycolide R |
| Laurencenyne | 3,7-dihydroxydihydrolaurene | 5-acetoxypalisadin B | Beta cryptoxanthin | Bromophycolide S |
| Trans-Laurencenyne | 3-alpha-Hydroxydebromoaplysin | 5-alpha-Hydroxyaplysistatin | Brassicasterol | Callicladol |
| 3-beta-Hydroperoxyaplysin | 9-hydroxy-3-epi-perforenone A | Bromophycoic acid A, D, E | Dehydrothyrsiferol | |
| 4-Hydroxy-1,8-epi-isotenerone | 11,14-dihydroaplysia-5,11,14,15-Tetrol | Bromophycolide A–D, F, J, M–O, Q, T, U | Isodehydrothyrsiferol | |
| 4-hydroxypalisidin C | 15-hydroxypalisadin A | Callophycoic acid C–E | Laurebiphenyl | |
| 5-acetoxyoxachamigrene | Acetylmajapolene A, B | Campesterol | Lithothamin A | |
| 5-epi-maneolactone | Aldingenin D | Cholest-4-en-3,6-dione | Mammeisin | |
| 7-acetyl-aplysiol | Aplysistatin | Dehydrovenustatriol | Thyrsenol A | |
| 7-hydroxylaurene | Aristolan-10-ol-9-one | Lactodehydrothyrsiferol | Thyrsenol B | |
| 8,10-dibromoisoaplysin | Aplysiodiol | Neurymenolide A, B | Thyrsiferol | |
| 9-Deoxyelatol | Barekoxide | Neoirietetraol | ||
| 10-Bromo-beta-chamigrene | beta-Sitosterol | Predehydrovenustatriolacetate | ||
| 10-Bromosoaplyin | Bromophycolide G, I | Prethyrsenol A | ||
| 10-hydroxyisolaurene | Caespitol | Pseudodehydrothyrsiferol | ||
| 10-Hydroxyaplysin | Callophycoic acid A, B, G, H, I, J | Stigmasterol | ||
| 12-hydroxy Isolaurene | Callophycol A, B | |||
| 12-Hydroxypalisadin B | Chamigrene Lactone | |||
| 15-Hydroxylaurene | Compositacin B, E, F, I, N | |||
| 15-Oxolaurene | Cholest-4-en-3-one | |||
| Almadioxide | Cholest-5-en-3alpha-ol | |||
| Aromadendrene | Cholest-5-en-3beta-ol | |||
| Aldingenin A, B, C | Debromoisocalenzanol | |||
| Allolaurinterol | Japonenyne A | |||
| Allolaurinterolacetate | Johnstonol | |||
| Aplysinol | Laurecomin A, D | |||
| Aristolan-8-en-1-one | Laurenokomarin | |||
| Aristolane | Laurepoxyene | |||
| Axinysone B | Laureacetal C | |||
| Aplysiol-7-one | Laurefurenyne B | |||
| Aplysiolic acid | Laurinterolacetate | |||
| (−)-BisezakyneA | Luzondiol | |||
| Brasilenol | Majapolene A | |||
| Bromocuparene | Oryzalexin S | |||
| Bromocyclococanol | Pacifenol | |||
| Bosseopentanoic acid | Palisadin D | |||
| Bromlaurenidificin | Perforenone A | |||
| Caespitane | Saringosterol | |||
| Caesspitenone | Tiomanene | |||
| Cycloeudesmol | ||||
| Cyclolaurenol | ||||
| Callophycoic acid F | ||||
| Callenzanol | ||||
| Chamigrene epoxide | ||||
| Chinzallene | ||||
| Compositacin A, C, D, G, H, J, K, L, M | ||||
| Cycloelatanene A, B | ||||
| Cycloisoallolaurinterol | ||||
| Chlorofucin | ||||
| Cupalaurenol | ||||
| Dactylyne | ||||
| Debromoaplysin | ||||
| Debromoepiaplysinol | ||||
| Debromolaurinterol | ||||
| Debromolaurinterolacetate | ||||
| Dendroidiol | ||||
| Dendroidone | ||||
| Deoxyprepacifenol | ||||
| Deschloroelatol | ||||
| Elatenyne | ||||
| Elatol | ||||
| Epiaplysinol | ||||
| Epibrasilenol | ||||
| Floridoside | ||||
| Filiformin | ||||
| Filiforminol | ||||
| Guimarediol | ||||
| Heterocladol | ||||
| Intricenyne | ||||
| Isocaespitol | ||||
| Isodihydrolaurene | ||||
| Isolaurallene | ||||
| Isolaureatin | ||||
| Isolaurene | ||||
| Itomanindole A | ||||
| Isolaurenidificin | ||||
| Isoafricanol | ||||
| Isoallolaurinterol | ||||
| Isoaplysin | ||||
| Isodactyloxene A | ||||
| Isodebromolaurinterol | ||||
| Isolaurenisol | ||||
| Isoobtusol | ||||
| Isopalisol | ||||
| Isorigidol | ||||
| Kumausallene | ||||
| Laurallene | ||||
| Laureacetal A, B | ||||
| Laurefurenyne A, C, D, E, F | ||||
| Laurencial | ||||
| Laurendecumallene A, B | ||||
| Laurendecumenyne A, B | ||||
| Laurenenyne | ||||
| Laurenisol | ||||
| Laurenone A | ||||
| Laureoxanyne | ||||
| Laurecomin B, C | ||||
| Laurencomposidiene | ||||
| Laurene | ||||
| Laurentristich-4-ol | ||||
| Laureperoxide | ||||
| Laurokamurene A-D | ||||
| Luzofuran | ||||
| Luzonenone | ||||
| Luzonensol | ||||
| Luzonensol acetate | ||||
| Laurepinnacin | ||||
| Laurinterol | ||||
| Ma’iliohydrin | ||||
| Mailione | ||||
| Majapolene B | ||||
| Microcladallene A-C | ||||
| Neoisoprelaurefucin | ||||
| Neolaurallene | ||||
| Nidificene | ||||
| Notoryne | ||||
| Octadecanedioic acid | ||||
| Obtusane | ||||
| Okamurene A-E | ||||
| Omaezallene | ||||
| Oxachamigrene | ||||
| Palisadin A-C | ||||
| Pannosane | ||||
| Pannosanol | ||||
| Perforatone | ||||
| Perforenol | ||||
| Prepacifenol | ||||
| Prelaureatin | ||||
| Rhodophytin | ||||
| Scopariol | ||||
| Seco-Laurokamurone | ||||
| Spirolaurenone | ||||
| Trans-Deacetylkumausyne | ||||
| Trans-Kumausyne | ||||
3.1.2. Validation of RdRp Catalytic Domain Docking Protocol
After being re-docked, the original inhibitor remdesivir triphosphate gave a binding affinity score of −7.1 kcal/mol, and the RMSD value for the superimposed crystal structures was calculated to be 1.51 Å. The computed RMSD value was below the threshold of 2.0 Å, indicating that the docking operation was conducted with good precision (see Figure 2).
3.1.3. Docking Catalytic Domain (RdRp): C-Terminal
After addressing active site 2 (N-terminal), we proceeded to move on to the active site 1 (C-terminal) to determine the applicability of already filtered compounds in step 1. This dual-step docking assisted in narrowing down the pool of candidates in a systematic but progressive fashion. In addition, the likelihood of discovering potent inhibitors that might bind and favorably interact with both AS1 and AS2 increased. The AS1 consists of two catalytic motifs. The first motif includes amino acid residues from 611–626. Out of which, the ASP: 618 is of paramount importance, as it is the most conserved residue. On the other hand, the second motif consists of residues ranging from 753–767. The three catalytic residues such as SER: 759, ASP: 760, and ASP: 761 are considered crucial for RdRp activity [198,199]. The highest docking score of −8.2 kcal/mol was seen for bromophycoic acid B and callicladol, whereas bromophycolide L obtained the lowest value, −7.0 kcal/mol. All results for the C-terminal docking are presented in Table S7, ESI of the supplemental information.
3.1.4. Docking nsp15
The docking process produced the output listed (in Table 2) for the endoribonuclease enzyme (nsp15) of SARS-CoV-2.
Seven NPs were found to have the lowest docking scores to the endoribonuclease protein. Halomon exhibited the lowest result, which was −4.2 kcal/mol. Bosseopentanoic and octadecanedioic acids had the strongest binding score in this category, at −4.9 kcal/mol (refer to Table S8, ESI). The second low-class subcategory had 110 NPs, 15 of which displayed the maximum docking values in this group. Nevertheless, at −5.0 kcal/mol, floridoside and laurefurenyne A had the lowest ability for binding (refer to Table S9, ESI). There were around 111 NPs in the first subcategory of the moderate class, and 41 of them displayed the lowest docking scores in the group. Nevertheless, the highest binding scores were demonstrated by five NPs, including allolaurinterol, aplysiodiol, callophycoic acid G, isolaurenisol, and mammeisin, with a −6.9 kcal/mol value (refer to Table S10, ESI). Additionally, 42 NPs made up the upper-moderate category. The absolute minimum affinity score for this group, −7.0 kcal/mol, was displayed by 11 NPs. The maximum binding value was displayed by two NPs, 15, 16-dehydrovenustatriol, and bromophycoic acid E, with a value of −7.9 kcal/mol (refer to Table S11, ESI). Lastly, the final category included the remaining 30 NPs (see Table 2). With −8.7 and −8.6 kcal/mol, respectively, bromophycolide T and 5 alpha-cholestane-3, 6-dione showed the maximum binding potential in this group. Ten NPs, however, showed docking scores of −8.0 kcal/mol only (refer to Table S12, ESI).

Table 2.
Docking output for nsp15 enzyme.
Table 2.
Docking output for nsp15 enzyme.
| Categorization of Ligands—nsp15 | ||||
|---|---|---|---|---|
| Low | Moderate | High | ||
| −4.0 to −4.9 kcal/mol | −5 to −5.9 kcal/mol | −6 to −6.9 kcal/mol | −7 to −7.9 kcal/mol | −8.0 kcal/mol and Above |
| 9-Octadecanoic acid | 1-Methyl-2,3,5-tribromoindole | (−)-3-(E)-Bromomethylidene-10-beta-bromo-beta-chamigrene | 2-Hydroxyluzofuranone A | 5-alpha-Cholestane-3,6-dione |
| 14-Methylpentadecanoic acid | 2-Bromospironippol | (+)-3-(Z)-Bromomethylidene-10-beta-bromo-beta-chamigrene | 3-alpha-Hydroperoxy-3-epiaplysin | 6-hydroxycholest-4-en-3-one |
| Bosseopentanoic acid | 3,4-epoxypalisadin B | (5S)-5-Acetoxy-beta-bisabolene | 5-alpha-Hydroxyaplysistatin | 15-Dehydroxythyrsenol A |
| Halomon | 3-alpha-Hydroxydebromoaplysin | (6R,9R,10S)-10-Bromo-9-hydroxychamigra-2,7(14)-diene | 10-epi-Dehydrothyrsiferol | 16Hydroxydehydrothyrsiferol |
| Hordenine | 4-Hydroxy-1,8-epi-isotenerone | (10R)-10-Bromo-alpha-chamigrene | 11,14-Dihydroaplysia-5,11,14,15-tetrol | Brassicasterol |
| Octadecanedioic acid | 5-Acetoxyoxachamigrene | 1,2-Dehydro-3,4-epoxypalisadin B | 13-Hydroxyprethyrsenol A | beta-sitosterol |
| Tiomanene | 5-Acetoxypalisadin B | 2-Hydroxyluzofuranone B | 15,16-Dehydrovenustatriol | Bromophycoic acid B |
| 6,8-Cycloeudesmane | 3,7-dihydroxydihydrolaurene | Aplysistatin | Bromophycoic acid C | |
| 7-Acetyl-aplysiol | 3-beta-Hydroperoxyaplysin | Beta cryptoxanthin | Bromophycolide G | |
| 9-Deoxyelatol | 3-Bromo-4,5-dihydroxybenzaldehyde | Bromophycoic acid A, D, E | Bromophycolide K | |
| 10-Bromo-beta-chamigrene | 3-epi-PerforenoneA | Bromophycolide A–F, H, I, J, M, N, R | Bromophycolide L | |
| Aristolan-8-en-1-one | 3R,4S-Luzonolone | Barekoxide | Bromophycolide O | |
| Aristolan-10-ol-9-one | 3S, 4R-Luzonolone | Bromophycolide S | Bromophycolide P | |
| Aristolane | (3Z)-Laurenyne | Callophycoic acid A, C, D–F, H, I | Bromophycolide Q | |
| Almadioxide | 3-Bromobarekoxide | Cholest-5-en-3-alpha-ol | Bromophycolide T | |
| Aromadendrene | 4-Hydroxypalisidin C | Dehydrovenustatriol | Bromophycolide U | |
| Aplysiol-7-one | 5-epi-Maneolactone | Isodehydrothyrsiferol | Callicladol | |
| Aplysiolic acid | 7-Hydroxylaurene | Lactodehydrothyrsiferol | Callophycoic acid B | |
| beta-Synderol | 8,10-Dibromoisoaplysin | Neurymenolide B | Campesterol | |
| Brasilenol | 9-Hydroxy-3-epi-perforenone A | Predehydrovenustatriolacetate | Cholest-4-en-3,6-dione | |
| (−)-BisezakyneA | 10-Acetoxyangasiol | Prethyrsenol A | Cholest-4-en-3-one | |
| Callenzanol | 10-Bromosoaplyin | Pseudodehydrothyrsiferol | Cholest-5-en-3-beta-ol | |
| Chamigrene epoxide | 10-Hydroxyaplysin | Saringosterol | Dehydrothyrsiferol | |
| Chinzallene | 10-hydroxyisolaurene | Laurebiphenyl | ||
| Compositacin A, D-H, L, M, N | 12-hydroxy Isolaurene | Lithothamin A | ||
| Cycloelatanene A, B | 12-Hydroxypalisadin B | Neurymenolide A | ||
| Cycloeudesmol | 15-hydroxypalisadin A | Stigmasterol | ||
| Cyclolaurenol | 15-Hydroxylaurene | Thyrsenol A | ||
| Dactylyne | 15-Oxolaurene | Thyrsenol B | ||
| Debromoepiaplysinol | Acetylmajapolene A | Thyrsiferol | ||
| Dendroidiol | Acetylmajapolene B | |||
| Dendroidone | Aldingenin A-D | |||
| Deoxyprepacifenol | Allolaurinterol | |||
| Deschloroelatol | Allolaurinterolacetate | |||
| Elatol | Aplysinol | |||
| Epibrasilenol | Axinysone B | |||
| Elatenyne | Aplysiodiol | |||
| Floridoside | Bromocyclococanol | |||
| Guimarediol | Bromlaurenidificin | |||
| Heterocladol | Bromocuparene | |||
| Isolaurenidificin | Callophycoic acid G, J | |||
| Isoafricanol | Callophycol A, B | |||
| Isoaplysin | Chamigrene Lactone | |||
| Isodactyloxene A | Compositacin B, C, I, J, K | |||
| Isoobtusol | Cycloisoallolaurinterol | |||
| Isopalisol | Chlorofucin | |||
| Isorigidol | Cupalaurenol | |||
| Intricenyne | Caespitane | |||
| Isolaurallene | Caespitol | |||
| Itomanindole A | Caesspitenone | |||
| Kumausallene | Debromoisocalenzanol | |||
| Laurecomin B, C | Debromolaurinterol | |||
| Laurenokomarin | Debromolaurinterolacetate | |||
| Luzondiol | Debromoaplysin | |||
| Luzonensin | Epiaplysinol | |||
| Luzonensol | Filiformin | |||
| Luzonensolacetate | Filiforminol | |||
| Laurallene | Isoallolaurinterol | |||
| Laureacetal B, C | Isodebromolaurinterol | |||
| Laurefurenyne A, B, D, E, F | Isolaurenisol | |||
| Laurencenyne | Isocaespitol | |||
| Laurencial | Isodihydrolaurene | |||
| Laurendecumallene A, B | Isolaureatin | |||
| Laurendecumenyne B | Isolaurene | |||
| Laurenenyne | Japonenyne A | |||
| Laurenisol | Johnstonol | |||
| Laureoxanyne | Laurecomin A, D | |||
| Laurepinnacin | Laurencomposidiene | |||
| Ma’iliohydrin | Laurene | |||
| Mailione | Laurentristich-4-ol | |||
| Microcladallene A-C | Laureperoxide | |||
| Neoisoprelaurefucin | Laurepoxyene | |||
| Neolaurallene | Laurinterolacetate | |||
| Nidificene | Laurokamurene A-D | |||
| Obtusane | Luzofuran | |||
| Okamurene C, E | Luzonenone | |||
| Omaezallene | Laurinterol | |||
| Pacifenol | Laureacetal A | |||
| Palisadin B, C, D | Laurefurenyne C | |||
| Pannosane | Laurendecumenyne A | |||
| Perforatone | Laurenone A | |||
| Perforenol | Majapolene A, B | |||
| Prelaureatin | Mammeisin | |||
| Rhodophytin | Neoirietetraol | |||
| Scopariol | Notoryne | |||
| trans-Deacetylkumausyne | Okamurene A, B, D | |||
| trans-Kumausyne | Oxachamigrene | |||
| trans-Laurencenyne | Oryzalexin S | |||
| Palisadin A | ||||
| Pannosanol | ||||
| Perforenone A | ||||
| Prepacifenol | ||||
| Seco-Laurokamurone | ||||
| Spirolaurenone | ||||
3.2. Selection of Best Interaction Complexes—RdRp and nsp15
High binding affinities must not be the only factor used to determine a ligand’s ability to inhibit druggable targets. Understanding the interactions between the residues is crucial to determine the potential success of the ligands in inhibiting the activity of RdRp and nsp15.
3.2.1. Selection Criteria
- It must interact with the majority of catalytic amino acids;
- It must form many hydrogen bonds with catalytic amino acids (RMSD ≤ 3.00);
- For electrostatic connections, the pre-specified threshold RMSD cutoff distance (Å ≤ 5.00) must not be exceeded.
3.2.2. Rejection Criteria
- No interactions with key amino acids;
- No hydrogen bonds are connected to important residues or exceeded RMSD ≥ 3.00;
- Mostly Van der Waals forces are involved in the interactions;
- For electrostatic bonding, the maximum pre-defined RMSD cutoff distance (Å ≤ 5.00) was surpassed.
3.3. Best Interaction Complexes—RdRp and nsp15
Following the aforementioned selection terms and conditions, only 10 docked complexes met our criteria for ideal residue interactions for RdRp and nsp15 out of a total of 618 protein–ligand complexes. Figure 3 showcases the selected ideal docked complexes.
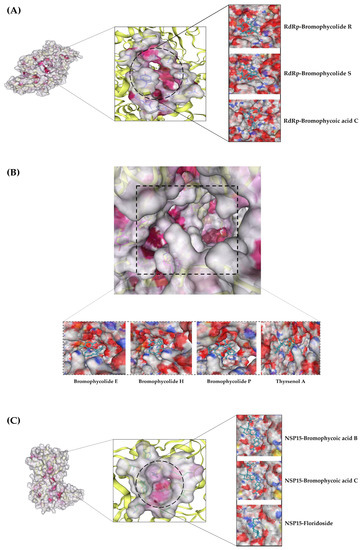
Figure 3.
All shortlisted NPs: (A) RdRp β-hairpin motif in complex with three NPs; (B) RdRp C-Terminal catalytic core in complex with four NPs; (C) Enzyme nsp15 in complex with three NPs.
Figure 3.
All shortlisted NPs: (A) RdRp β-hairpin motif in complex with three NPs; (B) RdRp C-Terminal catalytic core in complex with four NPs; (C) Enzyme nsp15 in complex with three NPs.
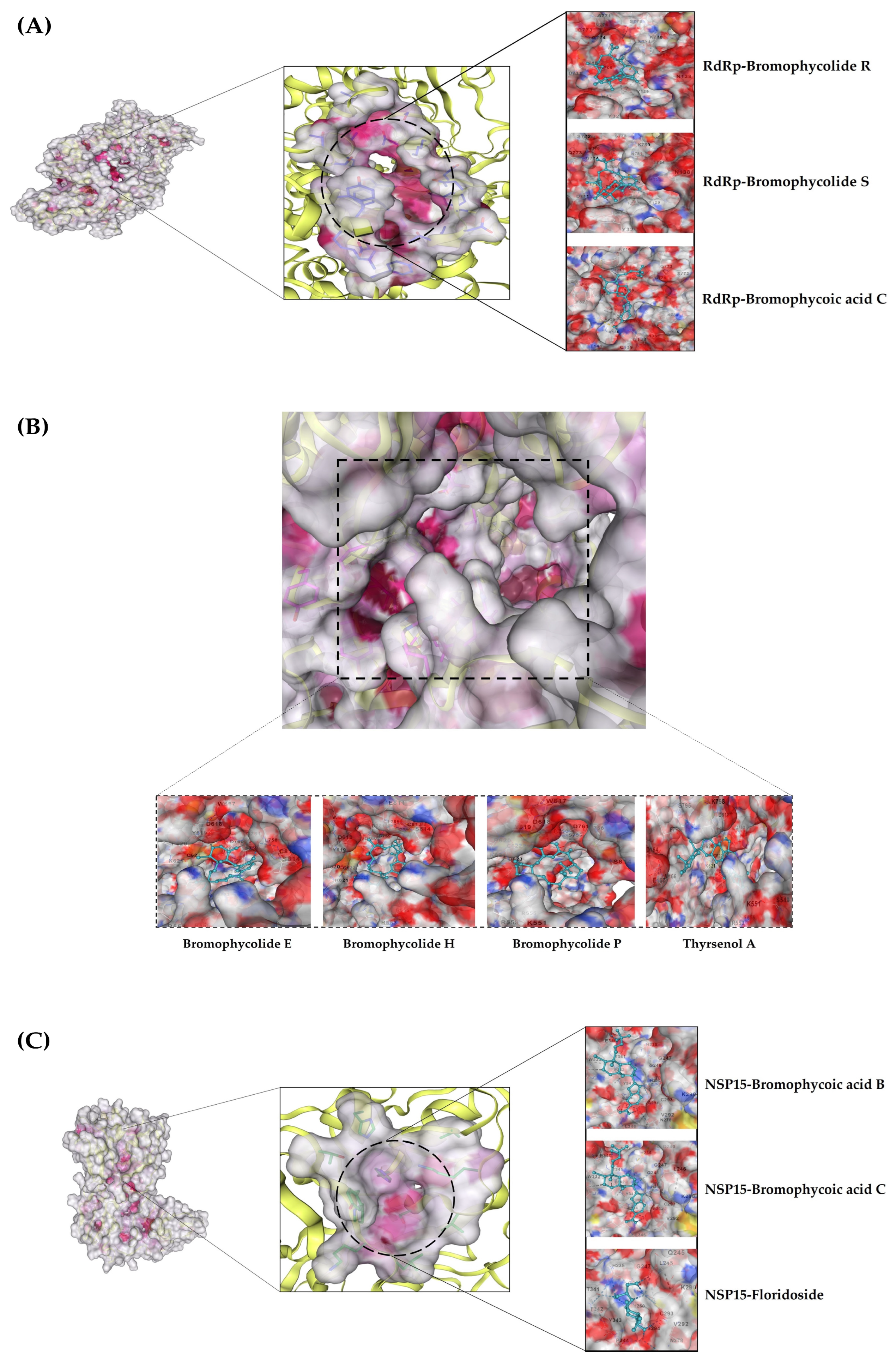
In the case of RdRp (β-hairpin motif), three NPs showed desirable interaction with most crucial amino acids. First, the bromophycolide R interacted strongly with seven key residues out of ten (see Figure 4 and Table 3). It interacted with SER: 709 (2.40 Å) and ASN: 781 (2.47 Å) by forming hydrogen bonds. Pi–Pi T-shaped bonds were formed with TYR: 129 (4.26 Å) and HIS: 133 (4.76 Å). Alkyl and pi–alkyl bonds were seen for TYR: 32 (4.20 Å) and LYS: 47 (3.95 Å). Van der Waals interaction was observed for ASN: 138 residue. Second, the bromophycolide S also interacted with seven residues of interest out of ten. Hydrogen bonds were formed with SER: 709 (2.00 Å) and ASN: 781 (2.66 Å). Pi–Pi T-shaped bond was formed only with TYR: 129 (4.33 Å). Furthermore, alkyl and pi–alkyl bonds were formed with two residues, TYR: 32 (4.20 Å) and LYS: 47 (3.95 Å). Lastly, HIS: 133 and ASN: 138 interacted via VdW forces. Ultimately, the third, bromophycoic acid C, showed interactions with six desired residues out of ten. Two residues, TYR: 129 formed a hydrogen bond (2.39 Å) and a Pi–Pi T-shaped connection (4.14 Å). However, SER: 709 (2.56 Å) formed hydrogen bonds. Pi–cation and pi–alkyl bonds were observed to be forming connections with LYS: 47 (Å = 4.33, 4.25 respectively). Lastly, with TYR: 32 residue, it formed two alkyl bonds (4.72, and 4.79 Å).
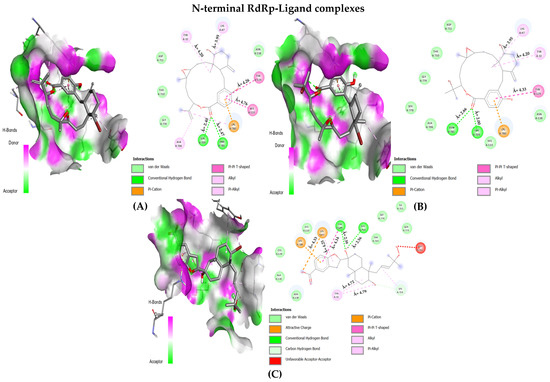
Figure 4.
The 2D interaction maps for phase 1 docking: (A) RdRp β-hairpin domain–bromophycolide R complex; (B) RdRp β-hairpin domain–bromophycolide S complex; (C) RdRp β-hairpin domain–bromophycoic acid C complex.
Figure 4.
The 2D interaction maps for phase 1 docking: (A) RdRp β-hairpin domain–bromophycolide R complex; (B) RdRp β-hairpin domain–bromophycolide S complex; (C) RdRp β-hairpin domain–bromophycoic acid C complex.
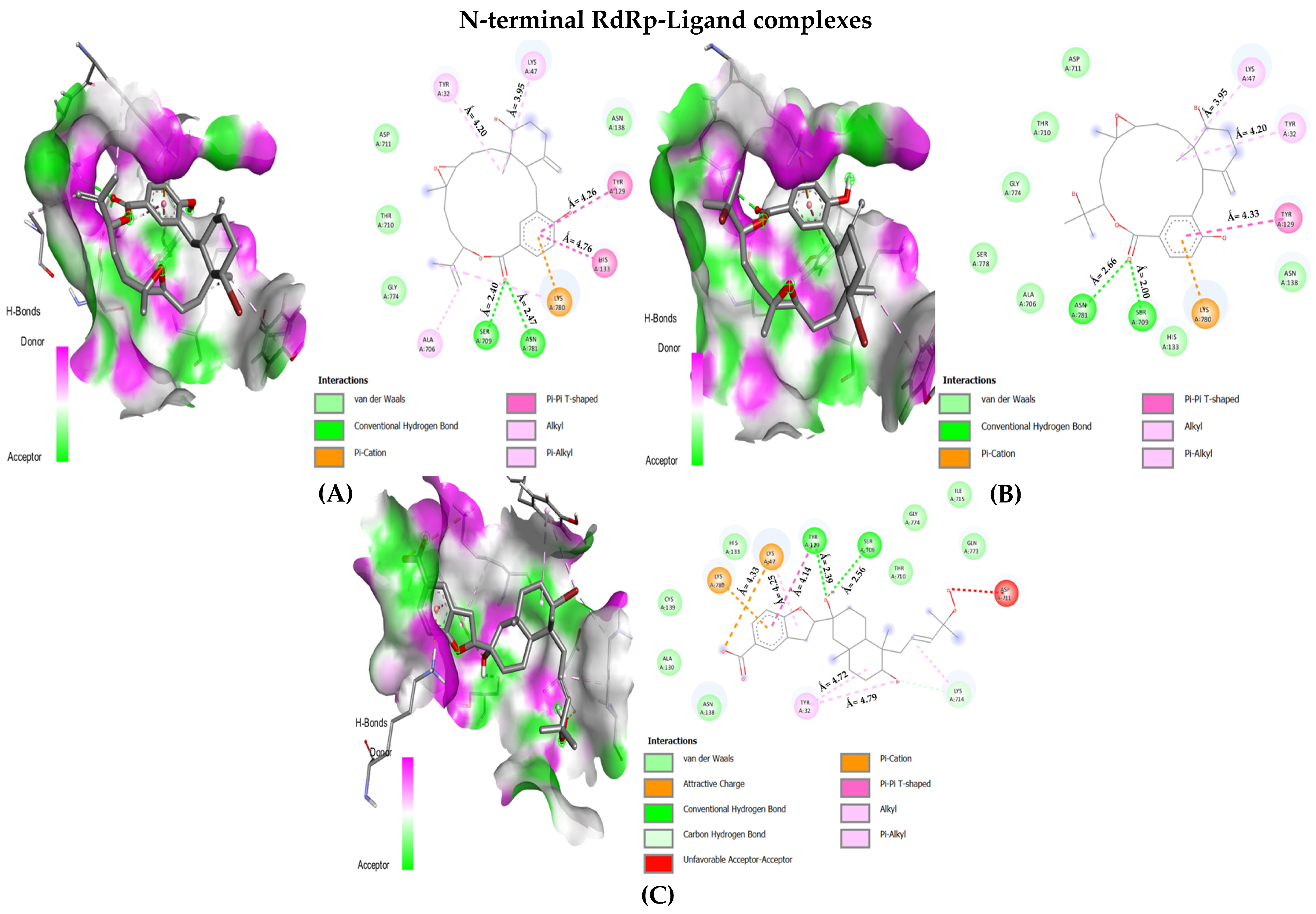

Table 3.
Amino acid interaction profile of filtered nine NPs.
Table 3.
Amino acid interaction profile of filtered nine NPs.
| Favourable Residue Interactions | |
|---|---|
| Filtered Ligands | Involved Amino Acids |
| RdRp (N-Terminal β-Hairpin Motif) | |
| Bromophycolide R | TYR32 LYS47 TYR129 HIS133 ASN138 ALA706 SER709 THR710 ASP711 GLY774 LYS780 ASN781 |
| Bromophycolide S | TYR32 LYS47 TYR129 HIS133 ASN138 ALA706 SER709 THR710 ASP711 GLY774 SER778 LYS780 ASN781 |
| Bromophycoic acid C | TYR32 LYS47 TYR129 ALA130 HIS133 ASN138 CYS139 SER709 THR710 ASP711 LYS714 ILE715 GLN773 GLY774 LYS780 |
| RdRp (C-terminal catalytic core) | |
| Bromophycolide E | ARG553 ARG555 ASP618 TYR619 PRO620 CYS622 ASP623 THR680 SER682 THR687 ALA688 ASN691 LEU758 SER759 ASP760 ASP761 CYS813 |
| Bromophycolide H | ARG553 ARG555 ASP618 LYS621 CYS622 ASP623 LEU758 SER759 ASP760 ASP761 CYS813 |
| Bromophycolide P | ARG553 ARG555 ASP618 TYR619 PRO620 LYS621 CYS622 ASP623 ASP760 ASP761 SER814 |
| Thyrsenol A | ARG555 ASP618 TYR619 PRO620 LYS621 CYS622 ASP623 THR687 ALA688 ASN691 LEU758 SER759 ASP760 ASP761 CYS813 |
| nsp15 enzyme | |
| Bromophycoic acid B | HIS235 GLY247 GLY248 HIS250 LYS290 VAL292 CYS293 SER294 TRP333 GLU340 THR341 TYR343 PRO344 LYS345 LEU346 |
| Bromophycoic acid C | HIS235 GLY247 GLY248 HIS250 LYS290 VAL292 CYS293 SER294 TRP333 GLU340 THR341 TYR343 PRO344 LYS345 LEU346 |
| Floridoside | HIS235 GLN245 LEU246 GLY247 GLY248 HIS250 LYS290 VAL292 CYS293 SER294 THR341 PHE342 TYR343 |
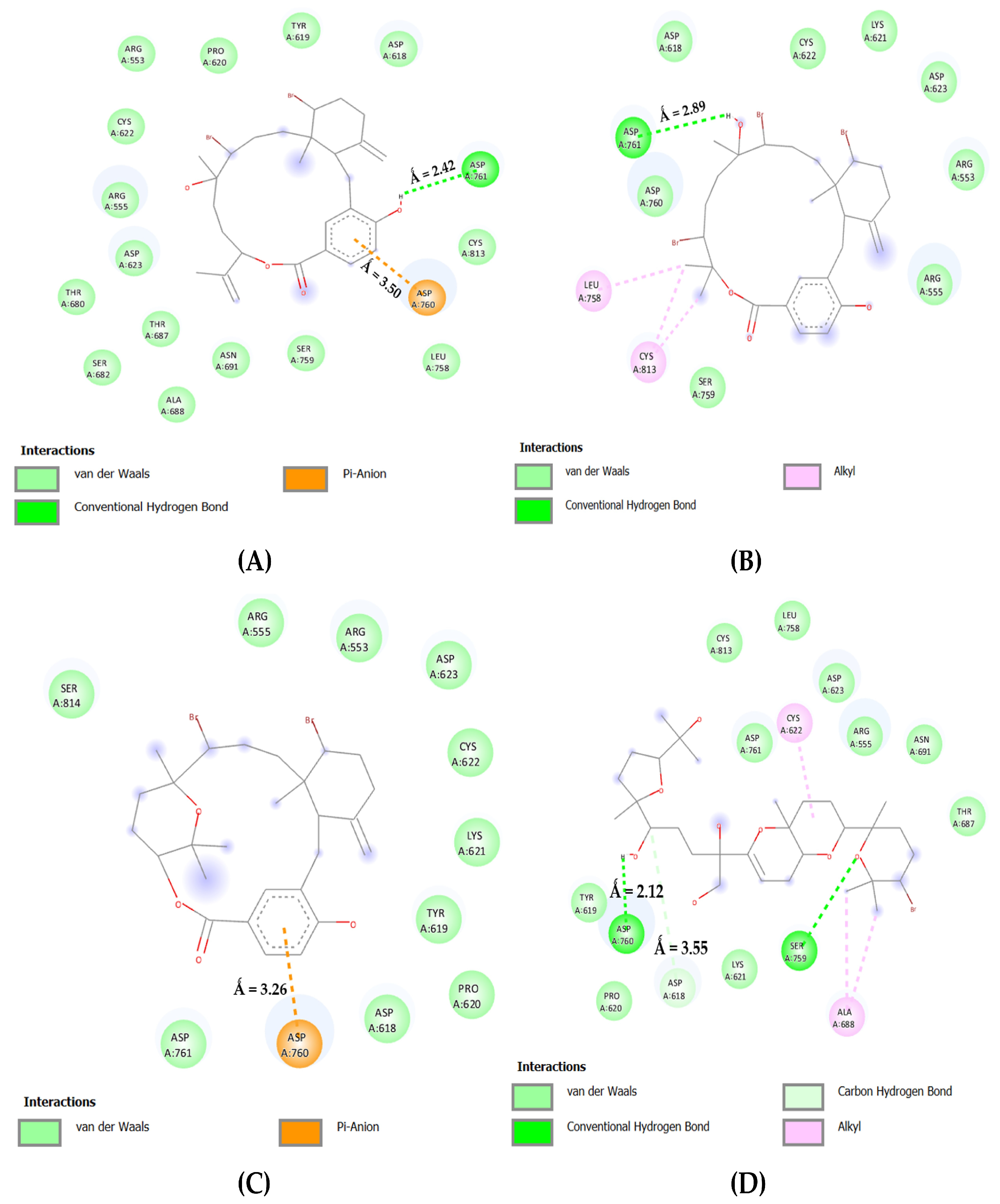
In Figure 5, the bromophycolide E in complex with the catalytic domain of RdRp interacted with ASP: 761 and 760 by forming a single hydrogen bond (2.42 Å) and an electrostatic connection (3.50 Å). In addition, ASP: 618 and SER: 759 interacted via Van der Waals forces. Furthermore, bromophycolide H formed a hydrogen bond with ASP: 761 (2.89 Å). Hydrophobic interactions were evident for the residues ASP: 618, 760, and SER: 759. Bromophycolide P interacted with ASP: 761 and 618 using hydrophobic interactions, whereas ASP: 760 interacted electrostatically, forming a pi–anion bond (3.26 Å). However, no interactions with amino acid SER: 759 occurred. Ultimately, thyrsenol A formed a couple of hydrogen bonds (2.12 Å, 2.69 Å) with ASP: 618 and SER: 759, respectively. A carbon–hydrogen link (3.55 Å) was observed with the residue ASP: 618, whereas ASP: 761 interacted via Van der Waals forces only.
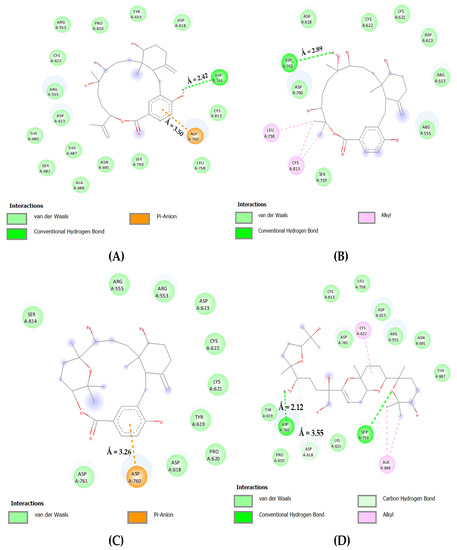
Figure 5.
The 2D interaction maps for phase 2 docking: (A) RdRp C-Terminal–bromophycolide E complex; (B) RdRp C-Terminal–bromophycolide H complex; (C) RdRp C-Terminal–bromophycolide P complex; (D) RdRp C-Terminal–Thyrsenol A complex.
Figure 5.
The 2D interaction maps for phase 2 docking: (A) RdRp C-Terminal–bromophycolide E complex; (B) RdRp C-Terminal–bromophycolide H complex; (C) RdRp C-Terminal–bromophycolide P complex; (D) RdRp C-Terminal–Thyrsenol A complex.
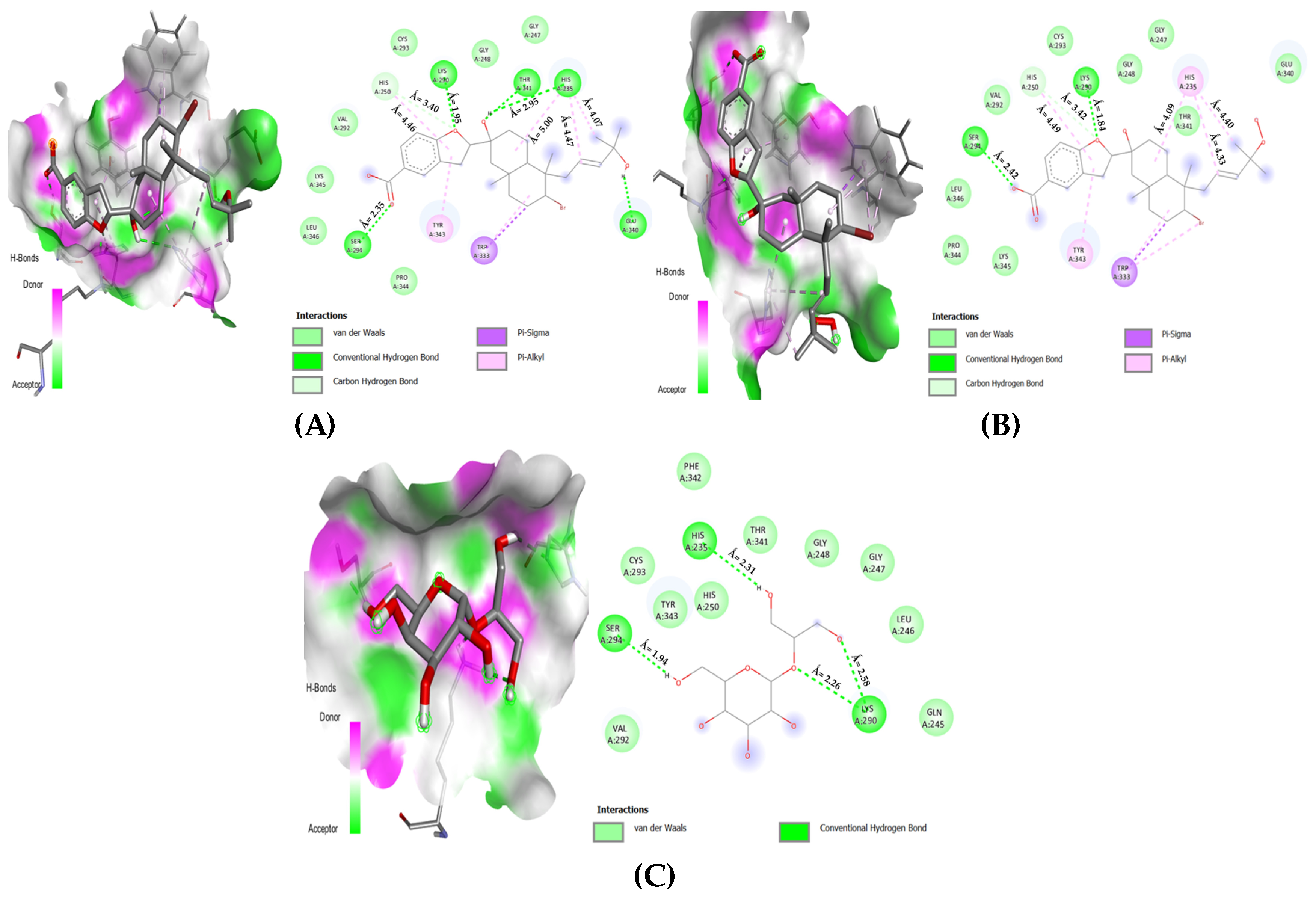
In Figure 6, for the nsp15 enzyme, three NPs showed the most favorable interactions by associating with four key amino acids out of eight (see Figure 3 and Table 5). First, the bromophycoic acid B formed hydrogen bonds with LYS: 290 (1.95 Å) and SER: 294 (2.35 Å). With HIS: 235, it formed one hydrogen (2.95 Å) and three pi–alkyl bonds (4.07, 4.47, and 5.00 Å). Moreover, HIS: 250 was connected with C-H bond (3.40 Å) and a pi–alkyl bond (4.46 Å). Second, the bromophycoic acid C formed hydrogen bonds with LYS: 290 (1.84 Å) and SER: 294 (2.42 Å). HIS: 235 was found to be linked by three pi–alkyl bonds (4.09, 4.33, and 4.40 Å), whereas HIS: 250 was connected by a C-H bond (3.42 Å) and a pi–alkyl bond (4.49 Å). Lastly, floridoside formed four hydrogen bonds with three residues, namely, LYS: 290 (2.26 and 2.58 Å), SER: 294 (1.94 Å), and HIS: 235 (2.31 Å). Van der Waals interactions were observed in the case of HIS: 250 residue.
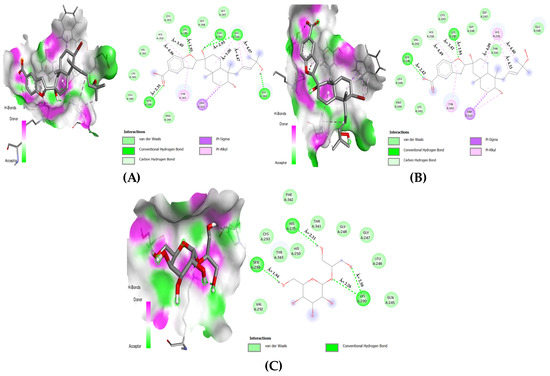
Figure 6.
The 2D interaction maps for nsp15 protein: (A) nsp15 endoribonuclease–bromophycoic acid B complex; (B) nsp15 endoribonuclease–bromophycoic acid C complex; (C) nsp15 endoribonuclease–floridoside complex.
Figure 6.
The 2D interaction maps for nsp15 protein: (A) nsp15 endoribonuclease–bromophycoic acid B complex; (B) nsp15 endoribonuclease–bromophycoic acid C complex; (C) nsp15 endoribonuclease–floridoside complex.
3.4. MD Simulations
The CABS-flex 2.0 tool displays good fit with NMR spectroscopic data on protein flexibility and is very efficient for fast simulations of protein residue flexibility.
MD Simulations for Selected RdRp and nsp15 Complexes
The aforementioned complexes were subjected to molecular dynamics simulations on the free to access CABS-flex 2.0 webserver. Multiple in silico studies concerning SARS-CoV-2 used this webserver for MD simulations, indicating its reliability [200,201,202,203].
In the case of bromophycolide R in complex with the N-terminal domain, RMSF score of 0.388 Å was obtained for the residue number ASN: 781, whereas the highest fluctuation was seen for ASN: 138 (1.606 Å). Similarly, for the complex of RdRp–bromophycolide S, the highest fluctuation (1.027 Å) was seen again for the amino acid ASN: 138. On the other hand, the lowest fluctuation was for HIS: 133 (0.159 Å). Likewise, the complex with bromophycoic acid C, TYR: 129 residue showed minimum fluctuation (0.169 Å). However, ASN: 138 fluctuated the highest (1.562 Å) of all residues (see Figure 7 and Table S13).
The C-terminal docking with bromophycolide E revealed that amino acid ASP: 761 had the lowest RMSF score of 0.139 Å, whereas the highest RMSF score of 0.7600 was seen for the ASP: 618 residue. In the case of bromophycolide H, it was observed that SER: 759 fluctuated the most (0.901 Å) and ASP: 761 residue fluctuated the least (0.540 Å). Finally, for both bromophycolide P and thyrsenol A, the least fluctuating amino acid was ASP: 761 (0.345 Å, 0.257 Å, respectively). On the other hand, the highest RMSF score of 0.594 Å and 0.498 Å for fluctuation was evident for SER: 759 residue (refer to Figure 8 and Table S14).
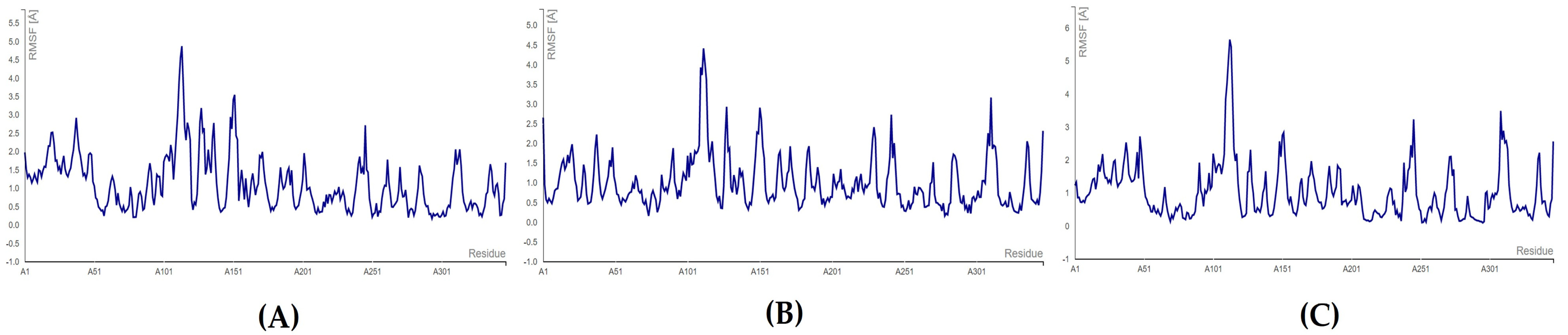
For the endoribonuclease enzyme (nsp15), the complexes formed with floridoside and bromophycoic acid B and C showed the highest fluctuating residue number HIS: 242. For bromophycoic acid C, the amino acid HIS: 242 fluctuated the most (2.726 Å), almost reaching the maximum limit of 3.0 Å. The lowest fluctuation was observed for the residue SER: 294 for the complexes of bromophycoic acid B (0.172 Å) and floridoside (0.134 Å). However, HIS: 249 was the lowest fluctuating residue (0.388 Å) for the nsp15–bromophycoic acid C. All complexes were concluded to be stable, as the threshold of 3.0 Å was not surpassed (see Figure 9 and Table S15). All RMSF values were well below the threshold of 3.0 Å. Thus, all 10 complexes were considered to form stable conformations.
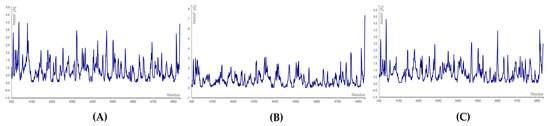
Figure 7.
RMSF plots for N-terminal docking of RdRp: (A) RdRp β-hairpin–bromophycolide R complex; (B) RdRp β-hairpin–bromophycolide S complex; (C) RdRp β-hairpin–bromophycoic acid C complex.
Figure 7.
RMSF plots for N-terminal docking of RdRp: (A) RdRp β-hairpin–bromophycolide R complex; (B) RdRp β-hairpin–bromophycolide S complex; (C) RdRp β-hairpin–bromophycoic acid C complex.
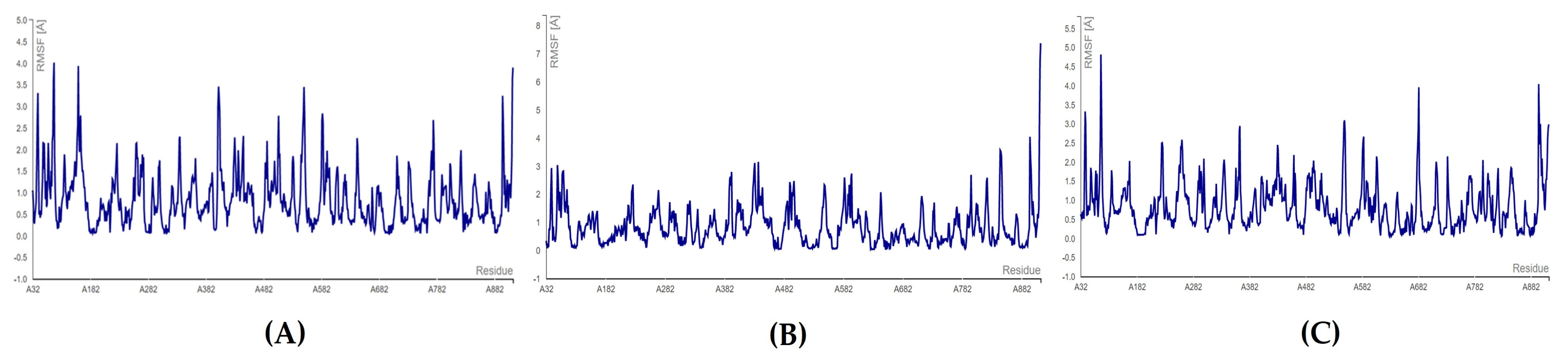
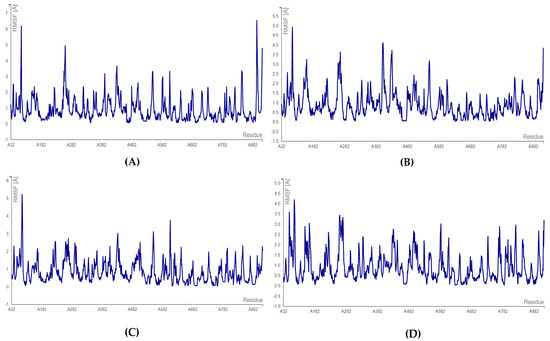
Figure 8.
RMSF plots for C-terminal docking of RdRp: (A) RdRp–bromophycolide E complex; (B) RdRp–bromophycolide H complex; (C) RdRp–bromophycolide P complex; (D) RdRp–thyrsenol A complex.
Figure 8.
RMSF plots for C-terminal docking of RdRp: (A) RdRp–bromophycolide E complex; (B) RdRp–bromophycolide H complex; (C) RdRp–bromophycolide P complex; (D) RdRp–thyrsenol A complex.
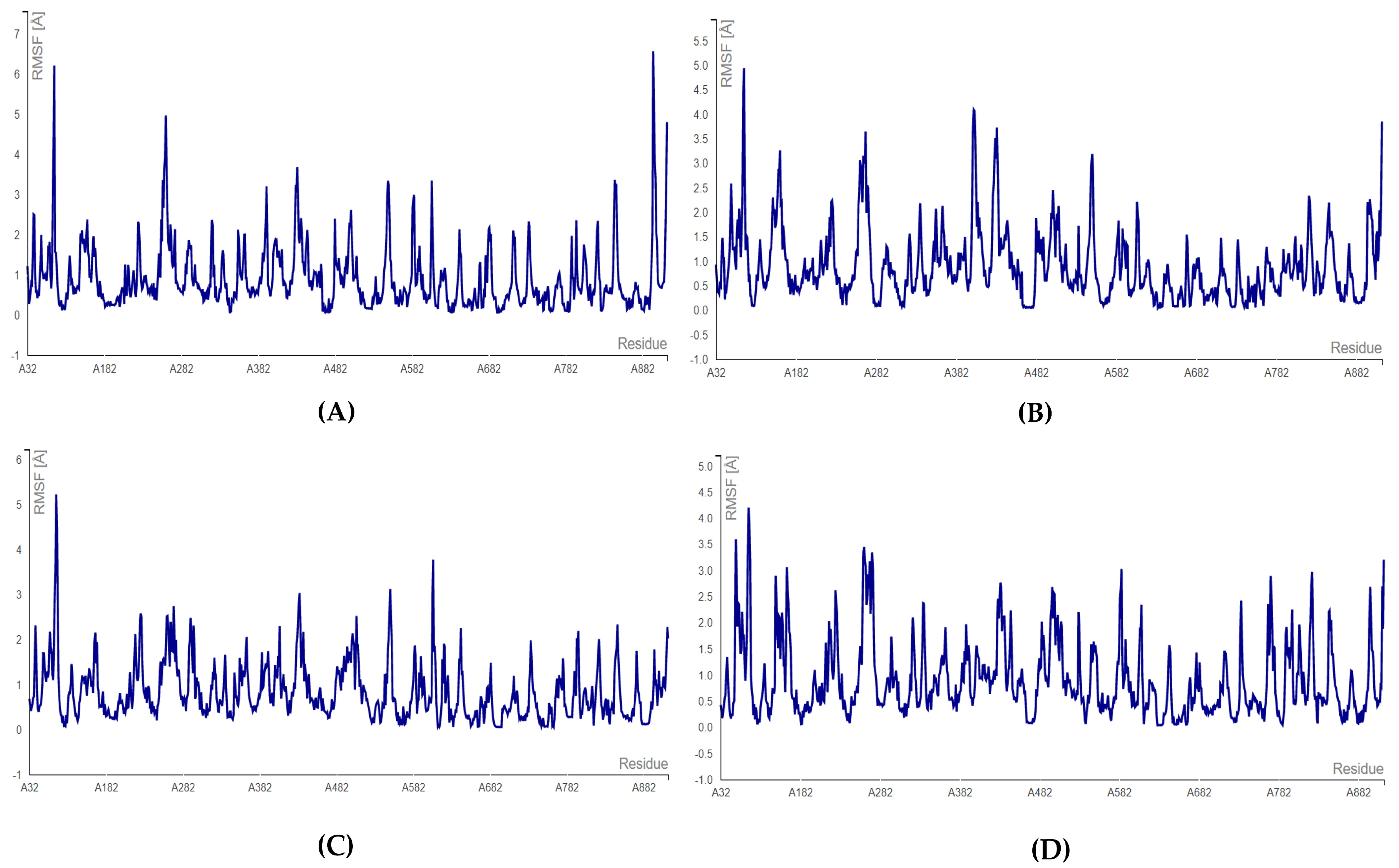
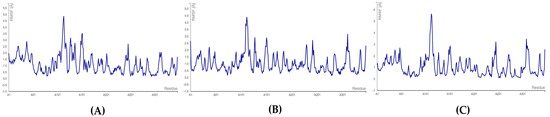
Figure 9.
RMSF plots for endoribonuclease protein: (A) nsp15–bromophycoic acid B; (B) nsp15–bromophycoic acid C; (C) nsp15–floridoside.
Figure 9.
RMSF plots for endoribonuclease protein: (A) nsp15–bromophycoic acid B; (B) nsp15–bromophycoic acid C; (C) nsp15–floridoside.
3.5. Toxicity Evaluation
Using the ProTox-II and StopTox webservers, the toxic characteristics of finalized natural compounds were studied.
3.5.1. ProTox-II Report
The ProTox-II algorithm divides ligands into six toxicity classes based on LD50 scores, which are estimates of the median lethal dosage, i.e., at which 50% of the test subjects pass away after oral exposure. In addition, the ability to cause genetic abnormalities as well as the danger of liver damage and cancer-causing potential were examined.
As members of class 2 (5 < LD50 < 50), thyrsenol A, bromophycoic acids B and C have the potential to be lethal upon oral consumption. The class 4 members included bromophycolide E, H, P, R and S (300 < LD50 < 2000). They are not hazardous, but oral consumption might be harmful. Ultimately, floridoside received the class 6 distinction (LD50 > 5000), which indicates a likelihood of non-toxicity. It was determined that none of the nine NPs were mutagenic, carcinogenic, or hepatotoxic. Bromophycolides and thyrsenol A, however, may cause immune toxicity (see Table 4).

Table 4.
ProTox-II results for shortlisted NPs.
Table 4.
ProTox-II results for shortlisted NPs.
| ProTox-II Toxicity | ||||||
|---|---|---|---|---|---|---|
| Top Ligands | Toxicity Values | Probability | ||||
| LD50 (mg/kg) | Toxicity Class | Hepatotoxicity | Carcinogenicity | Immunotoxicity | Mutagenicity | |
| Bromophycoic acid B | 12 | 2 | Inactive | Inactive | Inactive | Inactive |
| Bromophycoic acid C | 12 | 2 | Inactive | Inactive | Inactive | Inactive |
| Bromophycolide E | 1000 | 4 | Inactive | Inactive | Active | Inactive |
| Bromophycolide H | 1000 | 4 | Inactive | Inactive | Active | Inactive |
| Bromophycolide P | 1000 | 4 | Inactive | Inactive | Active | Inactive |
| Bromophycolide R | 1000 | 4 | Inactive | Inactive | Active | Inactive |
| Bromophycolide S | 1000 | 4 | Inactive | Inactive | Active | Inactive |
| Floridoside | 23,000 | 6 | Inactive | Inactive | Inactive | Inactive |
| Thyrsenol A | 7 | 2 | Inactive | Inactive | Active | Inactive |
3.5.2. StopTox Report
The StopTox server provided us with estimations of the likelihood that the selected NPs would cause acute toxicity.
Bromophycoic acids, bromophycolide E, H, P and floridoside have been indicated to be non-toxic when inhaled. Nevertheless, bromophycolide R and S could be harmful if inhaled. Bromophycoic acids, bromophycolide E and thyrsenol A were capable of showing the likelihood of oral harm. All nine NPs are unlikely to cause dermal toxicity, eye or skin irritation, or corrosion. Ultimately, the five bromophycolides may cause skin sensitivity (see Table 5).

Table 5.
StopTox results for shortlisted NPs.
Table 5.
StopTox results for shortlisted NPs.
| StopTox Acute Toxicity | |||||
|---|---|---|---|---|---|
| Top Ligands | Endpoints | ||||
| Inhalation | Oral | Dermal | Irritation and Corrosion | Skin Sensitization | |
| Bromophycoic acid B | Non-Toxic | Toxic | Non-Toxic | Eyes (-) Skin (-) | Non-sensitizer |
| Bromophycoic acid C | Non-Toxic | Toxic | Non-Toxic | Eyes (-) Skin (-) | Non-sensitizer |
| Bromophycolide E | Non-Toxic | Toxic | Non-Toxic | Eyes (-) Skin (-) | Sensitizer |
| Bromophycolide H | Non-Toxic | Non-Toxic | Non-Toxic | Eyes (-) Skin (-) | Sensitizer |
| Bromophycolide P | Non-Toxic | Non-Toxic | Non-Toxic | Eyes (-) Skin (-) | Sensitizer |
| Bromophycolide R | Toxic | Non-Toxic | Non-Toxic | Eyes (-) Skin (-) | Sensitizer |
| Bromophycolide S | Toxic | Non-Toxic | Non-Toxic | Eyes (-) Skin (-) | Sensitizer |
| Floridoside | Non-Toxic | Non-Toxic | Non-Toxic | Eyes (-) Skin (-) | Non-sensitizer |
| Thyrsenol A | Non-toxic | Toxic | Non-toxic | Eyes (-) Skin (-) | Non-sensitizer |
3.6. Pharmacokinetic Studies
3.6.1. Lipinski Framework (Ro5)
The Lipinski rule of five, commonly referred to as Pfizer’s rule, demonstrated the candidate NPs’ drug-like potency (see Table 6). With the exception of floridoside, all NPs violated the first criterion (MW ≤ 500). The criteria of maximum rotatable bonds (RB ≤ 10) and hydrogen bond acceptors (HBA ≤ 10) were both followed by all NPs. Only floridoside deviated from the HBD ≤ 5 limit for maximal hydrogen bond donors. The maximum Consensus log p-value was exceeded by all five bromophycolides (Log p ≤ 5), whereas floridoside received a negative score. The total number of NPs did not surpass the TPSA’s [(Å2) ≤ 140] upper limit.

Table 6.
Results for drug-likeness of the NPs.
Table 6.
Results for drug-likeness of the NPs.
| Drug-Likeness Assessment | ||||||
|---|---|---|---|---|---|---|
| Top Ligands | Mol. Weight (g/mol) MW ≤ 500 | Rotatable Bonds RB ≤ 10 | H Bond Acceptors HBA ≤ 10 | H Bond Donors HBD ≤ 5 | C Log p Log p ≤ 5 | TPSA (Å2) ≤ 140 |
| Bromophycoic acid B | 521.48 | 5 | 5 | 3 | 4.78 | 86.99 |
| Bromophycoic acid C | 537.48 | 6 | 6 | 3 | 4.77 | 96.22 |
| Bromophycolide E | 584.38 | 1 | 4 | 2 | 5.79 | 66.76 |
| Bromophycolide H | 665.29 | 0 | 4 | 2 | 6.10 | 66.76 |
| Bromophycolide P | 584.38 | 0 | 4 | 1 | 5.83 | 55.76 |
| Bromophycolide R | 503.47 | 1 | 4 | 1 | 5.54 | 59.06 |
| Bromophycolide S | 584.38 | 1 | 4 | 1 | 5.87 | 59.06 |
| Floridoside | 254.23 | 5 | 8 | 6 | −2.53 | 139.84 |
| Thyrsenol A | 619.63 | 8 | 8 | 4 | 3.51 | 117.84 |
3.6.2. Swiss-ADME
Both bromophycoic acids and thyrsenol A were found to be moderately soluble with the high bioavailability score. All five bromophycolides, on the other hand, were shown to have low water solubility. The bioavailability levels for them were the lowest. Lastly, floridoside has a 0.55 bioavailability value, making it potentially soluble. Just three NPs had a high likelihood of GI absorption; the likelihood was modest for the other five. The bromophycolide P had the greatest AB%, followed by the two bromophycoic acids R and S, with floridoside having the lowest (see Table 7).

Table 7.
Results for Swiss-ADME analysis.
Table 7.
Results for Swiss-ADME analysis.
| Swiss-ADME Output | |||||
|---|---|---|---|---|---|
| Top Ligands | Water Solubility | Bioavailability | GI Absorption | Absorption (%) | BBB Permeant |
| Bromophycoic acid B | Moderate | 0.56 | High | 78.98 | No |
| Bromophycoic acid C | Moderate | 0.56 | Low | 75.80 | No |
| Bromophycolide E | Poor | 0.17 | Low | 85.96 | No |
| Bromophycolide H | Poor | 0.17 | Low | 85.96 | No |
| Bromophycolide P | Poor | 0.17 | Low | 89.76 | No |
| Bromophycolide R | Poor | 0.17 | High | 88.62 | No |
| Bromophycolide S | Poor | 0.17 | Low | 88.62 | No |
| Floridoside | Soluble | 0.55 | Low | 60.75 | No |
| Thyrsenol A | Moderate | 0.55 | High | 68.34 | No |
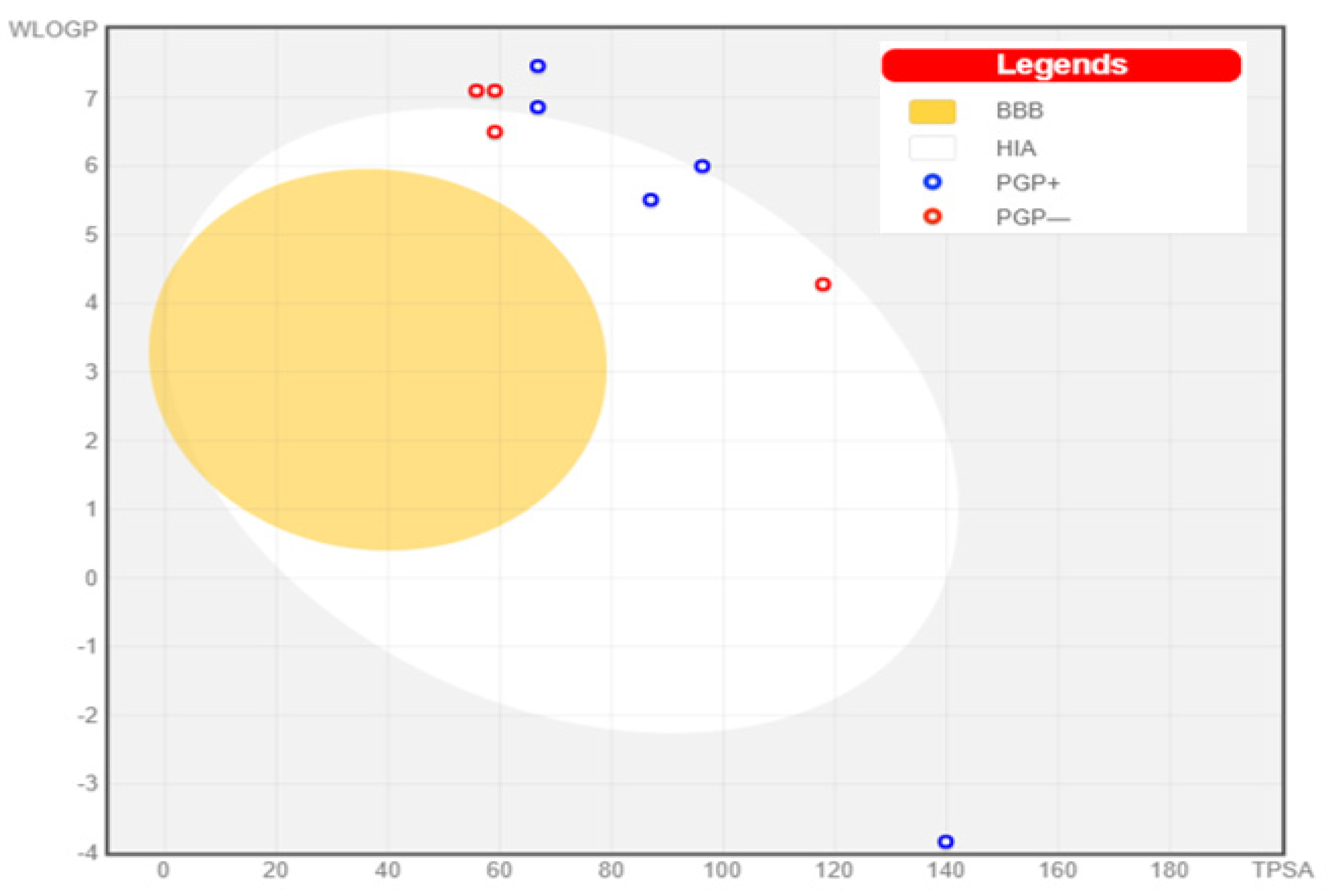
The GI tract was predicted not to be able to passively absorb six NPs, namely, bromophycoic acid C, bromophycolide E, H, P, and S. On the other hand, the GI tract may passively absorb bromophycoic acid B, bromophycolide R, and thyrsenol A. The blood–brain barrier was impermeable to all of the NPs of interest (see Figure 10).
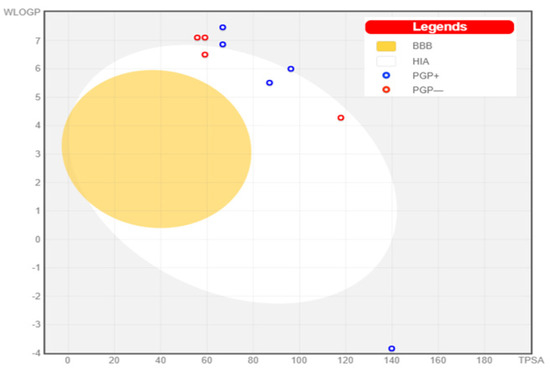
Figure 10.
Boiled egg graph for the nine NPs.
Figure 10.
Boiled egg graph for the nine NPs.
3.6.3. Bioactivity Evaluation
One of the most important steps in the pre-clinical drug-screening process is the assessment of bioactivity. The following NPs’ probable bioactivity was predicted by the PASS online program.
According to predictions, all NPs are potential antifungals that might aid in treating fungal infections linked to COVID-19. Antifungal qualities such as 1, 3-Beta-glucan synthase, beta glucuronidase, histidine kinase, rhizopuspepsin, and GRP78 expression inhibitors were linked to potential inhibition of COVID-19-associated mucormycosis (CAM) [204]. As possible histidine kinase inhibitors, bromophycoic acid B, bromophycolide E, H, P, R, and, S may block the important functional residues of Nsp15 protein, such as HIS: 234, 235, 242, 249, and 250. Four NPs, namely, bromophycoic acid C, the three bromophycolides, floridoside, and thyrsenol A were anticipated to have antiviral properties, indicating a higher likelihood of inhibiting the viral target—in this case, SARS-CoV-2. Depending on the stage of infection, additional relevant bioactivities may help to mitigate the effects of SARS-CoV-2 by either directly or indirectly influencing the immune system, including interferon antagonist, interferon gamma antagonist, transcription factor NF kappa B stimulant or inhibitor, immunostimulant, immunosuppressant, interleukin agonist and antagonist, cytokine release inhibitor, TNF expression, and JAK2 expression inhibitor [205,206]. It is interesting to note that floridoside was predicted to be effective in treating severe acute respiratory syndrome (see Table 8).

Table 8.
Potential bioactivity for selected NPs.
Table 8.
Potential bioactivity for selected NPs.
| PASS Online Program | |
|---|---|
| Top Ligands | Potential Bioactivity |
| Bromophycoic acid B | Antifungal, Histidine kinase inhibitor, Beta glucuronidase inhibitor, Antibacterial, Antioxidant, Antineoplastic, Interferon antagonist, Interferon gamma antagonist, Transcription factor NF kappa B stimulant, Immunostimulant |
| Bromophycoic acid C | Antifungal, Antiviral, Antioxidant, Anticancer, Antineoplastic, Beta glucuronidase inhibitor, Transcription factor NF kappa B stimulant |
| Bromophycolide E | Antineoplastic, Antibiotic, Antiviral, Antifungal, Beta glucuronidase inhibitor, Histidine kinase inhibitor, MMP9 expression inhibitor, Immunosuppressant, Respiratory analeptic, Antibacterial, Cytokine release inhibitor, Interleukin 10 antagonist, 1,3-Beta-glucan synthase inhibitor, Interferon gamma antagonist |
| Bromophycolide H | Antineoplastic, MMP9 expression inhibitor, Antifungal, Antibacterial, Antibiotic, Histidine kinase inhibitor, Cytokine release inhibitor, Interferon gamma antagonist, Interleukin 10 antagonist, 1,3-Beta-glucan synthase inhibitor |
| Bromophycolide P | Antineoplastic, Respiratory analeptic, Antibacterial, Antibiotic, Antifungal, MMP9 expression inhibitor, Beta glucuronidase inhibitor, Histidine kinase inhibitor, 1,3-Beta-glucan synthase inhibitor, Immunosuppressant, Interferon gamma antagonist, Cytokine release inhibitor, Expectorant, |
| Bromophycolide R | Antiviral, Antineoplastic alkaloid, Anticancer, Antifungal, Antibiotic, Antibacterial, Immunosuppressant, Transcription factor NF kappa B stimulant, Beta glucuronidase inhibitor, Histidine kinase inhibitor, 1,3-Beta-glucan synthase inhibitor, Respiratory analeptic, Anti-inflammatory, Cytokine release inhibitor, Interferon gamma antagonist, and an Interleukin 10 antagonist. |
| Bromophycolide S | Antiviral, Antineoplastic, Antibiotic, Anticancer, Antifungal, Immunosuppressant, Antibacterial, Transcription factor NF kappa B inhibitor, Histidine kinase inhibitor, Beta glucuronidase inhibitor, 1,3-Beta-glucan synthase inhibitor, Cytokine release inhibitor and an Interleukin 10 antagonist. |
| Floridoside | Antiviral, Anti-parasitic, Antifungal, Antifungal enhancer, Antibacterial, Anti-tuberculosic, Anticancer, Anti-infective, Antioxidant, Free radical scavenger, Anti-diabetic, Antineoplastic, Immunostimulant, Immunomodulator, Macrophage stimulant, Macrophage colony stimulating factor agonist, Respiratory analeptic, Transcription factor NF kappa B stimulant, Histamine release stimulant, Beta glucuronidase inhibitor, 1,3-Beta-glucan synthase inhibitor, Rhizopuspepsin inhibitor, GRP78 expression inhibitor, Mucolytic, Expectorant, Anti-inflammatory, Histamine release inhibitor, JAK2 expression inhibitor, Severe acute respiratory syndrome treatment, RdRp Inhibitor, Interferon gamma antagonist, Respiratory distress syndrome treatment, Cytokine release inhibitor, TNF expression inhibitor, Interleukin 2, 10, 12 agonist, Interleukin 1a, 4, 6 antagonist. |
| Thyrsenol A | Antineoplastic, Antifungal, Antiviral, Antiinflammatory, Antibacterial, Antibiotic |
4. Conclusions
In conclusion, in this work, we conducted in silico trials on natural compounds from the Laurencia genus to find potential candidates for the treatment of SARS-CoV-2’s RdRp and nsp15 enzymatic infections. Our research identified promising Laurencia red algal anti-SARS-CoV-2 products. The final seven candidates that were shortlisted for RdRp unmistakably exhibit strong interactions with the relevant residues that are crucial for the enzyme inhibition. Three of the seven ligands have the potential to inhibit RdRp when they bind to the N-terminal (AS2), whereas the remaining four ligands are effective RdRp inhibitors when they attach to the NTP entry channel at the C-terminal (AS1). Bromophycolide R displayed the best interaction with the β-hairpin motif region of RdRp (AS2). On the other hand, bromophycolide E and thyrsenol A demonstrated the most favorable residue interactions in the C-terminal (AS1) of RNA-dependent RNA polymerase. As a result of the correlation and required synergy (allosteric signaling) between the two active sites, the RdRp will ultimately be inhibited in both scenarios. The critical residues of nsp15 were interacted with best by bromophycoic acid B, C, and floridoside. Bromophycoic acid C had highly favorable amino acid interactions with both RdRp (N-terminal) and nsp15 proteins, making it a likely dual target inhibitor. Given the nature of the identified amino acid interactions, we propose additional research into the thyrsenol B, dehydrothyrsiferol, and thyrsiferol, against RdRp and nsp15 proteins. To increase the therapeutic prospects of these NPs, laboratory modifications are suggested by assessments of their toxicity and pharmacokinetics analyses.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/microbiolres14030069/s1, Table S1: A list of 300 natural products from the red algal Genus Laurencia involved in this work; Table S2: Natural products listed under subcategory 1 in low class for RdRp; Table S3: Natural products listed under subcategory 2 in low class for RdRp; Table S4: Natural products listed in lower-moderate class for RdRp; Table S5: Natural products listed in upper-moderate class for RdRp; Table S6: Natural products with highest binding affinity for RdRp; Table S7: Outcome of docking the C-terminal catalytic region of RdRp; Table S8: Natural products listed under subcategory 1 in low class for nsp15; Table S9: Natural products listed under sub-category 2 in low class for nsp15; Table S10: Natural products listed in lower-moderate class for nsp15; Table S11: Natural products listed in upper-moderate class for nsp15; Table S12: Natural products with highest binding affinity for nsp15; Table S13: MD Simulations for N-Terminal docked complexes; Table S14: MD Simulations for C-Terminal docked complexes; Table S15: MD Simulations for nsp15 docked complexes; Table S16: SMILES line notations for selected NPs.
Author Contributions
Conceptualization, O.P. and G.V.Z.; methodology, O.P.; software, O.P.; formal analysis, O.P.; investigation, O.P.; data curation, O.P. and H.A.; writing—original draft preparation, O.P.; writing—review and editing, O.P., H.A., G.V.Z. and M.V.T. supervision, G.V.Z. and M.V.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Ministry of Science and Higher Education of the Russian Federation (Reference # 075-15-2022-1118, dated 29 June 2022).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
Authors declare no conflict of interest.
References
- Lopez Bernal, J.; Andrews, N.; Gower, C.; Gallagher, E.; Simmons, R.; Thelwall, S.; Stowe, J.; Tessier, E.; Groves, N.; Dabrera, G.; et al. Effectiveness of COVID-19 Vaccines against the B.1.617.2 (Delta) Variant. N. Engl. J. Med. 2021, 385, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Puranik, A.; Lenehan, P.J.; Silvert, E.; Niesen, M.J.M.; Corchado-Garcia, J.; O’horo, J.C.; Virk, A.; Swift, M.D.; Halamka, J.; Badley, A.D.; et al. Comparison of two highly-effective mRNA vaccines for COVID-19 during periods of Alpha and Delta variant prevalence. medRxiv, 2021; preprint. [Google Scholar] [CrossRef]
- Callaway, E. COVID drug drives viral mutations—And now some want to halt its use. Nature 2023, 614, 399. [Google Scholar] [CrossRef]
- Phillips, N. The coronavirus is here to stay—here’s what that means. Nature 2021, 590, 382–384. [Google Scholar] [CrossRef]
- Min, J.S.; Kwon, S.; Jin, Y.-H. SARS-CoV-2 RdRp Inhibitors Selected from a Cell-Based SARS-CoV-2 RdRp Activity Assay System. Biomedicines 2021, 9, 996. [Google Scholar] [CrossRef]
- Ahn, D.-G.; Choi, J.-K.; Taylor, D.R.; Oh, J.-W. Biochemical characterization of a recombinant SARS coronavirus nsp12 RNA-dependent RNA polymerase capable of copying viral RNA templates. Arch. Virol. 2012, 157, 2095–2104. [Google Scholar] [CrossRef]
- De Farias, S.T.; Dos Santos, A.P.D.S., Jr.; Rêgo, T.G.D.; José, M.V. Origin and Evolution of RNA-Dependent RNA Polymerase. Front. Genet. 2017, 8, 125. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of COVID-19—Final Report. N. Engl. J. Med. 2020, 383, 1813–1826. [Google Scholar] [CrossRef]
- Jockusch, S.; Tao, C.; Li, X.; Chien, M.; Kumar, S.; Morozova, I.; Kalachikov, S.; Russo, J.J.; Ju, J. Sofosbuvir terminated RNA is more resistant to SARS-CoV-2 proofreader than RNA terminated by Remdesivir. Sci. Rep. 2020, 10, 16577. [Google Scholar] [CrossRef]
- Chien, M.; Anderson, T.K.; Jockusch, S.; Tao, C.; Li, X.; Kumar, S.; Russo, J.J.; Kirchdoerfer, R.N.; Ju, J. Nucleotide Analogues as Inhibitors of SARS-CoV-2 Polymerase, a Key Drug Target for COVID-19. J. Proteome Res. 2020, 19, 4690–4697. [Google Scholar] [CrossRef]
- Ju, J.; Li, X.; Kumar, S.; Jockusch, S.; Chien, M.; Tao, C.; Morozova, I.; Kalachikov, S.; Kirchdoerfer, R.N.; Russo, J.J. Nucleotide analogues as inhibitors of SARS-CoV Polymerase. Pharmacol. Res. Perspect. 2020, 8, e00674. [Google Scholar] [CrossRef] [PubMed]
- Aftab, S.O.; Ghouri, M.Z.; Masood, M.U.; Haider, Z.; Khan, Z.; Ahmad, A.; Munawar, N. Analysis of SARS-CoV-2 RNA-dependent RNA polymerase as a potential therapeutic drug target using a computational approach. J. Transl. Med. 2020, 18, 275. [Google Scholar] [CrossRef]
- Ahmad, J.; Ikram, S.; Ahmad, F.; Rehman, I.U.; Mushtaq, M. SARS-CoV-2 RNA Dependent RNA polymerase (RdRp)—A drug repurposing study. Heliyon 2020, 6, e04502. [Google Scholar] [CrossRef]
- Aktaş, A.; Tüzün, B.; Aslan, R.; Sayin, K.; Ataseven, H. New anti-viral drugs for the treatment of COVID-19 instead of favipiravir. J. Biomol. Struct. Dyn. 2020, 39, 7263–7273. [Google Scholar] [CrossRef]
- Alexpandi, R.; De Mesquita, J.F.; Pandian, S.K.; Ravi, A.V. Quinolines-Based SARS-CoV-2 3CLpro and RdRp Inhibitors and Spike-RBD-ACE2 Inhibitor for Drug-Repurposing Against COVID-19: An in silico Analysis. Front. Microbiol. 2020, 11, 1796. [Google Scholar] [CrossRef] [PubMed]
- Al-Masoudi, N.A.; Elias, R.S.; Saeed, B. Molecular Docking Studies of some Antiviral and Antimalarial Drugs Via Bindings to 3CL-Protease and Polymerase Enzymes of the Novel Coronavirus (SARS-CoV-2). Biointerface Res. Appl. Chem. 2020, 10, 6444–6459. [Google Scholar] [CrossRef]
- Ao, S.; Han, D.; Sun, L.; Wu, Y.; Liu, S.; Huang, Y. Identification of Potential Key Agents for Targeting RNA-Dependent RNA Polymerase of SARS-CoV-2 by Integrated Analysis and Virtual Drug Screening. Front. Genet. 2020, 11, 581668. [Google Scholar] [CrossRef] [PubMed]
- Calligari, P.; Bobone, S.; Ricci, G.; Bocedi, A. Molecular Investigation of SARS–CoV-2 Proteins and Their Interactions with Antiviral Drugs. Viruses 2020, 12, 445. [Google Scholar] [CrossRef]
- Da Silva, F.M.A.; Da Silva, K.P.A.; De Oliveira, L.P.M.; Costa, E.V.; Koolen, H.H.; Pinheiro, M.L.B.; De Souza, A.Q.L.; De Souza, A.D.L. Flavonoid glycosides and their putative human metabolites as potential inhibitors of the SARS-CoV-2 main protease (Mpro) and RNA-dependent RNA polymerase (RdRp). Mem. Instit. Oswaldo Cruz. 2020, 115, e200207. [Google Scholar] [CrossRef]
- da Silva, J.K.R.; Figueiredo, P.L.B.; Byler, K.G.; Setzer, W.N. Essential Oils as Antiviral Agents, Potential of Essential Oils to Treat SARS-CoV-2 Infection: An In-Silico Investigation. Int. J. Mol. Sci. 2020, 21, 3426. [Google Scholar] [CrossRef]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef] [PubMed]
- Elfiky, A.A. SARS-CoV-2 RNA dependent RNA polymerase (RdRp) targeting: An in silico perspective. J. Biomol. Struct. Dyn. 2021, 39, 3204–3212. [Google Scholar] [CrossRef] [PubMed]
- Indu, P.; Rameshkumar, M.R.; Arunagirinathan, N.; Al-Dhabi, N.A.; Arasu, M.V.; Ignacimuthu, S. Raltegravir, Indinavir, Tipranavir, Dolutegravir, and Etravirine against main protease and RNA-dependent RNA polymerase of SARS-CoV-2: A molecular docking and drug repurposing approach. J. Infect. Public. Health 2020, 13, 1856–1861. [Google Scholar] [CrossRef] [PubMed]
- Kandeel, M.; Kitade, Y.; Almubarak, A. Repurposing FDA-approved phytomedicines, natural products, antivirals and cell protectives against SARS-CoV-2 (COVID-19) RNA-dependent RNA polymerase. PeerJ 2020, 8, e10480. [Google Scholar] [CrossRef]
- Mohamed, T.A.; Elshamy, A.I.; Ibrahim, M.A.A.; Zellagui, A.; Moustafa, M.F.; Abdelrahman, A.H.M.; Ohta, S.; Pare, P.W.; Hegazy, M.-E.F. Carotane sesquiterpenes from Ferula vesceritensis: In silico analysis as SARS-CoV-2 binding inhibitors. RSC Adv. 2020, 10, 34541–34548. [Google Scholar] [CrossRef]
- Parvez, M.S.A.; Karim, M.A.; Hasan, M.; Jaman, J.; Karim, Z.; Tahsin, T.; Hasan, M.N.; Hosen, M.J. Prediction of potential inhibitors for RNA-dependent RNA polymerase of SARS-CoV-2 using comprehensive drug repurposing and molecular docking approach. Int. J. Biol. Macromol. 2020, 163, 1787–1797. [Google Scholar] [CrossRef]
- Pokhrel, R.; Chapagain, P.; Siltberg-Liberles, J. Potential RNA-dependent RNA polymerase inhibitors as prospective therapeutics against SARS-CoV-2. J. Med. Microbiol. 2020, 69, 864–873. [Google Scholar] [CrossRef]
- Ruan, Z.; Liu, C.; Guo, Y.; He, Z.; Huang, X.; Jia, X.; Yang, T. SARS-CoV-2 and SARS-CoV: Virtual screening of potential inhibitors targeting RNA-dependent RNA polymerase activity (NSP12). J. Med. Virol. 2021, 93, 389–400. [Google Scholar] [CrossRef]
- Tchesnokov, E.P.; Gordon, C.J.; Woolner, E.; Kocinkova, D.; Perry, J.K.; Feng, J.Y.; Porter, D.P.; Götte, M. Template-dependent inhibition of coronavirus RNA-dependent RNA polymerase by remdesivir reveals a second mechanism of action. J. Biol. Chem. 2020, 295, 16156. [Google Scholar] [CrossRef]
- Wu, C.; Liu, Y.; Yang, Y.; Zhang, P.; Zhong, W.; Wang, Y.; Wang, Q.; Xu, Y.; Li, M.; Li, X.; et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm. Sin. B 2020, 10, 766–788. [Google Scholar] [CrossRef]
- Ahmed, S.; Mahtarin, R.; Ahmed, S.S.; Akter, S.; Islam, S.; Al Mamun, A.; Islam, R.; Hossain, N.; Ali, A.; Sultana, M.U.C.; et al. Investigating the binding affinity, interaction, and structure-activity-relationship of 76 prescription antiviral drugs targeting RdRp and Mpro of SARS-CoV-2. J. Biomol. Struct. Dyn. 2021, 39, 6290–6305. [Google Scholar] [CrossRef] [PubMed]
- Barage, S.; Karthic, A.; Bavi, R.; Desai, N.; Kumar, R.; Kumar, V.; Lee, K.W. Identification and characterization of novel RdRp and Nsp15 inhibitors for SARS-COV2 using computational approach. J. Biomol. Struct. Dyn. 2022, 40, 2557–2574. [Google Scholar] [CrossRef] [PubMed]
- Borquaye, L.S.; Gasu, E.N.; Ampomah, G.B.; Kyei, L.K.; Amarh, M.A.; Mensah, C.N.; Nartey, D.; Commodore, M.; Adomako, A.K.; Acheampong, P.; et al. Alkaloids from Cryptolepis sanguinolenta as Potential Inhibitors of SARS-CoV-2 Viral Proteins: An In Silico Study. BioMed Res. Int. 2020, 2020, 532–560. [Google Scholar] [CrossRef] [PubMed]
- Dwarka, D.; Agoni, C.; Mellem, J.J.; Soliman, M.E.; Baijnath, H. Identification of potential SARS-CoV-2 inhibitors from South African medicinal plant extracts using molecular modelling approaches. S. Afr. J. Bot. 2020, 133, 273–284. [Google Scholar] [CrossRef] [PubMed]
- El Hassab, M.A.; Shoun, A.A.; Al-Rashood, S.T.; Al-Warhi, T.; Eldehna, W.M. Identification of a New Potential SARS-COV-2 RNA-Dependent RNA Polymerase Inhibitor via Combining Fragment-Based Drug Design, Docking, Molecular Dynamics, and MM-PBSA Calculations. Front. Chem. 2020, 8, 584894. [Google Scholar] [CrossRef] [PubMed]
- Gul, S.; Ozcan, O.; Asar, S.; Okyar, A.; Barıs, I.; Kavakli, I.H. In silico identification of widely used and well-tolerated drugs as potential SARS-CoV-2 3C-like protease and viral RNA-dependent RNA polymerase inhibitors for direct use in clinical trials. J. Biomol. Struct. Dyn. 2021, 39, 6772–6791. [Google Scholar] [CrossRef]
- Gutierrez-Villagomez, J.M.; Campos-García, T.; Molina-Torres, J.; López, M.G.; Vázquez-Martínez, J. Alkamides and Piperamides as Potential Antivirals against the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). J. Phys. Chem. Lett. 2020, 11, 8008–8016. [Google Scholar] [CrossRef]
- Kar, P.; Sharma, N.R.; Singh, B.; Sen, A.; Roy, A. Natural compounds from Clerodendrum spp. as possible therapeutic candidates against SARS-CoV-2: An in silico investigation. J. Biomol. Struct. Dyn. 2021, 39, 4774–4785. [Google Scholar] [CrossRef]
- Khan, A.; Khan, M.; Saleem, S.; Babar, Z.; Ali, A.; Khan, A.A.; Sardar, Z.; Hamayun, F.; Ali, S.S.; Wei, D.-Q. Phylogenetic Analysis and Structural Perspectives of RNA-Dependent RNA-Polymerase Inhibition from SARs-CoV-2 with Natural Products. Interdiscip. Sci. Comput. Life Sci. 2020, 12, 335–348. [Google Scholar] [CrossRef]
- Mutlu, O.; Ugurel, O.M.; Sariyer, E.; Ata, O.; Inci, T.G.; Ugurel, E.; Kocer, S.; Turgut-Balik, D. Targeting SARS-CoV-2 Nsp12/Nsp8 interaction interface with approved and investigational drugs: An in silico structure-based approach. J. Biomol. Struct. Dyn. 2022, 40, 918–930. [Google Scholar] [CrossRef]
- Narayanan, N.; Nair, D.T. Vitamin B12 may inhibit RNA-dependent-RNA polymerase activity of nsp12 from the SARS-CoV-2 virus. IUBMB Life 2020, 72, 2112–2120. [Google Scholar] [CrossRef]
- Ribaudo, G.; Ongaro, A.; Oselladore, E.; Zagotto, G.; Memo, M.; Gianoncelli, A. A computational approach to drug repurposing against SARS-CoV-2 RNA dependent RNA polymerase (RdRp). J. Biomol. Struct. Dyn. 2022, 40, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Vora, J.; Patel, D.; Sinha, S.; Jha, P.C.; Shrivastava, N. Identification of natural inhibitors against prime targets of SARS-CoV-2 using molecular docking, molecular dynamics simulation and MM-PBSA approaches. J. Biomol. Struct. Dyn. 2022, 40, 3296–3311. [Google Scholar] [CrossRef]
- Singh, J.; Malik, D.; Raina, A. Computational investigation for identification of potential phytochemicals and antiviral drugs as potential inhibitors for RNA-dependent RNA polymerase of COVID-19. J. Biomol. Struct. Dyn. 2022, 40, 3492–3507. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Sk, M.F.; Sonawane, A.; Kar, P.; Sadhukhan, S. Plant-derived natural polyphenols as potential antiviral drugs against SARS-CoV-2 via RNA-dependent RNA polymerase (RdRp) inhibition: An in-silico analysis. J. Biomol. Struct. Dyn. 2021, 39, 6249–6264. [Google Scholar] [CrossRef] [PubMed]
- Elkarhat, Z.; Charoute, H.; Elkhattabi, L.; Barakat, A.; Rouba, H. Potential inhibitors of SARS-CoV-2 RNA dependent RNA polymerase protein: Molecular docking, molecular dynamics simulations and MM-PBSA analyses. J. Biomol. Struct. Dyn. 2022, 40, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Brunt, D.; Lakernick, P.M.; Wu, C. Discovering new potential inhibitors to SARS-CoV-2 RNA dependent RNA polymerase (RdRp) using high throughput virtual screening and molecular dynamics simulations. Sci. Rep. 2022, 12, 19986. [Google Scholar] [CrossRef]
- Olender, S.A.; Perez, K.K.; Go, A.S.; Balani, B.; Price-Haywood, E.G.; Shah, N.S.; Wang, S.; Walunas, T.L.; Swaminathan, S.; Slim, J.; et al. Remdesivir for Severe Coronavirus Disease 2019 (COVID-19) Versus a Cohort Receiving Standard of Care. Clin. Infect. Dis. 2021, 73, e4166–e4174. [Google Scholar] [CrossRef] [PubMed]
- Al-Abdouh, A.; Bizanti, A.; Barbarawi, M.; Jabri, A.; Kumar, A.; Fashanu, O.E.; Khan, S.U.; Zhao, D.; Antar, A.A.; Michos, E.D. Remdesivir for the treatment of COVID-19: A systematic review and meta-analysis of randomized controlled trials. Contemp. Clin. Trials 2021, 101, 106272. [Google Scholar] [CrossRef]
- Dölken, L.; Stich, A.; Spinner, C.D. Remdesivir for Early COVID-19 Treatment of High-Risk Individuals Prior to or at Early Disease Onset—Lessons Learned. Viruses 2021, 13, 963. [Google Scholar] [CrossRef]
- Gottlieb, R.L.; Vaca, C.E.; Paredes, R.; Mera, J.; Webb, B.J.; Perez, G.; Oguchi, G.; Ryan, P.; Nielsen, B.U.; Brown, M.; et al. Early Remdesivir to Prevent Progression to Severe COVID-19 in Outpatients. N. Engl. J. Med. 2022, 386, 305–315. [Google Scholar] [CrossRef]
- Mahajan, L.; Singh, A. Gifty Clinical outcomes of using remdesivir in patients with moderate to severe COVID-19: A prospective randomised study. Indian J. Anaesth. 2021, 65 (Suppl. 1), S41–S46. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Binetti, E.; Borrazzo, C.; Cacciola, E.G.; Battistini, L.; Ceccarelli, G.; Mastroianni, C.M.; D’ettorre, G. Efficacy of Remdesivir-Containing Therapy in Hospitalized COVID-19 Patients: A Prospective Clinical Experience. J. Clin. Med. 2021, 10, 3784. [Google Scholar] [CrossRef] [PubMed]
- Yan, V.C.; Muller, F.L. Why Remdesivir Failed: Preclinical Assumptions Overestimate the Clinical Efficacy of Remdesivir for COVID-19 and Ebola. Antimicrob. Agents Chemother. 2021, 65, e0111721. [Google Scholar] [CrossRef] [PubMed]
- Ader, F.; Bouscambert-Duchamp, M.; Hites, M.; Peiffer-Smadja, N.; Poissy, J.; Belhadi, D.; Diallo, A.; Lê, M.-P.; Peytavin, G.; Staub, T.; et al. Remdesivir plus standard of care versus standard of care alone for the treatment of patients admitted to hospital with COVID-19 (DisCoVeRy): A phase 3, randomised, controlled, open-label trial. Lancet Infect. Dis. 2022, 22, 209–221. [Google Scholar] [CrossRef]
- Sun, D. Remdesivir for Treatment of COVID-19: Combination of Pulmonary and IV Administration May Offer Aditional Benefit. AAPS J. 2020, 22, 77, Erratum in AAPS J. 2020, 22, 102. [Google Scholar] [CrossRef]
- Martinez, D.R.; Schäfer, A.; Leist, S.R.; Li, D.; Gully, K.; Yount, B.; Feng, J.Y.; Bunyan, E.; Porter, D.P.; Cihlar, T.; et al. Prevention and therapy of SARS-CoV-2 and the B.1.351 variant in mice. Cell Rep. 2021, 36, 109450. [Google Scholar] [CrossRef]
- Taha, H.R.; Keewan, N.; Slati, F.; Al-Sawalha, N.A. Remdesivir: A Closer Look at Its Effect in COVID-19 Pandemic. Pharmacology 2021, 106, 462–468. [Google Scholar] [CrossRef]
- Matsuyama, S.; Kawase, M.; Nao, N.; Shirato, K.; Ujike, M.; Kamitani, W.; Shimojima, M.; Fukushi, S. The Inhaled Steroid Ciclesonide Blocks SARS-CoV-2 RNA Replication by Targeting the Viral Replication-Transcription Complex in Cultured Cells. J. Virol. 2020, 95, e01648-20. [Google Scholar] [CrossRef]
- Yang, J.-W.; Fan, L.-C.; Miao, X.-Y.; Mao, B.; Li, M.-H.; Lu, H.-W.; Liang, S.; Xu, J.-F. Corticosteroids for the treatment of human infection with influenza virus: A systematic review and meta-analysis. Clin. Microbiol. Infect. 2015, 21, 956–963. [Google Scholar] [CrossRef]
- Kim, Y.; Jedrzejczak, R.; Maltseva, N.I.; Wilamowski, M.; Endres, M.; Godzik, A.; Michalska, K.; Joachimiak, A. Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2. Protein Sci. 2020, 29, 1596–1605. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Fang, P.; Fang, L.; Hong, Y.; Zhu, X.; Wang, D.; Peng, G.; Xiao, S. Porcine deltacoronavirus nsp15 antagonizes interferon-β production independently of its endoribonuclease activity. Mol. Immunol. 2019, 114, 100–107. [Google Scholar] [CrossRef]
- El Sayed, K.A. Natural Products as Antiviral Agents. Stud. Nat. Prod. Chem. 2000, 24, 473–572. [Google Scholar] [CrossRef]
- Bzówka, M.; Mitusińska, K.; Raczyńska, A.; Samol, A.; Tuszyński, J.A.; Góra, A. Structural and Evolutionary Analysis Indicate That the SARS-CoV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design. Int. J. Mol. Sci. 2020, 21, 3099. [Google Scholar] [CrossRef]
- Kolarič, A.; Jukič, M.; Bren, U. Novel Small-Molecule Inhibitors of the SARS-CoV-2 Spike Protein Binding to Neuropilin 1. Pharmaceuticals 2022, 15, 165. [Google Scholar] [CrossRef]
- Yang, L.; Wang, Z. Natural Products, Alone or in Combination with FDA-Approved Drugs, to Treat COVID-19 and Lung Cancer. Biomedicines 2021, 9, 689. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, N.; Yang, L.; Song, X.-Q. Bioactive natural products in COVID-19 therapy. Front. Pharmacol. 2022, 13, 926507. [Google Scholar] [CrossRef]
- Pokharkar, O.; Lakshmanan, H.; Zyryanov, G.V.; Tsurkan, M.V. Antiviral Potential of Antillogorgia americana and elisabethae Natural Products against nsp16–nsp10 Complex, nsp13, and nsp14 Proteins of SARS-CoV-2: An In Silico Investigation. Microbiol. Res. 2023, 14, 993–1019. [Google Scholar]
- Wang, D.; Huang, J.; Yeung, A.W.K.; Tzvetkov, N.T.; Horbańczuk, J.O.; Willschke, H.; Gai, Z.; Atanasov, A.G. The Significance of Natural Product Derivatives and Traditional Medicine for COVID-19. Processes 2020, 8, 937. [Google Scholar] [CrossRef]
- Ismail, M.M.; Alotaibi, B.S.; El-Sheekh, M.M. Therapeutic Uses of Red Macroalgae. Molecules 2020, 25, 4411. [Google Scholar] [CrossRef] [PubMed]
- Reis, J.G.; Cadamuro, R.D.; Cabral, A.C.; Thaís da Silva, I.T.; Rodríguez-Lázaro, D.; Fongaro, G. Broad Spectrum Algae Compounds Against Viruses. Front. Microbiol. 2022, 12, 809296. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Riverol, A.; Piñón, R.A.; Morier, D.L.F.; Torres, L.Y.; Mendoza, L.D.; Barrio, A.G. Antiviral activity of an aqueous extract from the red alga Laurencia obtusa against influenza A and B viruses. Rev. Cub. Med. Trop. 2014, 66, 273–285. [Google Scholar]
- Gheda, S.F.; El-Adawi, H.I.; El-Deeb, N.M. Antiviral Profile of Brown and Red Seaweed Polysaccharides Against Hepatitis C Virus. Iran. J. Pharm. Res. IJPR 2016, 15, 483–491. [Google Scholar]
- Yu, X.-Q.; Jiang, C.-S.; Zhang, Y.; Sun, P.; Kurtán, T.; Mándi, A.; Li, X.-L.; Yao, L.-G.; Liu, A.-H.; Wang, B.; et al. Compositacins A–K: Bioactive chamigrane-type halosesquiterpenoids from the red alga Laurencia composita Yamada. Phytochemistry 2017, 136, 81–93. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Hoshino, M.; Fujita, M.; Urban, S. Cycloelatanene A and B: Absolute configuration determination and structural revision by the crystalline sponge method. Chem. Sci. 2017, 8, 1547–1550. [Google Scholar] [CrossRef]
- Shaaban, M.; Abou-El-Wafa, G.S.E.; Golz, C.; Laatsch, H. New Haloterpenes from the Marine Red Alga Laurencia papillosa: Structure Elucidation and Biological Activity. Mar. Drugs 2021, 19, 35. [Google Scholar] [CrossRef]
- Masuda, M.; Kogame, K.; Arisawa, S.; Suzuki, M. Morphology and Halogenated Secondary Metabolites of Three Gran Canarian Species of Laurencia (Ceramiales, Rhodophyta). Bot. Mar. 1998, 41, 265–277. [Google Scholar] [CrossRef]
- Sentíes, A.; Díaz-Larrea, J.; Cassano, V.; Gil-Rodríguez, M.C.; Fujii, M.T. Laurencia marilzae (Ceramiales, Rhodophyta) from the Mexican Caribbean: A New Record for the Tropical Western Atlantic. Bull. Mar. Sci. 2011, 87, 681–686. [Google Scholar] [CrossRef]
- Suzuki, M.; Vairappan, C.S. Halogenated secondary metabolites from Japanese species of the red algal genus Laurencia (Rhodomelaceae, Ceramiales). Curr. Top. Phytochem. 2005, 7, 1–34. [Google Scholar]
- Wang, B.-G.; Gloer, J.B.; Ji, N.-Y.; Zhao, J.-C. Halogenated Organic Molecules of Rhodomelaceae Origin: Chemistry and Biology. Chem. Rev. 2013, 113, 3632–3685. [Google Scholar] [CrossRef]
- Suzuki, M.; Takahashi, Y.; Nakano, S.; Abe, T.; Masuda, M.; Ohnishi, T.; Noya, Y.; Seki, K.-I. An experimental approach to study the biosynthesis of brominated metabolites by the red algal genus Laurencia. Phytochemistry 2009, 70, 1410–1415. [Google Scholar] [CrossRef] [PubMed]
- Young, D.N.; Howard, B.M.; Fenical, W. Subcellular localization of brominated secondary metabolites in the red alga Laurencia snyderae. J. Phycol. 1980, 16, 182–185. [Google Scholar] [CrossRef]
- Vairappan, C.S.; Suzuki, M.; Abe, T.; Masuda, M. Halogenated metabolites with antibacterial activity from the Okinawan Laurencia species. Phytochemistry 2001, 58, 517–523. [Google Scholar] [CrossRef] [PubMed]
- Cabrita, M.T.; Vale, C.; Rauter, A.P. Halogenated Compounds from Marine Algae. Mar. Drugs 2010, 8, 2301–2317. [Google Scholar] [CrossRef] [PubMed]
- Juagdan, E.G.; Kalidindi, R.; Scheuer, P. Two new chamigranes from an hawaiian red alga Laurencia cartilaginea. Tetrahedron 1997, 53, 521–528. [Google Scholar] [CrossRef]
- König, G.M.; Wright, A.D. Laurencia rigida: Chemical Investigations of Its Antifouling Dichloromethane Extract. J. Nat. Prod. 1997, 60, 967–970. [Google Scholar] [CrossRef]
- Alarif, W.M.; Al-Lihaibi, S.S.; Abdel-Lateff, A.; Ayyad, S.-E.N. New Antifungal Cholestane and Aldehyde Derivatives from the Red Alga Laurencia papillosa. Nat. Prod. Commun. 2011, 6, 1821–1824. [Google Scholar] [CrossRef]
- Li, Y.-X.; Li, Y.; Qian, Z.J.; Kim, M.M.; Kim, S.K. In Vitro Antioxidant Activity of 5-HMF Isolated from Marine Red Alga Laurencia undulata in Free-Radical-Mediated Oxidative Systems. J. Microbiol. Biotechnol. 2009, 19, 1319–1327. [Google Scholar] [CrossRef]
- Topcu, G.; Aydogmus, Z.; Imre, S.; Gören, A.C.; Pezzuto, J.M.; Clement, J.A.; Kingston, D.G.I. Brominated Sesquiterpenes from the Red Alga Laurencia obtusa. J. Nat. Prod. 2003, 66, 1505–1508. [Google Scholar] [CrossRef]
- Davyt, D.; Fernandez, R.; Suescun, L.; Mombrú, A.W.; Saldaña, J.; Domínguez, L.; Coll, J.; Fujii, M.T.; Manta, E. New Sesquiterpene Derivatives from the Red Alga Laurencia scoparia. Isolation, Structure Determination, and Anthelmintic Activity. J. Nat. Prod. 2001, 64, 1552–1555. [Google Scholar] [CrossRef]
- Jung, W.-K.; Choi, I.; Oh, S.; Park, S.-G.; Seo, S.-K.; Lee, S.-W.; Lee, D.-S.; Heo, S.-J.; Jeon, Y.-J.; Je, J.-Y.; et al. Anti-asthmatic effect of marine red alga (Laurencia undulata) polyphenolic extracts in a murine model of asthma. Food Chem. Toxicol. 2009, 47, 293–297. [Google Scholar] [CrossRef] [PubMed]
- Kurata, K.; Taniguchi, K.; Agatsuma, Y.; Suzuki, M. Diterpenoid feeding-deterrents from Laurencia saitoi. Phytochemistry 1998, 47, 363–369. [Google Scholar] [CrossRef]
- Dembitsky, V.M.; Tolstikov, G.A. Natural halogenated sesquiterpens from marine organisms, chemistry for sustainable development. Chem. Sustain. Dev. 2004, 12, 1–12. [Google Scholar]
- Suzuki, M.; Kurosawa, E.; Irie, T. Spirolaurenone, a new sesquiterpenoid containing bromine from Laurenciaglandulifera Kützing. Tetrahedron Lett. 1970, 11, 4995–4998. [Google Scholar] [CrossRef]
- Ji, N.-Y.; Li, X.-M.; Li, K.; Ding, L.-P.; Gloer, J.B.; Wang, B.-G. Diterpenes, Sesquiterpenes, and a C15-Acetogenin from the Marine Red Alga Laurencia mariannensis. J. Nat. Prod. 2007, 70, 1901–1905. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-D.; Miao, F.-P.; Li, K.; Ji, N.-Y. Sesquiterpenes and acetogenins from the marine red alga Laurencia okamurai. Fitoterapia 2012, 83, 518–522. [Google Scholar] [CrossRef]
- Liang, Y.; Li, X.-M.; Cui, C.-M.; Li, C.-S.; Sun, H.; Wang, B.-G. Sesquiterpene and Acetogenin Derivatives from the Marine Red Alga Laurencia okamurai. Mar. Drugs 2012, 10, 2817–2825. [Google Scholar] [CrossRef]
- Liang, Y.; Li, X.M.; Cui, C.M.; Li, C.S.; Wang, B.G. A new rearranged chamigrane sesquiterpene from Laurencia okamurai. Chin. Chem. Lett. 2009, 20, 190–192. [Google Scholar] [CrossRef]
- Li, X.-D.; Miao, F.-P.; Yin, X.-L.; Liu, J.-L.; Ji, N.-Y. Sesquiterpenes from the marine red alga Laurencia composita. Fitoterapia 2012, 83, 1191–1195. [Google Scholar] [CrossRef]
- Ji, N.-Y.; Li, X.-M.; Wang, B.-G. Sesquiterpenes and Other Metabolites from the Marine Red Alga Laurencia composita (Rhodomelaceae). Helv. Chim. Acta 2010, 93, 2281–2286. [Google Scholar] [CrossRef]
- Sims, J.J.; Lin, G.H.; Wing, R.M. Marine natural products X elatol, a halogenated sesquiterpene alcohol from the red alga Laurencia elata. Tetrahedron Lett. 1974, 15, 3487–3490. [Google Scholar] [CrossRef]
- Lhullier, C.; Donnangelo, A.; Caro, M.; Palermo, J.A.; Horta, P.A.; Falkenberg, M.; Schenkel, E.P. Isolation of elatol from Laurencia microcladia and its palatability to the sea urchin Echinometra lucunter. Biochem. Syst. Ecol. 2009, 37, 254–259. [Google Scholar] [CrossRef]
- González, A.; Darias, J.; Díaz, A.; Fourneron, J.; Martín, J.; Pérez, C. Evidence for the biogenesis of halogenated chamigrenes from the red alga Laurencia obtusa. Tetrahedron Lett. 1976, 17, 3051–3054. [Google Scholar] [CrossRef]
- Vairappan, C.S. Potent antibacterial activity of halogenated metabolites from Malaysian red algae, Laurencia majuscula (Rhodomelaceae, Ceramiales). Biomol. Eng. 2003, 20, 255–259. [Google Scholar] [CrossRef]
- Dos Santos, A.O.; Veiga-Santos, P.; Ueda-Nakamura, T.; Filho, B.P.D.; Sudatti, D.B.; Bianco, .M.; Pereira, R.C.; Nakamura, C.V. Effect of Elatol, Isolated from Red Seaweed Laurencia dendroidea, on Leishmania amazonensis. Mar. Drugs 2010, 8, 2733–2743. [Google Scholar] [CrossRef] [PubMed]
- Desoti, V.C.; Lazarin-Bidóia, D.; Sudatti, D.B.; Pereira, R.C.; Alonso, A.; Ueda-Nakamura, T.; Dias Filho, B.P.; Nakamura, C.V.; De Oliveira Silva, S. Trypanocidal Action of (−)-Elatol Involves an Oxidative Stress Triggered by Mitochondria Dysfunction. Mar. Drugs 2012, 10, 1631–1646. [Google Scholar] [CrossRef]
- Campos, A.; Souza, C.B.; Lhullier, C.; Falkenberg, M.; Schenkel, E.P.; Ribeiro-Do-Valle, R.M.; Siqueira, J.M. Anti-tumour effects of elatol, a marine derivative compound obtained from red algae Laurencia microcladia. J. Pharm. Pharmacol. 2012, 64, 1146–1154. [Google Scholar] [CrossRef] [PubMed]
- Dias, D.A.; Urban, S. Phytochemical studies of the southern Australian marine alga, Laurencia elata. Phytochemistry 2011, 72, 2081–2089. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2003, 20, 1–48. [Google Scholar] [CrossRef]
- Dorta, E.; Díaz-Marrero, A.R.; Cueto, M.; D’croz, L.; Maté, J.L.; Darias, J. Chamigrenelactone, a polyoxygenated sesquiterpene with a novel structural type and devoid of halogen from Laurencia obtusa. Tetrahedron Lett. 2004, 45, 7065–7068. [Google Scholar] [CrossRef]
- Brito, I.; Cueto, M.; Díaz-Marrero, A.R.; Darias, J.; San Martín, A. Oxachamigrenes, New Halogenated Sesquiterpenes from Laurencia obtusa. J. Nat. Prod. 2002, 65, 946–948. [Google Scholar] [CrossRef] [PubMed]
- Gonzaález, A.; Martín, J.; Martín, V.; Norte, M.; Pérez, R. Biomimetic approach to the synthesis of rhodolaureol and rhodolauradiol. Tetrahedron Lett. 1982, 23, 2395–2398. [Google Scholar] [CrossRef]
- Suzuki, M.; Kurosawa, E.; Furusaki, A. The Structure and Absolute Stereochemistry of a Halogenated Chamigrene Derivative from the Red Alga Laurencia Species. Bull. Chem. Soc. Jpn. 1988, 61, 3371–3373. [Google Scholar] [CrossRef]
- Ji, N.-Y.; Li, X.-M.; Li, K.; Wang, B.-G. Halogenated Sesquiterpenes from the Marine Red Alga Laurencia saitoi (Rhodomelaceae). Helv. Chim. Acta 2009, 92, 1873–1879. [Google Scholar] [CrossRef]
- Howard, B.M.; Fenical, W. Structures and chemistry of two new halogen-containing chamigrene derivatives from laurencia. Tetrahedron Lett. 1971, 16, 1687–1690. [Google Scholar] [CrossRef]
- Jongaramruong, J.; Blackman, A.J.; Skelton, B.W.; White, A.H. Chemical Relationships between the Sea Hare Aplysia parvula and the Red Seaweed Laurencia filiformis from Tasmania. Aust. J. Chem. 2002, 55, 275–280. [Google Scholar] [CrossRef]
- Suzuki, M.; Daitoh, M.; Vairappan, C.S.; Abe, T.; Masuda, M. Novel Halogenated Metabolites from the Malaysian Laurencia pannosa. J. Nat. Prod. 2001, 64, 597–602. [Google Scholar] [CrossRef]
- Suescun, L.; Mombrú, A.W.; Mariezcurrena, R.A.; Davyt, D.; Fernández, R.; Manta, E. Two natural products from the algae Laurencia scoparia. Acta Crystallogr. Sect. C Cryst. Struct. Commun. 2001, 57, 286–288. [Google Scholar] [CrossRef]
- Francisco, M.E.Y.; Erickson, K.L. Ma’iliohydrin, a Cytotoxic Chamigrene Dibromohydrin from a Philippine Laurencia Species. J. Nat. Prod. 2001, 64, 790–791. [Google Scholar] [CrossRef]
- Da Silva Machado, F.L.D.S.; Ventura, T.L.B.; Gestinari, L.M.d.S.; Cassano, V.; Resende, J.A.L.C.; Kaiser, C.R.; Lasunskaia, E.B.; Muzitano, M.F.; Soares, A.R. Sesquiterpenes from the Brazilian Red Alga Laurencia dendroidea. J. Agardh. Molecules 2014, 19, 3181–3192. [Google Scholar] [CrossRef]
- Rucker, R.; Kretzuschmar, U. 9-Aristolen-1a-ol and 1,2,9,10-tetradehydroaristolane, new aristolane type sesquiterpenes. Liebigs Ann. Chem. 1971, 748, 214–217. [Google Scholar] [CrossRef]
- Ji, N.-Y.; Li, X.-M.; Ding, L.-P.; Wang, B.-G. Aristolane Sesquiterpenes and Highly Brominated Indoles from the Marine Red Alga Laurencia similis (Rhodomelaceae). Helv. Chim. Acta 2007, 90, 385–391. [Google Scholar] [CrossRef]
- Li, C.-S.; Li, X.-M.; Cui, C.-M.; Wang, B.-G. Brominated Metabolites from the Marine Red Alga Laurencia similis. Z. Naturforsch. 2010, 65, 87–89. [Google Scholar] [CrossRef]
- Su, S.; Sun, W.-S.; Wang, B.; Cheng, W.; Liang, H.; Zhao, Y.-Y.; Zhang, Q.-Y.; Wu, J. A novel brominated cuparene-derived sesquiterpene ether from the red alga Laurencia sp. J. Asian Nat. Prod. Res. 2010, 12, 916–920. [Google Scholar] [CrossRef] [PubMed]
- Kamada, T.; Vairappan, C.S. New Bioactive Secondary Metabolites from Bornean Red Alga, Laurencia similis (Ceramiales). Nat. Prod. Commun. 2013, 8, 287–288. [Google Scholar] [CrossRef]
- Fraga, B.M. Natural sesquiterpenoids. Nat. Prod. Rep. 1997, 14, 145–162. [Google Scholar] [CrossRef]
- Kladi, M.; Vagias, C.; Papazafiri, P.; Furnari, G.; Serio, D.; Roussis, V. New sesquiterpenes from the red alga Laurencia microcladia. Tetrahedron 2007, 63, 7606–7611. [Google Scholar] [CrossRef]
- Yu, X.-Q.; He, W.-F.; Liu, D.-Q.; Feng, M.-T.; Fang, Y.; Wang, B.; Feng, L.-H.; Guo, Y.-W.; Mao, S.-C. A seco-laurane sesquiterpene and related laurane derivatives from the red alga Laurencia okamurai Yamada. Phytochemistry 2014, 103, 162–170. [Google Scholar] [CrossRef]
- Davyt, D.; Fernandez, R.; Suescun, L.; Mombrú, A.W.; Saldaña, J.; Dominguez, L.; Fujii, M.T.; Manta, E. Bisabolanes from the red alga Laurencia scoparia. J. Nat. Prod. 2006, 69, 1113–1116. [Google Scholar] [CrossRef]
- De Carvalho, L.T.; Fujii, M.T.; Roque, N.F.; Kato, M.J.; Lago, J.H.G. Aldingenin A, new brominated sesquiterpene from red algae Laurencia aldingensis. Tetrahedron Lett. 2003, 44, 2637–2640. [Google Scholar] [CrossRef]
- de Carvalho, L.R.; Fujii, M.T.; Roque, N.F.; Lago, J.H.G. Aldingenin derivatives from the red alga Laurencia aldingensis. Phytochemistry 2006, 67, 1331–1335. [Google Scholar] [CrossRef]
- Kladi, M.; Vagias, C.; Furnari, G.; Moreau, D.; Roussakis, C.; Roussis, V. Cytotoxic cuparene sesquiterpenes from Laurencia microcladia. Tetrahedron Lett. 2005, 46, 5723–5726. [Google Scholar] [CrossRef]
- Mao, S.-C.; Guo, Y.-W. A Laurane Sesquiterpene and Rearranged Derivatives from the Chinese Red Alga Laurencia okamurai Yamada. J. Nat. Prod. 2006, 69, 1209–1211. [Google Scholar] [CrossRef]
- Srikrishna, A.; Khan, I.; Babu, R.R.; Sajjanshetty, A. The first total synthesis of (±)-laurokamurene B. Tetrahedron 2007, 63, 12616–12620. [Google Scholar] [CrossRef]
- Irie, T.; Suzuki, T.; Yasunari, Y.; Kurosawa, E.; Masamune, T. Laurene, a sesquiterpene hydrocarbon from Laurencia species. Tetrahedron 1969, 25, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Wratten, S.J.; Faulkner, D.J. Metabolites of the red alga Laurencia subopposita. J. Org. Chem. 1977, 42, 3343–3349. [Google Scholar] [CrossRef]
- Alarif, W.M.; Al-Lihaibi, S.S.; Ayyad, S.-E.N.; Abdel-Rhman, M.H.; Badria, F.A. Laurene-type sesquiterpenes from the Red Sea red alga Laurencia obtusa as potential antitumor–antimicrobial agents. Eur. J. Med. Chem. 2012, 55, 462–466. [Google Scholar] [CrossRef]
- Yamamura, S.; Hirata, Y. Structures of aplysin and aplysinol, naturally occurring bromo-compounds. Tetrahedron 1963, 19, 1485–1496. [Google Scholar] [CrossRef]
- Elzen, G.W.; Williams, H.J.; Vinson, S.B. Isolation, identification and bioassay of cotton synomones mediating searching behavior by parasitoidCampoletis sonorensis. J. Chem. Ecol. 1984, 10, 1251–1264. [Google Scholar] [CrossRef] [PubMed]
- Mao, S.-C.; Guo, Y.-W. Cuparene-Derived Sesquiterpenes from the Chinese Red Alga Laurencia okamurai Yamada. Helv. Chim. Acta 2005, 88, 1034–1039. [Google Scholar] [CrossRef]
- Goldsmith, D.J.; John, T.K.; Kwong, C.D.; Painter, G.R. Preparation and rearrangement of trichothecane-like compounds. Synthesis of aplysin and filiformin. J. Org. Chem. 1980, 45, 3989–3993. [Google Scholar] [CrossRef]
- Shizuri, Y.; Yamada, K. Laurebiphenyl, a dimeric sesquiterpene of the cyclolaurane-type from the red alga Laurencia nidifica. Phytochemistry 1985, 24, 1385–1386. [Google Scholar] [CrossRef]
- Liang, Y.; Li, X.-M.; Li, C.-S.; Sun, H.; Wang, B.-G. Laurane-, Cyclolaurane-, and Cuparane-type Sesquiterpenes from the Marine Red Alga Laurencia okamurai. Nat. Prod. Commun. 2014, 9, 323–324. [Google Scholar] [CrossRef]
- Sun, J.; Han, L.; Shi, D.; Ma, M.; Li, S.; Wang, S.; Han, L.; Yang, Y.; Fan, X.; Shi, J.; et al. Sesquiterpenes from Red Alga Laurencia tristicha. J. Nat. Prod. 2008, 68, 915–919, Erratum in J. Nat. Prod. 2008, 71, 296. [Google Scholar] [CrossRef] [PubMed]
- Nemoto, H.; Nagamochi, M.; Ishibashi, H.; Fukumoto, K. A remarkable substituent effect on the enantioselectivity of tandem asymmetric epoxidation and enantiospecific ring expansion of cyclopropylidene alcohols: A new enantiocontrolled synthesis of (-)-debromoaplysin and (-)-aplysin. J. Org. Chem. 1994, 59, 74–79. [Google Scholar] [CrossRef]
- Ji, N.-Y.; Li, X.-M.; Li, K.; Ding, L.-P.; Wang, B.-G. Laurane-derived sesquiterpenes from the marine red alga Laurencia tristicha (Rhodomelaceae). Nat. Prod. Res. 2008, 22, 715–718. [Google Scholar] [CrossRef]
- Kazlauskas, R.; Murphy, P.; Quinn, R.; Wells, R. New laurene derivatives from Laurencia filiformis. Aust. J. Chem. 1976, 29, 2533–2539. [Google Scholar] [CrossRef]
- Dias, D.A.; White, J.M.; Urban, S. Laurencia Filiformis: Phytochemical Profiling by Conventional and HPLC-NMR Approaches. Nat. Prod. Commun. 2009, 4, 157–172. [Google Scholar] [CrossRef]
- Appleton, D.R.; Babcock, R.C.; Copp, B.R. Novel tryptophan-derived dipeptides and bioactive metabolites from the sea hare Aplysia dactylomela. Tetrahedron 2001, 57, 10181–10189. [Google Scholar] [CrossRef]
- König, G.M.; Wright, A.D. New C15 Acetogenins and Sesquiterpenes from the Red Alga Laurencia sp. cf. L. gracilis. J. Nat. Prod. 1994, 57, 477–485. [Google Scholar] [CrossRef]
- Kladi, M.; Xenaki, H.; Vagias, C.; Papazafiri, P.; Roussis, V. New cytotoxic sesquiterpenes from the red algae Laurencia obtusa and Laurencia microcladia. Tetrahedron 2006, 62, 182–189. [Google Scholar] [CrossRef]
- Iliopoulou, D.; Roussis, V.; Pannecouque, C.; De Clercq, E.; Vagias, C. Halogenated sesquiterpenes from the red alga Laurencia obtusa. Tetrahedron 2002, 58, 6749–6755. [Google Scholar] [CrossRef]
- Aydoğmuş, Z.; Imre, S.; Ersoy, L.; Wray, V. Halogenated secondary metabolites from Laurencia Obtusa. Nat. Prod. Res. 2004, 18, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Brennan, M.R.; Erickson, K.L. Austradiol acetate and austradiol diacetate, 4,6-dihydroxy-(+)-selinane derivatives from an Australian Laurencia sp. J. Org. Chem. 1982, 47, 3917–3921. [Google Scholar] [CrossRef]
- Kazlauskas, R.; Murphy, P.; Wells, R.; Daly, J.; Oberhansli, W. Heterocladol, a halogenated selinane sesquiterpene of biosynthetic significance from the red alga Laurencia filiformis: Its isolation, crystal structure andabsolute configuration. Aust. J. Chem. 1977, 30, 2679–2687. [Google Scholar] [CrossRef]
- Guella, G.; Skropeta, D.; Mancini, I.; Pietra, F. The First 6,8-Cycloeudesmane Sesquiterpene from a Marine Organism: The Red Seaweed Laurencia microcladia from the Baia di Calenzana, Elba Island. Z. Für Naturforschung B 2002, 57, 1147–1151. [Google Scholar] [CrossRef]
- Suzuki, M.; Takahashi, Y.; Mitome, Y.; Itoh, T.; Abe, T.; Masuda, M. Brominated metabolites from an Okinawan Laurencia intricata. Phytochemistry 2002, 60, 861–867. [Google Scholar] [CrossRef]
- Reward, B.M.; Fenical, W. α- and β-snyderol; new bromo-monocyclic sesquiterpenes from the seaweed Laurencia. Tetrahedron Lett. 1976, 17, 41–44. [Google Scholar] [CrossRef]
- Su, H.; Shi, D.-Y.; Li, J.; Guo, S.-J.; Li, L.-L.; Yuan, Z.-H.; Zhu, X.-B. Sesquiterpenes from Laurencia similis. Molecules 2009, 14, 1889–1897. [Google Scholar] [CrossRef]
- Kuniyoshi, M.; Marma, M.S.; Higa, T.; Bernardinelli, G.; Jefford, C.W. New Bromoterpenes from the Red Alga Laurencia luzonensis. J. Nat. Prod. 2001, 64, 696–700. [Google Scholar] [CrossRef]
- Kuniyoshi, M.; Wahome, P.G.; Miono, T.; Hashimoto, T.; Yokoyama, M.; Shrestha, K.L.; Higa, T. Terpenoids from Laurencia luzonensis. J. Nat. Prod. 2005, 68, 1314–1317. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Yuan, Z.-H.; Li, J.; Guo, S.-J.; Deng, L.-P.; Han, L.-J.; Zhu, X.-B.; Shi, D.-Y. Sesquiterpenes from the Marine Red Alga Laurencia saitoi. Helv. Chim. Acta 2009, 92, 1291–1297. [Google Scholar] [CrossRef]
- Makhanu, D.S.; Yokoyama, M.; Miono, T.; Maesato, T.; Maedomari, M.; Wisespongpand, P.; Kuniyoshi, M. New sesquiterpenes from the Okinawan red alga Laurencia luzonensis. Bull. Fac. Sci. Univ. Ryukyus. 2006, 81, 115–120. [Google Scholar]
- Vairappan, C.S.; Kamada, T.; Lee, W.-W.; Jeon, Y.-J. Anti-inflammatory activity of halogenated secondary metabolites of Laurencia snackeyi (Weber-van Bosse) Masuda in LPS-stimulated RAW 264.7 macrophages. J. Appl. Phycol. 2013, 25, 1805–1813. [Google Scholar] [CrossRef]
- Wijesinghe, W.; Kim, E.-A.; Kang, M.-C.; Lee, W.-W.; Lee, H.-S.; Vairappan, C.S.; Jeon, Y.-J. Assessment of anti-inflammatory effect of 5β-hydroxypalisadin B isolated from red seaweed Laurencia snackeyi in zebrafish embryo in vivo model. Environ. Toxicol. Pharmacol. 2014, 37, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Yuan, Z.; Li, J.; Guo, S.; Han, L.; Zhu, X.; Shi, D. [Studies on chemical constituents of Laurencia saitoi]. Zhongguo Zhong Yao Za Zhi 2009, 34, 871–874. [Google Scholar]
- Wright, A.D.; Goclik, E.; König, G.M. Three New Sesquiterpenes from the Red Alga Laurencia perforata. J. Nat. Prod. 2003, 66, 435–437. [Google Scholar] [CrossRef]
- Vairappan, C.S.; Suzuki, M.; Ishii, T.; Okino, T.; Abe, T.; Masuda, M. Antibacterial activity of halogenated sesquiterpenes from Malaysian Laurencia spp. Phytochemistry 2008, 69, 2490–2494. [Google Scholar] [CrossRef]
- Erickson, K.L.; Beutler, J.A.; Gray, G.N.; Cardellina, J.H.; Boyd, M.R. Majapolene A, a Cytotoxic Peroxide, and Related Sesquiterpenes from the Red Alga Laurencia majuscula. J. Nat. Prod. 1995, 58, 1848–1860. [Google Scholar] [CrossRef]
- Monde, K.; Taniguchi, T.; Miura, N.; Vairappan, C.S.; Suzuki, M. Absolute configurations of brominated sesquiterpenes determined by vibrational circular dichroism. Chirality 2006, 18, 335–339. [Google Scholar] [CrossRef]
- Da Silva, F.L.M.; Pacienza-Lima, W.; Rossi-Bergmann, B.; de Souza, L.M.G.; Fujii, M.T.; Campos de Paula, J.; Costa, S.S.; Lopes, N.P.; Kaiser, C.R.; Soares, A.R. Antileishmanial Sesquiterpenes from the Brazilian Red Alga Laurencia dendroidea. Planta Med. 2010, 77, 733–735. [Google Scholar] [CrossRef]
- Al-Massarani, S. Phytochemical and Biological Properties of Sesquiterpene Constituents From the Marine Red Seaweed Laurencia: A Review. Nat. Prod. Chem. Res. 2014, 2, 147. [Google Scholar] [CrossRef]
- Wang, B.; Svetlov, V.; Wolf, Y.I.; Koonin, E.V.; Nudler, E.; Artsimovitch, I. Allosteric Activation of SARS-CoV-2 RNA-Dependent RNA Polymerase by Remdesivir Triphosphate and Other Phosphorylated Nucleotides. mBio 2021, 12, e0142321. [Google Scholar] [CrossRef] [PubMed]
- National Center for Biotechnology Information Protein Database. Available online: http://www.ncbi.nlm.nih.gov/protein/ (accessed on 10 March 2023).
- Yin, W.; Mao, C.; Luan, X.; Shen, D.-D.; Shen, Q.; Su, H.; Wang, X.; Zhou, F.; Zhao, W.; Gao, M.; et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 2020, 368, 1499–1504. [Google Scholar] [CrossRef]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem substance and compound databases. Nucleic Acids Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef]
- Pence, H.E.; Williams, A. ChemSpider: An Online Chemical Information Resource. J. Chem. Educ. 2010, 87, 1123–1124. [Google Scholar] [CrossRef]
- Lyu, C.; Chen, T.; Qiang, B.; Liu, N.; Wang, H.; Zhang, L.; Liu, Z. CMNPD: A comprehensive marine natural products database towards facilitating drug discovery from the ocean. Nucleic Acids Res. 2020, 49, D509–D515. [Google Scholar] [CrossRef] [PubMed]
- Biovia, D.S. Discovery Studio. Available online: https://discover.3ds.com/discovery-studio-visualizer-download (accessed on 15 December 2022).
- Avogadro: An Open-Source Molecular Builder and Visualization Tool. Version 1.95. Available online: http://avogadro.cc/ (accessed on 23 November 2022).
- Hanwell, M.D.; Curtis, D.E.; Lonie, D.C.; Vandermeersch, T.; Zurek, E.; Hutchison, G.R. Avogadro: An advanced semantic chemical editor, visualization, and analysis platform. J. Cheminform. 2012, 4, 17. [Google Scholar] [CrossRef] [PubMed]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-Pdb Viewer: An environment for comparative protein modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A.F.; Forli, S. AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings. J. Chem. Inf. Model. 2021, 61, 3891–3898. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Kuriata, A.; Gierut, A.M.; Oleniecki, T.; Ciemny, M.P.; Kolinski, A.; Kurcinski, M.; Kmiecik, S. CABS-flex 2.0: A web server for fast simulations of flexibility of protein structures. Nucleic Acids Res. 2018, 46, W338–W343. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, P.; Eckert, A.O.; Schrey, A.K.; Preissner, R. ProTox-II: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2018, 46, W257–W263. [Google Scholar] [CrossRef] [PubMed]
- Borba, J.V.; Alves, V.M.; Braga, R.C.; Korn, D.R.; Overdahl, K.; Silva, A.C.; Hall, S.U.; Overdahl, E.; Braga, R.; Kleinstreuer, N.; et al. STopTox: An in Silico Alternative to Animal Testing for Acute Systemic and Topical Toxicity. Environ. Health Perspect. 2022, 130, 27012. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.H.; Abraham, M.H.; Le, J.; Hersey, A.; Luscombe, C.N.; Beck, G.; Sherborne, B.; Cooper, I. Rate-Limited Steps of Human Oral Absorption and QSAR Studies. Pharm. Res. 2002, 19, 1446–1457. [Google Scholar] [CrossRef]
- Husain, A.; Ahmad, A.; Alam Khan, S.; Asif, M.; Bhutani, R.; Al-Abbasi, F.A. Synthesis, molecular properties, toxicity and biological evaluation of some new substituted imidazolidine derivatives in search of potent anti-inflammatory agents. Saudi Pharm. J. 2016, 24, 104–114. [Google Scholar] [CrossRef]
- Filimonov, D.A.; Lagunin, A.A.; Gloriozova, T.A.; Rudik, A.V.; Druzhilovskii, D.S.; Pogodin, P.V.; Poroikov, V.V. Prediction of the Biological Activity Spectra of Organic Compounds Using the Pass Online Web Resource. Chem. Heterocycl. Compd. 2014, 50, 444–457. [Google Scholar] [CrossRef]
- Gao, Y.; Yan, L.; Huang, Y.; Liu, F.; Zhao, Y.; Cao, L.; Wang, T.; Sun, Q.; Ming, Z.; Zhang, L.; et al. Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 2020, 368, 779–782. [Google Scholar] [CrossRef]
- Begum, F.; Srivastava, A.K.; Ray, U. Repurposing nonnucleoside antivirals against SARS-CoV2 NSP12 (RNA dependent RNA polymerase): In silico-molecular insight. Biochem. Biophys. Res. Commun. 2021, 571, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Mosayebnia, M.; Bozorgi, A.H.; Rezaeianpour, M.; Kobarfard, F. In silico prediction of SARS-CoV-2 main protease and polymerase inhibitors: 3D-Pharmacophore modelling. J. Biomol. Struct. Dyn. 2022, 40, 6569–6586. [Google Scholar] [CrossRef] [PubMed]
- Tungary, E.; Wahjudi, M.; Kok, T. Secondary Metabolites of Various Indonesian Medicinal Plants as SARS-CoV-2 Inhibitors: In Silico Study. MPI Media Pharm. Indones. 2022, 4, 136–146. [Google Scholar] [CrossRef]
- Ikeguchi, M.; Ueno, J.; Sato, M.; Kidera, A. Protein Structural Change Upon Ligand Binding: Linear Response Theory. Phys. Rev. Lett. 2005, 94, 078102. [Google Scholar] [CrossRef]
- Agoni, C. The Binding of Remdesivir to SARS-CoV-2 RNA-Dependent RNA Polymerase May Pave The Way Towards the Design of Potential Drugs for COVID-19 Treatment. Curr. Pharm. Biotechnol. 2021, 22, 1520–1537. [Google Scholar] [CrossRef]
- Mishra, A.; Rathore, A.S. RNA dependent RNA polymerase (RdRp) as a drug target for SARS-CoV2. J. Biomol. Struct. Dyn. 2022, 40, 6039–6051. [Google Scholar] [CrossRef]
- Arora, S.; Lohiya, G.; Moharir, K.; Shah, S.; Yende, S. Identification of Potential Flavonoid Inhibitors of the SARS-CoV-2 Main Protease 6YNQ: A Molecular Docking Study. Digit. Chin. Med. 2020, 3, 239–248. [Google Scholar] [CrossRef]
- Umar, A.K. Flavonoid compounds of buah merah (Pandanus conoideus Lamk) as a potent SARS-CoV-2 main protease inhibitor: In silico approach. Futur. J. Pharm. Sci. 2021, 7, 158–159. [Google Scholar] [CrossRef]
- Ortega, J.T.; Jastrzebska, B.; Rangel, H.R. Omicron SARS-CoV-2 Variant Spike Protein Shows an Increased Affinity to the Human ACE2 Receptor: An In Silico Analysis. Pathogens 2021, 11, 45. [Google Scholar] [CrossRef]
- Katuwal, S.; Upadhyaya, S.R.; Marahatha, R.; Shrestha, A.; Regmi, B.P.; Khadayat, K.; Basnet, S.; Basnyat, R.C.; Parajuli, N. In Silico Study of Coumarins: Wedelolactone as a Potential Inhibitor of the Spike Protein of the SARS-CoV-2 Variants. J. Trop. Med. 2023, 2023, 4771745. [Google Scholar] [CrossRef]
- Pokharkar, O.; Lakshmanan, H.; Zyryanov, G.; Tsurkan, M. In Silico Evaluation of Antifungal Compounds from Marine Sponges against COVID-19-Associated Mucormycosis. Mar. Drugs 2022, 20, 215. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Liu, J.; Zhang, D.; Xu, Z.; Ji, J.; Wen, C. Cytokine Storm in COVID-19: The Current Evidence and Treatment Strategies. Front. Immunol. 2020, 11, 1708. [Google Scholar] [CrossRef] [PubMed]
- Cron, R.Q.; Caricchio, R.; Chatham, W.W. Calming the cytokine storm in COVID-19. Nat. Med. 2021, 27, 1674–1675. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).