Abstract
During chromatic acclimation (CA), cyanobacteria undergo shifts in their physiology and metabolism in response to changes in their light environment. Various forms of CA, which involves the tuning of light-harvesting accessory complexes known as phycobilisomes (PBS) in response to distinct wavelengths of light, have been recognized. Recently, a negative regulator of PBS abundance, PsoR, about which little was known, was identified. We used sequence analyses and bioinformatics to predict the role of PsoR in cyanobacteria and PBS regulation and to examine its presence in a diverse range of cyanobacteria. PsoR has sequence similarities to the β-CASP family of proteins involved in DNA and RNA processing. PsoR is a putative nuclease widespread across Cyanobacteria, of which over 700 homologs have been observed. Promoter analysis suggested that psoR is co-transcribed with upstream gene tcpA. Multiple transcription factors involved in global gene regulation and stress responses were predicted to bind to the psoR-tcpA promoter. The predicted protein–protein interactions with PsoR homologs included proteins involved in DNA and RNA metabolism, as well as a phycocyanin-associated protein predicted to interact with PsoR from Fremyella diplosiphon (FdPsoR). The widespread presence of PsoR homologs in Cyanobacteria and their ties to DNA- and RNA-metabolizing proteins indicated a potentially unique role for PsoR in CA and PBS abundance regulation.
1. Introduction
Cyanobacteria are oxygenic phototrophs that are responsible for 20–30% of global carbon fixation and the majority of global nitrogen fixation [1,2]. These prokaryotes serve as model organisms for studying photosynthesis and photoacclimation because of their fast growth and the tools available for the genetic manipulation of many species. In addition to research on the mechanisms governing their growth and fitness in natural contexts, there has been ongoing research into their use as a microbial chassis in the biotech industry for the production of biofuels, biopharmaceuticals, food supplements, bioremediation, and biofertilizers [3,4]. Due to the growing interest in this ancient phylogeny of bacteria, it is important to fully understand the genes and mechanisms that control their photosynthetic potential, productivity, and adaptive responses.
Cyanobacteria have evolved to survive in a wide range of ecological niches, including freshwater and marine environments, as well as in arid deserts, arctic tundra, and hot springs [5,6,7]. The dynamic conditions in these environments present various challenges for cyanobacteria, including nutrient limitations and fluctuating light levels, which impact organismal growth and fitness. To deal with variable photoenvironments, cyanobacteria have evolved a mechanism called chromatic acclimation (CA), which allows them to perceive and respond to a given light condition and adjust their physiology and metabolism accordingly to maximize productivity while minimizing the photodamage caused by excess light absorption [8,9,10].
A major cellular change that occurs during CA includes alterations to the pigment profile of phycobilisomes (PBS), which are accessory light-harvesting antennae that capture and transfer light energy to chlorophyll-containing photosystems for photosynthesis. PBSs are made up of phycobiliproteins that form a core with outward-radiating rods [11,12]. The PBS core is attached via a core-member linker to chlorophyll-containing reaction centers embedded in thylakoid membranes [13]. In addition to altering the rod pigments during CA in response to changes in the prevalence of red light (RL) vs. green light (GL) and the light intensity, PBSs undergo restructuring and degradation under high-stress conditions, such as in high light and with nutrient limitations [14,15,16,17,18,19]. As PBSs are light-harvesting accessory complexes, their structure changes during CA and in response to stresses, which has implications for pigment coupling and energy transfer rates [20,21], and ultimately, the overall photosynthetic potential and efficiency.
Three main types of CA have been discussed in the literature for nearly 50 years. Distinct CA types are largely categorized by whether and how RL-absorbing phycocyanin (PC) and GL-absorbing phycoerythrin (PE) phycobiliproteins are synthesized and incorporated into the rods of PBSs in response to the availability of external light [11,22]. Type I CA occurs when cyanobacteria do not alter the levels of PC or PE under different light conditions. In cyanobacteria that exhibit type II CA, PC levels remain unchanged, whereas PE levels are tuned in response to light cues, where PE is induced under conditions rich in GL wavelengths. In cyanobacteria with type III CA, which is also known as complementary chromatic acclimation (CCA), PE accumulates under GL, whereas inducible PC (PCi) differentially accumulates in the outer portion of PBS rods under RL [23]. Recently, additional forms of CA have been discovered in cyanobacteria that respond to other types of light, including type IV CA, in which organisms respond to blue and green wavelengths of light [24] and far-red light photoacclimation (FaRLiP), which results in cyanobacterial tuning of PBSs in response to RL and far-red (FR) light [25,26].
CCA, which is the form of CA that has been studied the most extensively in freshwater filamentous cyanobacterium Fremyella diplosiphon [14], is largely under the control of the photoreceptor RcaE [27]. RcaE works through two response regulators, RcaF and RcaC, to exert transcriptional regulation of the pigment content of PBSs in response to dynamic light signals [28,29,30]. RcaC is a DNA-binding transcriptional regulator that directly controls the expression of phycobiliprotein-encoding genes [31,32,33]. Under RL conditions, RcaC binds to and upregulates the cpc2 operon, which codes PCi proteins and upregulates pcyA, which in turn, codes an oxidoreductase that produces phycocyanobilin. The latter is the bilin that attaches to PC [31,34]. In RL, RcaC also represses the cpe operon, which produces PE proteins, and the pebAB operon, which encodes two oxidoreductases that produce phycoerythrobilin, which is the bilin that attaches to PE [31,33]. Under GL conditions, the repression of PE synthesis genes is relieved, resulting in PE accumulation [11]. In addition to RcaE, PE levels are also regulated by the Cgi system, a GL-dependent regulatory system [35].
An F. diplosiphon mutant with a spontaneous transposon insertion within the phycobilisome abundance regulator gene, psoR, displays disrupted pigmentation and growth [36]. Here, we describe bioinformatic analyses that were conducted to explore the presence of PsoR across cyanobacteria, as well as an assessment of conserved residues and putative interacting partners, with the aim of understanding its potential function. Sequence and phylogenetic analyses determined that PsoR was widespread throughout the cyanobacterial phylum but absent both outside of cyanobacteria and in PBS containing red algae. The results of the analyses of conserved protein sequences and amino residues suggested that PsoR functions as a β-CASP domain-containing ribonuclease within the metallo-β-lactamase fold superfamily, which may have direct implications for regulating pigment synthesis in vivo. Additionally, putative PsoR-interacting proteins further supported a potential role for the protein in regulating PBS abundance in response to external light cues and/or stress.
2. Materials and Methods
2.1. Sequence Analysis and Protein Modeling
The 554-amino-acid sequence of PsoR from F. diplosiphon was obtained from the National Center for Biotechnology Information (NCBI) database (GenBank ID: EKE97789.1) and analyzed using the Phyre2 Protein Fold Recognition Server [37] to predict the protein structure, including a search for conserved domains, sequence features, and prediction(s) of putative functions.
2.2. Sequence Homology and Phylogeny
A protein sequence of PsoR from F. diplosiphon was run through the position-specific iterated BLAST (PSI-BLAST) algorithm on the NCBI website [38]. Three successive iterations were performed to refine the results. Multiple sequence alignments of the obtained sequences were generated through the MEGA-7 program using MUSCLE [39]. A phylogenetic tree was created using the maximum likelihood method and the Jones-Taylor-Thornton matrix-based model in MEGA-X [40]. The likelihood log was-746644.20. An unrelated β-CASP protein from Gloeobacter violaceus (Gene ID: WP_011141021.1) was used to root the phylogenetic tree.
2.3. Promoter Analyses
The BPROM program [41] was used to analyze the potential promoter and transcription factor binding sites associated with the F. diplosiphon psoR gene sequence as well as the upstream and downstream areas flanking it. The analyzed sequence was retrieved from the NCBI database (accession ID: DQ286230.1).
2.4. Predicted Protein–Protein Interactions
We used the STRING program to search for potential proteins that may interact with PsoR from F. diplosiphon (FdPsoR) and its homologs [42]. The PsoR protein sequence (GenBank ID: EKE97789.1) from F. diplosiphon and homologs from Synechocystis sp. PCC 6803 (GenBank ID: sll0514) and Synechocystis sp. PCC 7002 (GenBank ID: ACA99583.1) were used.
3. Results
3.1. PsoR Is a Putative β-CASP Domain-Containing Ribonuclease
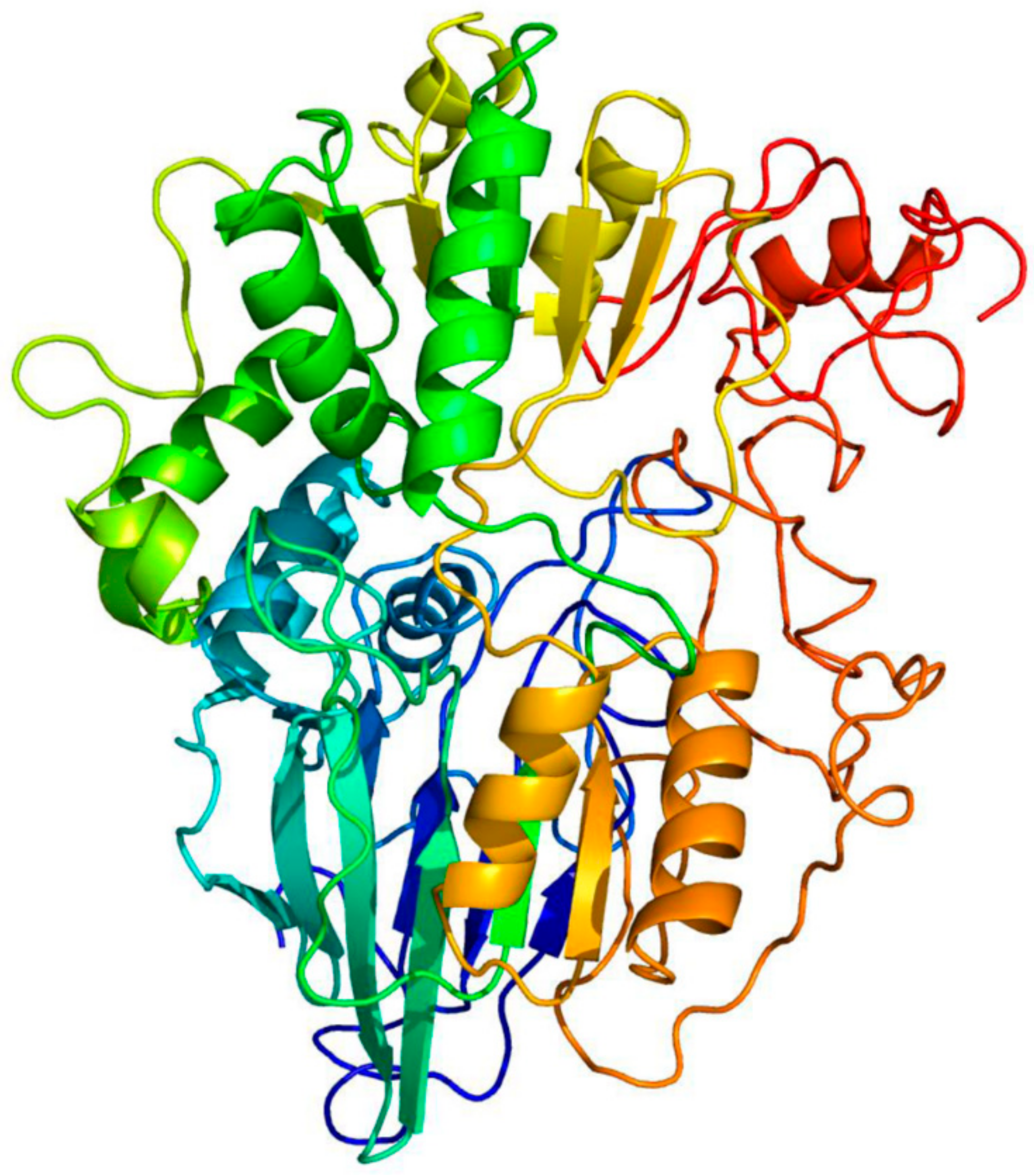
Using the Phyre2 server to predict the structure of FdPsoR, we found matches with β-CASP domain-containing proteins within the metallo-β-lactamase fold superfamily (Table 1). Based on the solved structures of known β-CASP proteins, Phyre2 was able to predict the overall structure of PsoR with 82% of residues modeled at >90% confidence (Figure 1). The β-lactamase superfamily contains proteins that act on a wide range of substrates, including DNA and RNA [43]. β-CASP domain-containing proteins are a subgroup that typically displays endo- and exonuclease activity on DNA and RNA substrates. These proteins also play a role in DNA repair and pre-mRNA maturation [44]. Although both DNA- and RNA-processing β-CASP nucleases have been found in eukaryotes, only β-CASP homologs acting on RNA have been found in bacteria and archaea to date [45]. The enzymes in this family include the Saccharomyces cerevisiae 3′-processing endonuclease Ysh1 and the cleavage and polyadenylation specificity factor MTH1203 from Methanothermobacter thermautotrophicus [46,47].

Table 1.
Phyre2 top hits for the PsoR sequence. The PsoR amino acid sequence from Fremyella diplosiphon was submitted to the Phyre2 Protein Fold Recognition Server. Multiple hits were retrieved with similarities to the Ysh1 superfamily of proteins. ID = identity.
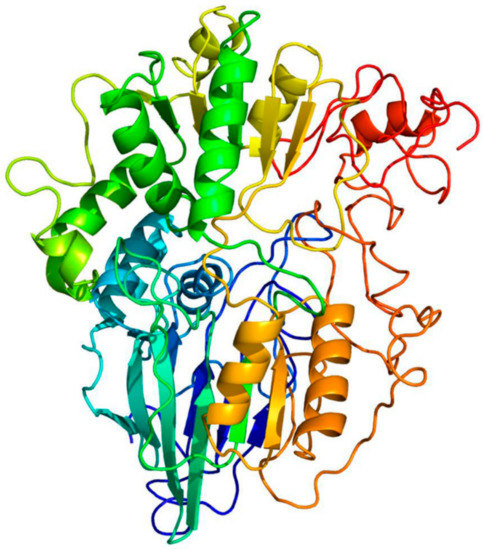
Figure 1.
Phyre2 modeling of the Fremyella diplosiphon PsoR protein. The structure of FdPsoR was modeled on the β-CASP containing enzyme MTH1203 from Methanothermobacter thermautotrophicus [46,47]; 82% of the FdPsoR residues were modeled at >90% confidence. The protein is rainbow-colored: red (N-terminus) to blue (C-terminus). This figure was made/generated by Alicia Layer.
The β-lactamase and β-CASP domain regions of FdPsoR span residues 20–430 (Figure 2). A C-terminal region followed from residue 431 to residue 554, and the protein contained a small n-terminal region comprising residues 1 to 19. β-CASP enzymes within prokaryotes typically function as dimers, with the C-terminal regions involved in dimerization [45,47,48,49]. Considering the 123-amino-acid C-terminal region of FdPsoR, this region may serve to dimerize or facilitate the interaction of PsoR with other proteins in vivo, although in this region, there were no sequence or structural similarities with putative interaction domains of other β-CASP proteins.

Figure 2.
Schematic of FdPsoR protein domains. The full-length PsoR sequence from Fremyella diplosiphon was analyzed using BLASTp [50]. Amino acids 19–430 of FdPsoR have sequence similarity to the Ysh1 superfamily of proteins (shown as a blue-labeled box).
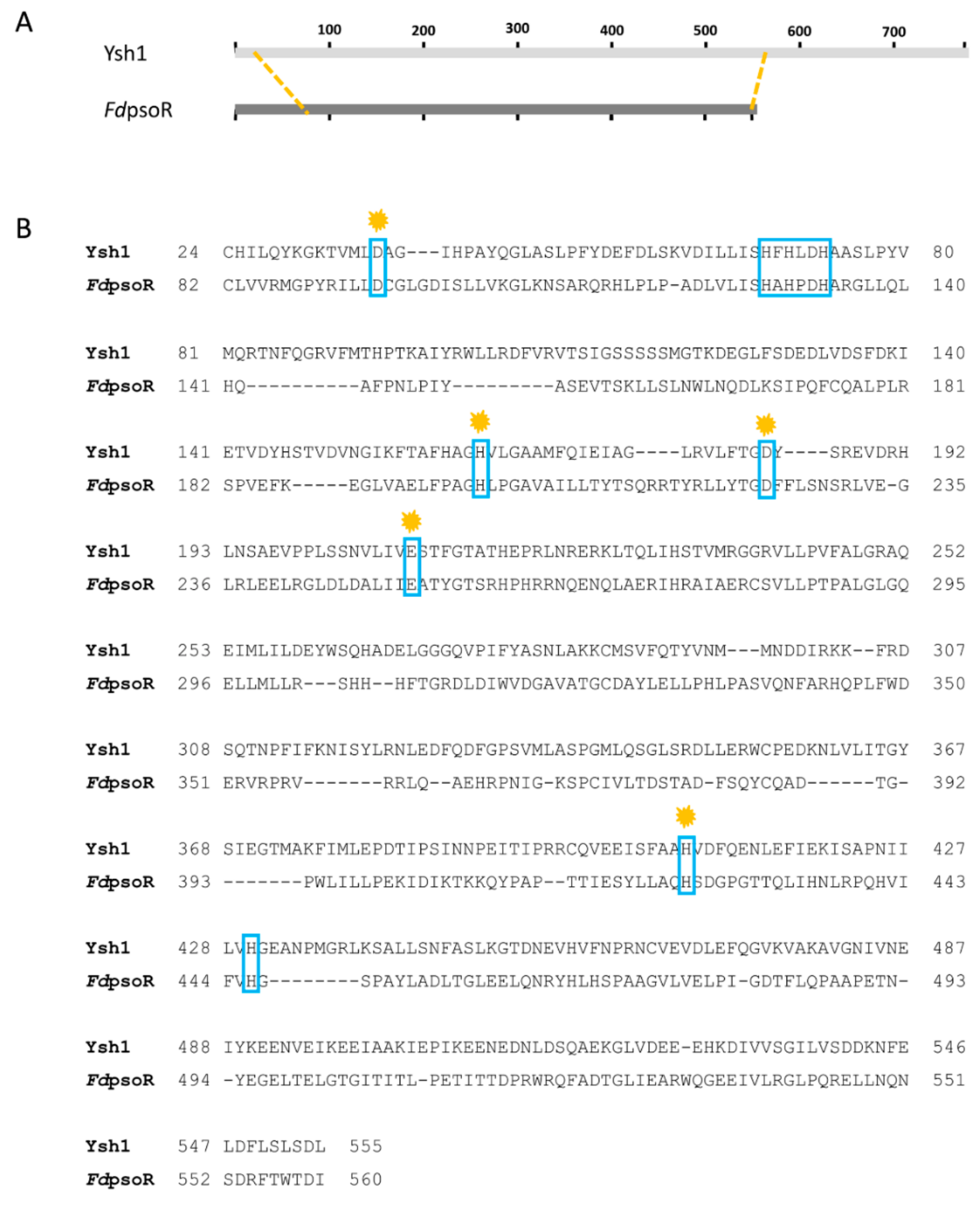
The β-CASP family of proteins has seven conserved residues that are involved in substrate binding and hydrolysis [44]. FdPsoR contains all seven conserved residues associated with enzymatic activity (Figure 3). The motifs contain aspartic acids, glutamic acids, and histidines, which are involved in binding metal ions, particularly zinc and magnesium (Table 2) [51].
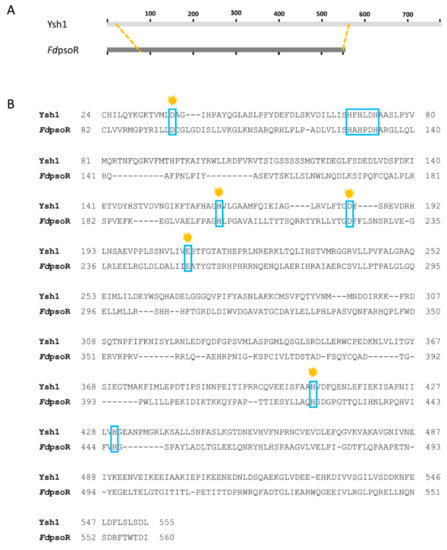
Figure 3.
Sequence alignment of Fremyella diplosiphon PsoR (FdPsoR) with mRNA endonuclease Ysh1 from Saccharomyces cerevisiae. Amino acid sequence alignment with FdPsoR and the mRNA endonuclease Ysh1 from S. cerevisiae using the NCBI Protein BLAST program: (A) Schematic showing the overall location of homology between FdPsoR and Ysh1. Orange dashed lines indicate where homology begins and ends. (B) Amino acid alignment of homologous regions between Ysh1 and FdPsoR. Blue boxes indicate conserved residues found in the seven motifs present in β-CASP proteins. Orange marks indicate areas where point mutations in Ysh1 resulted in a loss of endonuclease activity [51].

Table 2.
The seven conserved motifs of β-CASP proteins present in FdPsoR and Ysh1. Motifs 1–4 are found in the Metallo-β-lactamase superfamily of proteins involved in the hydrolysis of different substrates. In addition to these four motifs are motifs A, B, and C found in β-CASP domain-containing proteins involved in nucleolytic activity, using DNA and RNA as substrates. The motifs, in particular, are involved in metal binding, such as zinc and magnesium, to form the catalytic reaction center of the protein where cleavage occurs.
3.2. PsoR Is Widespread throughout the Cyanobacteria Phylum
A PSI-BLAST search resulted in the identification of 798 homologs found within the cyanobacteria phylum. Species containing a PsoR homolog had only one copy. No homologs were found outside the cyanobacteria. PsoR homologs were notably absent from red algae, another group of photosynthetic organisms containing PBSs.
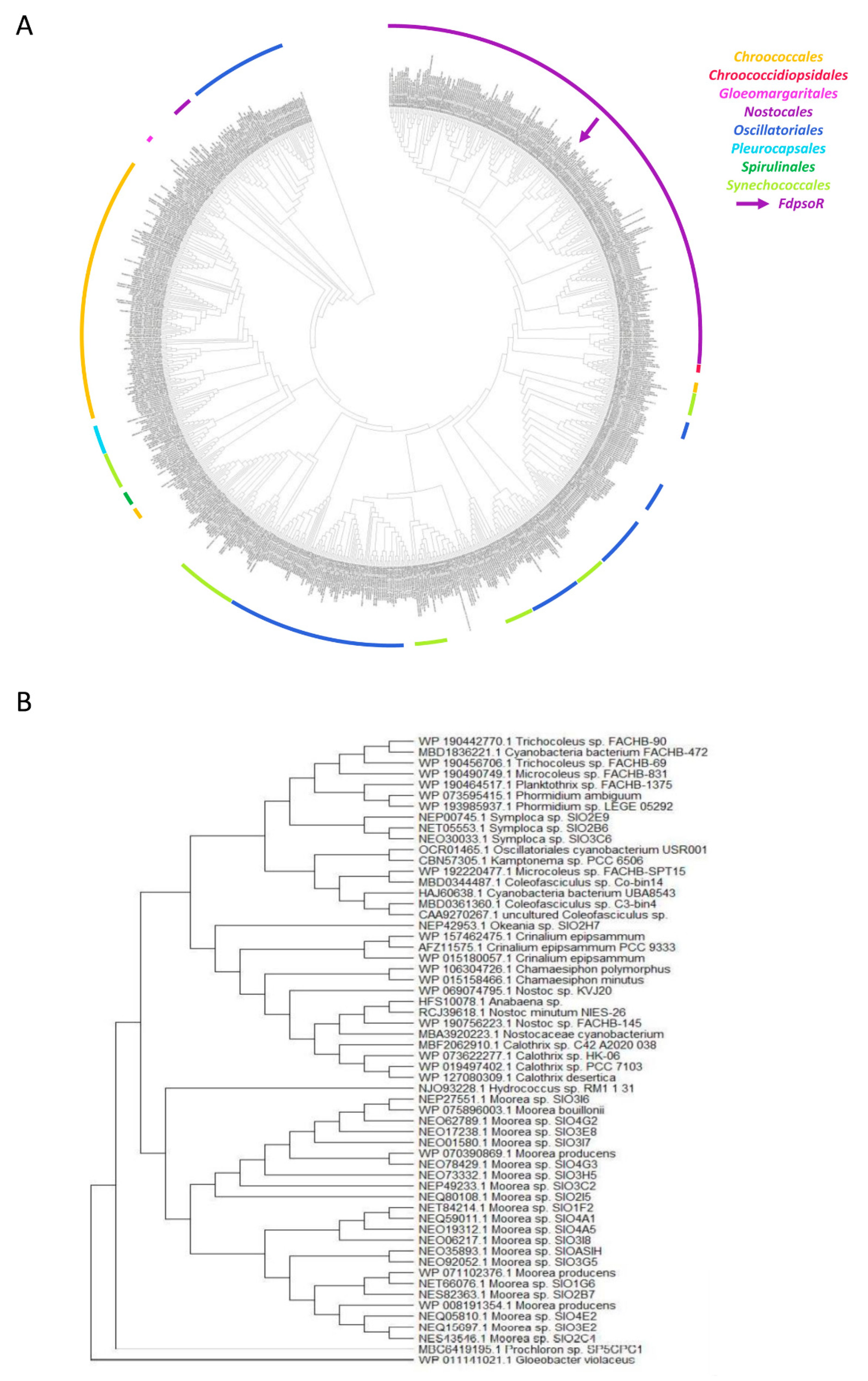
Homologs were found in cyanobacteria in all orders except Gloeobacter, the oldest extant group of cyanobacteria, which lacks thylakoid membranes and has atypical PBS structures [52,53]. A PsoR phylogenetic tree was generated and rooted using an unrelated β-CASP protein found in Gloeobacter (Figure 4A). PsoR homologs are generally grouped based on order and family clades, in which PsoR from F. diplosiphon is grouped with other homologs found in Nostocales species. The overall tree structure resembled that of a previous phylogenetic tree based on cyanobacterial genomes [54], suggesting that PsoR was vertically inherited. The majority of homologs (742 of 799) were the closest in size to FdPsoR, and the entire group ranged from 86 to 684 bp. These homologs contained one to three conserved motifs of β-CASP proteins. The shortest homologs in this group were not grouped in a single clade but were interspersed among the larger homologs. Despite their small size, these homologs were retained in the analysis because they may represent truncated forms of PsoR. In total, 57 homologs ranging in size from 344 to 918 bp, grouped in the phylogenetic tree closest to the outgroup, and found predominantly in Oscillatoriales (Figure 4B).
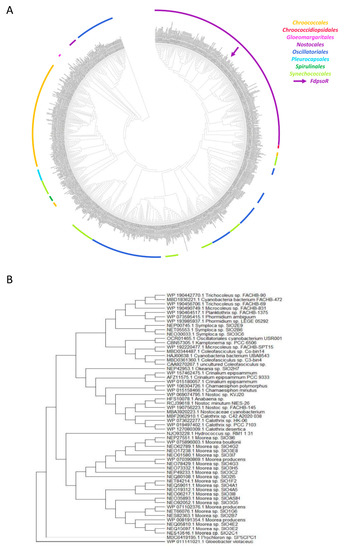
Figure 4.
Phylogenetic tree of PsoR within cyanobacteria. Protein sequences of PsoR homologs from cyanobacteria were identified using FdPsoR as the query sequence with the PSI-BLAST (Position-Specific Iterated BLAST) algorithm on the NCBI website [38]. The identified PsoR homologs were aligned through the MEGA-7 program using MUSCLE [39]. (A) A phylogenetic tree was created using the maximum likelihood method and Jones-Taylor-Thornton matrix-based model in MEGA-X [40]. The likelihood log was −746644.20. An unrelated β-CASP protein from Gloeobacter violaceus (Gene ID: WP_011141021.1) was used to root the phylogenetic tree. Colors indicate clusters of cyanobacterial orders. (B) Phylogenetic tree of the PsoR homolog branch closest to the Gloeobacter violaceus outgroup.
3.3. Promoter Analysis for psoR in Fremyella Diplosiphon
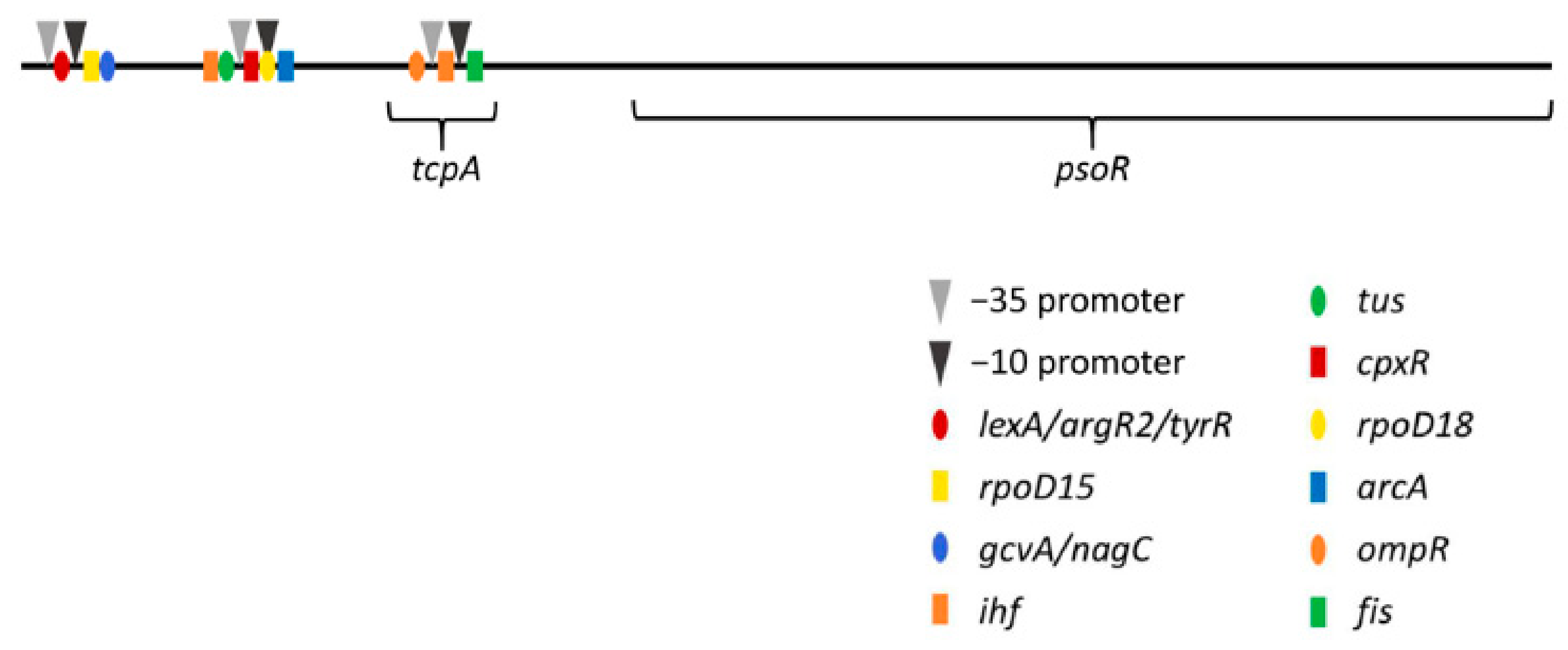
To determine the locations of potential promoter regions for psoR from F. diplosiphon and of possible transcriptional regulatory elements, a sequence-based promoter analysis was performed. Using the BPROM program from the Softberry website, three putative promoter regions and several transcription factor binding sites were located. Two of these promoter regions were upstream of a gene named tcpA, which is found upstream of psoR, while the third promoter region was located within tcpA itself (Figure 5). In addition to the −10 and −35 promoter regions, 13 transcription factor binding sites were found (Table 3). All but one of the factor binding sites were predicted to co-localize with one of the three promoter sites, and Ihf was predicted to bind two promoter regions; ihf and fis were found to encode global regulators that transcriptionally control hundreds of genes in response to environmental stimuli [55]. Another transcription factor binding site for OmpR was identified. Of note, the response regulator RcaC, which is involved in the regulation of CCA, is a member of the OmpR/PhoB family of DNA-binding proteins [33]. LexA encodes a transcriptional repressor involved in salt stress responses [56].
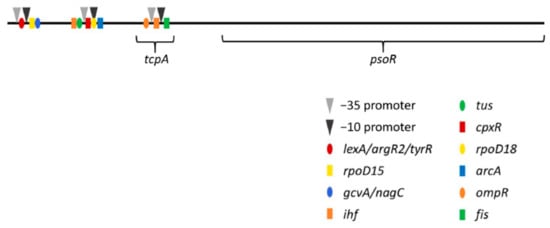
Figure 5.
Schematic of potential promoter and transcription factor binding sites upstream of psoR. Predicted promoter and transcriptional factor binding sites for psoR shown with upstream gene tcpA. The BPROM program [41] was used to analyze the potential promoter and transcription factor binding sites associated with the F. diplosiphon tcpA-psoR genomic region. The sequence analyzed was retrieved from the NCBI database (accession ID: DQ286230.1). In the figure, −10 and −35 promoter sites are indicated by arrowheads, while the predicted transcription factor binding sites are indicated by color-coded symbols.

Table 3.
List of potential promoter sites and transcription factor binding sites upstream of psoR in Fremyella diplosiphon. The sequence containing the upstream region of psoR had three predicted promoter regions. Potential transcription factor binding sites were also found, with binding sequences and locations indicated. Bp = base pairs; TF = transcription factor.
3.4. Protein–Protein Interactions of FdPsoR and Synechocystis PsoR Homologs
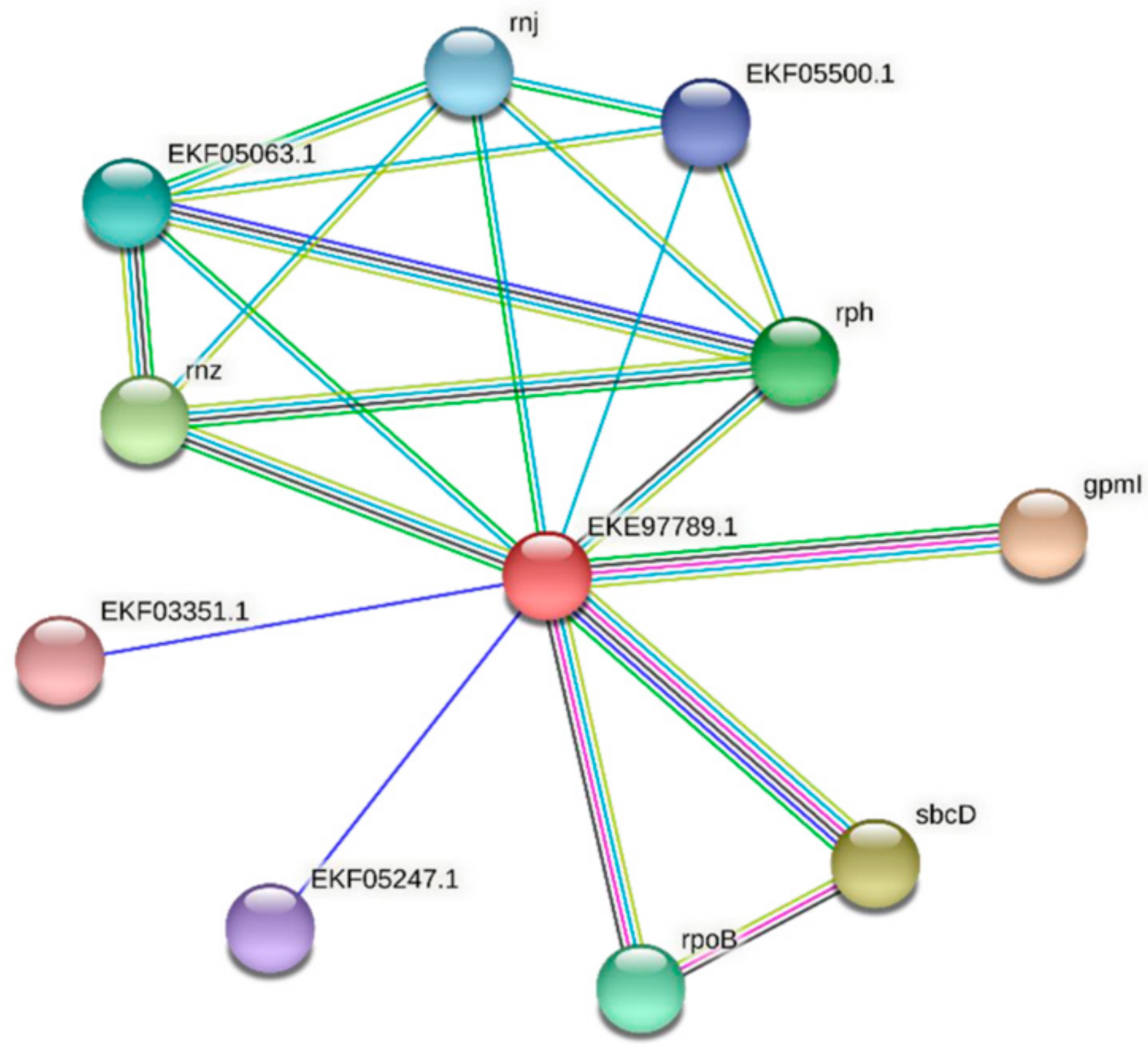
To predict whether FdPsoR and its homologs interact with other proteins in vivo, the STRING program was used to predict protein–protein interactions [42]. For FdPsoR, 10 potential interactors were determined (Figure 6). The majority of the proteins predicted to interact with FdPsoR were involved in DNA and RNA metabolism (Table 4). Of note, given the potential role of PsoR in regulating PBS abundance, one of the proteins identified as a potential interacting partner for FdPsoR was EKF03351.1, a phycocyanin-associated protein.
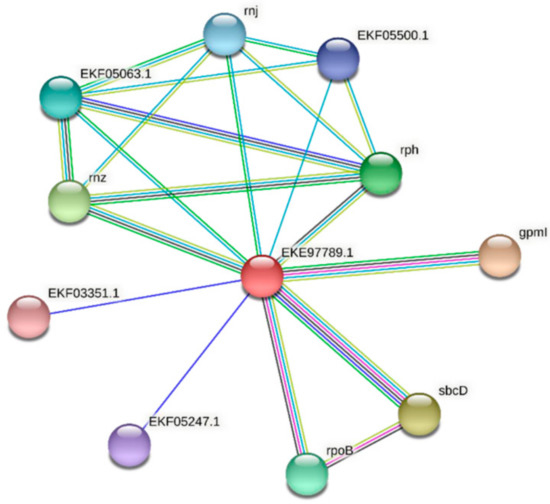
Figure 6.
Protein–protein interaction network for FdPsoR. A protein–protein interaction network was generated using STRING [42]. The PsoR protein sequence (GenBank ID: EKE97789.1) from F. diplosiphon was used as a query. Edges represent functional associations, and differently colored lines represent different types of interactions: green line, neighborhood of genes; blue line, co-occurrence across species; purple line, experimental evidence; yellow-green line, text mining; light blue line, database documentation; black line, co-expression data.

Table 4.
Proteins predicted to interact with FdPsoR from Fremyella diplosiphon using the STRING program.
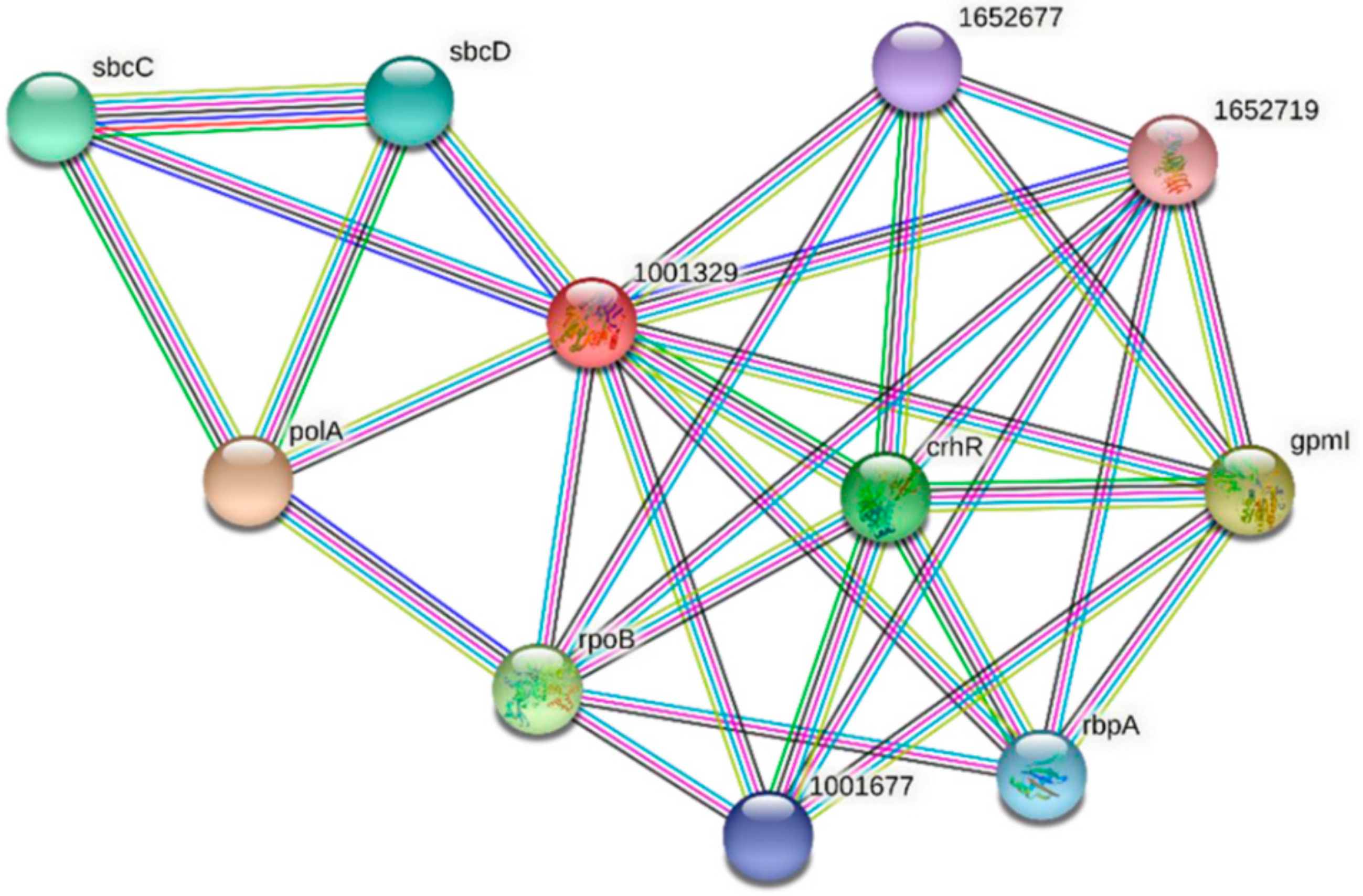
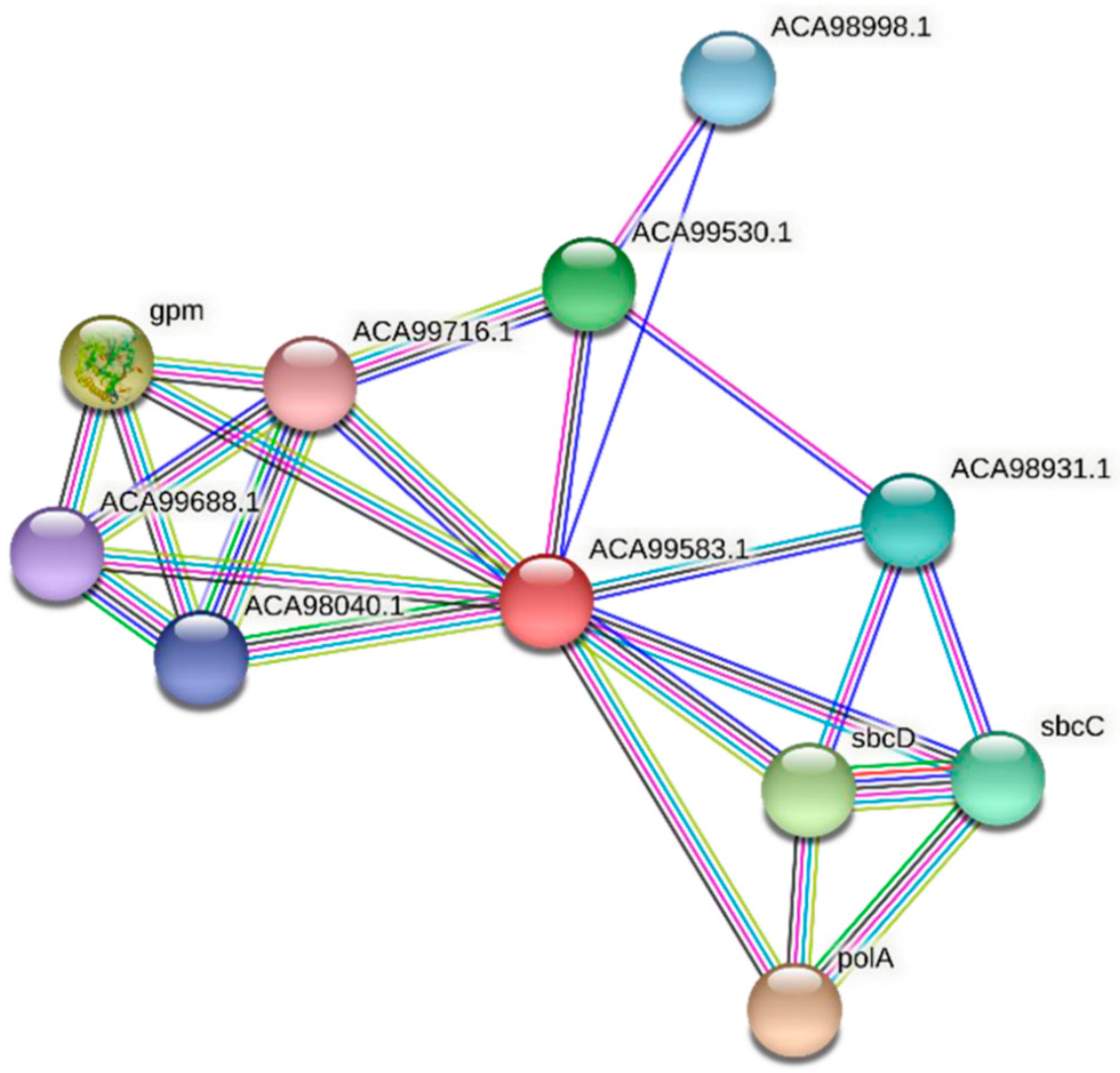
Two homologs of FdPsoR, Sll0514 from Synechocystis sp. PCC 6803 and ACA99583.1 from Synechocystis sp. PCC 7002, were also submitted to STRING. Ten proteins were predicted to interact with Sll0514 (Figure 7, Table 5), and nine proteins were predicted to interact with ACA99583.1 (Figure 8, Table 6). All three PsoR homologs were predicted to interact with Gpm, the 2,3-bisphosphoglycerate-independent phosphoglycerate mutase involved in glucose metabolism, and the components of SbcC/D, a heterodimer that cleaves DNA hairpins [57,58]. Both Synechocystis homologs were predicted to interact with PolA, which is DNA polymerase I [59].
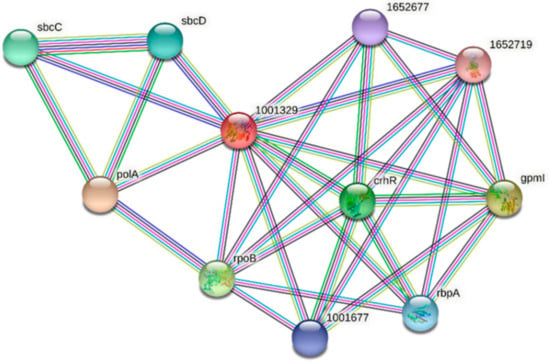
Figure 7.
Protein–protein interaction network for PsoR homolog Sll0514 in Synechocystis sp. PCC 6803. A protein–protein interaction network was generated using STRING [42]. The Sll0514 protein sequence (GenBank ID: sll0514) from Synechocystis sp. PCC 6803 was used as a query. Edges represent functional associations with differently colored lines representing different types of interactions: red line, fusion of genes; green line, neighborhood of genes; blue line, co-occurrence across species; purple line, experimental evidence; yellow-green line, text mining; light blue line, database documentation; black line, co-expression data.

Table 5.
Proteins predicted to interact with Sll0514 from Synechocystis sp. PCC 6803 using the STRING program.
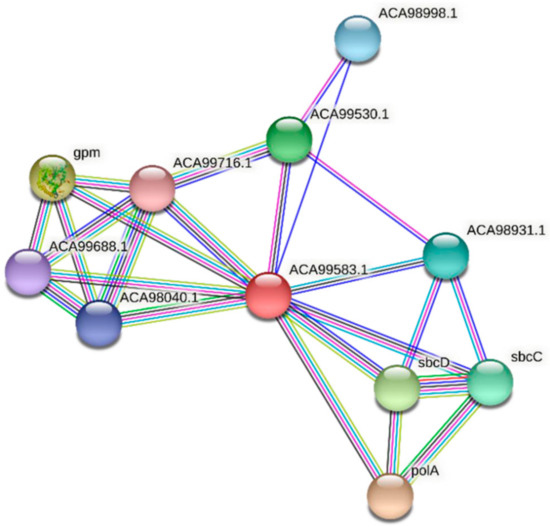
Figure 8.
Protein–protein interaction network for PsoR homolog ACA99583.1 in Synechocystis sp. PCC 7002. A protein–protein interaction network was generated using STRING [42]. The ACA99583.1 protein sequence (GenBank ID: ACA99583.1) from Synechocystis sp. PCC 7002 was used as a query. Edges represent functional associations with differently colored lines representing different types of interactions: red line, fusion of genes; green line, neighborhood of genes; blue line, co-occurrence across species; purple line, experimental evidence; yellow-green line, text mining; light blue line, database documentation; black line, co-expression data.

Table 6.
Proteins predicted to interact with ACA99583.1 from Synechocystis sp. PCC 7002 using the STRING program.
4. Discussion
Considering the critical role of light energy in photosynthesis, mechanisms that allow photosynthetic organisms, such as cyanobacteria, to perceive and respond to fluctuating light environments are vital for organisms to survive and thrive. In cyanobacteria, CCA involves a network of photoreceptors, effectors, and gene operons to finetune pigmentation, cell morphology, filament morphology, and metabolisms in response primarily to RL and GL. The psoR gene has been reported to encode a protein that negatively regulates PBS abundance [36]. Although significant insights have been gained into the regulation of structural genes needed for PBS synthesis, particularly in response to light cues during CA, the regulation of PBS abundance has not been extensively examined in cyanobacteria at the mechanistic level. To understand the roles of PsoR in CCA and cyanobacteria, we assessed its potential structure and function and searched for homologs across the cyanobacterial kingdom to determine when this gene may have arisen in these organisms.
The putative function of PsoR was investigated using the Phyre2 program. The protein encoded by psoR contains a predicted β-CASP domain, including a collection of conserved motifs/residues involved in the processing of DNA and RNA substrates. The β-CASP family of enzymes belongs to the metallo-β-lactamase superfamily, which includes proteins that process a wide variety of substrates in organisms. β-CASP proteins are found in all three domains of life, and they play a predominant role in pre-mRNA processing, although some β-CASP enzymes are involved in DNA repair [44,45,60]. The conserved motifs of PsoR are involved in metal binding, typically zinc ions, and in stabilizing nucleic acid substrates for cleavage. Due to its β-CASP domain and as all prokaryotic β-CASP enzymes identified thus far have been ribonucleases, the role of PsoR in cyanobacteria may be the regulation of mRNA, and it may specifically regulate genes involved in PBS abundance. Other β-CASP proteins tend to function as dimers or within larger protein complexes, usually binding through the C-terminal region of the β-CASP domain or a separate C-terminal region [45,47,48,49]. Given the presence of 123 residue C-terminal regions in FdPsoR, PsoR may also function as a dimer in vivo or may work through interactions with other proteins.
Homologs of PsoR from F. diplosiphon were found across the Cyanobacteria phylum, and only Gloeobacter lacked a homolog. Gloeobacter are an ancient clade of cyanobacteria that lack thylakoid membranes and have atypical PBSs. Instead of rods grouped around a hemisicoidal core, Gloeobacter PBSs are grouped in bundles of parallel rod-shaped structures attached to cytoplasmic membranes [52,53]. Gloeobacter also lacks the genes psbY, psbZ, and psb27, which encode subunits of photosystem II [53,61]. Homologs of PsoR were not found in red algae or other organisms, which suggests that PsoR homologs arose sometime after branching off from Gloeobacter, perhaps around the time that more complex PBSs evolved. Although red algae have PBSs similar to cyanobacteria, the lack of PsoR homologs may indicate a different enzyme or system for regulating PBS abundance in these distinct organisms.
The range of sizes found among PsoR homologs is notable. As the smallest homologs did not form a distinct branch but were found to be intermingled with homologs in sizes closer to FdPsoR, perhaps these shorter homologs are truncated forms of PsoR (Figure 4A). Whether they retain the functionality of PsoR needs to be explored experimentally. The 57 homologs that formed their own branch closest to the Gloeobacter outgroup contained the longest homologs found in our analysis, at as much as 918 bp (Figure 4B). Predominantly of the Oscillatoriales order, they could be extended forms of PsoR.
Three predicted promoter start sites and multiple transcription factor binding sites preceded the psoR gene; yet, one of the two regions also upstream of the neighboring tcpA gene may be a more likely start site of transcription (Figure 5). Previous work established that TcpA, or tetracontapeptide A, is a 40-amino-acid peptide encoded by a gene that appears to be co-transcribed with psoR and may also play a role in PBS abundance regulation [36]. As tcpA was reported to be found upstream of psoR in every cyanobacterial genome searched by Cobley et al. [36], the presence of these two promoter regions provides further evidence that the two genes are co-transcribed in cyanobacteria where they are both found, and they may even work together in some way to regulate PBS abundance in these bacteria.
The predicted protein–protein interactions indicated interactions with protein partners involved in gene and metabolism regulation. Since FdPsoR is a putative β-CASP nuclease, it is possible that FdPsoR interacts with other proteins involved in regulating CCA and metabolism in response to environmental stimuli. Of note, a PC-associated protein was among the proteins predicted to interact with FdPsoR.
Light fluctuations in terrestrial and aquatic environments can occur frequently, in which the light quality and quantity are altered by factors such as cloud coverage and shading by other objects and organisms. The ability to perceive and respond to changing light conditions is important for cyanobacteria to maximize productivity and minimize damage caused by excess light in these dynamic conditions. Due to the important role PBSs play in CA and photosynthesis, understanding how PBSs undergo remodeling or how their abundance is tuned to external cues could extend our understanding of how cyanobacteria survive and thrive in fluctuating light environments. Although significant insights into the light quality-dependent regulation of the pigment content have been obtained during 50 years of research on CCA, many aspects of PBS regulation remain to be elucidated. Here, PsoR has been reported to be widespread across cyanobacteria, where it is likely to play a critical role in cellular PBS regulation. Understanding how PsoR functions as a β-CASP protein and its role in CCA and cyanobacteria can help lead to a better understanding of how photosynthetic organisms fine-tune their photosynthetic machinery in response to changes in their light environment. This knowledge may contribute to improvements in using cyanobacteria for industrial and pharmaceutical purposes.
Author Contributions
Conceptualization, A.L. and B.L.M.; methodology, A.L. and B.L.M.; validation, A.L. and B.L.M.; formal analysis, A.L. and B.L.M.; data curation, A.L.; writing—original draft preparation, A.L. and B.L.M.; writing—review and editing, B.L.M.; visualization, A.L.; supervision, B.L.M.; project administration, B.L.M.; funding acquisition, B.L.M. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the National Science Foundation (MCB-1243983 to B.L.M.), the Office of Science of the U.S. Department of Energy (DE-FG02-91ER20021 to B.L.M.), and the Michigan State University Foundation.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We are grateful to members of the Montgomery Laboratory for assistance with experiments and discussion of the project.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
References
- Zwirglmaier, K.; Jardillier, L.; Ostrowski, M.; Mazard, S.; Garczarek, L.; Vaulot, D.; Not, F.; Massana, R.; Ulloa, O.; Scanlan, D.J. Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environ. Microbiol. 2008, 10, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Zehr, J.P. Nitrogen fixation by marine cyanobacteria. Trends Microbiol. 2011, 19, 162–173. [Google Scholar] [CrossRef] [PubMed]
- Santos-Merino, M.; Singh, A.K.; Ducat, D.C. New applications of synthetic biology tools for cyanobacterial metabolic engineering. Front. Bioeng. Biotechnol. 2019, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Hays, S.G.; Ducat, D.C. Engineering cyanobacteria as photosynthetic feedstock factories. Photosyn. Res. 2015, 123, 285–295. [Google Scholar] [CrossRef]
- Jungblut, A.D.; Hawes, I.; Mountfort, D.; Hitzfeld, B.; Dietrich, D.R.; Burns, B.P.; Neilan, B.A. Diversity within cyanobacterial mat communities in variable salinity meltwater ponds of McMurdo Ice Shelf, Antarctica. Environ. Microbiol. 2005, 7, 519–529. [Google Scholar] [CrossRef]
- Martinez, J.N.; Nishihara, A.; Lichtenberg, M.; Trampe, E.; Kawai, S.; Tank, M.; Kühl, M.; Hanada, S.; Thiel, V. Vertical distribution and diversity of phototrophic bacteria within a hot spring microbial mat (Nakabusa Hot Springs, Japan). Microbes Environ. 2019, 34, 374–387. [Google Scholar] [CrossRef]
- Samolov, E.; Baumann, K.; Büdel, B.; Jung, P.; Leinweber, P.; Mikhailyuk, T.; Karsten, U.; Glaser, K. Biodiversity of algae and cyanobacteria in biological soil crusts collected along a climatic gradient in Chile using an integrative approach. Microorganisms 2020, 8, 1047. [Google Scholar] [CrossRef]
- Montgomery, B.L. Mechanisms and fitness implications of photomorphogenesis during chromatic acclimation in cyanobacteria. J. Exp. Bot. 2016, 67, 4079–4090. [Google Scholar] [CrossRef]
- Ho, M.-Y.; Soulier, N.T.; Canniffe, D.P.; Shen, G.; Bryant, D.A. Light regulation of pigment and photosystem biosynthesis in cyanobacteria. Curr. Opin. Plant Biol. 2017, 37, 24–33. [Google Scholar] [CrossRef]
- Sanfilippo, J.E.; Garczarek, L.; Partensky, F.; Kehoe, D.M. Chromatic acclimation in cyanobacteria: A diverse and widespread process for optimizing photosynthesis. Annu. Rev. Microbiol. 2019, 73, 407–433. [Google Scholar] [CrossRef]
- Gutu, A.; Kehoe, D.M. Emerging perspectives on the mechanisms, regulation, and distribution of light color acclimation in cyanobacteria. Mol. Plant 2012, 5, 1–13. [Google Scholar] [CrossRef]
- Adir, N.; Dines, M.; Klartag, M.; McGregor, A.; Melamed-Frank, M. Assembly and disassembly of phycobilisomes. In Complex Intracellular Structures in Prokaryotes; Shively, J.M., Ed.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 47–77. [Google Scholar] [CrossRef]
- Zhao, K.H.; Su, P.; Bohm, S.; Song, B.; Zhou, M.; Bubenzer, C.; Scheer, H. Reconstitution of phycobilisome core-membrane linker, LCM, by autocatalytic chromophore binding to ApcE. Biochim. Biophys. Acta 2005, 1706, 81–87. [Google Scholar] [CrossRef]
- Bennett, A.; Bogorad, L. Complementary chromatic adaptation in a filamentous blue-green alga. J. Cell Biol. 1973, 58, 419–435. [Google Scholar] [CrossRef]
- Singh, A.K.; McIntyre, L.M.; Sherman, L.A. Microarray analysis of the genome-wide response to iron deficiency and iron reconstitution in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2003, 132, 1825–1839. [Google Scholar] [CrossRef]
- Tamary, E.; Kiss, V.; Nevo, R.; Adam, Z.; Bernát, G.; Rexroth, S.; Rögner, M.; Reich, Z. Structural and functional alterations of cyanobacterial phycobilisomes induced by high-light stress. Biochim. Biophys. Acta 2012, 1817, 319–327. [Google Scholar] [CrossRef]
- Collier, J.L.; Grossman, A.R. A small polypeptide triggers complete degradation of light-harvesting phycobiliproteins in nutrient-deprived cyanobacteria. EMBO J. 1994, 13, 1039–1047. [Google Scholar] [CrossRef]
- Nagarajan, A.; Zhou, M.; Nguyen, A.Y.; Liberton, M.; Kedia, K.; Shi, T.; Piehowski, P.; Shukla, A.; Fillmore, T.L.; Nicora, C.; et al. Proteomic insights into phycobilisome degradation, a selective and tightly controlled process in the fast-growing cyanobacterium Synechococcus elongatus UTEX 2973. Biomolecules 2019, 9, 374. [Google Scholar] [CrossRef]
- Six, C.; Joubin, L.; Partensky, F.; Holtzendorff, J.; Garczarek, L. UV-induced phycobilisome dismantling in the marine picocyanobacterium Synechococcus sp. WH8102. Photosynth. Res. 2007, 92, 75–86. [Google Scholar] [CrossRef]
- Chenu, A.; Keren, N.; Paltiel, Y.; Nevo, R.; Reich, Z.; Cao, J. Light adaptation in phycobilisome antennas: Influence on the rod length and structural arrangement. J. Phys. Chem. B 2017, 121, 9196–9202. [Google Scholar] [CrossRef]
- Kolodny, Y.; Zer, H.; Propper, M.; Yochelis, S.; Paltiel, Y.; Keren, N. Marine cyanobacteria tune energy transfer efficiency in their light-harvesting antennae by modifying pigment coupling. FEBS J. 2021, 288, 980–994. [Google Scholar] [CrossRef]
- Montgomery, B.L. Seeing new light: Recent insights into the occurrence and regulation of chromatic acclimation in cyanobacteria. Curr. Opin. Plant Biol. 2017, 37, 18–23. [Google Scholar] [CrossRef]
- Tandeau de Marsac, N. Occurrence and nature of chromatic adaptation in cyanobacteria. J. Bacteriol. 1977, 130, 82–91. [Google Scholar] [CrossRef]
- Everroad, C.; Six, C.; Partensky, F.; Thomas, J.-C.; Holtzendorff, J.; Wood, A.M. Biochemical bases of type IV chromatic adaptation in marine Synechococcus spp. J. Bacteriol. 2006, 188, 3345–3356. [Google Scholar] [CrossRef]
- Gan, F.; Shen, G.; Bryant, D.A. Occurrence of far-red light photoacclimation (FaRLiP) in diverse cyanobacteria. Life 2014, 5, 4. [Google Scholar] [CrossRef]
- Zhao, C.; Gan, F.; Shen, G.; Bryant, D.A. RfpA, RfpB, and RfpC are the master control elements of Far-Red Light Photoacclimation (FaRLiP). Front. Microbiol. 2015, 6, 1303. [Google Scholar] [CrossRef]
- Kehoe, D.M.; Grossman, A.R. Similarity of a chromatic adaptation sensor to phytochrome and ethylene receptors. Science 1996, 273, 1409–1412. [Google Scholar] [CrossRef]
- Kehoe, D.M.; Grossman, A.R. New classes of mutants in complementary chromatic adaptation provide evidence for a novel four-step phosphorelay system. J. Bacteriol. 1997, 179, 3914–3921. [Google Scholar] [CrossRef]
- Cobley, J.G.; Miranda, R.D. Mutations affecting chromatic adaptation in the cyanobacterium Fremyella diplosiphon. J. Bacteriol. 1983, 153, 1486–1492. [Google Scholar] [CrossRef]
- Terauchi, K.; Montgomery, B.L.; Grossman, A.R.; Lagarias, J.C.; Kehoe, D.M. RcaE is a complementary chromatic adaptation photoreceptor required for green and red light responsiveness. Mol. Microbiol. 2004, 51, 567–577. [Google Scholar] [CrossRef]
- Li, L.; Alvey, R.M.; Bezy, R.P.; Kehoe, D.M. Inverse transcriptional activities during complementary chromatic adaptation are controlled by the response regulator RcaC binding to red and green light-responsive promoters. Mol. Microbiol. 2008, 68, 286–297. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Kehoe, D.M. In vivo analysis of the roles of conserved aspartate and histidine residues within a complex response regulator. Mol. Microbiol. 2005, 55, 1538–1552. [Google Scholar] [CrossRef]
- Bezy, R.P.; Kehoe, D.M. Functional characterization of a cyanobacterial OmpR/PhoB class transcription factor binding site controlling light color responses. J. Bacteriol. 2010, 192, 5923–5933. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alvey, R.M.; Bezy, R.P.; Frankenberg-Dinkel, N.; Kehoe, D.M. A light regulated OmpR-class promoter element co-ordinates light-harvesting protein and chromophore biosynthetic enzyme gene expression. Mol. Microbiol. 2007, 64, 319–332. [Google Scholar] [CrossRef] [PubMed]
- Bezy, R.P.; Wiltbank, L.; Kehoe, D.M. Light-dependent attenuation of phycoerythrin gene expression reveals convergent evolution of green light sensing in cyanobacteria. Proc. Natl. Acad. Sci. USA 2011, 108, 18542–18547. [Google Scholar] [CrossRef]
- Cobley, J.; Khan, S.; Ahmad, H.; Bailey, S. The gene, tcpA, required for photoregulation of phycobilisome abundance and for heterotrophic growth in Fremyella diplosiphon, is a useful phylogenetic marker specific for the phylum, Cyanobacteria. In Proceedings of the 7th European Workshop on Molecular Biology of Cyanobacteria, České Budějovice, Czech Republic, 31 August–4 September 2008; p. 101. [Google Scholar]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Solovyev, V.; Salamov, A. Automatic annotation of microbial genomes and metagenomic sequences. In Metagenomics and Its Applications in Agriculture; Li, R.W., Ed.; Nova Science Publishers: Hauppauge, NY, USA, 2011; pp. 61–78. [Google Scholar]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Daiyasu, H.; Osaka, K.; Ishino, Y.; Toh, H. Expansion of the zinc metallo-hydrolase family of the beta-lactamase fold. FEBS Lett. 2001, 503, 1–6. [Google Scholar] [CrossRef]
- Callebaut, I.; Moshous, D.; Mornon, J.P.; de Villartay, J.P. Metallo-beta-lactamase fold within nucleic acids processing enzymes: The beta-CASP family. Nucleic Acids Res. 2002, 30, 3592–3601. [Google Scholar] [CrossRef] [PubMed]
- Dominski, Z.; Carpousis, A.J.; Clouet-d’Orval, B. Emergence of the β-CASP ribonucleases: Highly conserved and ubiquitous metallo-enzymes involved in messenger RNA maturation and degradation. Biochim. Biophys. Acta 2013, 1829, 532–551. [Google Scholar] [CrossRef] [PubMed]
- Mandel, C.R.; Kaneko, S.; Zhang, H.; Gebauer, D.; Vethantham, V.; Manley, J.L.; Tong, L. Polyadenylation factor CPSF-73 is the pre-mRNA 3′-end-processing endonuclease. Nature 2006, 444, 953–956. [Google Scholar] [CrossRef] [PubMed]
- Silva, A.P.; Chechik, M.; Byrne, R.T.; Waterman, D.G.; Ng, C.L.; Dodson, E.J.; Koonin, E.V.; Antson, A.A.; Smits, C. Structure and activity of a novel archaeal β-CASP protein with N-terminal KH domains. Structure 2011, 19, 622–632. [Google Scholar] [CrossRef]
- Mathy, N.; Hébert, A.; Mervelet, P.; Bénard, L.; Dorléans, A.; Li de la Sierra-Gallay, I.; Noirot, P.; Putzer, H.; Condon, C. Bacillus subtilis ribonucleases J1 and J2 form a complex with altered enzyme behaviour. Mol. Microbiol. 2010, 75, 489–498. [Google Scholar] [CrossRef]
- Phung, D.K.; Rinaldi, D.; Langendijk-Genevaux, P.S.; Quentin, Y.; Carpousis, A.J.; Clouet-d’Orval, B. Archaeal β-CASP ribonucleases of the aCPSF1 family are orthologs of the eukaryal CPSF-73 factor. Nucleic Acids Res. 2013, 41, 1091–1103. [Google Scholar] [CrossRef]
- States, D.J.; Gish, W. Combined use of sequence similarity and codon bias for coding region identification. J. Comput. Biol. 1994, 1, 39–50. [Google Scholar] [CrossRef]
- Ryan, K.; Calvo, O.; Manley, J.L. Evidence that polyadenylation factor CPSF-73 is the mRNA 3′ processing endonuclease. RNA 2004, 10, 565–573. [Google Scholar] [CrossRef]
- Krogmann, D.W.; Pérez-Gómez, B.; Gutiérrez-Cirlos, E.B.; Chagolla-López, A.; González de la Vara, L.; Gómez-Lojero, C. The presence of multidomain linkers determines the bundle-shape structure of the phycobilisome of the cyanobacterium Gloeobacter violaceus PCC 7421. Photosynth. Res. 2007, 93, 27–43. [Google Scholar] [CrossRef]
- Guglielmi, G.; Cohen-Bazire, G.; Bryant, D.A. The structure of Gloeobacter violaceus and its phycobilisomes. Arch. Microbiol. 1981, 129, 181–189. [Google Scholar] [CrossRef]
- Shih, P.M.; Wu, D.; Latifi, A.; Axen, S.D.; Fewer, D.P.; Talla, E.; Calteau, A.; Cai, F.; Tandeau de Marsac, N.; Rippka, R.; et al. Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 1053–1058. [Google Scholar] [CrossRef]
- Monteiro, L.M.O.; Sanches-Medeiros, A.; Westmann, C.A.; Silva-Rocha, R. Unraveling the complex interplay of Fis and IHF through synthetic promoter engineering. Front. Bioeng. Biotechnol. 2020, 8, 510. [Google Scholar] [CrossRef]
- Takashima, K.; Nagao, S.; Kizawa, A.; Suzuki, T.; Dohmae, N.; Hihara, Y. The role of transcriptional repressor activity of LexA in salt-stress responses of the cyanobacterium Synechocystis sp. PCC 6803. Sci. Rep. 2020, 10, 17393. [Google Scholar] [CrossRef]
- Nukui, M.; Mello, L.V.; Littlejohn, J.E.; Setlow, B.; Setlow, P.; Kim, K.; Leighton, T.; Jedrzejas, M.J. Structure and molecular mechanism of Bacillus anthracis cofactor-independent phosphoglycerate mutase: A crucial enzyme for spores and growing cells of Bacillus species. Biophys. J. 2007, 92, 977–988. [Google Scholar] [CrossRef]
- Connelly, J.C.; Kirkham, L.A.; Leach, D.R. The SbcCD nuclease of Escherichia coli is a structural maintenance of chromosomes (SMC) family protein that cleaves hairpin DNA. Proc. Natl. Acad. Sci. USA 1998, 95, 7969–7974. [Google Scholar] [CrossRef]
- Patel, P.H.; Suzuki, M.; Adman, E.; Shinkai, A.; Loeb, L.A. Prokaryotic DNA polymerase I: Evolution, structure, and “base flipping” mechanism for nucleotide selection. J. Mol. Biol. 2001, 308, 823–837. [Google Scholar] [CrossRef]
- Condon, C.; Gilet, L. The metallo-β-lactamase family of ribonucleases. In Ribonucleases; Nicholson, A.W., Ed.; Springer: Berlin, Germany, 2011; pp. 245–267. [Google Scholar] [CrossRef]
- Yokono, M.; Akimoto, S.; Koyama, K.; Tsuchiya, T.; Mimuro, M. Energy transfer processes in Gloeobacter violaceus PCC 7421 that possesses phycobilisomes with a unique morphology. Biochim. Biophys. Acta 2008, 1777, 55–65. [Google Scholar] [CrossRef][Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).