Non-Viral in Vitro Gene Delivery: It is Now Time to Set the Bar!
Abstract
1. Introduction
2. Non-Viral Gene Delivery Using Plasmid DNA and Cationic Polymers
2.1. Plasmid DNA
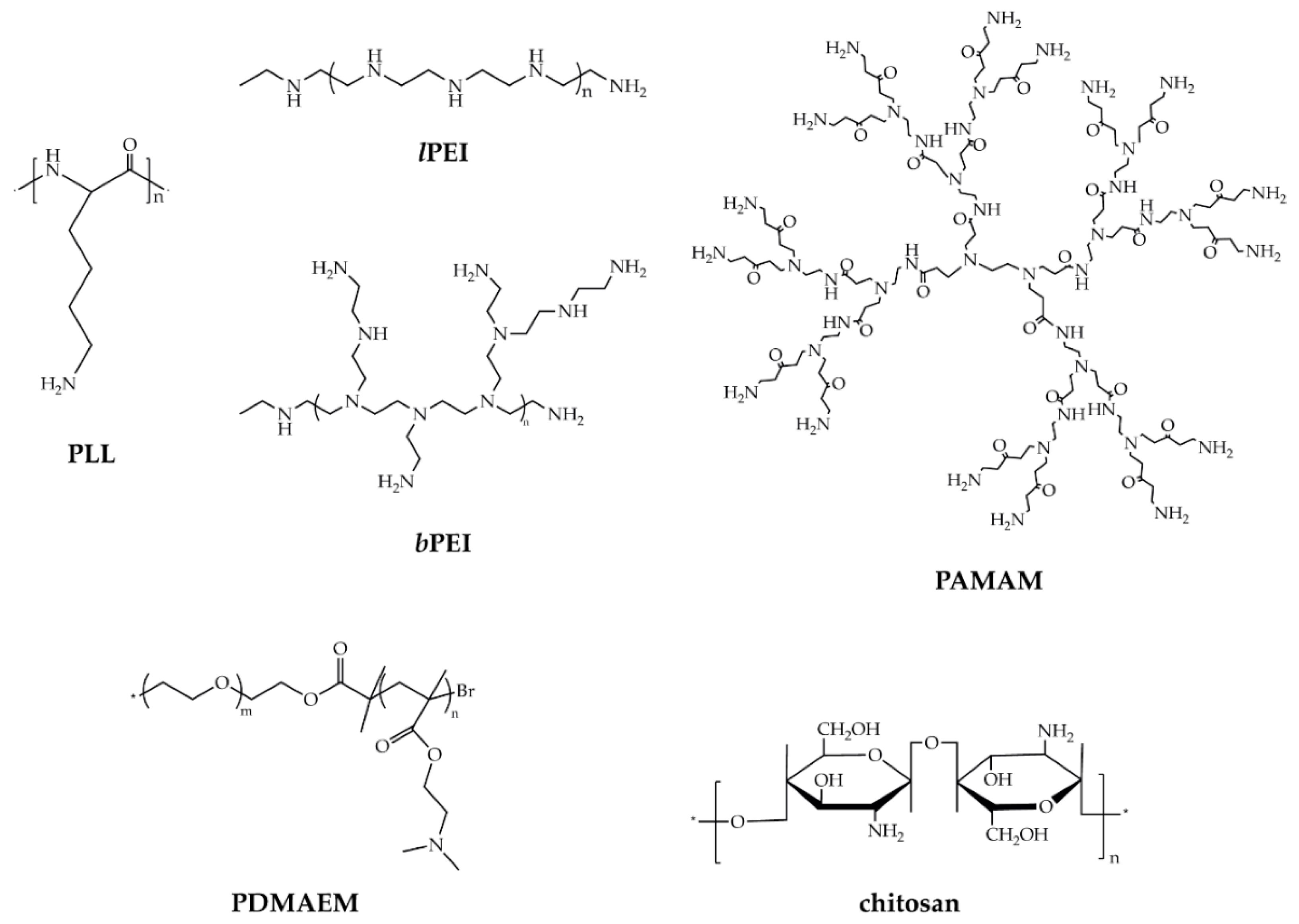
2.2. Cationic Polymers (CPs)
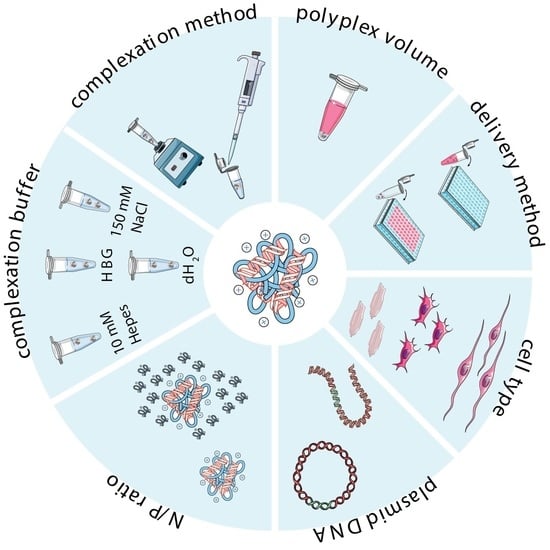
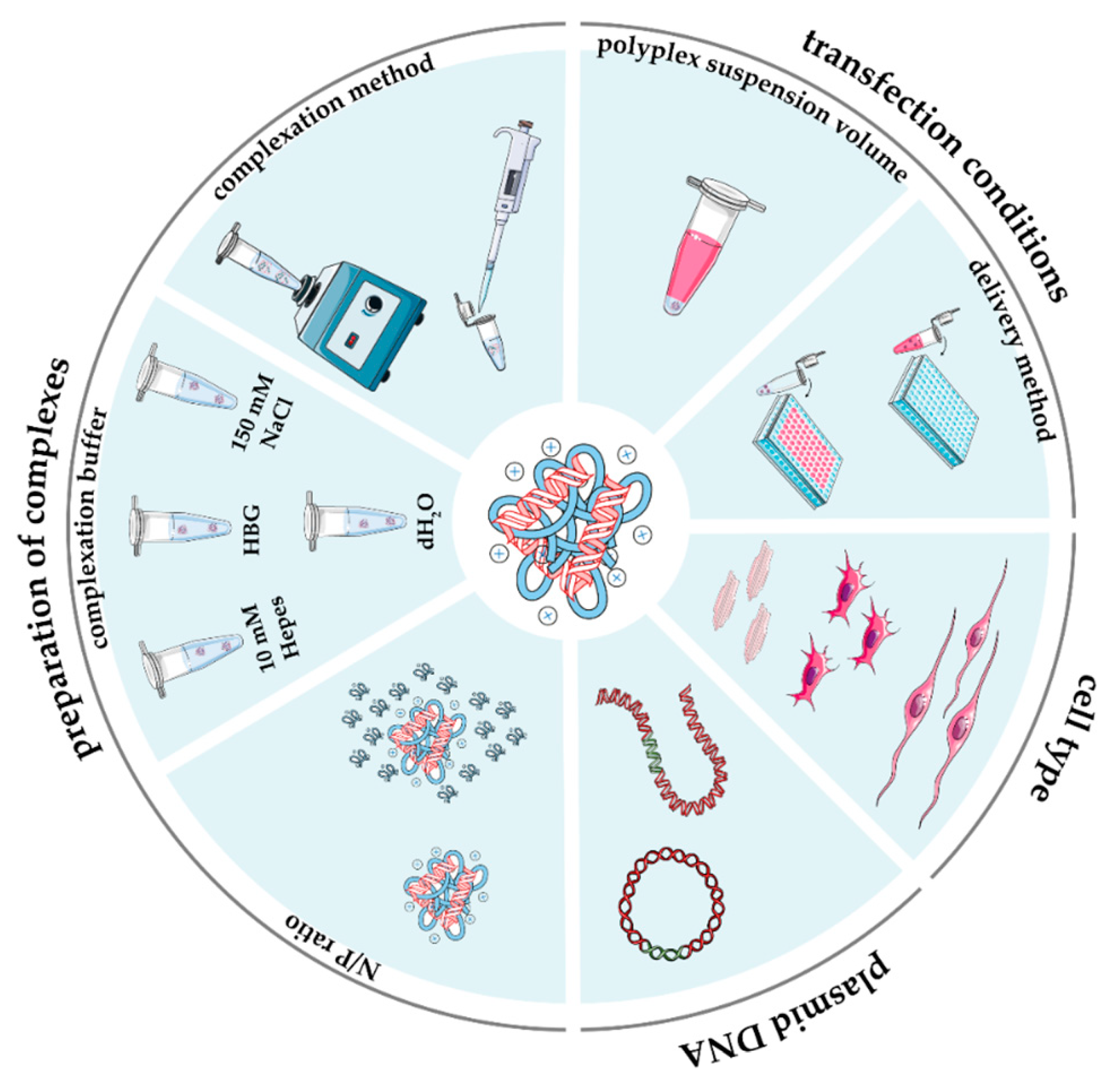
2.3. Preparation of Complexes
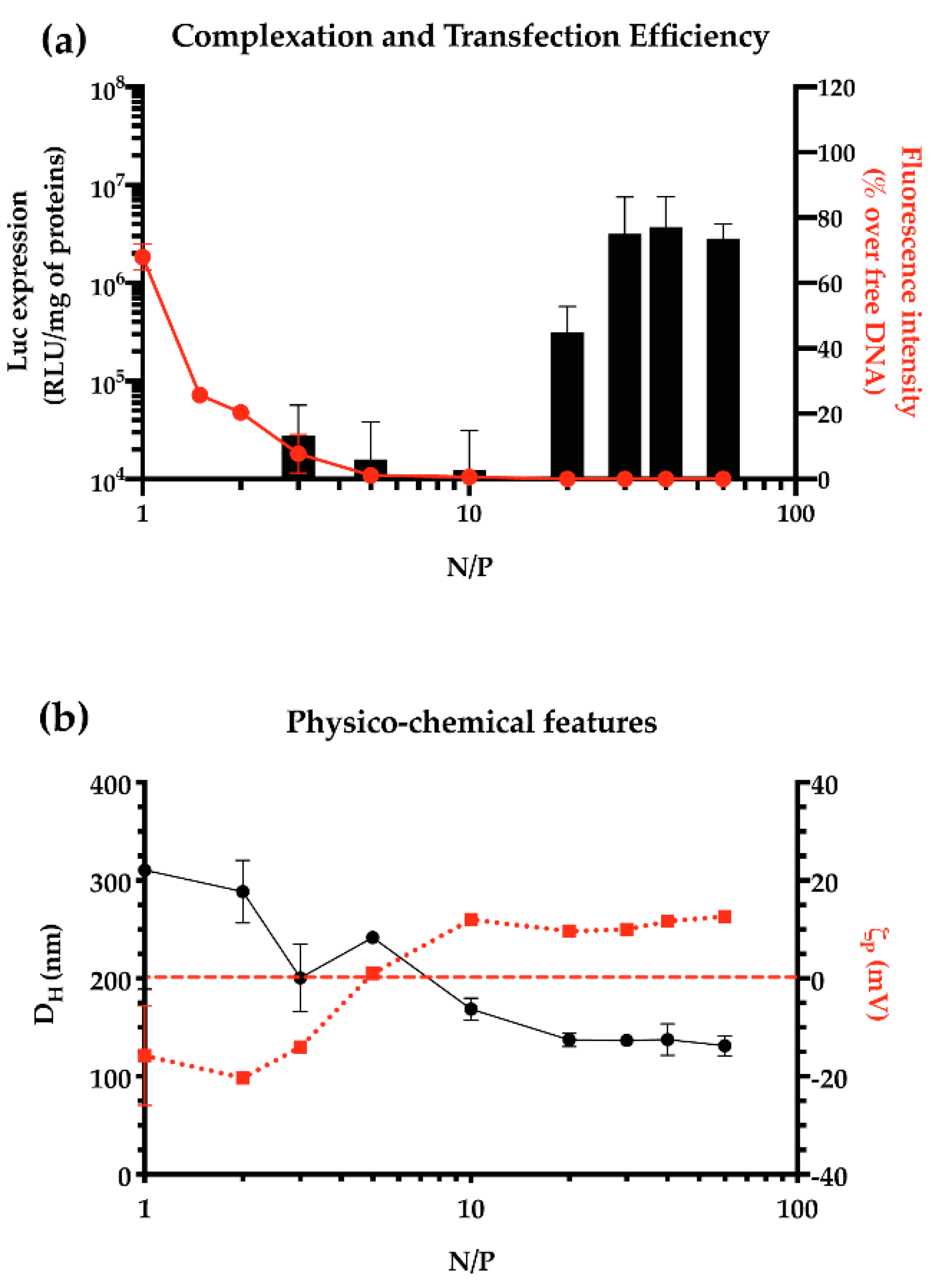
2.3.1. Cationic Polymer-to-Plasmid DNA Ratio
2.3.2. Polymer Solubilization and Complexation Buffer
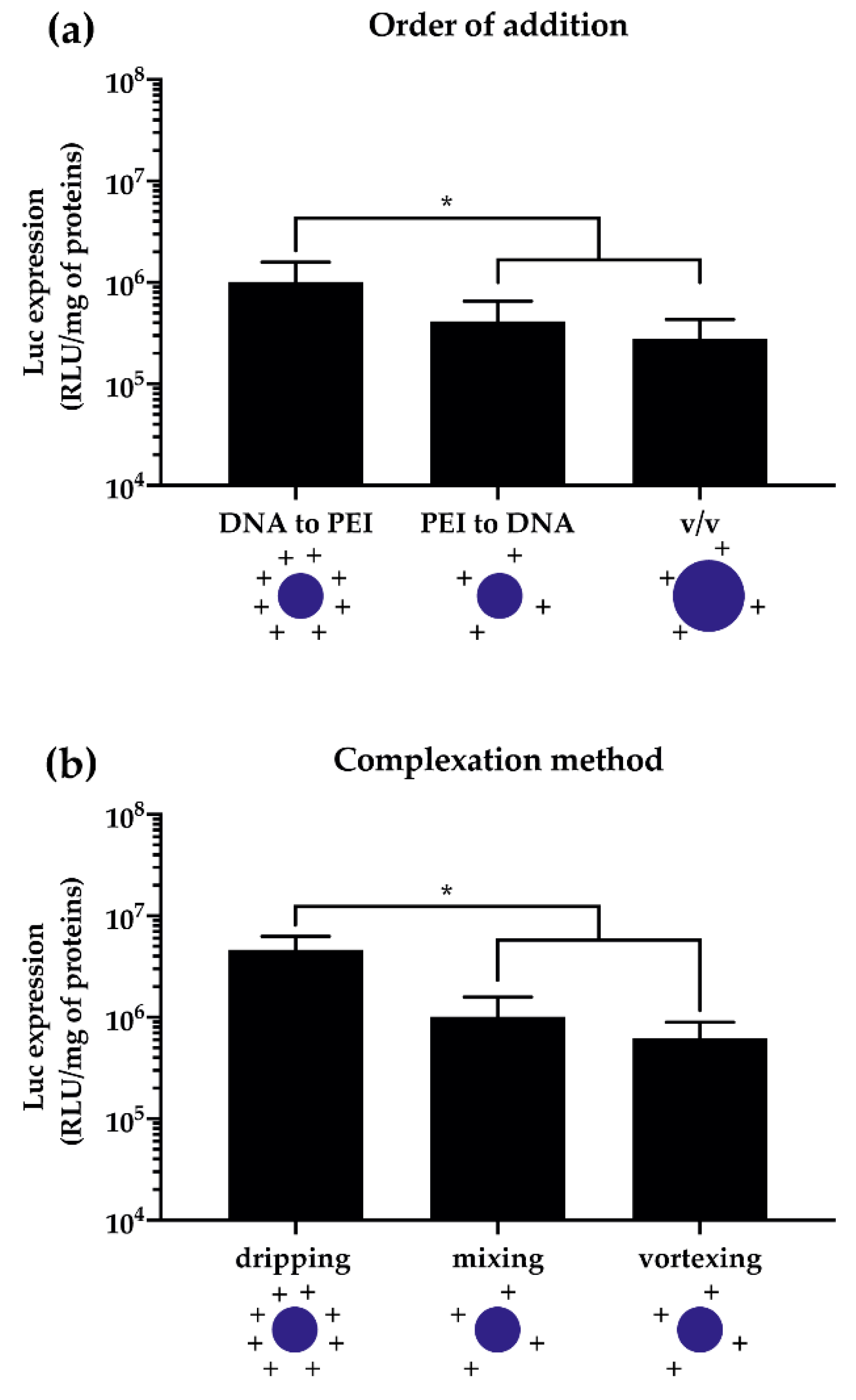
2.3.3. Complexation Method
3. Experimental Strategies and In Vitro Transfection Assays
3.1. Cell Type and Culture Conditions
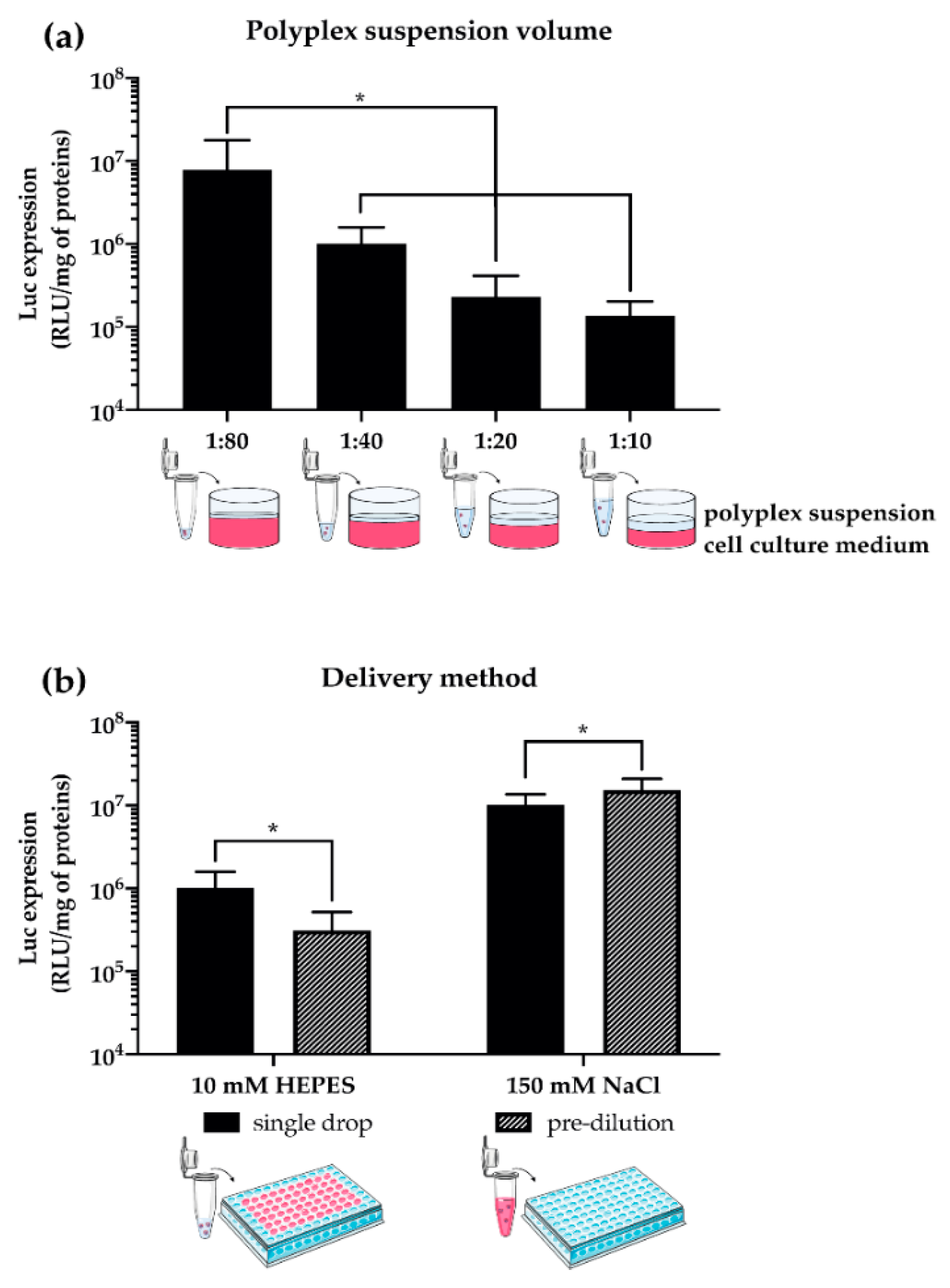
3.2. Transfection Conditions
3.3. Evaluation of Transfection Effectiveness: A Trade-off Between Transfection Efficiency and Cytotoxicity
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lostalé-Seijo, I.; Montenegro, J. Synthetic materials at the forefront of gene delivery. Nat. Rev. Chem. 2018, 2, 258–277. [Google Scholar] [CrossRef]
- Wirth, T.; Parker, N.; Ylä-Herttuala, S. History of gene therapy. Gene 2013, 525, 162–169. [Google Scholar] [CrossRef]
- Pan, J.; Zhang, S.; Chen, C.; Xiao, S.; Sun, Y.; Liu, C.; Su, X.; Li, D.; Xu, G.; Xu, B.; et al. Effect of recombinant adenovirus-p53 combined with radiotherapy on long-term prognosis of advanced nasopharyngeal carcinoma. J. Clin. Oncol. 2009, 27, 799–804. [Google Scholar] [CrossRef]
- Aied, A.; Greiser, U.; Pandit, A.; Wang, W. Polymer gene delivery: Overcoming the obstacles. Drug Discov. Today 2013, 18, 1090–1098. [Google Scholar] [CrossRef]
- Baldi, L.; Hacker, D.L.; Adam, M.; Wurm, F.M. Recombinant protein production by large-scale transient gene expression in mammalian cells: State of the art and future perspectives. Biotechnol. Lett. 2007, 29, 677–684. [Google Scholar] [CrossRef] [PubMed]
- Pham, P.L.; Kamen, A.; Durocher, Y. Large-scale transfection of mammalian cells for the fast production of recombinant protein. Mol. Biotechnol. 2006, 34, 225–237. [Google Scholar] [CrossRef]
- Bandaranayake, A.D.; Almo, S.C. Recent advances in mammalian protein production. FEBS Lett. 2014, 588, 253–260. [Google Scholar] [CrossRef]
- Wong, J.K.L.; Mohseni, R.; Hamidieh, A.A.; MacLaren, R.E.; Habib, N.; Seifalian, A.M. Will Nanotechnology Bring New Hope for Gene Delivery? Trends Biotechnol. 2017, 35, 434–451. [Google Scholar] [CrossRef]
- Lächelt, U.; Wagner, E. Nucleic Acid Therapeutics Using Polyplexes: A Journey of 50 Years (and Beyond). Chem. Rev. 2015, 115, 11043–11078. [Google Scholar] [CrossRef]
- Rinaldi, C.; Wood, M.J.A. Antisense oligonucleotides: The next frontier for treatment of neurological disorders. Nat. Rev. Neurol. 2018, 14, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Pezzoli, D.; Chiesa, R.; De Nardo, L.; Candiani, G. We still have a long way to go to effectively deliver genes! J. Appl. Biomater. Funct. Mater. 2012, 10, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Pezzoli, D.; Candiani, G. Non-viral gene delivery strategies for gene therapy: A “ménage à trois” among nucleic acids, materials, and the biological environment: Stimuli-responsive gene delivery vectors. J. Nanoparticle Res. 2013, 15, 1523. [Google Scholar]
- Thiel, C.; Nix, M. Efficient transfection of primary cells relevant for cardiovascular research by nucleofection. Methods Mol. Med. 2006, 129, 255–266. [Google Scholar] [PubMed]
- Lin, Y.C.; Li, M.; Wu, C.C. Simulation and experimental demonstration of the electric field assisted electroporation microchip for in vitro gene delivery enhancement. Lab Chip 2004, 4, 104–108. [Google Scholar] [CrossRef]
- Iversen, N.; Birkenes, B.; Torsdalen, K.; Djurovic, S. Electroporation by nucleofactor is the best nonviral transfection technique in human endothelial and smooth muscle cells. Genet. Vaccines Ther. 2005, 3. [Google Scholar] [CrossRef]
- Liao, Z.K.; Tsai, K.C.; Wang, H.T.; Tseng, S.H.; Deng, W.P.; Chen, W.S.; Hwang, L.H. Sonoporation-mediated anti-angiogenic gene transfer into muscle effectively regresses distant orthotopic tumors. Cancer Gene Ther. 2012, 19, 171–180. [Google Scholar] [CrossRef][Green Version]
- Li, Y.S.; Davidson, E.; Reid, C.N.; McHale, A.P. Optimising ultrasound-mediated gene transfer (sonoporation) in vitro and prolonged expression of a transgene in vivo: Potential applications for gene therapy of cancer. Cancer Lett. 2009, 273, 62–69. [Google Scholar] [CrossRef]
- Mignet, N.; Marie, C.; Delalande, A.; Manta, S.; Bureau, M.F.; Renault, G.; Scherman, D.; Pichon, C. Microbubbles for nucleic acid delivery in liver using mild sonoporation. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2019; Volume 1943, pp. 377–387. [Google Scholar]
- Scherer, F.; Anton, M.; Schillinger, U.; Henke, J.; Bergemann, C.; Krüger, A.; Gänsbacher, B.; Plank, C. Magnetofection: Enhancing and targeting gene delivery by magnetic force in vitro and in vivo. Gene Ther. 2002, 9, 102–109. [Google Scholar] [CrossRef]
- Krötz, F.; Sohn, H.-Y.; Gloe, T.; Plank, C.; Pohl, U. Magnetofection Potentiates Gene Delivery to Cultured Endothelial Cells. J. Vasc. Res. 2003, 40, 425–434. [Google Scholar] [CrossRef]
- Davis, A.A.; Farrar, M.J.; Nishimura, N.; Jin, M.M.; Schaffer, C.B. Optoporation and genetic manipulation of cells using femtosecond laser pulses. Biophys. J. 2013, 105, 862–871. [Google Scholar] [CrossRef]
- Schneckenburger, H.; Hendinger, A.; Sailer, R.; Strauss, W.S.L.; Schmitt, M. Laser-assisted optoporation of single cells. J. Biomed. Opt. 2002, 7, 410. [Google Scholar] [CrossRef]
- O’Brien, J.A.; Lummis, S.C.R. Biolistic transfection of neuronal cultures using a hand-held gene gun. Nat. Protoc. 2006, 1, 977–981. [Google Scholar] [CrossRef] [PubMed]
- Chow, Y.T.; Chen, S.; Wang, R.; Liu, C.; Kong, C.W.; Li, R.A.; Cheng, S.H.; Sun, D. Single cell transfection through precise microinjection with quantitatively controlled injection volumes. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- King, R. Gene delivery to mammalian cells by microinjection. Methods Mol. Biol. 2004, 245, 167–174. [Google Scholar] [PubMed]
- Chenuet, S.; Derouazi, M.; Hacker, D.; Wurm, F. DNA delivery by microinjection for the generation of recombinant mammalian cell lines. Methods Mol. Biol. 2009, 518, 99–112. [Google Scholar]
- Mehier-Humbert, S.; Guy, R.H. Physical methods for gene transfer: Improving the kinetics of gene delivery into cells. Adv. Drug Deliv. Rev. 2005, 57, 733–753. [Google Scholar] [CrossRef]
- Du, X.; Wang, J.; Zhou, Q.; Zhang, L.; Wang, S.; Zhang, Z.; Yao, C. Advanced physical techniques for gene delivery based on membrane perforation. Drug Deliv. 2018, 25, 1516–1525. [Google Scholar] [CrossRef]
- Mellott, A.J.; Forrest, M.L.; Detamore, M.S. Physical non-viral gene delivery methods for tissue engineering. Ann. Biomed. Eng. 2013, 41, 446–468. [Google Scholar] [CrossRef]
- Lungwitz, U.; Breunig, M.; Blunk, T.; Göpferich, A. Polyethylenimine-based non-viral gene delivery systems. Eur. J. Pharm. Biopharm. 2005, 60, 247–266. [Google Scholar]
- Ogris, M.; Wagner, E. Targeting tumors with non-viral gene delivery systems. Drug Discov. Today 2002, 7, 479–485. [Google Scholar] [CrossRef]
- Tomlinson, E.; Rolland, A.P. Controllable gene therapy pharmaceutics of non-viral gene delivery systems. J. Control. Release 1996, 39, 357–372. [Google Scholar] [CrossRef]
- De Laporte, L.; Cruz Rea, J.; Shea, L.D. Design of modular non-viral gene therapy vectors. Biomaterials 2006, 27, 947–954. [Google Scholar] [CrossRef] [PubMed]
- Finer, M.; Glorioso, J. A brief account of viral vectors and their promise for gene therapy. Gene Ther. 2017, 24, 1–2. [Google Scholar] [CrossRef]
- Keeler, A.M.; ElMallah, M.K.; Flotte, T.R. Gene Therapy 2017: Progress and Future Directions. Clin. Transl. Sci. 2017, 10, 242–248. [Google Scholar] [CrossRef] [PubMed]
- Ginn, S.L.; Amaya, A.K.; Alexander, I.E.; Edelstein, M.; Abedi, M.R. Gene therapy clinical trials worldwide to 2017: An update. J. Gene Med. 2018, 20, e3015. [Google Scholar] [CrossRef] [PubMed]
- van der Loo, J.C.M.; Wright, J.F. Progress and challenges in viral vector manufacturing. Hum. Mol. Genet. 2016, 25, R42–R52. [Google Scholar] [CrossRef]
- Kotterman, M.A.; Chalberg, T.W.; Schaffer, D.V. Viral Vectors for Gene Therapy: Translational and Clinical Outlook. Annu. Rev. Biomed. Eng. 2015, 17, 63–89. [Google Scholar] [CrossRef]
- Thomas, C.E.; Ehrhardt, A.; Kay, M.A. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 2003, 4, 346–358. [Google Scholar] [CrossRef]
- Nayerossadat, N.; Ali, P.; Maedeh, T. Viral and nonviral delivery systems for gene delivery. Adv. Biomed. Res. 2012, 1, 27. [Google Scholar] [CrossRef]
- Sung, Y.K.; Kim, S.W. Recent advances in the development of gene delivery systems. Biomater. Res. 2019, 23. [Google Scholar] [CrossRef]
- Jinturkar, K.A.; Rathi, M.N.; Misra, A. Gene Delivery Using Physical Methods. In Challenges in Delivery of Therapeutic Genomics and Proteomics; Elsevier: Amsterdan, The Netherlands, 2011; pp. 83–126. [Google Scholar]
- Lundstrom, K. Viral Vectors in Gene Therapy. Diseases 2018, 6, 42. [Google Scholar] [CrossRef]
- Milone, M.C.; O’Doherty, U. Clinical use of lentiviral vectors. Leukemia 2018, 32, 1529–1541. [Google Scholar] [CrossRef] [PubMed]
- Park, T.G.; Jeong, J.H.; Kim, S.W. Current status of polymeric gene delivery systems. Adv. Drug Deliv. Rev. 2006, 58, 467–486. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.H.; Chen, C.K.; Ravikrishnan, A.; Rane, S.; Pfeifer, B.A. Overcoming nonviral gene delivery barriers: Perspective and future. Mol. Pharm. 2013, 10, 4082–4098. [Google Scholar] [CrossRef] [PubMed]
- Dizaj, S.; Jafari, S.; Khosroushahi, A. A sight on the current nanoparticle-based gene delivery vectors. Nanoscale Res. Lett. 2014, 9, 252. [Google Scholar] [CrossRef] [PubMed]
- Levingstone, T.J.; Herbaj, S.; Redmond, J.; McCarthy, H.O.; Dunne, N.J. Calcium Phosphate Nanoparticles-Based Systems for RNAi Delivery: Applications in Bone Tissue Regeneration. Nanomaterials 2020, 10, 146. [Google Scholar] [CrossRef] [PubMed]
- Graham, F.L.; van der Eb, A.J. A new technique for the assay of infectivity of human adenovirus 5 DNA. Virology 1973, 52, 456–467. [Google Scholar] [CrossRef]
- Wang, W.; Li, W.; Ma, N.; Steinhoff, G. Non-Viral Gene Delivery Methods. Curr. Pharm. Biotechnol. 2013, 14, 46–60. [Google Scholar]
- Thapa, B.; Narain, R. Mechanism, current challenges and new approaches for non viral gene delivery. In Polymers and Nanomaterials for Gene Therapy; Woodhead Publishing: Cambridge, UK, 2016; ISBN 9780081005217. [Google Scholar]
- Patil; Gao; Lin; Li; Dang; Tian; Zhang; Jiang; Qadir; Qian The Development of Functional Non-Viral Vectors for Gene Delivery. Int. J. Mol. Sci. 2019, 20, 5491. [CrossRef]
- Godbey, W.T.; Mikos, A.G. Recent progress in gene delivery using non-viral transfer complexes. J. Control. Release 2001, 72, 115–125. [Google Scholar] [CrossRef]
- Davis, M.E. Non-viral gene delivery systems. Curr. Opin. Biotechnol. 2002, 13, 128–131. [Google Scholar] [CrossRef]
- Bonetta, L. The inside scoop - Evaluating gene delivery methods. Nat. Methods 2005, 2, 875–882. [Google Scholar] [CrossRef]
- Felgner, P.L.; Gadek, T.R.; Holm, M.; Roman, R.; Chan, H.W.; Wenz, M.; Northrop, J.P.; Ringold, G.M.; Danielsen, M. Lipofection: A highly efficient, lipid-mediated DNA-transfection procedure. Proc. Natl. Acad. Sci. USA 1987, 84, 7413–7417. [Google Scholar] [CrossRef]
- Bono, N.; Pennetta, C.; Sganappa, A.; Giupponi, E.; Sansone, F.; Volonterio, A.; Candiani, G. Design and synthesis of biologically active cationic amphiphiles built on the calix[4]arene scaffold. Int. J. Pharm. 2018, 549, 436–445. [Google Scholar] [CrossRef]
- Hoekstra, D.; Rejman, J.; Wasungu, L.; Shi, F.; Zuhorn, I. Gene delivery by cationic lipids: In and out of an endosome. Biochem. Soc. Trans. 2007, 35, 68–71. [Google Scholar] [CrossRef]
- Simões, S.; Filipe, A.; Faneca, H.; Mano, M.; Penacho, N.; Düzgünes, N.; de Lima, M.P. Cationic liposomes for gene delivery. Expert Opin. Drug Deliv. 2005, 2, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Candiani, G.; Pezzoli, D.; Ciani, L.; Chiesa, R.; Ristori, S. Bioreducible liposomes for gene delivery: From the formulation to the mechanism of action. PLoS One 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Pezzoli, D.; Kajaste-Rudnitski, A.; Chiesa, R.; Candiani, G. Lipid-based nanoparticles as nonviral gene delivery vectors. Methods Mol. Biol. 2013, 1025, 269–279. [Google Scholar] [PubMed]
- Malloggi, C.; Pezzoli, D.; Magagnin, L.; De Nardo, L.; Mantovani, D.; Tallarita, E.; Candiani, G. Comparative evaluation and optimization of off-the-shelf cationic polymers for gene delivery purposes. Polym. Chem. 2015, 6, 6325–6339. [Google Scholar] [CrossRef]
- Pezzoli, D.; Tsekoura, E.K.; Remant Bahadur, K.C.; Candiani, G.; Mantovani, D.; Uludağ, H. Hydrophobe-substituted bPEI derivatives: Boosting transfection on primary vascular cells. Sci. China Mater. 2017, 60, 529–542. [Google Scholar] [CrossRef]
- Bono, N.; Pennetta, C.; Bellucci, M.C.; Sganappa, A.; Malloggi, C.; Tedeschi, G.; Candiani, G.; Volonterio, A. Role of Generation on Successful DNA Delivery of PAMAM-(Guanidino)Neomycin Conjugates. ACS Omega 2019, 4, 6796–6807. [Google Scholar] [CrossRef]
- Boussif, O.; Lezoualc’h, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.P. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: Polyethylenimine. Proc. Natl. Acad. Sci. 1995, 92, 7297–7301. [Google Scholar] [CrossRef]
- Von Harpe, A.; Petersen, H.; Li, Y.; Kissel, T. Characterization of commercially available and synthesized polyethylenimines for gene delivery. J. Control. Release 2000, 69, 309–322. [Google Scholar] [CrossRef]
- Xu, Q.; Wang, C.-H.; Wayne Pack, D. Polymeric Carriers for Gene Delivery: Chitosan and Poly(amidoamine) Dendrimers. Curr. Pharm. Des. 2010, 16, 2350–2368. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.B.; Chen, M.; Chen, C.K.; Pfeifer, B.A.; Jones, C.H. Overcoming gene-delivery hurdles: Physiological considerations for nonviral vectors. Trends Biotechnol. 2016, 34, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Mintzer, M.A.; Simanek, E.E. Nonviral vectors for gene delivery. Chem. Rev. 2009, 109, 259–302. [Google Scholar] [CrossRef]
- Gigante, A.; Li, M.; Junghänel, S.; Hirschhäuser, C.; Knauer, S.; Schmuck, C. Non-viral transfection vectors: Are hybrid materials the way forward? Medchemcomm 2019, 10, 1692–1718. [Google Scholar]
- Slivac, I.; Guay, D.; Mangion, M.; Champeil, J.; Gaillet, B. Non-viral nucleic acid delivery methods. Expert Opin. Biol. Ther. 2017, 17, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Bespalov, A.; Steckler, T. Lacking quality in research: Is behavioral neuroscience affected more than other areas of biomedical science? J. Neurosci. Methods 2018, 300, 4–9. [Google Scholar] [CrossRef]
- Brazma, A.; Hingamp, P.; Quackenbush, J.; Sherlock, G.; Spellman, P.; Stoeckert, C.; Aach, J.; Ansorge, W.; Ball, C.A.; Causton, H.C.; et al. Minimum information about a microarray experiment (MIAME) - Toward standards for microarray data. Nat. Genet. 2001, 29, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Le Novère, N.; Finney, A.; Hucka, M.; Bhalla, U.S.; Campagne, F.; Collado-Vides, J.; Crampin, E.J.; Halstead, M.; Klipp, E.; Mendes, P.; et al. Minimum information requested in the annotation of biochemical models (MIRIAM). Nat. Biotechnol. 2005, 23, 1509–1515. [Google Scholar] [CrossRef]
- Taylor, C.F.; Field, D.; Sansone, S.A.; Aerts, J.; Apweiler, R.; Ashburner, M.; Ball, C.A.; Binz, P.A.; Bogue, M.; Booth, T.; et al. Promoting coherent minimum reporting guidelines for biological and biomedical investigations: The MIBBI project. Nat. Biotechnol. 2008, 26, 889–896. [Google Scholar] [CrossRef] [PubMed]
- Faria, M.; Björnmalm, M.; Thurecht, K.J.; Kent, S.J.; Parton, R.G.; Kavallaris, M.; Johnston, A.P.R.; Gooding, J.J.; Corrie, S.R.; Boyd, B.J.; et al. Minimum information reporting in bio–nano experimental literature. Nat. Nanotechnol. 2018, 13, 777–785. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.; Uludaǧ, H. Effects of size and topology of DNA molecules on intracellular delivery with non-viral gene carriers. BMC Biotechnol. 2008, 8, 23. [Google Scholar] [CrossRef] [PubMed]
- Remaut, K.; Symens, N.; Lucas, B.; Demeester, J.; De Smedt, S.C. Cell division responsive peptides for optimized plasmid DNA delivery: The mitotic window of opportunity? J. Control. Release 2014, 179, 1–9. [Google Scholar] [CrossRef]
- Cherng, J.Y.; Schuurmans-Nieuwenbroek, N.M.E.; Jiskoot, W.; Talsma, H.; Zuidam, N.J.; Hennink, W.E.; Crommelin, D.J.A. Effect of DNA topology on the transfection efficiency of poly((2-dimethylamino)ethyl methacrylate)-plasmid complexes. J. Control. Release 1999, 60, 343–353. [Google Scholar] [CrossRef]
- Weintraub, H.; Cheng, P.F.; Conrad, K. Expression of transfected DNA depends on DNA topology. Cell 1986, 46, 115–122. [Google Scholar] [CrossRef]
- Tudini, E.; Burke, L.J.; Whiley, P.J.; Sevcik, J.; Spurdle, A.B.; Brown, M.A. Caution: Plasmid DNA topology affects luciferase assay reproducibility and outcomes. Biotechniques 2019, 67, 94–96. [Google Scholar] [CrossRef]
- Mairhofer, J.; Grabherr, R. Rational vector design for efficient non-viral gene delivery: Challenges facing the use of plasmid DNA. Mol. Biotechnol. 2008, 39, 97–104. [Google Scholar] [CrossRef]
- Kreiss, P.; Cameron, B.; Rangara, R.; Mailhe, P.; Aguerre-Charriol, O.; Airiau, M.; Scherman, D.; Crouzet, J.; Pitard, B. Plasmid DNA size does not affect the physicochemical properties of lipoplexes but modulates gene transfer efficiency. Nucleic Acids Res. 1999, 27, 3792–3798. [Google Scholar] [CrossRef]
- Shaimardanova, A.A.; Kitaeva, K.V.; Abdrakhmanova, I.I.; Chernov, V.M.; Rutland, C.S.; Rizvanov, A.A.; Chulpanova, D.S.; Solovyeva, V.V. Production and application of multicistronic constructs for various human disease therapies. Pharmaceutics 2019, 11, 580. [Google Scholar] [CrossRef]
- Chan, K.K.-K.; Wu, S.M.; Nissom, P.M.; Oh, S.K.W.; Choo, A.B.H. Generation of high-level stable transgene expressing human embryonic stem cell lines using Chinese hamster elongation factor-1 alpha promoter system. Stem Cells Dev. 2008, 17, 825–836. [Google Scholar] [CrossRef] [PubMed]
- Tokushige, K.; Moradpour, D.; Wakita, T.; Geissler, M.; Hayashi, N.; Wands, J.R. Comparison between cytomegalovirus promoter and elongation factor-1 alpha promoter-driven constructs in the establishment of cell lines expressing hepatitis C virus core protein. J. Virol. Methods 1997, 64, 73–80. [Google Scholar] [CrossRef]
- Gopalkrishnan, R. Use of the human EF-1alpha promoter for expression can significantly increase success in establishing stable cell lines with consistent expression: A study using the tetracycline-inducible system in human cancer cells. Nucleic Acids Res. 1999, 27, 4775–4782. [Google Scholar] [CrossRef]
- Raup, A.; Jérôme, V.; Freitag, R.; Synatschke, C.V.; Müller, A.H.E. Promoter, transgene, and cell line effects in the transfection of mammalian cells using PDMAEMA-based nano-stars. Biotechnol. Reports 2016, 11, 53–61. [Google Scholar] [CrossRef]
- Chung, S.; Andersson, T.; Sonntag, K.-C.; Björklund, L.; Isacson, O.; Kim, K.-S. Analysis of Different Promoter Systems for Efficient Transgene Expression in Mouse Embryonic Stem Cell Lines. Stem Cells 2002, 20, 139–145. [Google Scholar] [CrossRef]
- Irobalieva, R.N.; Fogg, J.M.; Catanese, D.J.; Sutthibutpong, T.; Chen, M.; Barker, A.K.; Ludtke, S.J.; Harris, S.A.; Schmid, M.F.; Chiu, W.; et al. Structural diversity of supercoiled DNA. Nat. Commun. 2015, 6, 8440. [Google Scholar] [CrossRef]
- Peng, L.; Wagner, E. Polymeric Carriers for Nucleic Acid Delivery: Current Designs and Future Directions. Biomacromolecules 2019, 20, 3613–3626. [Google Scholar] [CrossRef]
- Lai, W.F.; Wong, W.T. Design of Polymeric Gene Carriers for Effective Intracellular Delivery. Trends Biotechnol. 2018, 36, 713–728. [Google Scholar] [CrossRef]
- Wightman, L.; Kircheis, R.; Wagner, E. Polymer-Based gene delivery systems. In Pharmaceutical Gene Delivery Systems; CRC Press: Boca Raton, FL, USA, 2003; pp. 109–135. ISBN 9780203912331. [Google Scholar]
- Zhang, X.; Oulad-Abdelghani, M.; Zelkin, A.N.; Wang, Y.; Haîkel, Y.; Mainard, D.; Voegel, J.-C.; Caruso, F.; Benkirane-Jessel, N. Poly(L-lysine) nanostructured particles for gene delivery and hormone stimulation. Biomaterials 2010, 31, 1699–1706. [Google Scholar] [CrossRef]
- Mandal, H.; Katiyar, S.S.; Swami, R.; Kushwah, V.; Katare, P.B.; Kumar Meka, A.; Banerjee, S.K.; Popat, A.; Jain, S. ε-Poly-l-Lysine/plasmid DNA nanoplexes for efficient gene delivery in vivo. Int. J. Pharm. 2018, 542, 142–152. [Google Scholar] [CrossRef]
- Pezzoli, D.; Giupponi, E.; Mantovani, D.; Candiani, G. Size matters for in vitro gene delivery: Investigating the relationships among complexation protocol, transfection medium, size and sedimentation. Sci. Rep. 2017, 7, 44134. [Google Scholar] [CrossRef] [PubMed]
- Zakeri, A.; Kouhbanani, M.A.J.; Beheshtkhoo, N.; Beigi, V.; Mousavi, S.M.; Hashemi, S.A.R.; Karimi Zade, A.; Amani, A.M.; Savardashtaki, A.; Mirzaei, E.; et al. Polyethylenimine-based nanocarriers in co-delivery of drug and gene: A developing horizon. Nano Rev. Exp. 2018, 9, 1488497. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Xin, X.; Qiu, L.; Li, W.; Guan, G.; Li, G.; Qiao, M.; Zhao, X.; Hu, H.; Chen, D. Co-delivery of p53 and MDM2 inhibitor RG7388 using a hydroxyl terminal PAMAM dendrimer derivative for synergistic cancer therapy. Acta Biomater. 2019, 100, 118–131. [Google Scholar] [CrossRef]
- Ghilardi, A.; Pezzoli, D.; Bellucci, M.C.; Malloggi, C.; Negri, A.; Sganappa, A.; Tedeschi, G.; Candiani, G.; Volonterio, A. Synthesis of multifunctional PAMAM-aminoglycoside conjugates with enhanced transfection efficiency. Bioconjug. Chem. 2013, 24, 1928–1936. [Google Scholar] [CrossRef]
- Dong, Y.; Yu, T.; Ding, L.; Laurini, E.; Huang, Y.; Zhang, M.; Weng, Y.; Lin, S.; Chen, P.; Marson, D.; et al. A Dual Targeting Dendrimer-Mediated siRNA Delivery System for Effective Gene Silencing in Cancer Therapy. J. Am. Chem. Soc. 2018, 140, 16264–16274. [Google Scholar] [CrossRef]
- Chen, C.; Posocco, P.; Liu, X.; Cheng, Q.; Laurini, E.; Zhou, J.; Liu, C.; Wang, Y.; Tang, J.; Col, V.D.; et al. Mastering Dendrimer Self-Assembly for Efficient siRNA Delivery: From Conceptual Design to In Vivo Efficient Gene Silencing. Small 2016, 12, 3667–3676. [Google Scholar] [CrossRef]
- Agarwal, S.; Zhang, Y.; Maji, S.; Greiner, A. PDMAEMA based gene delivery materials. Mater. Today 2012, 15, 388–393. [Google Scholar] [CrossRef]
- Köping-Höggård, M.; Vårum, K.M.; Issa, M.; Danielsen, S.; Christensen, B.E.; Stokke, B.T.; Artursson, P. Improved chitosan-mediated gene delivery based on easily dissociated chitosan polyplexes of highly defined chitosan oligomers. Gene Ther. 2004, 11, 1441–1452. [Google Scholar] [CrossRef] [PubMed]
- Chuan, D.; Jin, T.; Fan, R.; Zhou, L.; Guo, G. Chitosan for gene delivery: Methods for improvement and applications. Adv. Colloid Interface Sci. 2019, 268, 25–38. [Google Scholar] [CrossRef]
- Strand, S.P.; Lelu, S.; Reitan, N.K.; de Lange Davies, C.; Artursson, P.; Vårum, K.M. Molecular design of chitosan gene delivery systems with an optimized balance between polyplex stability and polyplex unpacking. Biomaterials 2010, 31, 975–987. [Google Scholar] [CrossRef]
- Rahmani, S.; Hakimi, S.; Esmaeily, A.; Samadi, F.Y.; Mortazavian, E.; Nazari, M.; Mohammadi, Z.; Tehrani, N.R.; Tehrani, M.R. Novel chitosan based nanoparticles as gene delivery systems to cancerous and noncancerous cells. Int. J. Pharm. 2019, 560, 306–314. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Cui, S.; Zhao, Y.; Zhang, C.; Zhang, S.; Peng, X. Grafting chitosan with polyethylenimine in an ionic liquid for efficient gene delivery. PLoS One 2015, 10, e0121817. [Google Scholar] [CrossRef]
- Pezzoli, D.; Tarsini, P.; Melone, L.; Candiani, G. RGD-derivatized PEI-PEG copolymers: Influence of the degree of substitution on the targeting behavior. J. Drug Deliv. Sci. Technol. 2017, 37, 115–122. [Google Scholar] [CrossRef]
- Bansal, R.; Seth, B.; Tiwari, S.; Jahan, S.; Kumari, M.; Pant, A.B.; Chaturvedi, R.K.; Kumar, P.; Gupta, K.C. Hexadecylated linear PEI self-assembled nanostructures as efficient vectors for neuronal gene delivery. Drug Deliv. Transl. Res. 2018, 8, 1436–1449. [Google Scholar] [CrossRef] [PubMed]
- Thapa, B.; Plianwong, S.; Remant Bahadur, K.; Rutherford, B.; Uludağ, H. Small hydrophobe substitution on polyethylenimine for plasmid DNA delivery: Optimal substitution is critical for effective delivery. Acta Biomater. 2016, 33, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Pack, D.W.; Hoffman, A.S.; Pun, S.; Stayton, P.S. Design and development of polymers for gene delivery. Nat. Rev. Drug Discov. 2005, 4, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Bahadur, K.C.R.; Uludağ, H. PEI and its derivatives for gene therapy. In Polymers and Nanomaterials for Gene Therapy; Elsevier Inc.: Amsterdam, The Netherlands, 2016; ISBN 9780081005217. [Google Scholar]
- Dai, Z.; Gjetting, T.; Mattebjerg, M.A.; Wu, C.; Andresen, T.L. Elucidating the interplay between DNA-condensing and free polycations in gene transfection through a mechanistic study of linear and branched PEI. Biomaterials 2011, 32, 8626–8634. [Google Scholar] [CrossRef] [PubMed]
- Godbey, W.T.; Wu, K.K.; Mikos, A.G. Size matters: Molecular weight affects the efficiency of poly(ethylenimine) as a gene delivery vehicle. J. Biomed. Mater. Res. 1999, 45, 268–275. [Google Scholar] [CrossRef]
- Fischer, D.; Bieber, T.; Li, Y.; Elsässer, H.P.; Kissel, T. A novel non-viral vector for DNA delivery based on low molecular weight, branched polyethylenimine: Effect of molecular weight on transfection efficiency and cytotoxicity. Pharm. Res. 1999, 16, 1273–1279. [Google Scholar] [CrossRef]
- Bonner, D.K.; Zhao, X.; Buss, H.; Langer, R.; Hammond, P.T. Crosslinked linear polyethylenimine enhances delivery of DNA to the cytoplasm. J. Control. Release 2013, 167, 101–107. [Google Scholar] [CrossRef]
- Yue, Y.; Jin, F.; Deng, R.; Cai, J.; Chen, Y.; Lin, M.C.M.; Kung, H.F.; Wu, C. Revisit complexation between DNA and polyethylenimine - Effect of uncomplexed chains free in the solution mixture on gene transfection. J. Control. Release 2011, 155, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Melone, L.; Rossi, B.; Pastori, N.; Panzeri, W.; Mele, A.; Punta, C. TEMPO-Oxidized Cellulose Cross-Linked with Branched Polyethyleneimine: Nanostructured Adsorbent Sponges for Water Remediation. Chempluschem 2015, 80, 1408–1415. [Google Scholar] [CrossRef] [PubMed]
- ISO/Guide 30:2015(en), Reference Materials—Selected Terms and Definitions. Available online: https://www.iso.org/obp/ui/#iso:std:iso:guide:30:ed-3:v1:en (accessed on 20 January 2020).
- ISO 17034:2016(en), General Requirements for the Competence of Reference Material Producers. Available online: https://www.iso.org/obp/ui/#iso:std:iso:17034:ed-1:v1:en (accessed on 20 January 2020).
- Plant, A.L.; Locascio, L.E.; May, W.E.; Gallagher, P.D. Improved reproducibility by assuring confidence in measurements in biomedical research. Nat. Methods 2014, 11, 895–898. [Google Scholar] [CrossRef]
- Ogris, M.; Steinlein, P.; Kursa, M.; Mechtler, K.; Kircheis, R.; Wagner, E. The size of DNA/transferrin-PEI complexes is an important factor for gene expression in cultured cells. Gene Ther. 1998, 5, 1425–1433. [Google Scholar] [CrossRef]
- Zhang, C.; Yadava, P.; Hughes, J. Polyethylenimine strategies for plasmid delivery to brain-derived cells. Methods 2004, 33, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Lucotti, A.; Tommasini, M.; Pezzoli, D.; Candiani, G. Molecular interactions of DNA with transfectants: A study based on infrared spectroscopy and quantum chemistry as aids to fluorescence spectroscopy and dynamic light scattering analyses. RSC Adv. 2014, 4, 49620–49627. [Google Scholar] [CrossRef]
- Breunig, M.; Lungwitz, U.; Liebl, R.; Klar, J.; Obermayer, B.; Blunk, T.; Goepferich, A. Mechanistic insights into linear polyethylenimine-mediated gene transfer. Biochim. Biophys. Acta - Gen. Subj. 2007, 1770, 196–205. [Google Scholar] [CrossRef]
- Breunig, M.; Lungwitz, U.; Liebl, R.; Fontanari, C.; Klar, J.; Kurtz, A.; Blunk, T.; Goepferich, A. Gene delivery with low molecular weight linear polyethylenimines. J. Gene Med. 2005, 7, 1287–1298. [Google Scholar] [CrossRef]
- Wiseman, J.W.; Goddard, C.A.; McLelland, D.; Colledge, W.H. A comparison of linear and branched polyethylenimine (PEI) with DCChol/DOPE liposomes for gene delivery to epithelial cells in vitro and in vivo. Gene Ther. 2003, 10, 1654–1662. [Google Scholar] [CrossRef]
- Zhang, H.; Chen, Z.; Du, M.; Li, Y.; Chen, Y. Enhanced gene transfection efficiency by low-dose 25 kDa polyethylenimine by the assistance of 1.8 kDa polyethylenimine. Drug Deliv. 2018, 25, 1740–1745. [Google Scholar] [CrossRef]
- Wightman, L.; Kircheis, R.; Rössler, V.; Garotta, S.; Ruzicka, R.; Kursa, M.; Wagner, E. Different behavior of branched and linear polyethylenimine for gene delivery in vitro and in vivo. J. Gene Med. 2001, 3, 362–372. [Google Scholar] [CrossRef] [PubMed]
- Forest, V.; Pourchez, J. Preferential binding of positive nanoparticles on cell membranes is due to electrostatic interactions: A too simplistic explanation that does not take into account the nanoparticle protein corona. Mater. Sci. Eng. C 2017, 70, 889–896. [Google Scholar] [CrossRef] [PubMed]
- Gallops, C.E.; Yu, C.; Ziebarth, J.D.; Wang, Y. Effect of the Protonation Level and Ionic Strength on the Structure of Linear Polyethyleneimine. ACS Omega 2019, 4, 7255–7264. [Google Scholar] [CrossRef]
- Lynch, I.; Dawson, K.A. Protein-nanoparticle interactions. Nano Today 2008, 3, 40–47. [Google Scholar] [CrossRef]
- Maiolo, D.; Colombo, J.; Beretta, J.; Malloggi, C.; Candiani, G.; Baldelli Bombelli, F. The polyplex, protein corona, cell interplay: Tips and drawbacks. Colloids Surfaces B Biointerfaces 2018, 168, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Troiber, C.; Kasper, J.C.; Milani, S.; Scheible, M.; Martin, I.; Schaubhut, F.; Küchler, S.; Rädler, J.; Simmel, F.C.; Friess, W.; et al. Comparison of four different particle sizing methods for siRNA polyplex characterization. Eur. J. Pharm. Biopharm. 2013, 84, 255–264. [Google Scholar] [CrossRef]
- Bhattacharjee, S. DLS and zeta potential - What they are and what they are not? J. Control. Release 2016, 235, 337–351. [Google Scholar] [CrossRef]
- Bertin, A. Polyelectrolyte complexes of DNA and polycations as gene delivery vectors. Adv. Polym. Sci. 2014, 256, 103–196. [Google Scholar]
- Van Gaal, E.V.B.; Van Eijk, R.; Oosting, R.S.; Kok, R.J.; Hennink, W.E.; Crommelin, D.J.A.; Mastrobattista, E. How to screen non-viral gene delivery systems in vitro? J. Control. Release 2011, 154, 218–232. [Google Scholar] [CrossRef]
- Wilke, M.; Fortunati, E.; van den Broek, M.; Hoogeveen, A.T.; Scholte, B.J. Efficacy of a peptide-based gene delivery system depends on mitotic activity. Gene Ther. 1996, 3, 1133–1142. [Google Scholar]
- Brunner, S.; Sauer, T.; Carotta, S.; Cotten, M.; Saltik, M.; Wagner, E. Cell cycle dependence of gene transfer by lipoplex polyplex and recombinant adenovirus. Gene Ther. 2000, 7, 401–407. [Google Scholar] [CrossRef] [PubMed]
- Grellier, M.; Bordenave, L.; Amédée, J. Cell-to-cell communication between osteogenic and endothelial lineages: Implications for tissue engineering. Trends Biotechnol. 2009, 27, 562–571. [Google Scholar] [CrossRef]
- Giupponi, E.; Visone, R.; Occhetta, P.; Colombo, F.; Rasponi, M.; Candiani, G. Development of a microfluidic platform for high-throughput screening of non-viral gene delivery vectors. Biotechnol. Bioeng. 2018, 115, 775–784. [Google Scholar] [CrossRef]
- Chernov, V.M.; Chernova, O.A.; Sanchez-Vega, J.T.; Kolpakov, A.I.; Ilinskaya, O.N. Mycoplasma contamination of cell cultures: Vesicular traffic in bacteria and control over infectious agents. Acta Naturae 2014, 6, 41–51. [Google Scholar] [CrossRef]
- Nikfarjam, L.; Farzaneh, P. Prevention and detection of mycoplasma contamination in cell culture. Cell J. 2012, 13, 203–212. [Google Scholar] [PubMed]
- Rottem, S.; Kosower, N.S.; Kornspan, J.D. Contamination of Tissue Cultures by Mycoplasmas. In Biomedical Tissue Culture; InTech: London, UK, 2012. [Google Scholar]
- Yin, Z.; Zhang, Y.; Liang, S.; Zhao, S.; Du, J.; Cheng, B. Mycoplasma contamination-mediated attenuation of plasmid DNA transfection efficiency is augmented via l-arginine deprivation in HEK-293 cells. J. Zhejiang Univ. B 2019, 20, 1021–1026. [Google Scholar] [CrossRef] [PubMed]
- Gedye, C.; Cardwell, T.; Dimopoulos, N.; Tan, B.S.; Jackson, H.; Svobodová, S.; Anaka, M.; Behren, A.; Maher, C.; Hofmann, O.; et al. Mycoplasma Infection Alters Cancer Stem Cell Properties in Vitro. Stem Cell Rev. Reports 2016, 12, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Freedman, L.P.; Gibson, M.C.; Ethier, S.P.; Soule, H.R.; Neve, R.M.; Reid, Y.A. Reproducibility: Changing the policies and culture of cell line authentication. Nat. Methods 2015, 12, 493–497. [Google Scholar] [CrossRef] [PubMed]
- Hopert, A.; Uphoff, C.C.; Wirth, M.; Hauser, H.; Drexler, H.G. Mycoplasma detection by PCR analysis. In Vitro Cell. Dev. Biol. Anim. 1993, 29A, 819–821. [Google Scholar] [CrossRef]
- Tang, J.; Hu, M.; Lee, S.; Roblin, R. A polymerase chain reaction based method for detecting Mycoplasma/Acholeplasma contaminants in cell culture. J. Microbiol. Methods 2000, 39, 121–126. [Google Scholar] [CrossRef]
- Dobrovolny, P.L.; Bess, D. Optimized PCR-based detection of mycoplasma. J. Vis. Exp. 2011. [Google Scholar] [CrossRef]
- Huh, S.H.; Do, H.J.; Lim, H.Y.; Kim, D.K.; Choi, S.J.; Song, H.; Kim, N.H.; Park, J.K.; Chang, W.K.; Chung, H.M.; et al. Optimization of 25 kDa linear polyethylenimine for efficient gene delivery. Biologicals 2007, 35, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Deng, R.; Yue, Y.; Jin, F.; Chen, Y.; Kung, H.F.; Lin, M.C.M.; Wu, C. Revisit the complexation of PEI and DNA - How to make low cytotoxic and highly efficient PEI gene transfection non-viral vectors with a controllable chain length and structure? J. Control. Release 2009, 140, 40–46. [Google Scholar] [CrossRef]
- Tripathi, S.K.; Goyal, R.; Kumar, P.; Gupta, K.C. Linear polyethylenimine-graft-chitosan copolymers as efficient DNA/siRNA delivery vectors in vitro and in vivo. Nanomedicine Nanotechnology, Biol. Med. 2012, 8, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Pezzoli, D.; Olimpieri, F.; Malloggi, C.; Bertini, S.; Volonterio, A.; Candiani, G. Chitosan-Graft-Branched Polyethylenimine Copolymers: Influence of Degree of Grafting on Transfection Behavior. PLoS One 2012, 7, e34711. [Google Scholar] [CrossRef]
- Nishikawa, M.; Huang, L. Nonviral vectors in the new millennium: Delivery barriers in gene transfer. Hum. Gene Ther. 2001, 12, 861–870. [Google Scholar] [CrossRef] [PubMed]
- Faruqui, N.; Kummrow, A.; Fu, B.; Divieto, C.; Rojas, F.; Kisulu, F.; Cavalcante, J.J.V.; Wang, J.; Campbell, J.; Martins, J.L.; et al. Cellular metrology: Scoping for a value proposition in extra-and intracellular measurements. Front. Bioeng. Biotechnol. 2019, 7, 456. [Google Scholar] [CrossRef]
- Roth, C.M. Quantitative Measurements and Rational Materials Design for Intracellular Delivery of Oligonucleotides. Biotechnol. Prog. 2008, 24, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Varga, C.M.; Hong, K.; Lauffenburger, D.A. Quantitative analysis of synthetic gene delivery vector design properties. Mol. Ther. 2001, 4, 438–446. [Google Scholar] [CrossRef]
- Welsh, S.; Kay, S.A. Reporter gene expression for monitoring gene transfer. Curr. Opin. Biotechnol. 1997, 8, 617–622. [Google Scholar] [CrossRef]
- Collins, S.; Morrissey, D.; Rajendran, S.; Casey, G.; Scallan, M.; Harrison, P.; OSullivan, G.; Tangney, M. Comparison of DNA Delivery and Expression Using Frequently Used Delivery Methods. In Gene Therapy—Developments and Future Perspectives; InTechOpen: London, UK, 2011. [Google Scholar]
- Marjanovič, I.; Kandušer, M.; Miklavčič, D.; Keber, M.M.; Pavlin, M. Comparison of Flow Cytometry, Fluorescence Microscopy and Spectrofluorometry for Analysis of Gene Electrotransfer Efficiency. J. Membr. Biol. 2014, 247, 1259–1267. [Google Scholar] [CrossRef] [PubMed]
- Rathenberg, J.; Nevian, T.; Witzemann, V. High-efficiency transfection of individual neurons using modified electrophysiology techniques. J. Neurosci. Methods 2003, 126, 91–98. [Google Scholar] [CrossRef]
- Peng, L.; Xiong, W.; Cai, Y.; Chen, Y.; He, Y.; Yang, J.; Jin, J.; Li, H. A simple, rapid method for evaluation of transfection efficiency based on fluorescent dye. Bioengineered 2017, 8, 225–231. [Google Scholar] [CrossRef]
- Goedhart, J.; van Weeren, L.; Adjobo-Hermans, M.J.W.; Elzenaar, I.; Hink, M.A.; Gadella, T.W.J. Quantitative Co-expression of proteins at the single cell level - application to a multimeric FRET sensor. PLoS One 2011, 6, e27321. [Google Scholar] [CrossRef] [PubMed]
- Armknecht, S.; Boutros, M.; Kiger, A.; Nybakken, K.; Mathey-Prevot, B.; Perrimon, N. High-throughput RNA interference screens in Drosophila tissue culture cells. In Methods Enzymology; Elsevier Inc.: Amsterdam, The Netherlands, 2005; Volume 392, pp. 55–73. [Google Scholar]
- Yun, C.; Dasgupta, R. Luciferase reporter assay in Drosophila and mammalian tissue culture cells. Curr. Protoc. Chem. Biol. 2014, 6, 7–23. [Google Scholar] [CrossRef] [PubMed]
- Carter, M.; Shieh, J. Biochemical Assays and Intracellular Signaling. In Guide to Research Techniques in Neuroscience; Elsevier: Amsterdam, The Netherlands, 2015; pp. 311–343. [Google Scholar]
- Mosaad, E.O.; Futrega, K.; Seim, I.; Gloss, B.; Chambers, K.F.; Clements, J.A.; Doran, M.R. Constraints to counting bioluminescence producing cells by a commonly used transgene promoter and its implications for experimental design. Sci. Rep. 2019, 9, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.Q.; Chen, J.L.; Lv, T.F.; He, C.X.; Tang, G.P.; Liang, W.Q.; Tabata, Y.; Gao, J.Q. N/P ratio significantly influences the transfection efficiency and cytotoxicity of a polyethylenimine/chitosan/DNA complex. Biol. Pharm. Bull. 2009, 32, 706–710. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wagner, E. History of Polymeric Gene Delivery Systems. Top. Curr. Chem. 2017, 375, 1–39. [Google Scholar]
- il Kim, T.; Bai, C.Z.; Nam, K.; Park, J. Comparison between arginine conjugated PAMAM dendrimers with structural diversity for gene delivery systems. J. Control. Release 2009, 136, 132–139. [Google Scholar] [CrossRef]
- Kroll, A.; Pillukat, M.H.; Hahn, D.; Schnekenburger, J. Current in vitro methods in nanoparticle risk assessment: Limitations and challenges. Eur. J. Pharm. Biopharm. 2009, 72, 370–377. [Google Scholar] [CrossRef]
- Schulze, C.; Kroll, A.; Lehr, C.M.; Schäfer, U.F.; Becker, K.; Schnekenburger, J.; Schulze Isfort, C.; Landsiedel, R.; Wohlleben, W. Not ready to use—Overcoming pitfalls when dispersing nanoparticles in physiological media. Nanotoxicology 2008, 2, 51–61. [Google Scholar] [CrossRef]
- Hamid, R.; Rotshteyn, Y.; Rabadi, L.; Parikh, R.; Bullock, P. Comparison of alamar blue and MTT assays for high through-put screening. Toxicol. Vitr. 2004, 18, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Salem, A.K.; Searson, P.C.; Leong, K.W. Multifunctional nanorods for gene delivery. Nat. Mater. 2003, 2, 668–671. [Google Scholar] [CrossRef]
- Xiao, Y.P.; Zhang, J.; Liu, Y.H.; Zhang, J.H.; Yu, Q.Y.; Huang, Z.; Yu, X.Q. Low molecular weight PEI-based fluorinated polymers for efficient gene delivery. Eur. J. Med. Chem. 2019, 602–611. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, Y.; Wang, M.; Xiao, J.; Cheng, Y. Fluorinated poly(propylenimine) dendrimers as gene vectors. Biomaterials 2014, 35, 5407–5413. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Chen, J.X.; Sun, Y.X.; Deng, J.Z.; Li, C.; Zhang, X.Z.; Zhuo, R.X. Construction of cell penetrating peptide vectors with N-terminal stearylated nuclear localization signal for targeted delivery of DNA into the cell nuclei. J. Control. Release 2011, 155, 26–33. [Google Scholar] [CrossRef] [PubMed]

| Nucleic Acid | Description | Site of Action | Applications/Pathway |
|---|---|---|---|
| pDNAs (also called chimeras) [1,9] | large circular dsDNAs (<10 kbp) | nucleus | nuclear localization followed by transgene expression under specific promoters to induce protein expression |
| mRNAs [1,9] | large ssRNAs (<10 kbp) | cytosol | positive regulation of protein expression |
| short regulatory RNAs (siRNAs/miRNAs/shRNA) [1,9] | short regulatory RNA (15–30 nt) | cytosol | RNA interference mechanisms to shorten mRNA half-life and downregulate translation |
| ASOs [10] | short DNA, RNA or analogs (15–30 nt) | cytosol and nucleus | RNA alteration to reduce, restore, or modify protein expression |
| Strategy | Description | Pros | Cons | |
|---|---|---|---|---|
| Physical/mechanical methods [27,29,42] | electroporation | application of an electric field by voltage pulses to induce transient cell membrane poration | high efficiency; low costs; high reproducibility; ability to transfer large size DNA | tissue/cell damage; invasiveness; some DNA instability |
| sonoporation | use of highly-focused ultrasounds to trigger transient cell membrane poration | non-invasiveness; possibility to be used in combination with microbubbles/non-viral vectors | low efficiency; low reproducibility; tissue/cell damage | |
| optoporation | use of short ultra-focused laser pulses to induce transient cell membrane poration | high efficiency; high spatial precision | tissue/cell damage; low irradiation area; poor penetration of the laser pulses | |
| magnetofection | application of a magnetic field to ease the transfer of NAs-coated paramagnetic particles into cells | high efficiency; non-invasiveness; possibility to be used in combination with non-viral vectors | poor efficiency with naked DNA; possible agglomeration of magnetic particles | |
| microinjection | direct injection of NAs into single cells by means of a needle | high efficiency; simplicity; reproducibility; low cytotoxicity; ability to transfer large size DNA | time consuming; inability to transfect large number of cells | |
| gene gun | propulsion of NAs-coated particles towards the target site | high efficiency; safety | tissue/cell damage; poor penetration of particles | |
| Viral vectors [2,39,43,44] | adenoviruses (AdVs) | non-enveloped dsDNA–virus able to carry ≤8 kbp DNA | efficient in a broad range of host cells | high immunogenicity; transient expression |
| adeno-associated viruses (AAVs) | non-enveloped recombinant ssDNA–virus with a small carrying capacity (≤4 kbp) | efficient in a broad range of host cells; non-inflammatory/pathogenic | small carrying capacity | |
| retroviruses | enveloped ssRNA-carrying virus with ≤8 kbp RNA capacity | long-term expression | limited tropism to dividing cells; random integration | |
| lentiviruses | enveloped ssRNA-carrying virus with ≤8 kbp RNA capacity | efficient in a broad range of host cells; long-term expression | potential oncogenic responses | |
| herpes simplex viruses (HSV)-1 | enveloped dsDNA–virus with >30 kbp carrying capacity | large packing capacity; efficient in a broad range of host cells | potential inflammatory responses; transient expression | |
| Non-viral vectors [11,45,46,47,48] | inorganic nanoparticles | metal-based nanoparticles of different size and shapes | possibility of functionalization; low cytotoxicity | instability; toxicity |
| cation lipids | lipids able to self-assemble with NAs to give lipoplexes | tunable features; safety; low cytotoxicity | low transfection efficiency | |
| cationic polymers | polymers able to self-assemble with NAs to give polyplexes | tunable features; possibility of functionalization; mild cytotoxicity; stability in protein-rich media; low cytotoxicity | low transfection efficiency | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bono, N.; Ponti, F.; Mantovani, D.; Candiani, G. Non-Viral in Vitro Gene Delivery: It is Now Time to Set the Bar! Pharmaceutics 2020, 12, 183. https://doi.org/10.3390/pharmaceutics12020183
Bono N, Ponti F, Mantovani D, Candiani G. Non-Viral in Vitro Gene Delivery: It is Now Time to Set the Bar! Pharmaceutics. 2020; 12(2):183. https://doi.org/10.3390/pharmaceutics12020183
Chicago/Turabian StyleBono, Nina, Federica Ponti, Diego Mantovani, and Gabriele Candiani. 2020. "Non-Viral in Vitro Gene Delivery: It is Now Time to Set the Bar!" Pharmaceutics 12, no. 2: 183. https://doi.org/10.3390/pharmaceutics12020183
APA StyleBono, N., Ponti, F., Mantovani, D., & Candiani, G. (2020). Non-Viral in Vitro Gene Delivery: It is Now Time to Set the Bar! Pharmaceutics, 12(2), 183. https://doi.org/10.3390/pharmaceutics12020183