Engineered Disease Resistance in Cotton Using RNA-Interference to Knock down Cotton leaf curl Kokhran virus-Burewala and Cotton leaf curl Multan betasatellite Expression
Abstract
:1. Introduction
2. Materials and Methods
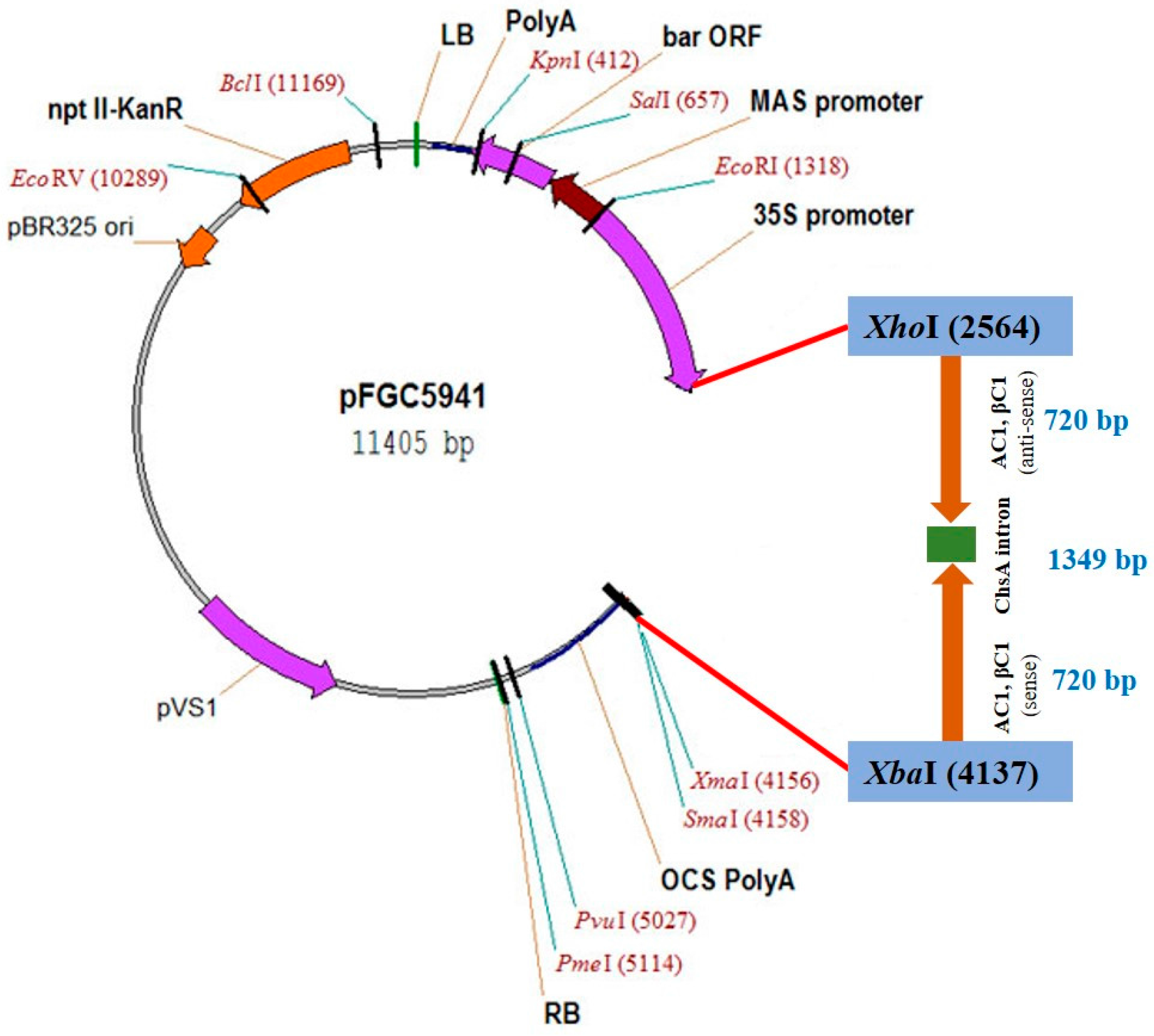
2.1. Plasmid Construction for siRNA/Intron Spliced Hairpin RNA Generation
2.2. Cotton Plant Transformation
2.3. Molecular Analysis of Transgenic Cotton Plants
2.4. Challenge-Inoculation of Transgenic Plants
2.5. Real-time Polymerase Chain Reaction Analysis
2.6. Fluorescence In Situ Hybridization
2.7. In Silico Analysis of dsRNA Hairpin Structure
3. Results
3.1. Genetic Transformation of Cotton
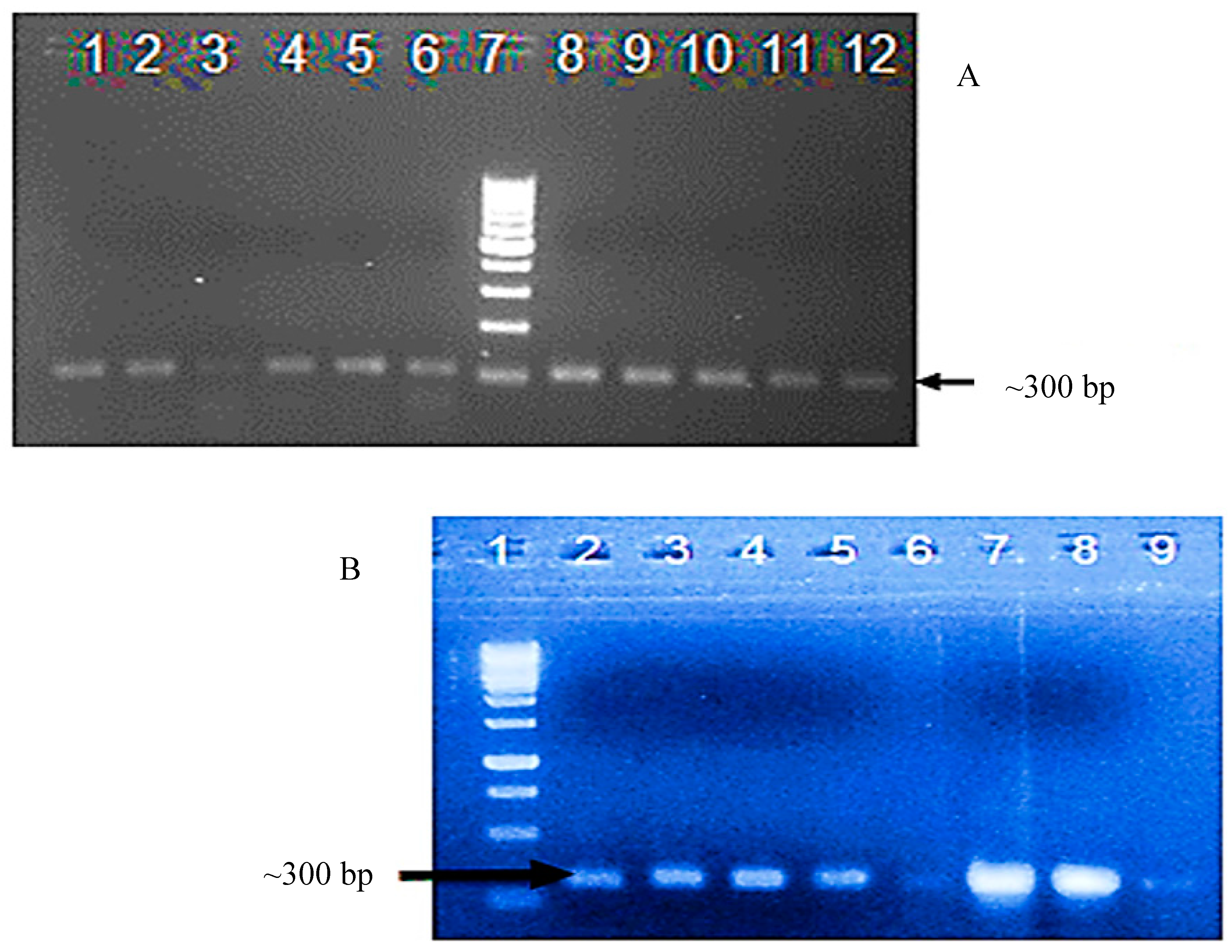
3.2. Confirmation of Transgene Presence in T0 and T1 Plants
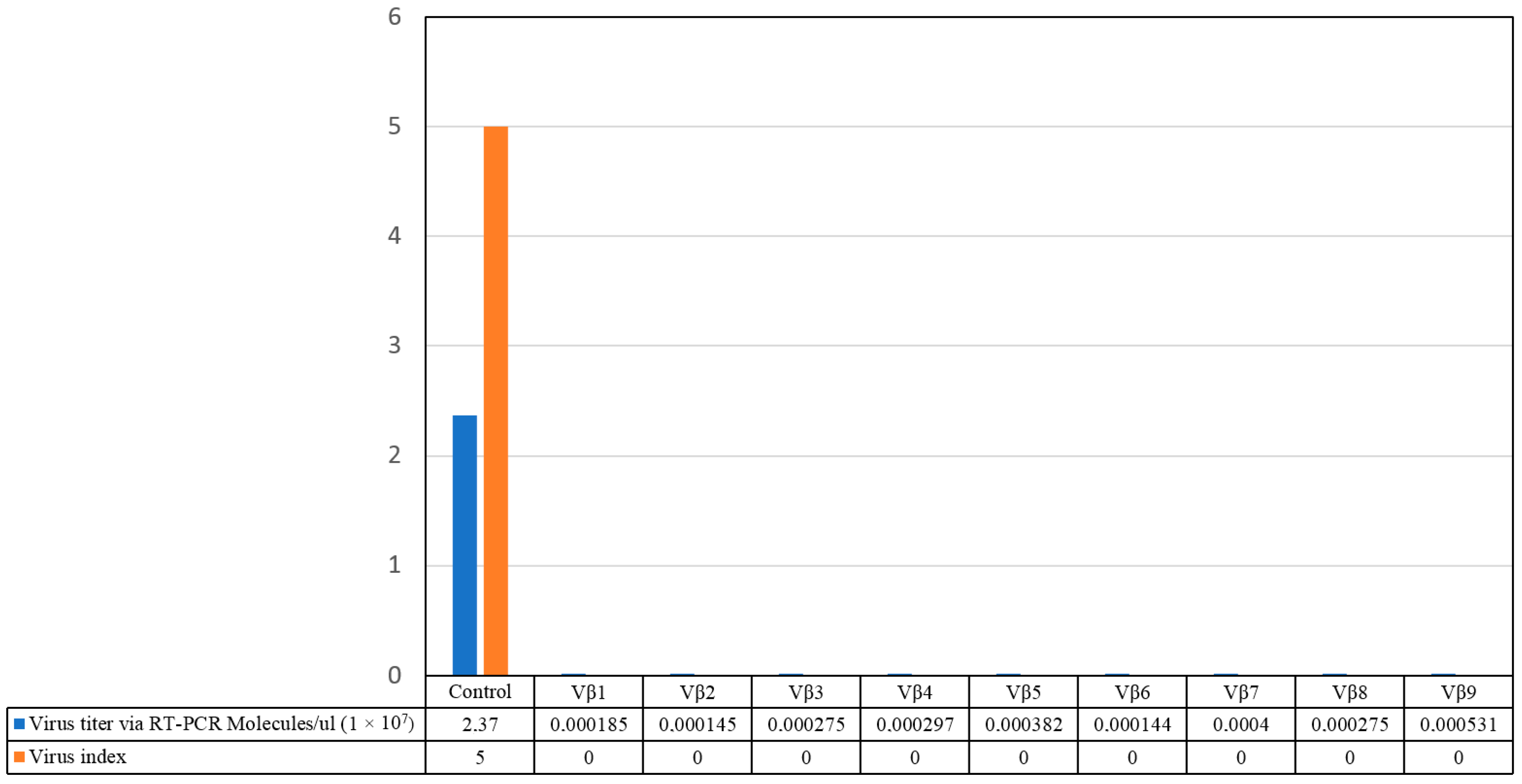
3.3. Challenge Inoculation of Transgenic Plants with CLCuKoV-Bu/CLCuMB by Whitefly Inoculation
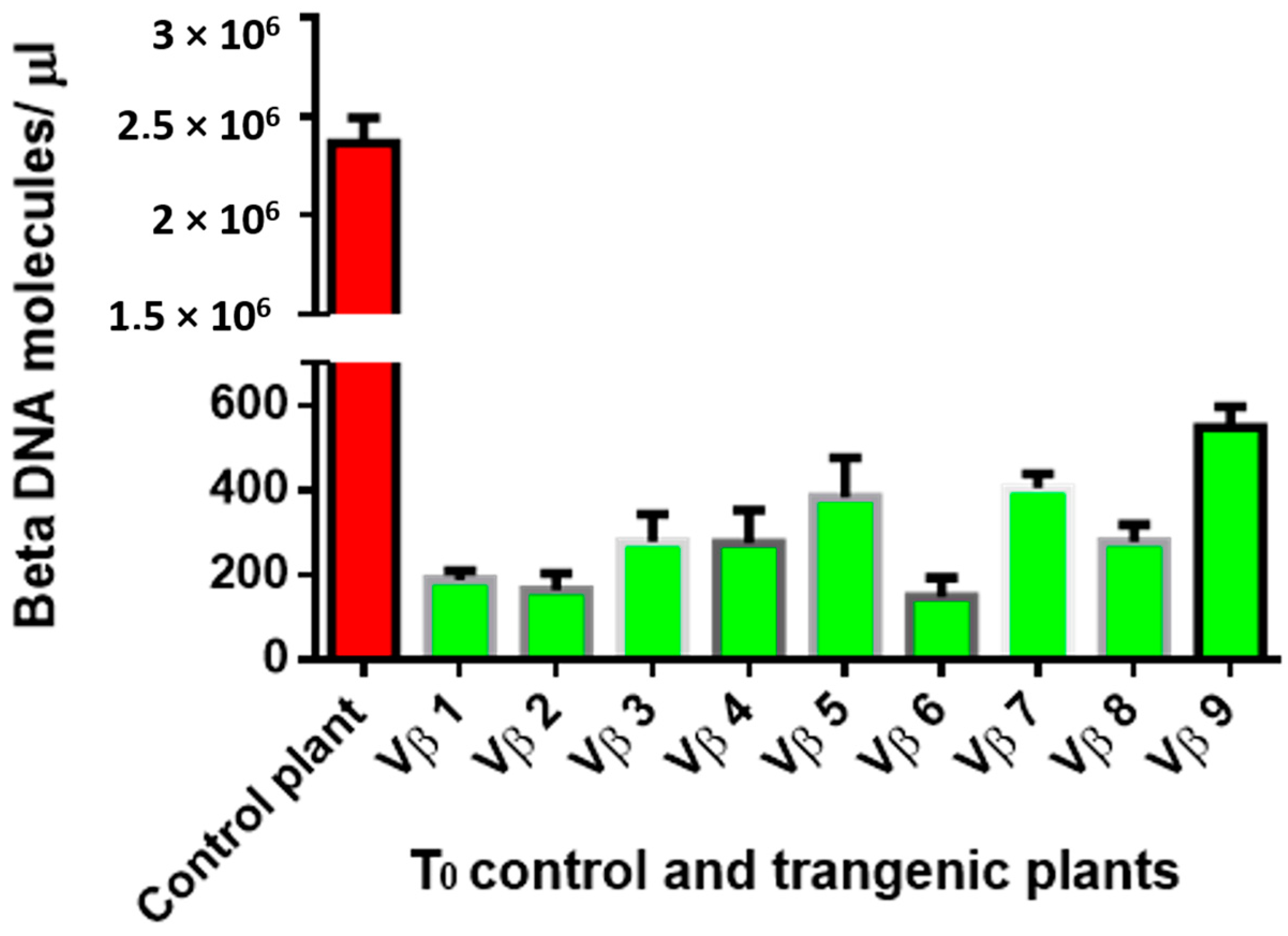
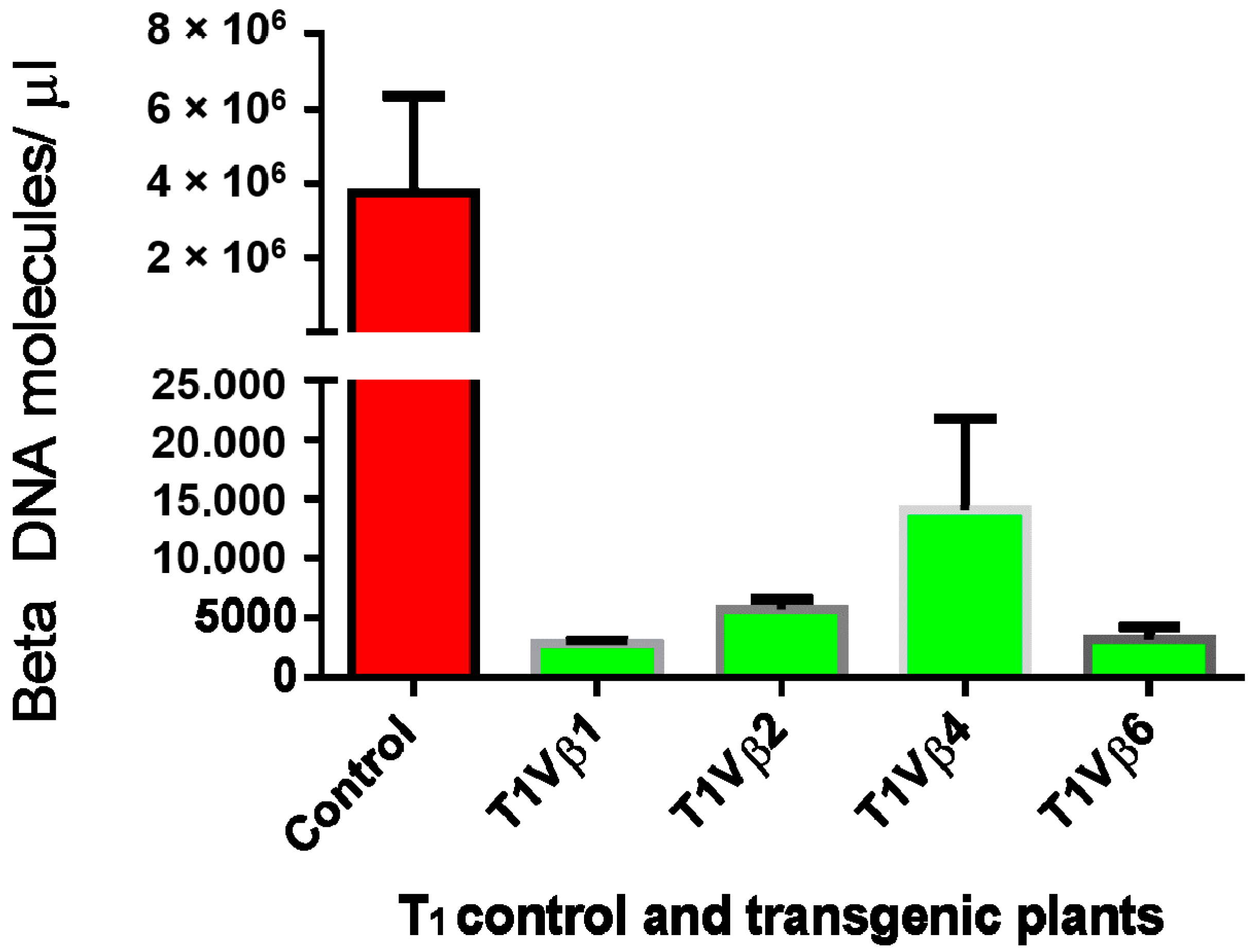
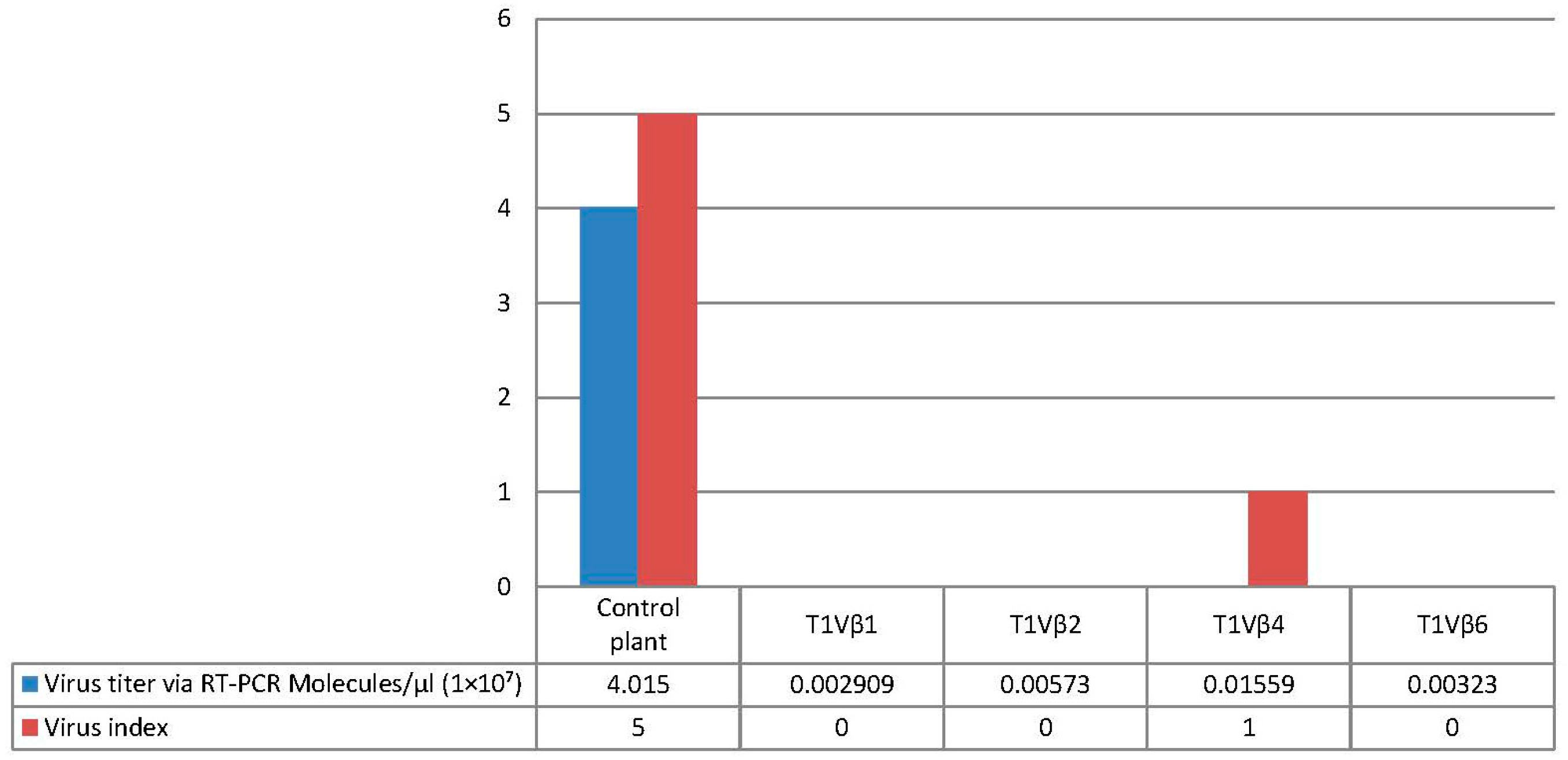
3.4. Betasatellite Accumulation in Transgenic and Non-Transgenic Cotton Plants, Post-Virus Inoculation Using Viruliferous Whiteflies
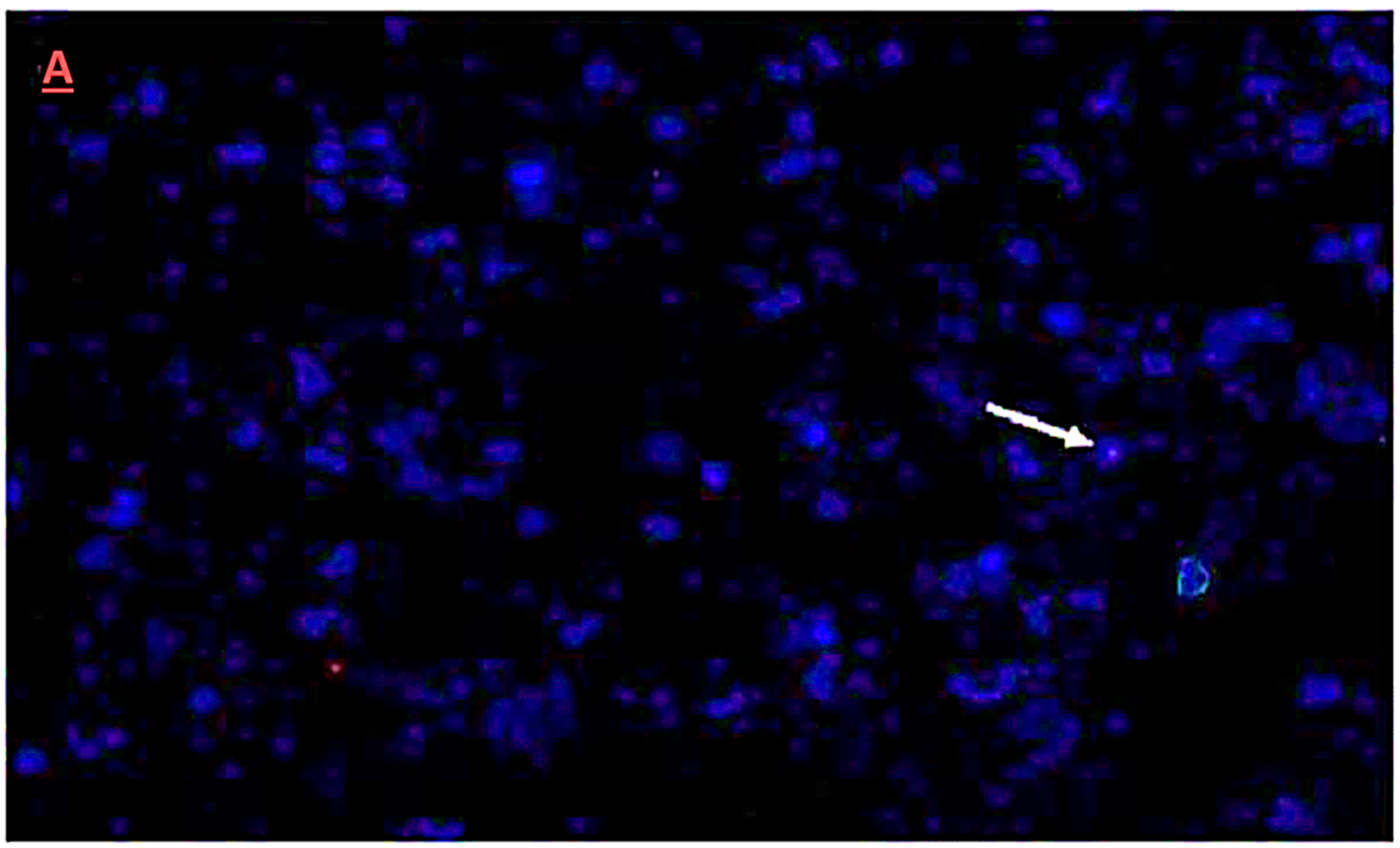
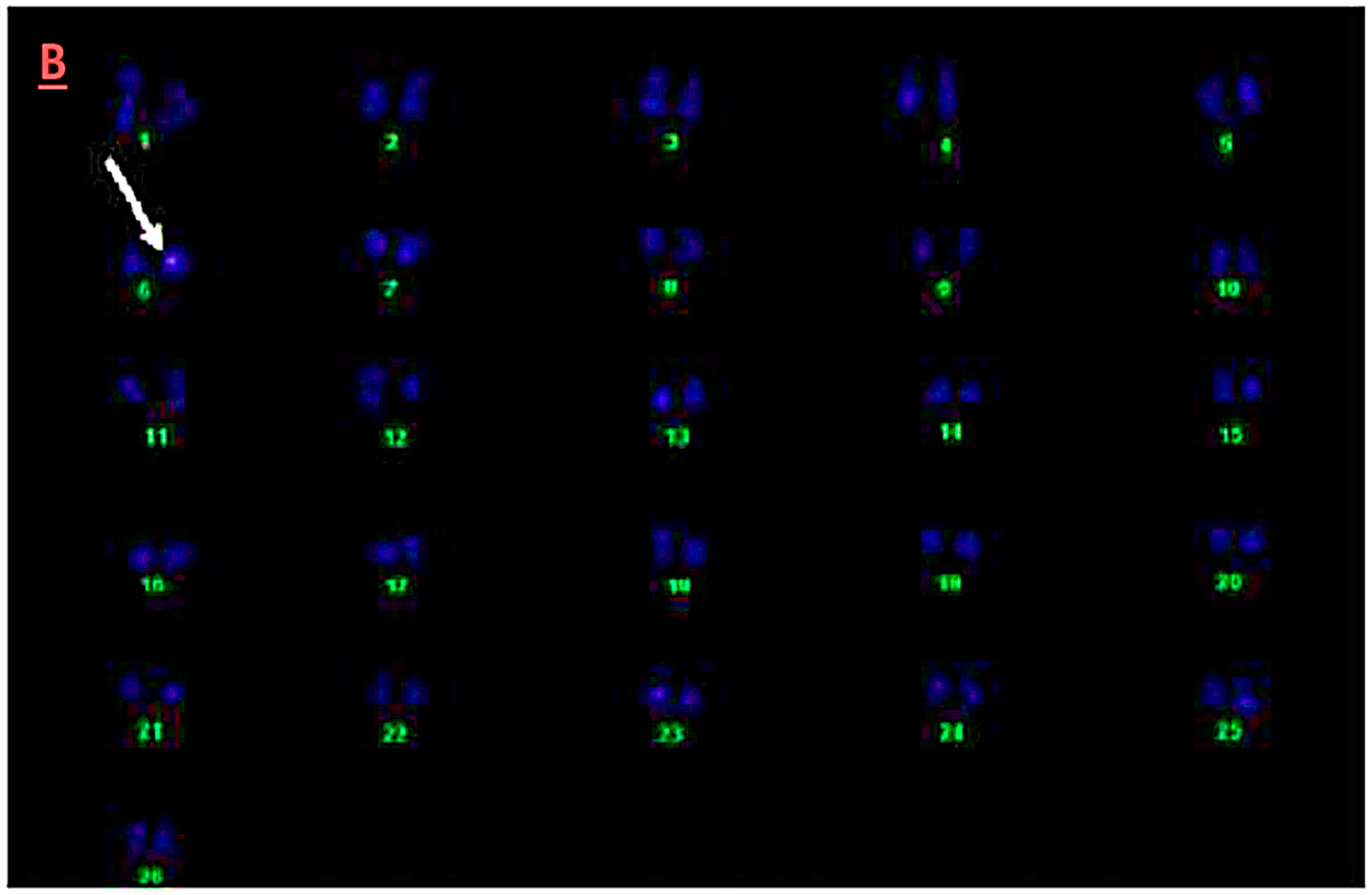
3.5. Fluorescence In-Situ Hybridization Analysis
3.6. Bioinformatic Analysis of the RNAi Construct
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Briddon, R.W.; Markham, P. Cotton leaf curl virus disease. Virus Res. 2000, 71, 151–159. [Google Scholar] [CrossRef]
- Farooq, A.; Farooq, J.; Mahmood, A.; Batool, A.; Rehman, A.; Shakeel, A.; Mehboob, S. An overview of cotton leaf curl virus disease (CLCuD) a serious threat to cotton productivity. Aust. J. Crop Sci. 2011, 5, 1823–1831. [Google Scholar]
- Farooq, J.; Farooq, A.; Riaz, M.; Shahid, M.R.; Saeed, F.; Iqbal, M.S.; Hussain, T.; Batool, A.; Mahmood, A. Cotton leaf curl virus disease a principle cause of decline in cotton productivity in Pakistan (a mini review). Can. J. Plant Prot. 2014, 2, 9–16. [Google Scholar]
- Ali, A.; Abdulai, A. The adoption of genetically modified cotton and poverty reduction in Pakistan. J. Agric. Econ. 2010, 61, 175–192. [Google Scholar] [CrossRef]
- Kerschen, A.; Napoli, C.A.; Jorgensen, R.A.; Müller, A.E. Effectiveness of RNA interference in transgenic plants. FEBS Lett. 2004, 566, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Kirthi, N.; Savithri, H.S. A conserved zinc finger motif in the coat protein of Tomato leaf curl Bangalore virus is responsible for binding to ssDNA. Arch. Virol. 2003, 148, 2369–2380. [Google Scholar] [CrossRef] [PubMed]
- Klahre, U.; Crete, P.; Leuenberger, S.A.; Iglesias, V.A.; Meins, F. High molecular weight RNAs and small interfering RNAs induce systemic posttranscriptional gene silencing in plants. Proc. Natl. Acad. Sci. USA 2002, 99, 11981–11986. [Google Scholar] [CrossRef] [PubMed]
- Hannon, G.J. RNA interference. Nature 2002, 418, 244–251. [Google Scholar] [CrossRef] [PubMed]
- Aragão, F.J.; Faria, J.C. First transgenic geminivirus-resistant plant in the field. Nat. Biotechnol. 2009, 27, 1086–1088. [Google Scholar] [CrossRef] [PubMed]
- Asad, S.; Haris, W.A.; Bashir, A.; Zafar, Y.; Malik, K.A.; Malik, N.N.; Lichtenstein, C.P. Transgenic tobacco expressing geminiviral RNAs are resistant to the serious viral pathogen causing cotton leaf curl disease. Arch. Virol. 2003, 148, 2341–2352. [Google Scholar] [CrossRef] [PubMed]
- Baulcombe, D. RNA silencing. Trends Biochem. Sci. 2005, 30, 290–293. [Google Scholar] [CrossRef] [PubMed]
- Shepherd, D.N.; Martin, D.P.; Thomson, J.A. Transgenic strategies for developing crops resistant to geminiviruses. Plant Sci. 2009, 176, 1–11. [Google Scholar] [CrossRef]
- Sohrab, S.S.; Kamal, M.A.; Ilah, A.; Husen, A.; Bhattacharya, P.S.; Rana, D. Development of Cotton leaf curl virus resistant transgenic cotton using antisense βC1 gene. Saudi J. Biol. Sci. 2014, 23, 358–362. [Google Scholar] [CrossRef] [PubMed]
- Muzaffar, A.; Kiani, S.; Khan, M.A.; Rao, A.Q.; Ali, A.; Awan, M.F.; Iqbal, A.; Nasir, I.A.; Shahid, A.A.; Husnain, T. Chloroplast localization of Cry1Ac and Cry2A protein-an alternative way of insect control in cotton. Biol. Res. 2015, 48, 14. [Google Scholar] [CrossRef] [PubMed]
- Malik, H.J.; Raza, A.; Amin, I.; Scheffler, J.A.; Scheffler, B.E.; Brown, J.K.; Mansoor, S. RNAi-mediated mortality of the whitefly through transgenic expression of double stranded RNA homologous to acetylcholinesterase and ecdysone receptor in tobacco plants. Sci. Rep. 2016, 6, 38469. [Google Scholar] [CrossRef] [PubMed]
- Vyas, M.; Raza, A.; Ali, A.M.; Ashraf, M.A.; Mansoor, S.; Ahmad, A.S.; Brown, J.K. Knock down of whitefly gut gene expression and mortality by orally delivered gut gene-specific dsRNA. PLoS ONE 2017, 12, e0168921. [Google Scholar] [CrossRef] [PubMed]
- Amin, I.; Hussain, K.; Akbergenov, R.; Yadav, J.S.; Qazi, J.; Mansoor, S.; Briddon, R.W. Suppressors of RNA silencing encoded by the components of the Cotton leaf curl begomovirus-Betasatellite complex. Mol. Plant Microbe Interact. 2011, 24, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Briddon, R.W.; Stanley, J. Subviral agents associated with plant single-stranded DNA viruses. Virology 2006, 344, 198–210. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Li, G.; Wang, D.; Hu, D.; Zhou, X. A begomovirus DNAβ-encoded protein binds DNA, functions as a suppressor of RNA silencing, and targets the cell nucleus. J. Virol. 2005, 79, 10764–10775. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.J.; Orozco, B.M.; Roe, J.L.; Nagar, S.; Ou, S.; Feiler, H.S.; Hanley-Bowdoin, L. A geminivirus replication protein interacts with the retinoblastoma protein through a novel domain to determine symptoms and tissue specificity of infection in plants. J. EMBO. 2000, 19, 3485–3495. [Google Scholar] [CrossRef] [PubMed]
- Laufs, J.; Schumacher, S.; Geisler, N.; Jupin, I.; Gronenborn, B. Identification of the nicking tyrosine of geminivirus Rep protein. FEBS Lett. 1995, 377, 258–262. [Google Scholar] [CrossRef]
- Saeed, M.; Behjatnia, S.A.; Mansoor, S.; Zafar, Y.; Hasnain, S.; Rezaian, M.A. A single complementary-sense transcript of a geminiviral DNA β satellite is determinant of pathogenicity. Mol. Plant Microbe Interact. 2005, 18, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.; Zafar, Y.; Randles, J.W.; Rezaian, M.A. A monopartite begomovirus-associated DNA β satellite substitutes for the DNA B of a bipartite begomovirus to permit systemic infection. J. Gen. Virol. 2007, 88, 2881–2889. [Google Scholar] [CrossRef] [PubMed]
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 1900, 2, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Rao, A.Q.; Bakhsh, A.; Kiani, S.; Shahzad, K.; Shahid, A.A.; Husnain, T.; Riazuddin, S. The myth of plant transformation. Biotechnol. Adv. 2009, 27, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Rao, A.Q.; Irfan, M.; Saleem, Z.; Husnain, T.; Riazuddin, S. Phytochrome B Gene of Arabidopsis thaliana increases the yield of cotton (Gossypiumhirsutum). J. Zhejiang Univ. Sci. B. 2011, 12, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Akhtar, K.P.; Ullah, R.; Khan, I.A.; Saeed, M.; Sarwar, N.; Mansoor, S. First symptomatic evidence of infection of Gossypiumarboreum with Cotton leaf curl Burewala virus through grafting. Int. J. Agric. Biol. 2013, 15, 157–160. [Google Scholar]
- Rahman, M.; Noreen, S.; Husnain, T.; Riazuddin, S. Fast and efficient method to determine the position of alien genes in transgenic plants. Emir. J. Food Agric. 2010, 22, 223–231. [Google Scholar]
- Rahman, N.; Khatoon, A.; Rahman, H. Studies on the development of spectrophotometric method for the determination of haloperidol in pharmaceutical preparations. Quím. Nova 2012, 35, 392–397. [Google Scholar] [CrossRef]
- In-Silico Project Support for Life Sciences. Available online: http://in-silico.net/tools/biology/sequence_conversion (accessed on 23 June 2016).
- Mathews, D.H.; Turner, D.H.; Watson, R.M. RNA secondary structure prediction. Curr. Protoc. Nucleic Acid Chem. 2007. [Google Scholar] [CrossRef]
- Predict a Secondary Structure Web Server. Available online: http://rna.urmc.rochester.edu/RNAstructureWeb/Servers/Predict1/Predict1.html (accessed on 23 January 2016).
- Akhtar, K.P.; Khan, A.I.; Hussain, M.; Khan, M.S.I. Comparison of resistance levels to Cotton leaf curl virus (CLCuV) among newly developed cotton mutants and commercial cultivars. J. Plant Pathol. 2002, 18, 179–186. [Google Scholar] [CrossRef]
- Ammara, U.; Mansoor, S.; Saeed, M.; Amin, I.; Briddon, R.; Al-Sadi, A. RNA interference-based resistance in transgenic tomato plants against-Oman (TYLCV-OM) and its associated betasatellite. J. Virol. 2015, 12, 38. [Google Scholar] [CrossRef] [PubMed]
| Experiment Number | Seeds (n) | Embryos (n) | Embryos Transferred to MS Media (n) | Embryos Transferred to Glass Tube (n) | Plants Transferred to Greenhouse (n) | Plants in the Field (n) |
|---|---|---|---|---|---|---|
| 1 | 20 g | 10 | 10 | 0 | 0 | 0 |
| 2 | 20 g | 30 | 28 | 28 | 5 | 1 |
| 3 | 20 g | 45 | 38 | 20 | 6 | 1 |
| 4 | 20 g | 70 | 70 | 12 | 1 | 0 |
| 5 | 40 g | 100 | 95 | 15 | 8 | 3 |
| 6 | 20 g | 35 | 30 | 0 | 0 | 0 |
| 7 | 40 g | 132 | 132 | 10 | 7 | 4 |
| 8 | 20 g | 10 | 10 | 10 | 2 | 1 |
| 9 | 20 g | 33 | 33 | 14 | 5 | 3 |
| Total | 220 g | 465 | 465 | 109 | 34 | 13 |
| Control | C-l | C-2 | C-3 | C-4 | C-5 | C-6 | C-7 | C-8 | C-9 |
| 6 | 6 | 4 | 4 | 4 | 5 | 5 | 6 | 5 | |
| T0 | Vβ1 | Vβ2 | Vβ3 | Vβ4 | Vβ5 | Vβ6 | Vβ7 | Vβ8 | Vβ9 |
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Control | C1-1 | C2-1 | C4-1 | C6-1 | |||||
| 5 | 5 | 5 | 5 | ||||||
| T1 | T1Vβ1 | T1Vβ2 | T1Vβ4 | T1Vβ6 | |||||
| 0 | 0 | 1 | 0 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahmad, A.; Zia-Ur-Rehman, M.; Hameed, U.; Qayyum Rao, A.; Ahad, A.; Yasmeen, A.; Akram, F.; Bajwa, K.S.; Scheffler, J.; Nasir, I.A.; et al. Engineered Disease Resistance in Cotton Using RNA-Interference to Knock down Cotton leaf curl Kokhran virus-Burewala and Cotton leaf curl Multan betasatellite Expression. Viruses 2017, 9, 257. https://doi.org/10.3390/v9090257
Ahmad A, Zia-Ur-Rehman M, Hameed U, Qayyum Rao A, Ahad A, Yasmeen A, Akram F, Bajwa KS, Scheffler J, Nasir IA, et al. Engineered Disease Resistance in Cotton Using RNA-Interference to Knock down Cotton leaf curl Kokhran virus-Burewala and Cotton leaf curl Multan betasatellite Expression. Viruses. 2017; 9(9):257. https://doi.org/10.3390/v9090257
Chicago/Turabian StyleAhmad, Aftab, Muhammad Zia-Ur-Rehman, Usman Hameed, Abdul Qayyum Rao, Ammara Ahad, Aneela Yasmeen, Faheem Akram, Kamran Shahzad Bajwa, Jodi Scheffler, Idrees Ahmad Nasir, and et al. 2017. "Engineered Disease Resistance in Cotton Using RNA-Interference to Knock down Cotton leaf curl Kokhran virus-Burewala and Cotton leaf curl Multan betasatellite Expression" Viruses 9, no. 9: 257. https://doi.org/10.3390/v9090257
APA StyleAhmad, A., Zia-Ur-Rehman, M., Hameed, U., Qayyum Rao, A., Ahad, A., Yasmeen, A., Akram, F., Bajwa, K. S., Scheffler, J., Nasir, I. A., Shahid, A. A., Iqbal, M. J., Husnain, T., Haider, M. S., & Brown, J. K. (2017). Engineered Disease Resistance in Cotton Using RNA-Interference to Knock down Cotton leaf curl Kokhran virus-Burewala and Cotton leaf curl Multan betasatellite Expression. Viruses, 9(9), 257. https://doi.org/10.3390/v9090257





