Emerging Artificial Intelligence (AI) Technologies Used in the Development of Solid Dosage Forms
Abstract
1. Introduction
2. Commonly Used Databases
3. Data Processing Methods
3.1. Tabular Data Processing
3.2. Molecular Representation Methods for APIs and Excipients
4. Overview of AI Algorithms in Solid Dosage Forms Development
Advantages and Disadvantages of Different Algorithms
5. Model Predictive Performance Evaluation and Explainability
6. Applications of AI in Solid Dosage Forms
6.1. Overview of Solid Dosage Formulations Designed by AI
6.2. Tablets
6.2.1. Predicting Drug Release
6.2.2. Developing 3D-Printed Tablets Using AI
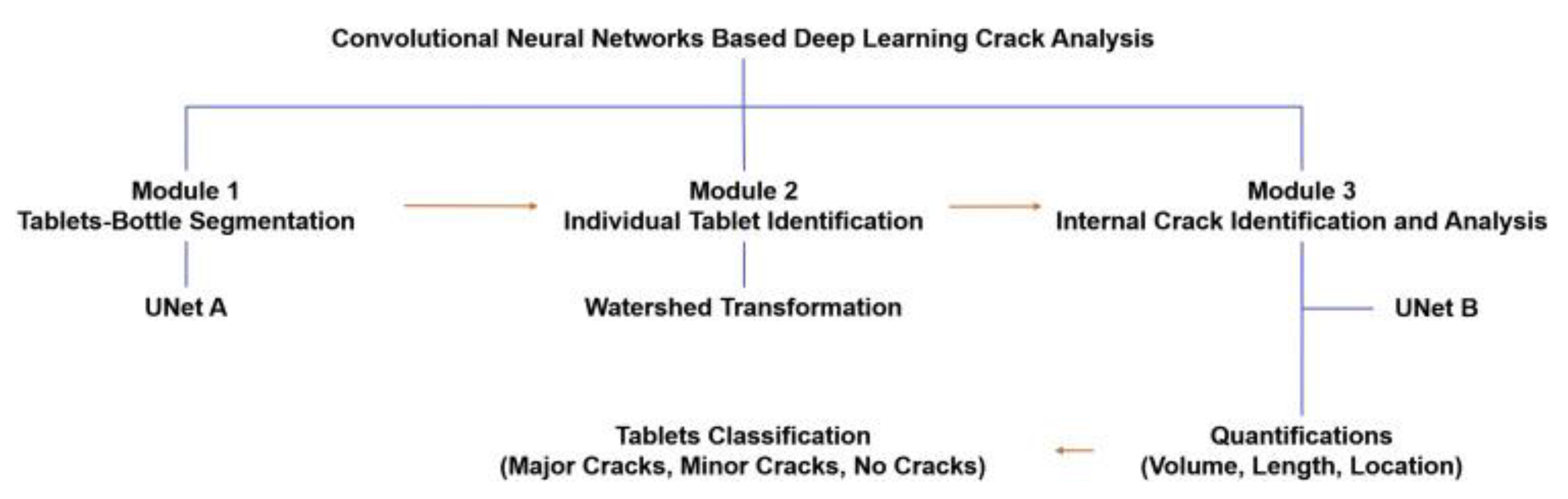
6.2.3. Detecting Tablet Defects
6.3. Powders
6.3.1. Applications of AI in Process Control during Powder Engineering
6.3.2. Applications of AI in Designing Dry Powder for Inhalation
6.4. Capsules
6.5. Granules
6.6. Solid Dispersions
6.6.1. Predicting Physical or Chemical Stability
6.6.2. Predicting Dissolution Rates and Profiles
6.7. AI Applications in Pharmaceutical Image Analysis
6.7.1. Image Pre-Processing Methods
6.7.2. Case Studies of AI-Based Image Analysis
7. Prospects
8. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Davies, P. Oral Solid Dosage Forms. In Pharmaceutical Preformulation and Formulation; CRC Press: Boca Raton, FL, USA, 2016; pp. 379–442. [Google Scholar] [CrossRef]
- Shaikh, R.; O’Brien, D.P.; Croker, D.M.; Walker, G.M. The development of a pharmaceutical oral solid dosage forms. Comput. Aided Chem. Eng. 2018, 41, 27–65. [Google Scholar] [CrossRef]
- Chow, K.; Tong, H.H.Y.; Lum, S.; Chow, A.H.L. Engineering of Pharmaceutical Materials: An Industrial Perspective. J. Pharm. Sci. 2008, 97, 2855–2877. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; Chen, Y.; Zhang, G.; Yu, L.; Mantri, R. Developing Solid Oral Dosage Forms: Pharmaceutical Theory and Practice. 2016. Available online: https://books.google.com/books?hl=en&lr=&id=lk1ODAAAQBAJ&oi=fnd&pg=PP1&dq=Developing+Solid+Oral+Dosage+Forms+Pharmaceutical+Theory+and+Practice&ots=fer2FYISJi&sig=iQQMeuSM5xOpk39zMzRuHulN95k (accessed on 5 August 2022).
- Challenges and Opportunities in Oral Formulation Development-Google Scholar. Available online: https://scholar.google.com/scholar?hl=en&as_sdt=0%2C39&inst=9599013809589351610&q=Challenges+and+Opportunities+in+Oral+Formulation+Development&btnG= (accessed on 5 August 2022).
- Loftsson, T.; Brewster, M.E. Pharmaceutical Applications of Cyclodextrins: Basic Science and Product Development. J. Pharm. Pharmacol. 2010, 62, 1607–1621. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lubricants, Y.W. Lubricants in pharmaceutical solid dosage forms. Lubricants 2014, 2, 21–43. [Google Scholar] [CrossRef]
- Benet, L.Z.; Goyan, J.E. Bioequivalence and Narrow Therapeutic Index Drugs. Pharmacother. J. Hum. Pharmacol. Drug Ther. 1995, 15, 433–440. [Google Scholar] [CrossRef]
- Surasarang, S.H.; Keen, J.M.; Huang, S.; Zhang, F.; McGinity, J.W.; Williams, R.O., III. Hot melt extrusion versus spray drying: Hot melt extrusion degrades albendazole. Taylor Fr. 2016, 43, 797–811. [Google Scholar] [CrossRef] [PubMed]
- Bannigan, P.; Aldeghi, M.; Bao, Z.; Häse, F.; Aspuru-Guzik, A.; Allen, C. Machine learning directed drug formulation development. Adv. Drug Deliv. Rev. 2021, 175, 113806. [Google Scholar] [CrossRef] [PubMed]
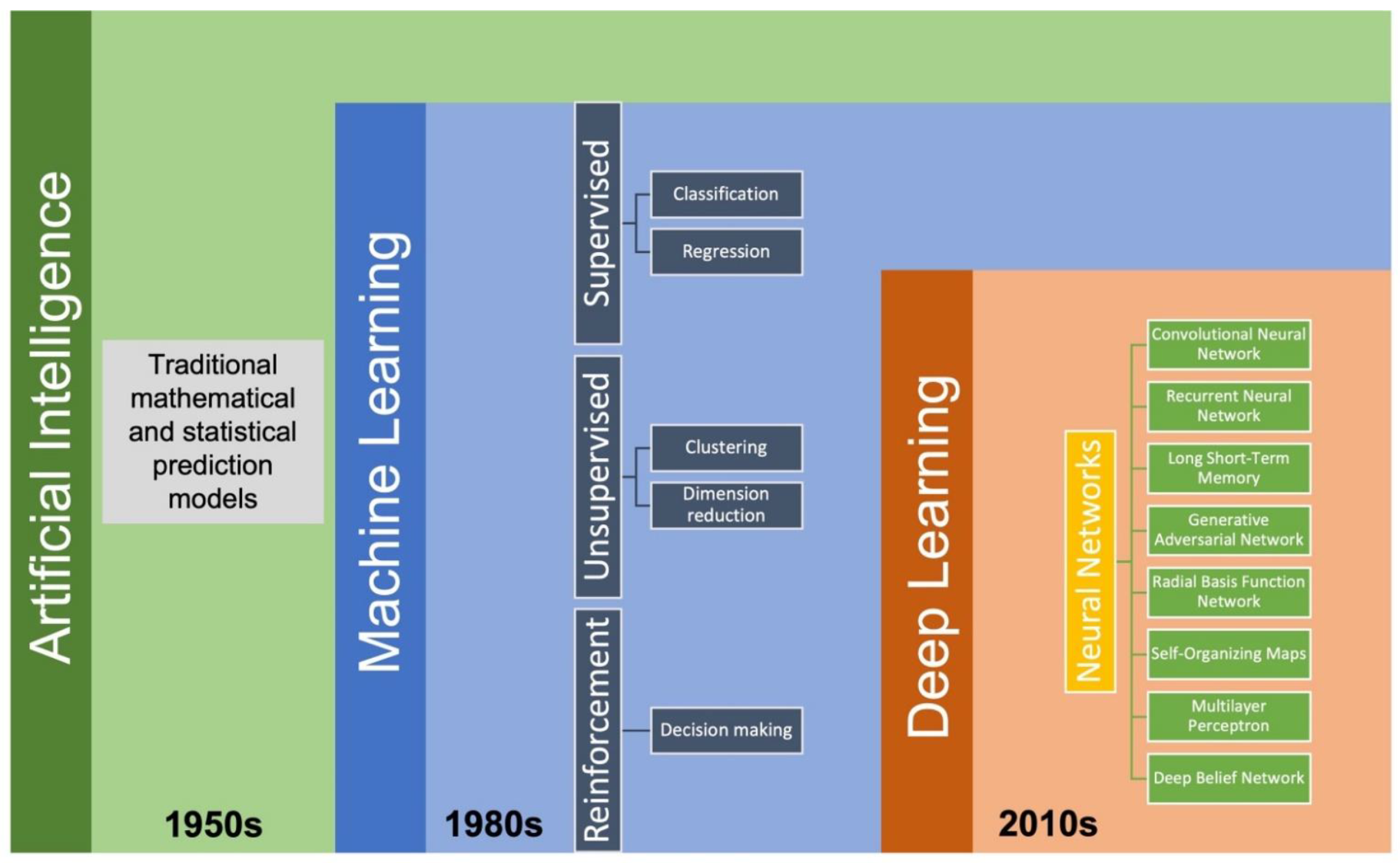
- McCarthy, J.; Minsky, M.; Rochester, N.; Magazine, C.S.A. 2006 Undefined. A Proposal for the Dartmouth Summer Research Project on Artificial Intelligence, August 31, 1955. 2006. Available online: https://ojs.aaai.org/index.php/aimagazine/article/view/1904 (accessed on 4 August 2022).
- 4 Basic Steps in Implementing an AI-Driven Design Workflow-EDN. Available online: https://www.edn.com/four-basic-steps-in-implementing-an-ai-driven-design-workflow/ (accessed on 4 October 2021).
- Machine Learning-Google Books. Available online: https://www.google.com/books/edition/Machine_Learning/ylE4DQAAQBAJ?hl=en&gbpv=1&dq=AI+machine+learning&pg=PR5&printsec=frontcover (accessed on 5 August 2022).
- Zain Amin, M.; Ali, A. Performance Evaluation of Supervised Machine Learning Classifiers for Predicting Healthcare Operational Decisions; Technical Report; University of California: Irvine, CA, USA, 2017. [Google Scholar] [CrossRef]
- Berk, R. An impact assessment of machine learning risk forecasts on parole board decisions and recidivism. J. Exp. Criminol. 2017, 13, 193–216. [Google Scholar] [CrossRef]
- Berk, R.A.; Sorenson, S.B.; Barnes, G. Forecasting Domestic Violence: A Machine Learning Approach to Help Inform Arraignment Decisions. J. Empir. Leg. Stud. 2016, 13, 94–115. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y. Deep learning. Nature 2015, 521, 436–444. Available online: https://idp.nature.com/authorize/casa?redirect_uri=https://www.nature.com/articles/nature14539&casa_token=ytaPO_BVoo0AAAAA:1bw1c5ZJZYvzg8zP0G_iOcKro4uMBvBuY6ZnZM8sUXo4RAxznZrDmU4TR0-3rv-wIBWs6GLIefCxfKo (accessed on 5 August 2022). [CrossRef] [PubMed]
- Affonso, C.; Rossi, A.L.D.; Vieira, F.H.A.; de Carvalho, A.C.P.d.L.F. Deep Learning for Biological Image Classification. Expert Syst. Appl. 2017, 85, 114–122. [Google Scholar] [CrossRef]
- Liu, L.; Ouyang, W.; Wang, X.; Fieguth, P.; Chen, J.; Liu, X.; Pietikäinen, M. Deep Learning for Generic Object Detection: A Survey. Int. J. Comput. Vis. 2020, 128, 261–318. [Google Scholar] [CrossRef]
- Garcia-Garcia, A.; Orts-Escolano, S.; Oprea, S.; Villena-Martinez, V.; Garcia-Rodriguez, J. A review on deep learning techniques applied to semantic segmentation. arXiv 2017, arXiv:1704.06857. [Google Scholar]
- Xu, Z.; Sun, J. Model-driven deep-learning. Natl. Sci. Rev. 2018, 5, 22–24. [Google Scholar] [CrossRef]
- Chan, H.P.; Samala, R.K.; Hadjiiski, L.M.; Zhou, C. Deep Learning in Medical Image Analysis. Adv. Exp. Med. Biol. 2020, 1213, 3–21. [Google Scholar] [CrossRef] [PubMed]
- Steinwandter, V.; Borchert, D.; Herwig, C. Data science tools and applications on the way to Pharma 4.0. Drug Discov. Today 2019, 24, 1795–1805. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Ye, Z.; Gao, H.; Ouyang, D. Computational pharmaceutics-A new paradigm of drug delivery. J. Control. Release 2021, 338, 119–136. [Google Scholar] [CrossRef] [PubMed]
- AI in Pharma Global Market Report. 2022. Available online: https://www.prnewswire.com/news-releases/ai-in-pharma-global-market-report-2022-301542906.html (accessed on 5 August 2022).
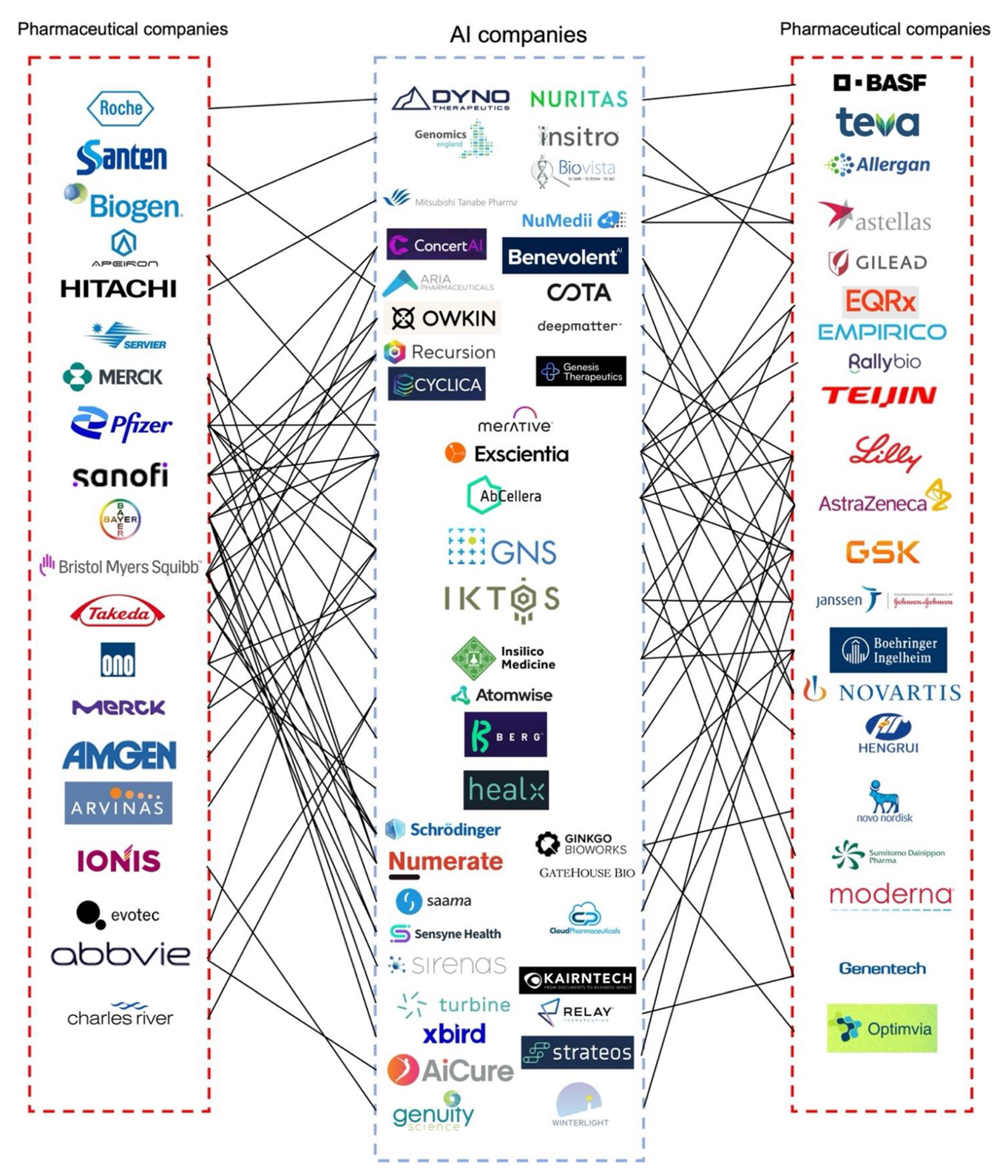
- Mak, K.K.; Pichika, M.R. Artificial intelligence in drug development: Present status and future prospects. Drug Discov. Today 2019, 24, 773–780. [Google Scholar] [CrossRef] [PubMed]
- MLPDS–Machine Learning for Pharmaceutical Discovery and Synthesis Consortium. Available online: https://mlpds.mit.edu/ (accessed on 18 September 2022).
- AstraZeneca Links with Alibaba and Tencent in China Push | Reuters. Available online: https://www.reuters.com/article/us-astrazeneca-china/astrazeneca-links-with-alibaba-and-tencent-in-china-push-idUSKBN1FM1FM (accessed on 17 October 2022).
- Proposed Regulatory Framework for Modifications to Artificial Intelligence/Machine Learning (AI/ML)-Based Software as a Medical Device (SaMD)-Discussion Paper and Request for Feedback. Available online: https://www.fda.gov/downloads/medicaldevices/deviceregulationandguidance/guidancedocuments/ucm514737.pdf (accessed on 1 August 2022).
- Zhao, L.; Kim, M.J.; Zhang, L.; Lionberger, R. Generating Model Integrated Evidence for Generic Drug Development and Assessment. Clin. Pharmacol. Ther. 2019, 105, 338–349. [Google Scholar] [CrossRef] [PubMed]
- Marshall, S.; Madabushi, R.; Manolis, E.; Krudys, K.; Staab, A.; Dykstra, K.; Visser, S.A. Model-Informed Drug Discovery and Development: Current Industry Good Practice and Regulatory Expectations and Future Perspectives. CPT Pharmacomet. Syst. Pharmacol. 2019, 8, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.X.; Raw, A.; Wu, L.; Capacci-Daniel, C.; Zhang, Y.; Rosencrance, S. FDA’s new pharmaceutical quality initiative: Knowledge-aided assessment & structured applications. Int. J. Pharm. X 2019, 1, 100010. [Google Scholar] [CrossRef] [PubMed]
- Solid Dose: Under-Hyped but Not Under-Represented. Available online: https://www.pharmamanufacturing.com/articles/2019/solid-dose-under-hyped-but-not-under-represented/ (accessed on 4 October 2021).
- Lou, H.; Lian, B.; Hageman, M.J. Applications of Machine Learning in Solid Oral Dosage Form Development. J. Pharm. Sci. 2021, 110, 3150–3165. [Google Scholar] [CrossRef]
- Hicks, C.R. Fundamental Concepts in the Design of Experiments. Available online: https://philpapers.org/rec/HICFCI (accessed on 18 September 2022).
- U.S. Pharmacopeia. Available online: https://www.usp.org/ (accessed on 6 October 2021).
- Kim, S. Getting the Most out of PubChem for Virtual Screening. Expert Opin. Drug Discov. 2016, 11, 843. [Google Scholar] [CrossRef]
- The Cambridge Structural Database (CSD)—The Cambridge Crystallographic Data Centre (CCDC). Available online: https://www.ccdc.cam.ac.uk/solutions/csd-core/components/csd/ (accessed on 6 October 2021).
- Gabrielson, S.W. SciFinder. J. Med. Libr. Assoc. 2018, 106, 588. [Google Scholar] [CrossRef]
- The Merck Index Online-Chemicals, Drugs and Biologicals. Available online: https://www.rsc.org/merck-index (accessed on 6 October 2021).
- Inactive Ingredient Search for Approved Drug Products. Available online: https://www.accessdata.fda.gov/scripts/cder/iig/index.cfm (accessed on 17 October 2022).
- Drugs@FDA: FDA-Approved Drugs. Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm (accessed on 17 October 2022).
- Orange Book: Approved Drug Products with Therapeutic Equivalence Evaluations. Available online: https://www.fda.gov/drugs/drug-approvals-and-databases/approved-drug-products-therapeutic-equivalence-evaluations-orange-book (accessed on 17 October 2022).
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Dissolution Methods. Available online: https://www.accessdata.fda.gov/scripts/cder/dissolution/dsp_getallData.cfm (accessed on 6 October 2021).
- MedlinePlus-Health Information from the National Library of Medicine. Available online: https://medlineplus.gov/ (accessed on 6 October 2021).
- Drug Information Portal-U.S. National Library of Medicine-Quick Access to Quality Drug Information. Available online: https://druginfo.nlm.nih.gov/drugportal/jsp/drugportal/about.jsp (accessed on 6 October 2021).
- Liu, H.; Shah, S.; Jiang, W. On-line outlier detection and data cleaning. Comput. Chem. Eng. 2004, 28, 1635–1647. [Google Scholar] [CrossRef]
- Zhu, J.; Ge, Z.; Song, Z.; Gao, F. Review and big data perspectives on robust data mining approaches for industrial process modeling with outliers and missing data. Annu. Rev. Control. 2018, 46, 107–133. [Google Scholar] [CrossRef]
- Palo, H.K.; Sahoo, S.; Subudhi, A.K. Dimensionality Reduction Techniques: Principles, Benefits, and Limitations. Data Anal. Bioinform. Mach. Learn. Perspect. 2021, 77–107. [Google Scholar] [CrossRef]
- Abd Elrahman, S.M.; Abraham, A. A Review of Class Imbalance Problem. J. Netw. Innov. Comput. 2013, 1, 332–340. Available online: https://www.mirlabs.net/jnic/index.html (accessed on 19 September 2022).
- Lee, H.; Kim, J.; Kim, S.; Yoo, J.; Choi, G.J.; Jeong, Y.S. Deep Learning-Based Prediction of Physical Stability considering Class Imbalance for Amorphous Solid Dispersions. J. Chem. 2022, 2022. [Google Scholar] [CrossRef]
- Jeni, L.A.; Cohn, J.F.; de La Torre, F. Facing imbalanced data-Recommendations for the use of performance metrics. In Proceedings of the 2013 Humaine Association Conference on Affective Computing and Intelligent Interaction, ACII 2013, Geneva, Switzerland, 2–5 September 2013; pp. 245–251. [Google Scholar] [CrossRef]
- Raghunathan, S.; Priyakumar, U.D. Molecular representations for machine learning applications in chemistry. Int. J. Quantum Chem. 2022, 122, e26870. [Google Scholar] [CrossRef]
- Wigh, D.S.; Goodman, J.M.; Lapkin, A.A. A review of molecular representation in the age of machine learning. WIREs Comput. Mol. Sci. 2022, e1603. [Google Scholar] [CrossRef]
- Dong, J.; Gao, H.; Ouyang, D. PharmSD: A novel AI-based computational platform for solid dispersion formulation design. Int. J. Pharm. 2021, 604, 120705. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Liu, Y.; Cheng, J.; Li, Y.; Liu, S.; Duan, Y.; Zhang, L.; Luo, S. An Ensemble Structure and Physicochemical (SPOC) Descriptor for Machine-Learning Prediction of Chemical Reaction and Molecular Properties. ChemPhysChem 2022, 23, e202200255. [Google Scholar] [CrossRef]
- RDKit. Available online: https://www.rdkit.org/ (accessed on 12 July 2022).
- Han, R.; Xiong, H.; Ye, Z.; Yang, Y.; Huang, T.; Jing, Q.; Lu, J.; Pan, H.; Ren, F.; Ouyang, D. Predicting physical stability of solid dispersions by machine learning techniques. J. Control. Release 2019, 311–312, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Kazemi, P.; Khalid, M.H.; Szlek, J.; Mirtič, A.; Reynolds, G.; Jachowicz, R.; Mendyk, A. Computational intelligence modeling of granule size distribution for oscillating milling. Powder Technol. 2016, 301, 1252–1258. [Google Scholar] [CrossRef]
- Ye, Z.; Yang, W.; Yang, Y.; Ouyang, D. Interpretable machine learning methods for in vitro pharmaceutical formulation development. Food Front. 2021, 2, 195–207. [Google Scholar] [CrossRef]
- Commonly Used Machine Learning Algorithms | Data Science. Available online: https://www.analyticsvidhya.com/blog/2017/09/common-machine-learning-algorithms/ (accessed on 4 October 2021).
- Deep Learning vs. Machine Learning—What’s the Difference? | Flatiron School. Available online: https://flatironschool.com/blog/deep-learning-vs-machine-learning (accessed on 5 October 2021).
- Ma, X.; Kittikunakorn, N.; Sorman, B.; Xi, H.; Chen, A.; Marsh, M.; Mongeau, A.; Piché, N.; Williams, R.O.; Skomski, D. Application of Deep Learning Convolutional Neural Networks for Internal Tablet Defect Detection: High Accuracy, Throughput, and Adaptability. J. Pharm. Sci. 2020, 109, 1547–1557. [Google Scholar] [CrossRef]
- Westphal, E.; Seitz, H. A machine learning method for defect detection and visualization in selective laser sintering based on convolutional neural networks. Addit. Manuf. 2021, 41, 101965. [Google Scholar] [CrossRef]
- Hesse, R.; Krull, F.; Antonyuk, S. Prediction of Random Packing Density and Flowability for Non-Spherical Particles by Deep Convolutional Neural Networks and Discrete Element Method Simulations. Powder Technol. 2021, 393, 559–581. [Google Scholar] [CrossRef]
- Goh, W.Y.; Lim, C.P.; Peh, K.K.; Subari, K. Application of a recurrent neural network to prediction of drug dissolution profiles. Neural Comput. Appl. 2002, 10, 311–317. [Google Scholar] [CrossRef]
- Top 8 Programming Languages for Artificial Intelligence Projects | Ksolves. Available online: https://www.ksolves.com/blog/artificial-intelligence/top-8-programming-languages-for-artificial-intelligence-projects (accessed on 5 October 2021).
- 10 Best Artificial Intelligence Software (AI Software Reviews in 2021). Available online: https://www.softwaretestinghelp.com/artificial-intelligence-software/ (accessed on 5 October 2021).
- Ieracitano, C.; Pantó, F.; Mammone, N.; Paviglianiti, A.; Frontera, P.; Morabito, F.C. Toward an Automatic Classification of SEM Images of Nanomaterials via a Deep Learning Approach. Smart Innov. Syst. Technol. 2020, 151, 61–72. [Google Scholar] [CrossRef]
- Castro, B.M.; Elbadawi, M.; Ong, J.J.; Pollard, T.; Song, Z.; Gaisford, S.; Pérez, G.; Basit, A.W.; Cabalar, P.; Goyanes, A. Machine learning predicts 3D printing performance of over 900 drug delivery systems. J. Control. Release 2021, 337, 530–545. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Li, L.; Zhang, S.; Lomeo, J.; Zhu, A.; Chen, J.; Barrett, S.; Koynov, A.; Forster, S.; Wuelfing, P.; et al. Correlative Image-Based Release Prediction and 3D Microstructure Characterization for a Long Acting Parenteral Implant. Pharm. Res. 2021, 38, 1915–1929. [Google Scholar] [CrossRef] [PubMed]
- Padilla, R.; Passos, W.L.; Dias, T.L.B.; Netto, S.L.; da Silva, E.A.B. A Comparative Analysis of Object Detection Metrics with a Companion Open-Source Toolkit. Electronics 2021, 10, 279. [Google Scholar] [CrossRef]
- Taha, A.A.; Hanbury, A. Metrics for evaluating 3D medical image segmentation: Analysis, selection, and tool. BMC Med. Imaging 2015, 15, 29. [Google Scholar] [CrossRef]
- Casalicchio, G.; Molnar, C.; Bischl, B. Visualizing the Feature Importance for Black Box Models. In Joint European Conference on Machine Learning and Knowledge Discovery in Databases; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2018; Volume 11051, pp. 655–670. [Google Scholar] [CrossRef]
- Huynh-Thu, V.A.; Saeys, Y.; Wehenkel, L.; Geurts, P. Statistical interpretation of machine learning-based feature importance scores for biomarker discovery. Bioinformatics 2012, 28, 1766–1774. [Google Scholar] [CrossRef] [PubMed]
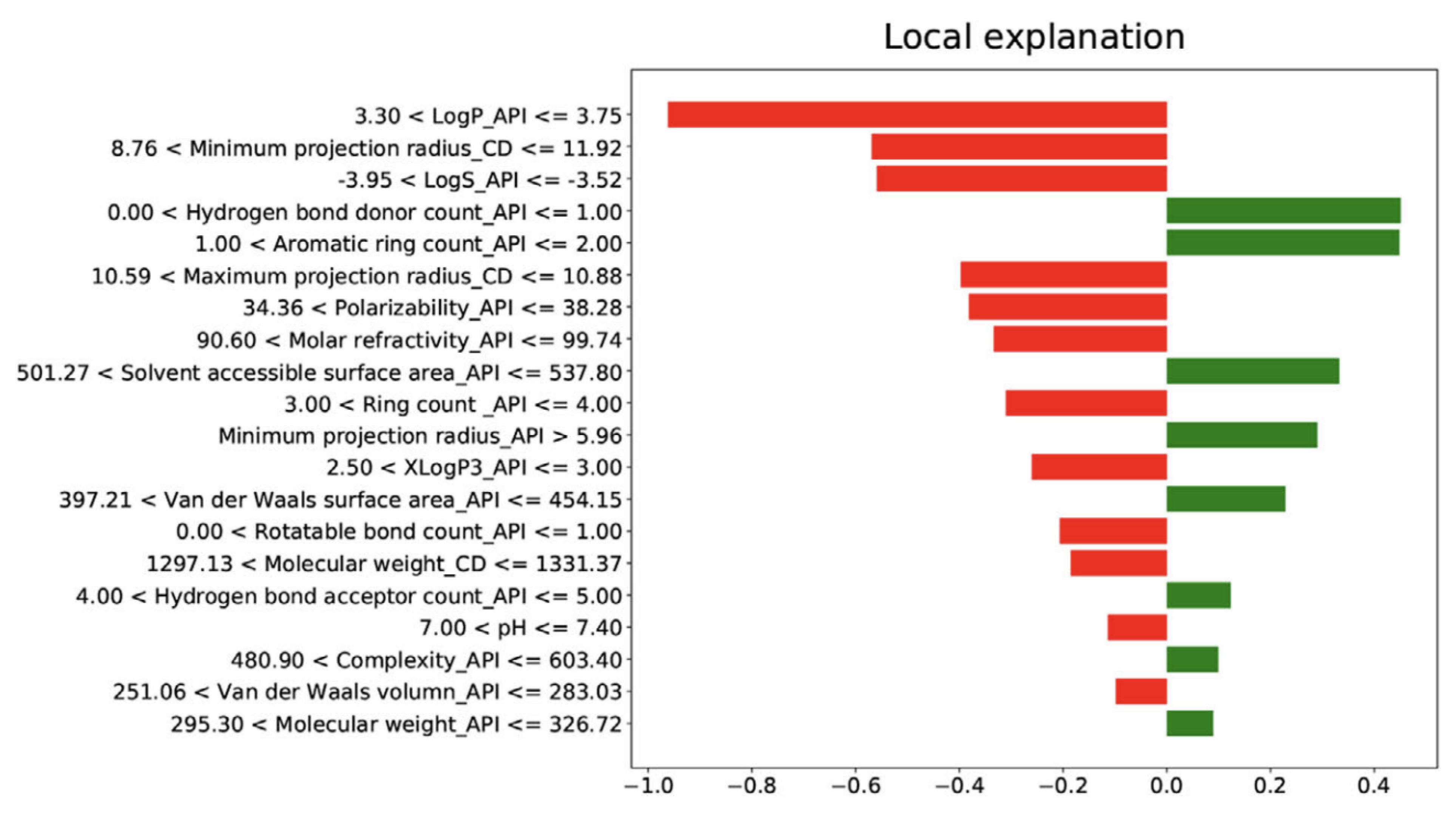
- LIME-Local Interpretable Model-Agnostic Explanations—Marco Tulio Ribeiro. Available online: https://homes.cs.washington.edu/~marcotcr/blog/lime/ (accessed on 18 July 2022).
- Welcome to the SHAP Documentation—SHAP Latest Documentation. Available online: https://shap.readthedocs.io/en/latest/index.html (accessed on 17 March 2022).
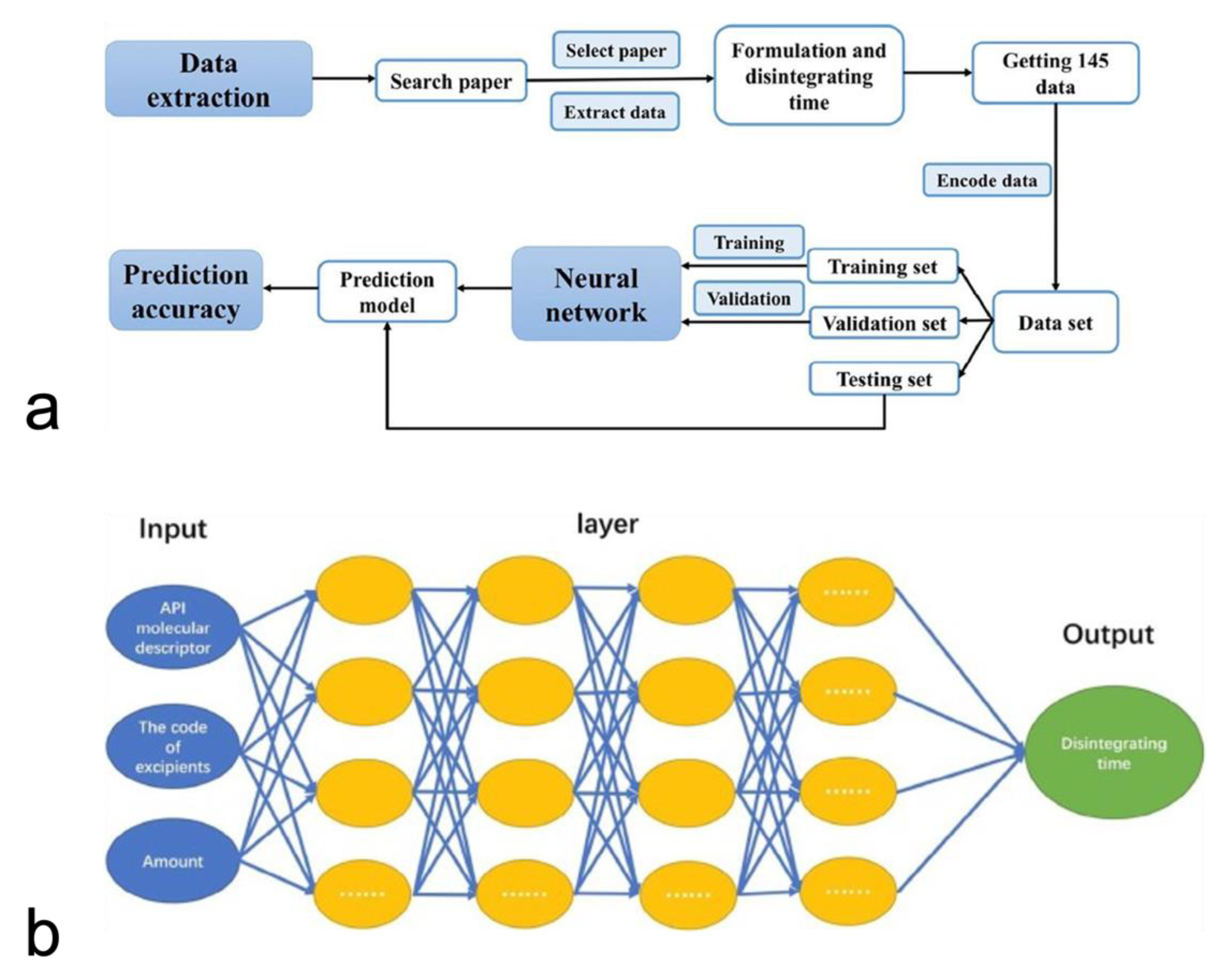
- Szlęk, J.; Khalid, M.H.; Pacławski, A.; Czub, N.; Mendyk, A. Puzzle out Machine Learning Model-Explaining Disintegration Process in ODTs. Pharmaceutics 2022, 14, 859. [Google Scholar] [CrossRef] [PubMed]
- Galata, D.L.; Könyves, Z.; Nagy, B.; Novák, M.; Mészáros, L.A.; Szabó, E.; Farkas, A.; Marosi, G.; Nagy, Z.K. Real-time release testing of dissolution based on surrogate models developed by machine learning algorithms using NIR spectra, compression force and particle size distribution as input data. Int. J. Pharm. 2021, 597, 120338. [Google Scholar] [CrossRef]
- Salem, S.; Byrn, S.R.; Smith, D.T.; Gurvich, V.J.; Hoag, S.W.; Zhang, F.; Williams, R.O.; Clase, K.L. Impact Assessment of the Variables Affecting the Drug Release and Extraction of Polyethylene Oxide Based Tablets. J. Drug Deliv. Sci. Technol. 2022, 71, 103337. [Google Scholar] [CrossRef]
- Obeid, S.; Madžarević, M.; Krkobabić, M.; Ibrić, S. Predicting drug release from diazepam FDM printed tablets using deep learning approach: Influence of process parameters and tablet surface/volume ratio. Int. J. Pharm. 2021, 601, 120507. [Google Scholar] [CrossRef] [PubMed]
- Ficzere, M.; Mészáros, L.A.; Kállai-Szabó, N.; Kovács, A.; Antal, I.; Nagy, Z.K.; Galata, D.L. Real-time coating thickness measurement and defect recognition of film coated tablets with machine vision and deep learning. Int. J. Pharm. 2022, 623, 121957. [Google Scholar] [CrossRef]
- Floryanzia, S.; Ramesh, P.; Mills, M.; Kulkarni, S.; Chen, G.; Shah, P.; Lavrich, D. Disintegration testing augmented by computer Vision technology. Int. J. Pharm. 2022, 619, 121668. [Google Scholar] [CrossRef]
- Mészáros, L.A.; Farkas, A.; Madarász, L.; Bicsár, R.; Galata, D.L.; Nagy, B.; Nagy, Z.K. UV/VIS imaging-based PAT tool for drug particle size inspection in intact tablets supported by pattern recognition neural networks. Int. J. Pharm. 2022, 620, 121773. [Google Scholar] [CrossRef]
- Chauhan, S.; O’Callaghan, S.; Wall, A.; Pawlak, T.; Doyle, B.; Adelfio, A.; Trajkovic, S.; Gaffney, M.; Khaldi, N. Using Peptidomics and Machine Learning to Assess Effects of Drying Processes on the Peptide Profile within a Functional Ingredient. Processes 2021, 9, 425. [Google Scholar] [CrossRef]
- Farizhandi, A.A.K.; Alishiri, M.; Lau, R. Machine learning approach for carrier surface design in carrier-based dry powder inhalation. Comput. Chem. Eng. 2021, 151, 107367. [Google Scholar] [CrossRef]
- Jiang, J.; Peng, H.-H.; Yang, Z.; Ma, X.; Sahakijpijarn, S.; Moon, C.; Ouyang, D.; Iii, R.O.W. The applications of Machine learning (ML) in designing dry powder for inhalation by using thin-film-freezing technology. Int. J. Pharm. 2022, 626, 122179. [Google Scholar] [CrossRef] [PubMed]
- Xi, H.; Zhu, A.; Klinzing, G.R.; Zhou, L.; Zhang, S.; Gmitter, A.J.; Ploeger, K.; Sundararajan, P.; Mahjour, M.; Xu, W. Characterization of Spray Dried Particles Through Microstructural Imaging. J. Pharm. Sci. 2020, 109, 3404–3412. [Google Scholar] [CrossRef] [PubMed]
- Lou, H.; Chung, J.I.; Kiang, Y.H.; Xiao, L.Y.; Hageman, M.J. The application of machine learning algorithms in understanding the effect of core/shell technique on improving powder compactability. Int. J. Pharm. 2019, 555, 368–379. [Google Scholar] [CrossRef]
- Sinha, K.; Murphy, E.; Kumar, P.; Springer, K.A.; Ho, R.; Nere, N.K. A Novel Computational Approach Coupled with Machine Learning to Predict the Extent of Agglomeration in Particulate Processes. AAPS PharmSciTech 2022, 23, 18. [Google Scholar] [CrossRef]
- Zhou, J.; He, J.; Li, G.; Liu, Y. Identifying Capsule Defect Based on an Improved Convolutional Neural Network. Shock. Vib. 2020, 2020, 8887723. [Google Scholar] [CrossRef]
- Doerr, F.J.S.; Florence, A.J. A micro-XRT image analysis and machine learning methodology for the characterisation of multi-particulate capsule formulations. Int. J. Pharm. X 2020, 2, 100041. [Google Scholar] [CrossRef]
- Landin, M. Artificial Intelligence Tools for Scaling Up of High Shear Wet Granulation Process. J. Pharm. Sci. 2017, 106, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Medarević, D.P.; Kleinebudde, P.; Djuriš, J.; Djurić, Z.; Ibrić, S. Drug Development and Industrial Pharmacy Combined application of mixture experimental design and artificial neural networks in the solid dispersion development Combined application of mixture experimental design and artificial neural networks in the solid dispersion development. Drug Dev. Ind. Pharm. 2015, 42, 389–402. [Google Scholar] [CrossRef] [PubMed]
- Ghourichay, M.P.; Kiaie, S.H.; Nokhodchi, A.; Javadzadeh, Y. Formulation and Quality Control of Orally Disintegrating Tablets (ODTs): Recent Advances and Perspectives. Biomed Res. Int. 2021, 2021. [Google Scholar] [CrossRef] [PubMed]
- Jivraj, M.; Martini, L.G.; Thomson, C.M. An overview of the different excipients useful for the direct compression of tablets. Pharm. Sci. Technol. Today 2000, 3, 58–63. [Google Scholar] [CrossRef]
- Petrović, J.; Ibrić, S.; Betz, G.; Urić, Z. Optimization of Matrix Tablets Controlled Drug Release Using Elman Dynamic Neural Networks and Decision Trees. Int. J. Pharm. 2012, 428, 57–67. [Google Scholar] [CrossRef]
- Han, R.; Yang, Y.; Li, X.; Ouyang, D. Predicting oral disintegrating tablet formulations by neural network techniques. Asian J. Pharm. Sci. 2018, 13, 336–342. [Google Scholar] [CrossRef]
- Alhijjaj, M.; Nasereddin, J.; Belton, P.; Pharmaceutics, S.Q. 2019 undefined. Impact of processing parameters on the quality of pharmaceutical solid dosage forms produced by fused deposition modeling (FDM). Pharmaceutics 2019, 11, 633. [Google Scholar] [CrossRef]
- Vaz, V.M.; Kumar, L. 3D Printing as a Promising Tool in Personalized Medicine. AAPS PharmSciTech 2021, 22, 49. [Google Scholar] [CrossRef]
- Yost, E.; Chalus, P.; Zhang, S.; Peter, S.; Narang, A.S. Quantitative X-Ray Microcomputed Tomography Assessment of Internal Tablet Defects. J. Pharm. Sci. 2019, 108, 1818–1830. [Google Scholar] [CrossRef] [PubMed]
- Pharmaceutical Powder: An Overview-Pharmapproach.com. Available online: https://www.pharmapproach.com/pharmaceutical-powder-an-overview/ (accessed on 10 October 2021).
- Pharmaceutical Crystals: Science and Engineering-Tonglei Li, Alessandra Mattei-Google Books. Available online: https://books.google.com/books?id=KHhsDwAAQBAJ&pg=PA316&lpg=PA316&dq=powders+10nm+to+1000µm.&source=bl&ots=DttDf0IhVU&sig=ACfU3U1DFb94jn3f1ZV6ibv5zvyTA50BPA&hl=en&sa=X&ved=2ahUKEwihiJSercDzAhWSlmoFHceXBUwQ6AF6BAgDEAM#v=onepage&q=powders10nmto1000µm.&f=false (accessed on 10 October 2021).
- Need for Particle Engineering Increases. Available online: https://www.pharmtech.com/view/need-particle-engineering-increases (accessed on 10 October 2021).
- Optimization of Aerosol Drug Delivery-Google Books. Available online: https://books.google.com/books?id=JipsHpMQHPAC&pg=PA92&lpg=PA92&dq=pulmonary+powder+1um+5um&source=bl&ots=qnov3W2EIR&sig=ACfU3U1MP003bMtT1k5COKonCzg5yJmmsw&hl=en&sa=X&ved=2ahUKEwjI1PO6xsDzAhVcmWoFHdEDBuEQ6AF6BAgTEAM#v=onepage&q=pulmonarypowder1um5um&f=false (accessed on 10 October 2021).
- Giry, K.; Péan, J.M.; Giraud, L.; Marsas, S.; Rolland, H.; Wüthrich, P. Drug/lactose co-micronization by jet milling to improve aerosolization properties of a powder for inhalation. Int. J. Pharm. 2006, 321, 162–166. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, H.; Danjo, K. Application of supercritical fluid to preparation of powders of high-molecular weight drugs for inhalation. Adv. Drug Deliv. Rev. 2008, 60, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, M.; Baptista, B.; Lopes, J.A.; Sarraguça, M.C. Pharmaceutical cocrystallization techniques. Advances and challenges. Int. J. Pharm. 2018, 547, 404–420. [Google Scholar] [CrossRef]
- Moura, C.; Neves, F.; Costa, E. Impact of jet-milling and wet-polishing size reduction technologies on inhalation API particle properties. Powder Technol. 2016, 298, 90–98. [Google Scholar] [CrossRef]
- Keskes, S.; Hanini, S.; Hentabli, M.; Laidi, M. Artificial Intelligence and Mathematical Modelling of the Drying Kinetics of Pharmaceutical Powders. Kem. U Ind. 2020, 69, 137–152. [Google Scholar] [CrossRef]
- Aghbashlo, M.; Mobli, H.; Rafiee, S.; Madadlou, A. The use of artificial neural network to predict exergetic performance of spray drying process: A preliminary study. Comput. Electron. Agric. 2012, 88, 32–43. [Google Scholar] [CrossRef]
- Lavorini, F.; Pistolesi, M.; Usmani, O.S. Recent advances in capsule-based dry powder inhaler technology. Multidiscip. Respir. Med. 2017, 12, 11. [Google Scholar] [CrossRef]
- Mitchell, J.P.; Nagel, M.W.; Wiersema, K.J.; Doyle, C.C. Aerodynamic particle size analysis of aerosols from pressurized metered-dose inhalers: Comparison of Andersen 8-stage cascade impactor, next generation pharmaceutical impactor, and model 3321 Aerodynamic Particle Sizer aerosol spectrometer. AAPS PharmSciTech 2003, 4, 425–433. [Google Scholar] [CrossRef]
- Chrominfo: Advantages and Disadvantages of Granules Dosage Form. Available online: https://chrominfo.blogspot.com/2020/12/Advantages-and-disadvantages-of-granules-dosage-form.html (accessed on 9 October 2021).
- Zhao, J.; Tian, G.; Qiu, Y.; Qu, H. Rapid quantification of active pharmaceutical ingredient for sugar-free Yangwei granules in commercial production using FT-NIR spectroscopy based on machine learning techniques. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2021, 245, 118878. [Google Scholar] [CrossRef]
- Huang, Y.; Dai, W.G. Fundamental aspects of solid dispersion technology for poorly soluble drugs. Acta Pharm. Sin. B 2014, 4, 18–25. [Google Scholar] [CrossRef]
- Nikghalb, L.A.; Singh, G.; Singh, G.; Kahkeshan, K.F. Solid Dispersion: Methods and Polymers to increase the solubility of poorly soluble drugs. J. Appl. Pharm. Sci. 2012, 2, 170–175. [Google Scholar] [CrossRef]
- Shanbhag, A.; Rabel, S.; Nauka, E.; Casadevall, G.; Shivanand, P.; Eichenbaum, G.; Mansky, P. Method for screening of solid dispersion formulations of low-solubility compounds—Miniaturization and automation of solvent casting and dissolution testing. Int. J. Pharm. 2008, 351, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Sun, H.; Dong, J.; Ouyang, D. PharmDE: A new expert system for drug-excipient compatibility evaluation. Int. J. Pharm. 2021, 607, 120962. [Google Scholar] [CrossRef]
- Sun, D.D.; Lee, P.I. Evolution of supersaturation of amorphous pharmaceuticals: The effect of rate of supersaturation generation. Mol. Pharm. 2013, 10, 4330–4346. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Su, Y.; Wang, W.; Xiong, W.; Sun, X.; Ji, Y.; Yu, H.; Li, H.; Ouyang, D. Integrated computer-aided formulation design: A case study of andrographolide/cyclodextrin ternary formulation. Asian J. Pharm. Sci. 2021, 16, 494–507. [Google Scholar] [CrossRef] [PubMed]
- Farkas, D.; Madarász, L.; Nagy, Z.; Antal, I. Pharmaceutics NKS, 2021 undefined. Image analysis: A versatile tool in the manufacturing and quality control of pharmaceutical dosage forms. Pharmaceutics 2021, 13, 685. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Mikołajczyk, A.; Grochowski, M. Data augmentation for improving deep learning in image classification problem. In Proceedings of the 2018 International Interdisciplinary PhD Workshop (IIPhDW), Świnoujście, Poland, 9–12 May 2018; pp. 117–122. [Google Scholar]
- Umri, B.K.; Akhyari, M.W.; Kusrini, K. Detection of COVID-19 in Chest X-ray Image Using CLAHE and Convolutional Neural Network. Available online: https://ieeexplore.ieee.org/abstract/document/9320806/?casa_token=Ywp_llxzq3oAAAAA:IDvMLID0Iko1sh_zVzxN4Edg-By10X1RTaLlHop5mqOahC__KGBn7XoqoGh2j_J2zWEesPvK (accessed on 26 July 2022).
- Pitaloka, D.A.; Wulandari, A.; Basaruddin, T.; Liliana, D.Y. Enhancing CNN with preprocessing stage in automatic emotion recognition. Procedia Comput. Sci. 2017, 116, 523–529. [Google Scholar] [CrossRef]
- Kojima, R.; Ishida, S.; Ohta, M.; Iwata, H.; Honma, T.; Okuno, Y. KGCN: A graph-based deep learning framework for chemical structures. J. Cheminform. 2020, 12, 32. [Google Scholar] [CrossRef]
- Blanchard, A.E.; Stanley, C.; Bhowmik, D. Using GANs with adaptive training data to search for new molecules. J. Cheminform. 2021, 13, 14. [Google Scholar] [CrossRef] [PubMed]



| Some Popular Databases of Solid Dosage Formulations | ||||
|---|---|---|---|---|
| Name | Size | Publisher | Reference | |
| APIs/Chemicals | US Pharmacopoeia | >5000 | US Pharmacopoeia Convention | [36] |
| PubChem | >111 million | National Center for Biotechnology Information (NCBI) | [37] | |
| Cambridge Structure Database | >900,000 | University of Cambridge | [38] | |
| SciFinder | 142 million | Chemical Abstracts Service (CAS) | [39] | |
| Merck Index | >10,000 | Royal Society of Chemistry (RSC) | [40] | |
| Excipients | Inactive Ingredient Search for Approved Drug Products | 9438 | U.S. FDA | [41] |
| Formulations | Drugs@FDA (FDA-Approved Drugs) | >20,000 | U.S. FDA | [42] |
| Orange Book (Approved Drug Products with Therapeutic Equivalence Evaluations) | N/A | U.S. FDA | [43] | |
| DrugBank | >500,000 | University of Alberta | [44] | |
| Dissolution Methods | 1388 | U.S. FDA | [45] | |
| MedlinePlus® | ∼1500 | National Institute of Health | [46] | |
| Drug Information Portal | >49,000 | National Institute of Health | [47] | |
| Advantages and Disadvantages of Different AI Algorithms | |||
|---|---|---|---|
| Algorithms | Advantages | Disadvantages | |
| Regression | Linear regression | Easy to implement Efficient to train Performs well for linearly separable data | Prone to overfitting and noise The assumption of linearity of dependent and independent variables |
| Lasso regression | Performs shrinkage and variable selection Good prediction and interpretation | Model selection is unstable | |
| Ridge regression | Can avoid overfitting Performs well when having high-dimension data Does not require unbiased estimators | Unable to perform feature selection Shrinks the coefficient towards 0 Trades off bias for variance | |
| Classification | K-Nearest Neighbors | No training periods Easy to implement | Sensitive to missing values and outliers Does not work well for high-dimensional data Poor performance when having large databases |
| Support Vector Machines | Performs well when classes are separable Performs well in higher dimensions Outliers are less impactful | Slow processing speed Poor performance when having overlapped classes Challenging to select appropriate hyperparameters | |
| Random Forrest | Good performance when having imbalanced data Minimizes errors Can deal with massive databases Good handling of missing data Less impact of outliers | Easier for overfitting Relatively low accuracy Black box algorithm | |
| Naïve Bayes | Scalable databases Real-time and fast predictions Compatible with high-dimensional data | Poor performance of the estimator The assumption that variables are independent is not always true | |
| Clustering | K-means clustering | Easy to implement Fast computation time with huge variables Can recover from failure automatically | Sensitive to noisy data and outliers Need to specify the number of clusters (k) in advance |
| Density-Based Spatial Clustering of Applications with Noise clustering | Does not require specification of the number of clusters in advance Performs well with arbitrary shaped clusters Robust to outliers | Poor performance when data has high dimensions Fails when having varying cluster density | |
| Mean shift clustering | It can be used for complex clusters Robust to outliers It only needs bandwidth to determine the number of clusters | Poor performance when having high-dimensional data Slow implementation time | |
| Deep learning | ANN | Can store information on the entire network Exhibits fault tolerance Has distributed memory Gradual corruption Can perform multi-tasking simultaneously | A relatively high requirement in terms of hardware Poor explainability Challenging to determine ANN structure Unknown duration of the networks |
| CNN | High accuracy once CNN is fine-tuned Can detect the important features or patterns in the images | Requires higher computational power, especially GPU Large training data required | |
| RNN | Can model a collection of records The assumption that each pattern is dependent on the previous ones It can be coupled with convolutional layers to extend the pixel neighborhood | Vanishing gradient Difficult to train RNN Slow computation time | |
| Summary of Different Machine Learning Evaluation Metrics | ||
|---|---|---|
| Regression Metrics | Classification Metrics | Image Analysis |
|
|
|
| Applications of AI in Solid Dosages Forms (Since 2015) | |||
| Dosage Forms | Applications | Algorithms | Reference |
|---|---|---|---|
| Tablet | Predicting drug release | ANN, SVM, Ensemble of Regression Trees, and decision tree | [80,81] |
| Developing 3D-printed tablets | ANN, self-organizing maps, RF, SVM, and CNN | [65,71,82] | |
| Detecting tablet defects | CNN, You Only Look Once v5 (YOLOv5) | [83,64,65] | |
| Estimation of disintegration rate | RF, XGBoost, ANN, and CNN | [79,84] | |
| Drug particle size inspection | Pattern recognition neural network | [85] | |
| Powders | Process control of powder engineering | ANN | [86] |
| Designing dry powder for inhalation | RF, XGBoost, LightGBM, SVM, KNN, ANN, and CNN | [87,88] | |
| Predicting particle size distribution of spray-dried powder | Unspecified | [89] | |
| Improving spray-dried powder compatibility | SVM and ANN | [90] | |
| Predicting the extent of agglomeration | SVM, RF, and partial least squares regression | [91] | |
| Capsules | Identifying capsule defects | KNN, SVM, and CNN | [92] |
| Detecting the defects of the pellets within the capsules | SVM | [93] | |
| Granules | Granulation process control | Neuro-fuzzy logic and genetic programming | [94] |
| Predicting particle size distribution | ANN, multiple linear regression, and genetic programming | [60] | |
| Solid dispersions | Predicting physical or chemical stabilities | ANN, SVM, RF, LightGBM, KNN, and naïve Bayes | [52,59] |
| Predicting dissolution rates and profiles | RF, SVM, LightGBM, and XGBoost | [56,95] | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, J.; Ma, X.; Ouyang, D.; Williams, R.O., III. Emerging Artificial Intelligence (AI) Technologies Used in the Development of Solid Dosage Forms. Pharmaceutics 2022, 14, 2257. https://doi.org/10.3390/pharmaceutics14112257
Jiang J, Ma X, Ouyang D, Williams RO III. Emerging Artificial Intelligence (AI) Technologies Used in the Development of Solid Dosage Forms. Pharmaceutics. 2022; 14(11):2257. https://doi.org/10.3390/pharmaceutics14112257
Chicago/Turabian StyleJiang, Junhuang, Xiangyu Ma, Defang Ouyang, and Robert O. Williams, III. 2022. "Emerging Artificial Intelligence (AI) Technologies Used in the Development of Solid Dosage Forms" Pharmaceutics 14, no. 11: 2257. https://doi.org/10.3390/pharmaceutics14112257
APA StyleJiang, J., Ma, X., Ouyang, D., & Williams, R. O., III. (2022). Emerging Artificial Intelligence (AI) Technologies Used in the Development of Solid Dosage Forms. Pharmaceutics, 14(11), 2257. https://doi.org/10.3390/pharmaceutics14112257








