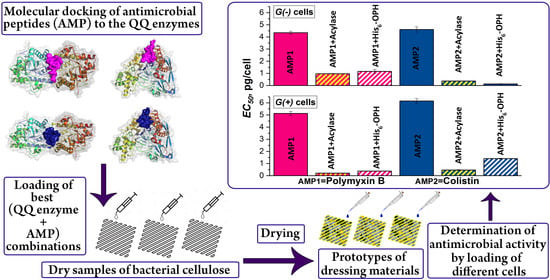
Bacterial Cellulose Containing Combinations of Antimicrobial Peptides with Various QQ Enzymes as a Prototype of an “Enhanced Antibacterial” Dressing: In Silico and In Vitro Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Antibacterial Combinations Based on Antibacterial Agents and Enzymes Loaded onto Bacterial Cellulose (BC)
2.3. Water Absorption and Scanning Electron Microscopy (SEM) Analysis
2.4. Computational Methods
3. Results
3.1. Enzymes for Hydrolysis of G(−) Bacteria Signal Molecules
3.1.1. Physical–Chemical Interactions in Simulated Models of Enzyme–AMP Combinations
3.1.2. Antimicrobial Activity of AMP Combinations with AHL-Hydrolyzing Enzymes
3.2. Enzymes for Hydrolysis of G(+) Bacteria Signal Molecules in Combination with AMPs
3.2.1. Physical–Chemical Interactions in Simulated Models of AM Combinations with Peptidases
3.2.2. Antimicrobial Activity of BC Samples Loaded with AMPs in Combination with Peptidases
3.3. Absorption Capacity of BC Samples
3.4. Scanning Electron Microscopy of the Surface of BC Samples
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AHL | N-acyl homoserine lactone |
| AIP | Autoinducing peptide |
| AMP | Antimicrobial peptide |
| BC | Bacterial cellulose |
| CFU | Colony-forming unit |
| His6-OPH | Hexahistidine-tagged organophosphorus hydrolase |
| Quorum quenching | |
| PON2 | Paraoxonase 2 |
References
- Picheth, G.F.; Pirich, C.L.; Sierakowski, M.R.; Woehl, M.A.; Sakakibara, C.N.; de Souza, C.F.; Martin, A.A.; da Silva, R.; de Freitas, R.A. Bacterial cellulose in biomedical applications: A review. Int. J. Biol. Macromol. 2017, 104, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Portela, R.; Leal, C.R.; Almeida, P.L.; Sobral, R.G. Bacterial cellulose: A versatile biopolymer for wound dressing applications. Microb. Biotechnol. 2019, 12, 586–610. [Google Scholar] [CrossRef] [PubMed]
- Gorgieva, S.; Trček, J. Bacterial Cellulose: Production, Modification and Perspectives in Biomedical Applications. Nanomaterials 2019, 9, 1352. [Google Scholar] [CrossRef]
- Lemnaru, G.M.; Truşcă, R.D.; Ilie, C.I.; Țiplea, R.E.; Ficai, D.; Oprea, O.; Stoica-Guzun, A.; Ficai, A.; Dițu, L.M. Antibacterial activity of bacterial cellulose loaded with bacitracin and amoxicillin: In vitro studies. Molecules 2020, 25, 4069. [Google Scholar] [CrossRef]
- Fürsatz, M.; Skog, M.; Sivlér, P.; Palm, E.; Aronsson, C.; Skallberg, A.; Greczynski, G.; Khalaf, H.; Bengtsson, T.; Aili, D. Functionalization of bacterial cellulose wound dressings with the antimicrobial peptide ε-poly-L-Lysine. Biomed. Mater. 2018, 13, 025014. [Google Scholar] [CrossRef] [PubMed]
- Pasaribu, K.M.; Gea, S.; Ilyas, S.; Tamrin, T.; Radecka, I. Characterization of Bacterial Cellulose-Based Wound Dressing in Different Order Impregnation of Chitosan and Collagen. Biomolecules 2020, 10, 1511. [Google Scholar] [CrossRef]
- Savitskaya, I.S.; Shokatayeva, D.H.; Kistaubayeva, A.S.; Ignatova, L.V.; Digel, I.E. Antimicrobial and wound healing properties of a bacterial cellulose based material containing B. subtilis cells. Heliyon 2019, 5, e02592. [Google Scholar] [CrossRef]
- World Health Organization. Antibacterial Agents in Clinical Development: An Analysis of the Antibacterial Clinical Development Pipeline, Including Tuberculosis (WHO/EMP/IAU/2017.11). Available online: https://www.who.int/medicines/areas/rational_use/antibacterial_agents_clinical_development/en/ (accessed on 9 September 2020).
- Tobias, N.J.; Brehm, J.; Kresovic, D.; Brameyer, S.; Bode, H.B.; Heermann, R. New vocabulary for bacterial communication. ChemBioChem 2020, 21, 759–768. [Google Scholar] [CrossRef]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588. [Google Scholar] [CrossRef] [PubMed]
- Monnet, V.; Gardan, R. Quorum-sensing regulators in Gram-positive bacteria: ‘Cherchez le Peptide’. Mol. Microbiol. 2015, 97, 181–184. [Google Scholar] [CrossRef]
- Vogel, J.; Quax, W.J. Enzymatic Quorum Quenching in Biofilms. In Quorum Sensing: Molecular Mechanism and Biotechnological Application; Tommonaro, G., Ed.; Academic Press: London, UK, 2019; pp. 173–193. [Google Scholar]
- Fetzner, S. Quorum quenching enzymes. J. Biotechnol. 2015, 201, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Sirotkina, M.; Efremenko, E.N. Rhodococcus lactonase with organophosphate hydrolase (OPH) activity and His6-tagged OPH with lactonase activity: Evolutionary proximity of the enzymes and new possibilities in their application. Appl. Microbiol. Biotechnol. 2014, 98, 2647–2656. [Google Scholar] [CrossRef]
- Aslanli, A.; Lyagin, I.; Efremenko, E. Novel approach to quorum quenching: Rational design of antibacterials in combination with hexahistidine-tagged organophosphorus hydrolase. Biol. Chem. 2018, 399, 869–879. [Google Scholar] [CrossRef]
- Maslova, O.; Aslanli, A.; Stepanov, N.; Lyagin, I.; Efremenko, E. Catalytic characteristics of new antibacterials based on hexahistidine-containing organophosphorus hydrolase. Catalysts 2017, 7, 271. [Google Scholar] [CrossRef]
- Aslanli, A.; Efremenko, E. Simultaneous molecular docking of different ligands to His6-tagged organophosphorus hydrolase as an effective tool for assessing their effect on the enzyme. PeerJ 2019, 7, e7684. [Google Scholar] [CrossRef]
- Efremenko, E.N.; Lyagin, I.V.; Klyachko, N.L.; Bronich, T.; Zavyalova, N.V.; Jiang, Y.; Kabanov, A.V. A simple and highly effective catalytic nanozyme scavenger for organophosphorus neurotoxins. J. Control. Release 2017, 247, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Lyagin, I.V.; Efremenko, E.N. Biomolecular engineering of biocatalysts hydrolyzing neurotoxic organophosphates. Biochimie 2018, 144, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Aslanli, A.; Lyagin, I.; Efremenko, E. Charges’ interaction in polyelectrolyte (nano)complexing of His6-OPH with peptides: Unpredictable results due to imperfect or useless concept? Int. J. Biol. Macromol. 2019, 140, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Aslanli, A.; Stepanov, N.; Razheva, T.; Podorozhko, E.A.; Lyagin, I.; Lozinsky, V.I.; Efremenko, E.N. enzymatically functionalized composite materials based on nanocellulose and poly(vinyl alcohol) cryogel and possessing antimicrobial activity. Materials 2019, 12, 3619. [Google Scholar] [CrossRef]
- Efremenko, E.; Votchitseva, Y.; Plieva, F.; Galaev, I.; Mattiasson, B. Purification of His6-organophosphate hydrolase using monolithic supermacroporous polyacrylamide cryogels developed for immobilized metal affinity chromatography. Appl. Microbiol. Biotechnol. 2006, 70, 558–563. [Google Scholar] [CrossRef] [PubMed]
- Stepanov, N.A.; Efremenko, E.N. “Deceived” concentrated immobilized cells as biocatalyst for intensive bacterial cellulose production from various sources. Catalysts 2018, 8, 33. [Google Scholar] [CrossRef]
- Votchitseva, Y.A.; Efremenko, E.N.; Aliev, T.K.; Varfolomeyev, S.D. Properties of hexahistidine-tagged organophosphate hydrolase. Biochemistry (Moscow) 2006, 71, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Hashida, Y.; Inouye, K. Molecular mechanism of the inhibitory effect of cobalt ion on thermolysin activity and the suppressive effect of calcium ion on the cobalt ion-dependent inactivation of thermolysin. J. Biochem. 2007, 141, 879–888. [Google Scholar] [CrossRef] [PubMed]
- Frolov, G.; Lyagin, I.; Senko, O.; Stepanov, N.; Pogorelsky, I.; Efremenko, E. Metal nanoparticles for improving bactericide functionality of usual fibers. Nanomaterials 2020, 10, 1724. [Google Scholar] [CrossRef]
- Stepanov, N.; Senko, O.; Perminova, I.; Efremenko, E. A new approach to assess the effect of various humic compounds on the metabolic activity of cells participating in methanogenesis. Sustainability 2019, 11, 3158. [Google Scholar] [CrossRef]
- Ismayilov, I.T.; Stepanov, N.A.; Efremenko, E.N.; Abbasov, V.M. Evaluation of biocidal properties of vegetable oil-based corrosion inhibitors using bioluminescent enzymatic method. Moscow Univ. Chem. Bull. 2015, 70, 197–201. [Google Scholar] [CrossRef]
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc. 2010, 5, 725–738. [Google Scholar] [CrossRef] [PubMed]
- Mandrich, L.; Cerreta, M.; Manco, G. An engineered version of human PON2 opens the way to understand the role of its post-translational modifications in modulating catalytic activity. PLoS ONE 2015, 10, e0144579. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.A.; Sept, D.; Joseph, S.; Holst, M.J.; McCammon, J.A. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 2001, 98, 10037–10041. [Google Scholar] [CrossRef]
- Dolinsky, T.J.; Czodrowski, P.; Li, H.; Nielsen, J.E.; Jensen, J.H.; Klebe, G.; Baker, N.A. PDB2PQR: Expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 2007, 35 (suppl. 2), W522–W525. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and Auto-DockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Sadovnichy, V.; Tikhonravov, A.; Voevodin, V.; Opanasenko, V. University. In Contemporary High Performance Computing: From Petascale toward Exascale; Vetter, J.S., Ed.; CRC Press: Boca Raton, FL, USA, 2013; pp. 283–307. [Google Scholar]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Kusada, H.; Tamaki, H.; Kamagata, Y.; Hanada, S.; Kimura, N. A novel quorum-quenching N-acylhomoserine lactone acylase from Acidovorax sp. strain MR-S7 mediates antibiotic resistance. Appl. Environ. Microbiol. 2017, 83, e00080-17. [Google Scholar] [CrossRef] [PubMed]
- Tapeinou, A.; Matsoukas, M.T.; Simal, C.; Tselios, T. Review cyclic peptides on a merry-go-round; towards drug design. Biopolymers 2015, 104, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Kaye, K.S.; Pogue, J.M.; Tran, T.B.; Nation, R.L.; Li, J. Agents of last resort: Polymyxin resistance. Infect. Dis. Clin. N. Am. 2016, 30, 391–414. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Salvesen, G. Handbook of Proteolytic Enzymes, 3rd ed.; Academic Press: London, UK, 2013. [Google Scholar] [CrossRef]
- Brown, P.; Abdulle, O.; Boakes, S.; Duperchy, E.; Moss, S.; Simonovic, M.; Stanway, S.; Wilson, A.; Dawson, M.J. Direct modifications of the cyclic peptide polymyxin B leading to analogues with enhanced in vitro antibacterial activity. Bioorg. Med. Chem. Lett. 2020, 30, 127163. [Google Scholar] [CrossRef] [PubMed]






| Antimicrobial Agent | Enzyme | Affinity, kJ × mol−1 | Occupied Area, % | ||
|---|---|---|---|---|---|
| Mean | Median | Total | Near Active Sites | ||
| Dermicidin | Acylase | −20.4 | −20.1 ± 0.9 | 13.5 | 0.6 |
| Lactonase | −14.6 | −15.1 ± 1.0 | 30.7 | 0.7 | |
| PON2 | −14.7 | −14.6 ± 0.5 | 16.8 | 0 | |
| His6-OPH | −15.2 | −15.1 ± 0.9 | 10.0 | 0.1 | |
| Polymyxin B | Acylase | −25.3 | −24.7 ± 1.5 | 3.0 | 0 |
| Lactonase | −28.6 | −27.8 ± 1.9 | 11.8 | 0.5 | |
| PON2 | −25.9 | −26.1 ± 1.0 | 7.6 | 0 | |
| His6-OPH | −23.0 | −22.6 ± 1.1 | 7.6 | 0.4 | |
| Colistin | Acylase | −25.2 | −24.5 ± 1.4 | 7.2 | 0.2 |
| Lactonase | −22.9 | −22.8 ± 1.4 | 12.9 | 0.5 | |
| PON2 | −19.5 | −19.2 ± 0.8 | 16.3 | 0 | |
| His6-OPH | −21.5 | −21.3 ± 1.0 | 9.6 | 0.1 | |
| Oritavancin | Acylase | −41.6 | −41.8 ± 1.5 | 11.4 | 0.6 |
| Lactonase | −35.5 | −34.9 ± 2.2 | 18.9 | 0.6 | |
| PON2 | −29.4 | −29.1 ± 0.7 | 20.1 | 0 | |
| His6-OPH | −33.7 | −33.5 ± 0.9 | 7.5 | 0.1 | |
| Indolicidin | Acylase | −32.4 | −32.2 ± 1.0 | 10.2 | 0.5 |
| Lactonase | −31.7 | −31.6 ± 1.2 | 15.1 | 0.7 | |
| PON2 | −28.7 | −28.2 ± 1.0 | 19.5 | 0 | |
| His6-OPH | −32.3 | −32.0 ± 1.2 | 7.8 | 0.6 | |
| Temporin A | Acylase | −25.3 | −25.3 ± 0.4 | 9.2 | 0.6 |
| Lactonase | −26.1 | −25.9 ± 0.6 | 15.8 | 0.7 | |
| PON2 | −26.6 | −26.6 ± 0.5 | 16.5 | 0 | |
| His6-OPH | −25.2 | −25.1 ± 0.2 | 9.6 | 0.1 | |
| G(+) Bacterial Cells:Bacillus subtilisB-522 | ||||
| QQ Enzyme | Colistin | Polymyxin B | Indolicidin | Temporin A |
| None | 6.16 ± 0.19 | 5.13 ± 0.16 | 66.72 ± 0.43 | 7.90 ± 0.06 |
| His6-OPH | 1.41 ± 0.03 | 0.38 ± 0.01 | 12.24 ± 0.23 | 5.31 ± 0.01 |
| Penicillin acylase | 0.47 ± 0.03 | 0.21 ± 0.01 | ||
| Thermolysin | 614 ± 13 | 364 ± 12 | ||
| Carboxypeptidase A | 14,166 ± 94 | 1684 ± 53 | ||
| G(−) Bacterial Cells: Pseudomonas sp. 78G | ||||
| None | 4.61 ± 0.22 | 4.35 ± 0.11 | 6.93 ± 0.18 | 2.24 ± 0.03 |
| His6-OPH | 0.13 ± 0.02 | 1.16 ± 0.16 | 2.61 ± 0.24 | 1.18 ± 0.02 |
| Penicillin acylase | 0.37 ± 0.02 | 0.97 ± 0.02 | ||
| Thermolysin | 17.28 ± 0.49 | 3.14 ± 0.32 | ||
| Carboxypeptidase A | 7.05 ± 0.19 | 33.28 ± 1.39 | ||
| Peptidase (Source) [39] | Molar Mass; Optimal pH | * Preferable Peptide Bond for Cleavage |
|---|---|---|
| Coccolysin (Enterococcus faecalis) | 31.5 kDa; pH 6–8 | P1’ = Leu, Phe, Ile or Ala |
| Griselysin (Streptomyces griseus) | 35.1 kDa; pH ~7 | P1, P1’ = hydrophobic residues |
| Stearolysin (Geobacillus stearothermophilus) | 34 kDa; neutral pH | P1, P1’ = hydrophobic residues |
| Mycolysin (Streptomyces sp.) | 37.1 kDa; pH 7–8.5 | P1’ = hydrophobic residues |
| Microcystinase (Sphingomonadaceae sp.) | 36 kDa; pH 6.5–8.4 | Arg–Adda bond ** |
| Carboxypeptidase A (bovine pancreas) | 34.6 kDa; pH 7–9 | C-terminal hydrophobic residues (incl. acylated ones) |
| Antimicrobial Agent | Enzyme | Affinity, kJ × mol−1 | Occupied Area, % | ||
|---|---|---|---|---|---|
| Mean | Median | Total | Near Active Sites | ||
| Dermicidin | Coccolysin | −14.4 | −14.2 ± 0.6 | 32.5 | 0.1 |
| Griselysin | −13.1 | −13.0 ± 0.6 | 16.4 | 0 | |
| Stearolysin | −16.4 | −16.3 ± 0.3 | 10.8 | 0 | |
| Mycolysin | −12.3 | −12.3 ± 0.5 | 14.8 | 0.7 | |
| Microcystinase | −13.4 | −13.4 ± 1.0 | 21.7 | 0 | |
| Carboxypeptidase A | −18.7 | −18.4 ± 1.2 | 35.6 | 1.9 | |
| Polymyxin B | Coccolysin | −29.1 | −29.1 ± 1.5 | 9.1 | 0.3 |
| Griselysin | −30.5 | −30.3 ± 1.0 | 17.3 | 0.2 | |
| Stearolysin | −35.0 | −35.1 ± 0.6 | 6.9 | 0 | |
| Mycolysin | −32.3 | −32.2 ± 1.4 | 10.8 | 0.4 | |
| Microcystinase | −26.8 | −26.8 ± 0.5 | 11.0 | 0.5 | |
| Carboxypeptidase A | −26.3 | −26.6 ± 2.1 | 13.5 | 1.9 | |
| Colistin | Coccolysin | −27.5 | −27.2 ± 1.4 | 9.5 | 0.3 |
| Griselysin | −28.2 | −27.6 ± 1.9 | 18.8 | 0.2 | |
| Stearolysin | −31.7 | −31.6 ± 1.2 | 5.4 | 0 | |
| Mycolysin | −26.3 | −26.1 ± 0.6 | 12.6 | 0.5 | |
| Microcystinase | −27.6 | −27.0 ± 2.1 | 10.3 | 0.6 | |
| Carboxypeptidase A | −21.7 | −21.7 ± 0.9 | 7.6 | 1.9 | |
| Oritavancin | Coccolysin | −39.0 | −38.5 ± 1.7 | 15.2 | 0.4 |
| Griselysin | −39.0 | −38.9 ± 1.9 | 18.9 | 0.2 | |
| Stearolysin | −47.4 | −47.1 ± 2.5 | 11.2 | 0.1 | |
| Mycolysin | −34.7 | −34.3 ± 1.0 | 10.8 | 0.5 | |
| Microcystinase | −38.0 | −37.6 ± 0.5 | 13.9 | 0.2 | |
| Carboxypeptidase A | −36.7 | −36.6 ± 1.6 | 11.2 | 1.7 | |
| Indolicidin | Coccolysin | −36.3 | −36.6 ± 1.1 | 12.0 | 0.4 |
| Griselysin | −29.3 | −29.3 ± 0.9 | 18.6 | 0.3 | |
| Stearolysin | −43.9 | −43.7 ± 0.8 | 10.2 | 0 | |
| Mycolysin | −33.4 | −33.0 ± 0.9 | 6.9 | 0 | |
| Microcystinase | −36.8 | −36.8 ± 0.8 | 13.0 | 0.6 | |
| Carboxypeptidase A | −30.5 | −30.1 ± 1.3 | 14.0 | 1.9 | |
| Temporin A | Coccolysin | −26.3 | −26.3 ± 1.4 | 9.8 | 0.3 |
| Griselysin | −26.6 | −26.8 ± 0.4 | 11.4 | 0 | |
| Stearolysin | −29.8 | −29.9 ± 0.6 | 9.2 | 0 | |
| Mycolysin | −29.5 | −29.5 ± 1.6 | 8.7 | 0 | |
| Microcystinase | −31.5 | −31.4 ± 1.1 | 11.6 | 0.6 | |
| Carboxypeptidase A | −27.6 | −27.6 ± 0.8 | 12.1 | 1.9 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aslanli, A.; Lyagin, I.; Stepanov, N.; Presnov, D.; Efremenko, E. Bacterial Cellulose Containing Combinations of Antimicrobial Peptides with Various QQ Enzymes as a Prototype of an “Enhanced Antibacterial” Dressing: In Silico and In Vitro Data. Pharmaceutics 2020, 12, 1155. https://doi.org/10.3390/pharmaceutics12121155
Aslanli A, Lyagin I, Stepanov N, Presnov D, Efremenko E. Bacterial Cellulose Containing Combinations of Antimicrobial Peptides with Various QQ Enzymes as a Prototype of an “Enhanced Antibacterial” Dressing: In Silico and In Vitro Data. Pharmaceutics. 2020; 12(12):1155. https://doi.org/10.3390/pharmaceutics12121155
Chicago/Turabian StyleAslanli, Aysel, Ilya Lyagin, Nikolay Stepanov, Denis Presnov, and Elena Efremenko. 2020. "Bacterial Cellulose Containing Combinations of Antimicrobial Peptides with Various QQ Enzymes as a Prototype of an “Enhanced Antibacterial” Dressing: In Silico and In Vitro Data" Pharmaceutics 12, no. 12: 1155. https://doi.org/10.3390/pharmaceutics12121155
APA StyleAslanli, A., Lyagin, I., Stepanov, N., Presnov, D., & Efremenko, E. (2020). Bacterial Cellulose Containing Combinations of Antimicrobial Peptides with Various QQ Enzymes as a Prototype of an “Enhanced Antibacterial” Dressing: In Silico and In Vitro Data. Pharmaceutics, 12(12), 1155. https://doi.org/10.3390/pharmaceutics12121155