Perspectives on the Evolution of Porcine Parvovirus
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection, Extraction of Viral DNA, Detection, Sequencing, and Isolation of Porcine Parvovirus
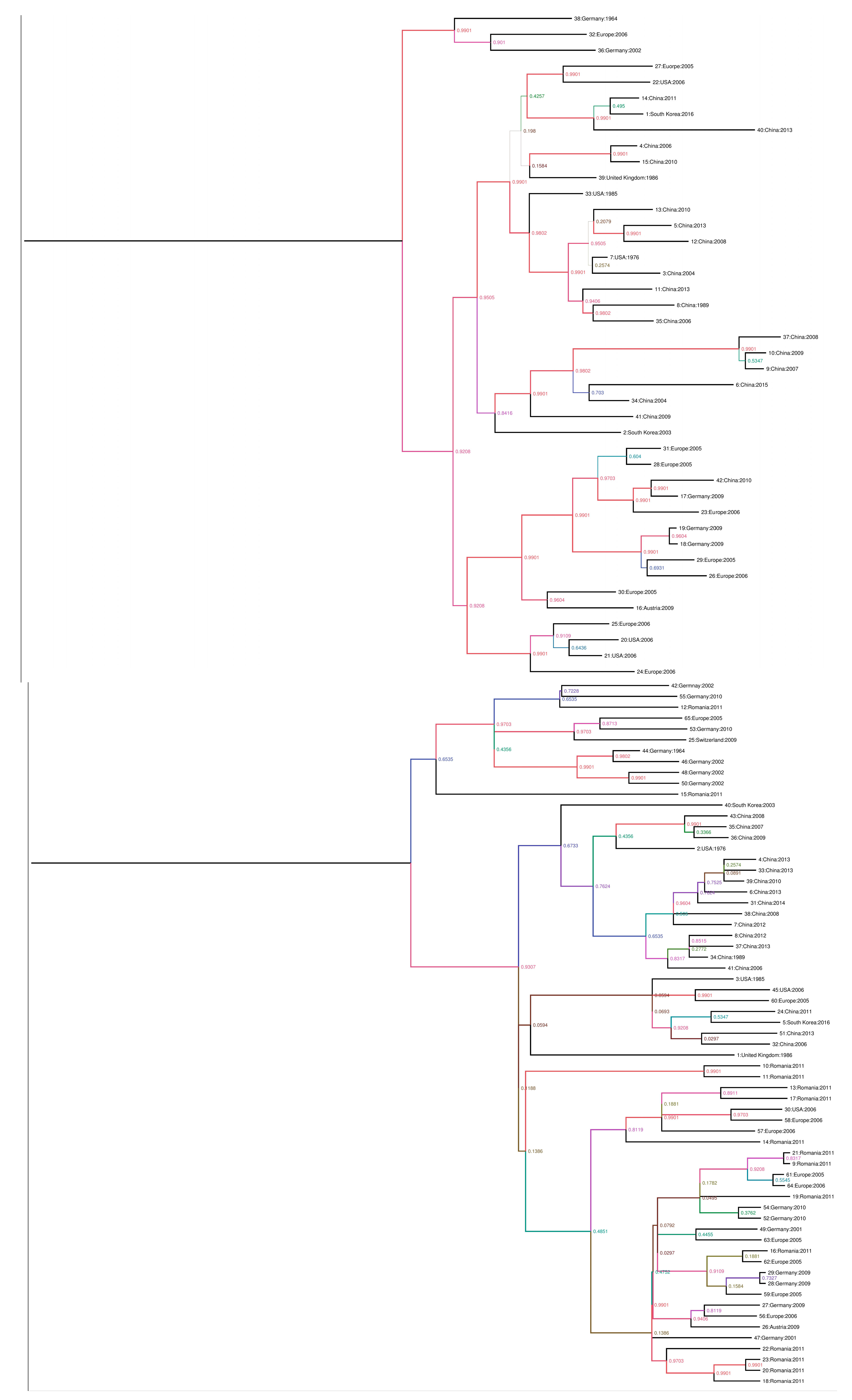
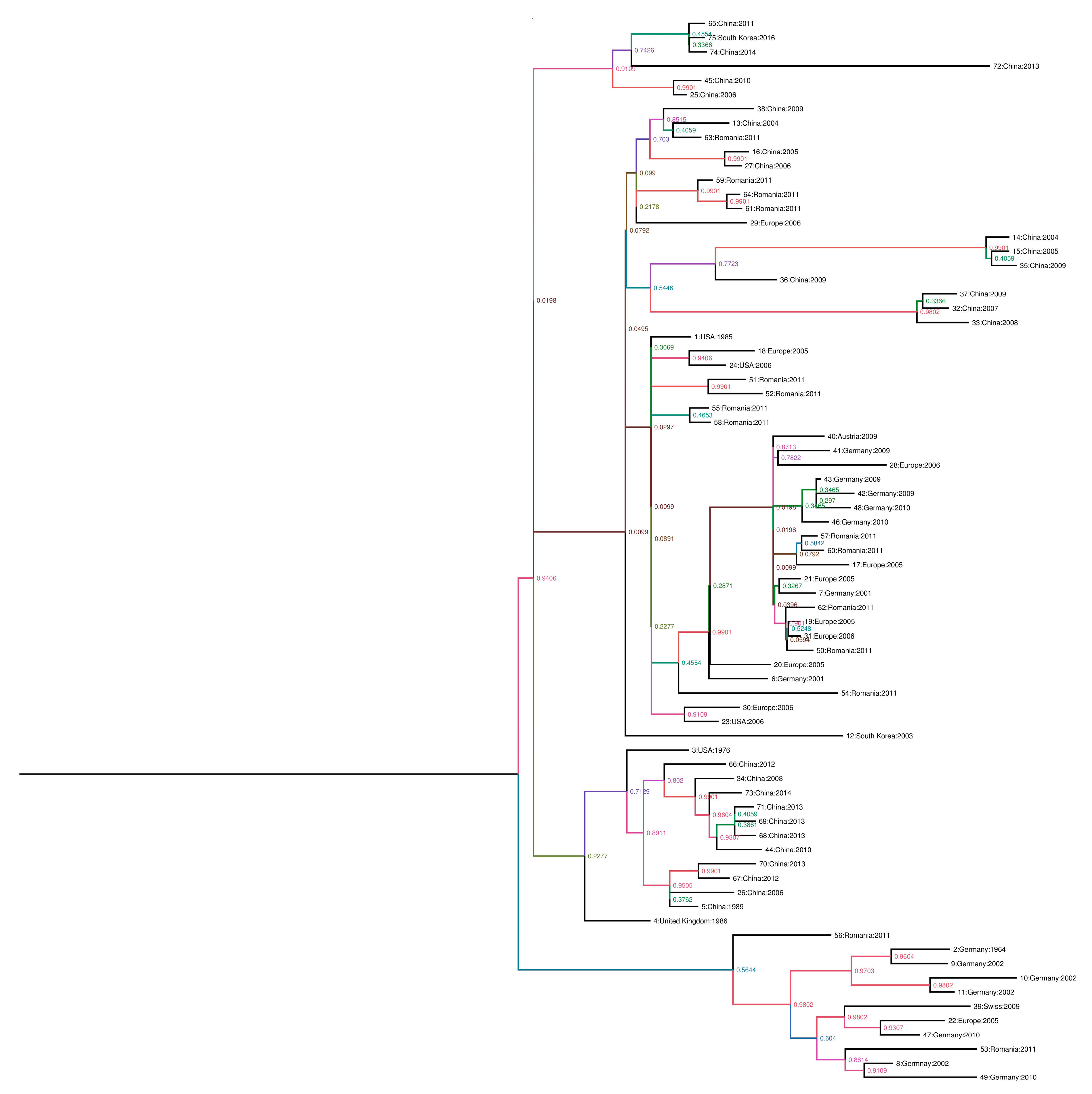
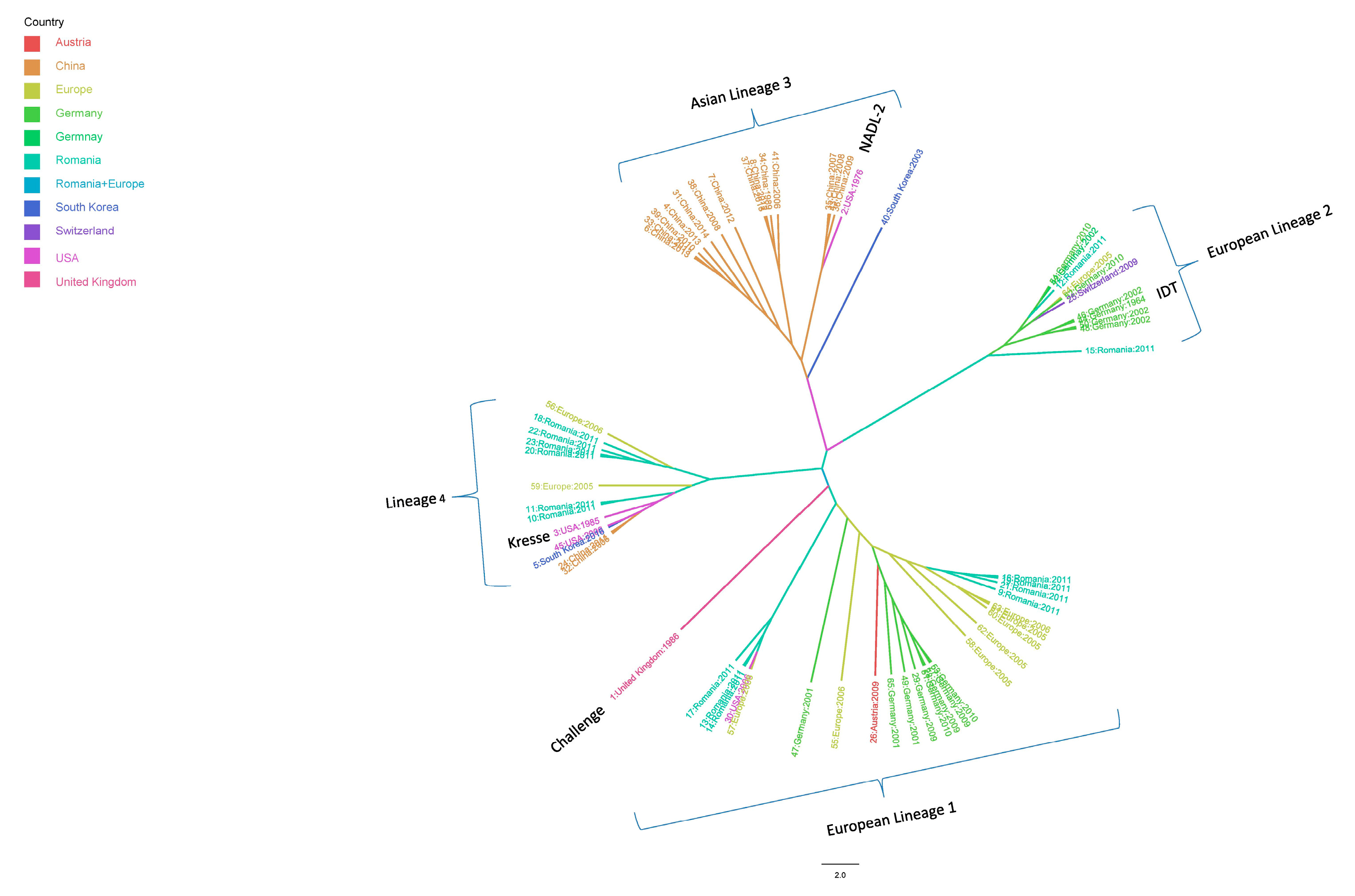
2.2. Phylogenic Analysis and Evolutionary Rate Estimation
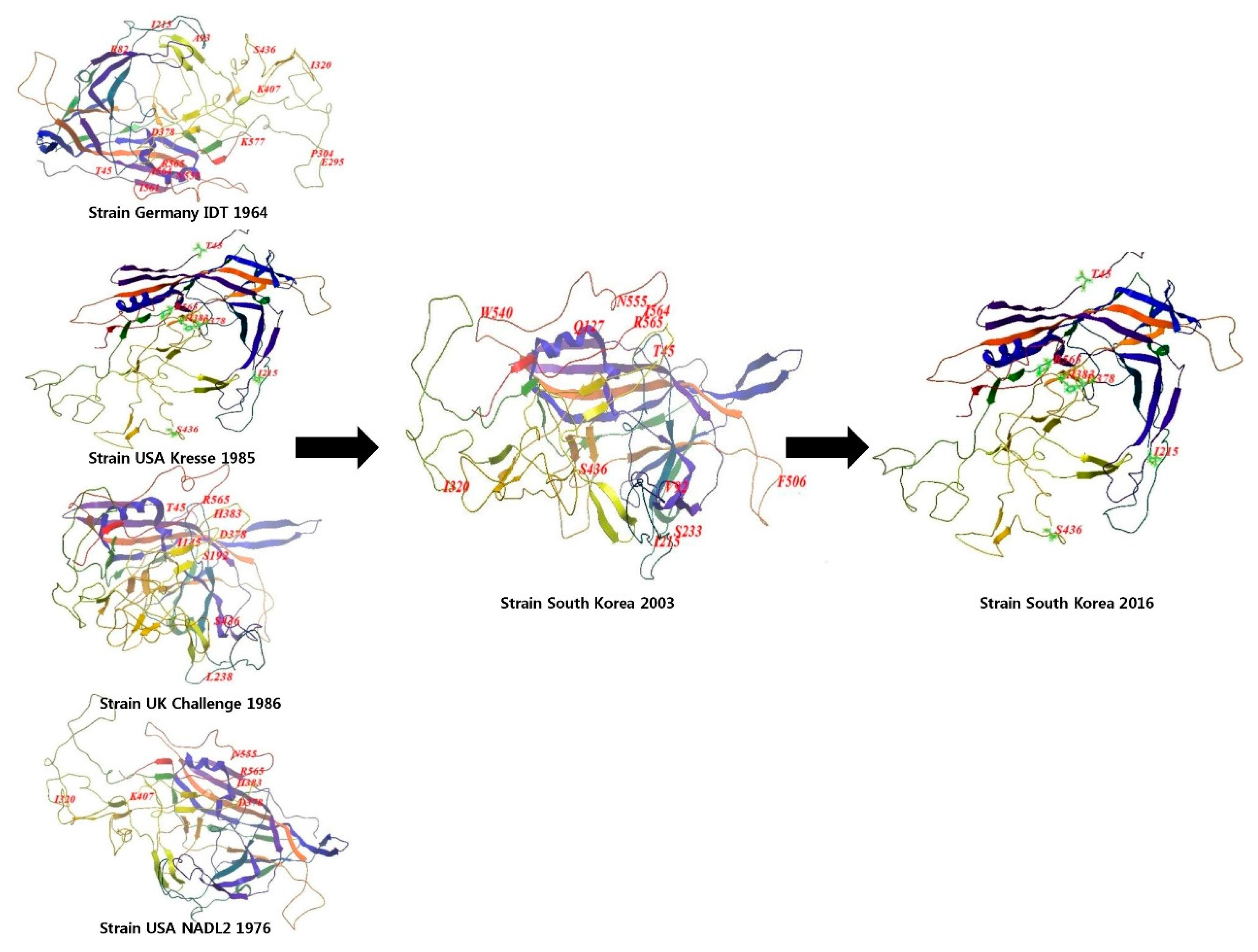
2.3. Molecular Structure of the T142_South Korea Strain
2.4. Recombination Analysis and Estimates of Amino Acid Mutation
3. Results and Discussion
3.1. Nucleic Acid Detection of Porcine Parvovirus and Phylogenetic Analysis
3.2. Isolation and Characterization of Strain T142_South Korea
3.3. Recombination and Structural Analysis of Porcine Parvovirus Strain, T142
3.4. Evolution Rates in Recent Porcine Pravoviruses Including T142 Strain.
3.5. Homology Comparison between Porcine Parvovirus Strains before and after the 21th Century
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Streck, A.F.; Canal, C.W.; Truyen, U. Molecular epidemiology and evolution of porcine parvoviruses. Infect. Genet. Evol. 2015, 36, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Shackelton, L.A.; Hoelzer, K.; Parrish, C.R.; Holmes, E.C. Comparative analysis reveals frequent recombination in the parvoviruses. J. Gen. Virol. 2007, 88, 3294–3301. [Google Scholar] [CrossRef] [PubMed]
- Streck, A.F.; Bonatto, S.L.; Homeier, T.; Souza, C.K.; Goncalves, K.R.; Gava, D.; Canal, C.W.; Truyen, U. High rate of viral evolution in the capsid protein of porcine parvovirus. J. Gen. Virol. 2011, 92, 2628–2636. [Google Scholar] [CrossRef] [PubMed]
- Cadar, D.; Dan, A.; Tombacz, K.; Lorincz, M.; Kiss, T.; Becskei, Z.; Spinu, M.; Tuboly, T.; Csagola, A. Phylogeny and evolutionary genetics of porcine parvovirus in wild boars. Infect. Genet. Evol. 2012, 12, 1163–1171. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, S.; Boisvert, M.; Tijssen, P. Genetic elements in the VP region of porcine parvovirus are critical to replication efficiency in cell culture. J. Virol. 2011, 85, 3025–3029. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Ritzmann, M.; Selbitz, H.J.; Heinritzi, K.; Truyen, U. VP1 sequences of German porcine parvovirus isolates define two genetic lineages. J. Gen. Virol. 2006, 87, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Tijssen, P.; Bergeron, J.; Dubuc, R.; Hébert, B. Minor genetic changes among porcine parvovirus groups are responsible for major distinguishing biological properties. Semin. Virol. 1995, 6, 319–328. [Google Scholar] [CrossRef]
- Kresse, J.I.; Taylor, W.D.; Stewart, W.W.; Eernisse, K.A. Parvovirus infection in pigs with necrotic and vesicle-like lesions. Vet. Microbiol. 1985, 10, 525–531. [Google Scholar] [CrossRef]
- Ren, X.; Tao, Y.; Cui, J.; Suo, S.; Cong, Y.; Tijssen, P. Phylogeny and evolution of porcine parvovirus. Virus Res. 2013, 178, 392–397. [Google Scholar] [CrossRef] [PubMed]
- Martins Soares, R.; Cortez, A.; Heinemann, M.B.; Sakamoto, S.M.; Martins, V.G.; Bacci, M., Jr.; De Campos Fernandes, F.M.; Richtzenhain, L.J. Genetic variability of porcine parvovirus isolates revealed by analysis of partial sequences of the structural coding gene VP2. J. Gen. Virol. 2003, 84, 1505–1515. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Chae, C. Simultaneous detection of porcine circovirus 2 and porcine parvovirus in naturally and experimentally coinfected pigs by double in situ hybridization. J. Vet. Diagn. Investig. 2002, 14, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.; Zimmermann, P.; Selbitz, H.J.; Truyen, U. Real-time PCR protocol for the detection of porcine parvovirus in field samples. J. Virol. Methods 2006, 134, 257–260. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In Nucleic Acids Symposium Series; Information Retrieval Ltd.: London, UK, January 1999; Volume 41, pp. 95–98. [Google Scholar]
- Rambaut, A.; Lam, T.T.; Max Carvalho, L.; Pybus, O.G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2016, 2, vew007. [Google Scholar] [CrossRef] [PubMed]
- Lemey, P.; Rambaut, A.; Drummond, A.J.; Suchard, M.A. Bayesian phylogeography finds its roots. PLoS Comput. Biol. 2009, 5, e1000520. [Google Scholar] [CrossRef] [PubMed]
- Drummond, A.J.; Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 2007, 7, 214. [Google Scholar] [CrossRef] [PubMed]
- Simpson, A.A.; Hébert, B.; Sullivan, G.M.; Parrish, C.R.; Zádori, Z.; Tijssen, P.; Rossmann, M.G. The structure of porcine parvovirus: Comparison with related viruses. J. Mol. Biol. 2002, 315, 1189–1198. [Google Scholar] [CrossRef] [PubMed]



| Primer Name | Sequence | Product Size (Binding Position to Accession Number KY994646) |
|---|---|---|
| PPV START F | GGCTGAAAAGAGGCGGGAAATT | 503 bp (1–503) |
| PPV R | CAGTAGCAGTCACTTGGACTTAG | |
| PPV NS1 F1 | ATGGCAGCGGGAAACACTTACT | 1005 bp (160–1165) |
| PPV NS1 R1 | TGTTCTTGCTAGAGTAAGAGTTG | |
| PPV NS1 F2 | ACCGGAGGAGAAAATTTAATCA | 1020 bp (1100–2120) |
| PPV NS1 R2 | TGCACAGTTTTCACCAAAGCAGG | |
| PPV VP1 F1 | TTGGTCGGAAATAGAAACCGACATA | 1040 bp (2070–3110) |
| PPV VP1 R1 | TTGTTCAAAACTAACTAAGTTT | |
| PPV VP1 F2 | GCAGTTAATATCCAACAACATG | 1020 bp (3050–4070) |
| PPV VP1 R2 | CCCATATTTGACCATTTGGAAATA | |
| PPV VP1 F3 | ACCATTAACAGCACTAAACAATAC | 400 bp (4020–4420) |
| PPV VP1 R3 | CTAGTATAATTTTCTTGGTATAA | |
| PPV END | CTAAAGACATAAGGTCATATAAGT | 742 bp (4020–4762) |
| PPV RT F | AGGTAAGAAGATCGCCGAGAAA | 110 bp (2550–2660) |
| PPV RT R | AGATGTCCCTTTAGCTTTTTTTTTAGC | |
| PPV P1 | ATACAATTCTATTTCATGGG | 330 bp (1330–1660) |
| PPV P6 | TATGTTCTGGTCTTTCCTCG |
| Amino Acid Location | Europe Group (n = 42) | Asia Group (n = 29) |
|---|---|---|
| 20 | A (10) T (32) | T (29) |
| 82 | K (10) R (32) | R (29) |
| 144 | E (42) | A (6) E (23) |
| 215 | I (1) T (41) | I (15) V (3) T (11) |
| 228 | E (20) Q (22) | Q (29) |
| 304 | T (10) P (32) | P (29) |
| 378 | G (42) | D (9) G(20) |
| 383 | H (1) Q (41) | H (16) Q (13) |
| 414 | S (18) A(24) | S (1) A(28) |
| 419 | Q (20) E(22) | E (29) |
| 436 | T (23) A (4) P(15) | S (12) A(9) H(1) P(7) |
| 555 | N (40) K (2) | N (21) K (8) |
| 565 | K (41) E (1) | K (20) R (9) |
| Dataset | Number of Sequence Used | Clock Model | Mean Rate | 95% HPD Interval |
|---|---|---|---|---|
| NS1 complete | 71 | UCLD | 9.71 × 10−6 | 2.30 × 10−5 |
| 8.72 × 10−9 | ||||
| VP1 complete | 65 | UCLD | 3.27 × 10−5 | 7.06 × 10−5 |
| 5.83 × 10−7 | ||||
| VP2 complete | 75 | UCLD | 5.47 × 10−5 | 1.05 × 10−4 |
| 1.49 × 10−5 | ||||
| NS1, VP1,VP2 complete | 42 | UCLD | 4.25 × 10−5 | 8.01 × 10−5 |
| 7.51 × 10−6 |
| Average Total Variant Site Numbers/Strain | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ~2000 (n = 4) | 2001~2005 (n = 17) | 2006~2010 (n = 27) | 2011~2016 (n = 26) | |||||||||||
| Variant Number/Strain | 5.0/strain | 5.88/strain | 4.70/strain | 3.28/strain | ||||||||||
| Number of Strains Variance Occurred by Region | ||||||||||||||
| Location | 20 | 45 | 82 | 93 | 101 | 144 | 215 | 226 | 228 | 233 | 238 | 304 | 320 | 366 |
| Number | 10 | 8 | 10 | 4 | 3 | 6 | 19 | 3 | 20 | 17 | 4 | 10 | 21 | 7 |
| Location | 378 | 383 | 391 | 407 | 414 | 419 | 436 | 439 | 521 | 550 | 555 | 564 | 565 | 570 |
| Number | 9 | 17 | 9 | 16 | 19 | 20 | 54 | 3 | 5 | 4 | 10 | 7 | 10 | 7 |
| Average Total Variant Site Numbers/Strain | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ~2000 (n = 4) | 2001~2005 (n = 17) | 2006~2010 (n = 27) | 2011~2016 (n = 26) | ||||||||||||
| Variant Number/Strain | 6.5/strain | 9.529/strain | 7.9629/strain | 6.4615/strain | |||||||||||
| Number of Strains Variance Occurred by Region | |||||||||||||||
| Location | 20 | 45 | 82 | 93 | 101 | 144 | 215 | 226 | 228 | 233 | 238 | 304 | 320 | ||
| Number | 9 | 66 | 9 | 4 | 3 | 6 | 59 | 3 | 10 | 17 | 4 | 11 | 56 | ||
| Location | 366 | 378 | 383 | 391 | 407 | 414 | 419 | 436 | 439 | 521 | 550 | 555 | 564 | 565 | 570 |
| Number | 7 | 9 | 17 | 9 | 61 | 18 | 20 | 61 | 3 | 5 | 4 | 63 | 7 | 10 | 7 |
| Similarity | Strains 6–22 | Strains 23–49 | Strains 50–75 |
|---|---|---|---|
| Nucleotide strains (1–5) | 98.9853% | 99.0555% | 99.1969% |
| Amino acid strains (1–5) | 98.0223% | 98.2933% | 98.5731% |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oh, W.-T.; Kim, R.-Y.; Nguyen, V.-G.; Chung, H.-C.; Park, B.-K. Perspectives on the Evolution of Porcine Parvovirus. Viruses 2017, 9, 196. https://doi.org/10.3390/v9080196
Oh W-T, Kim R-Y, Nguyen V-G, Chung H-C, Park B-K. Perspectives on the Evolution of Porcine Parvovirus. Viruses. 2017; 9(8):196. https://doi.org/10.3390/v9080196
Chicago/Turabian StyleOh, Woo-Taek, Ri-Yeon Kim, Van-Giap Nguyen, Hee-Chun Chung, and Bong-Kyun Park. 2017. "Perspectives on the Evolution of Porcine Parvovirus" Viruses 9, no. 8: 196. https://doi.org/10.3390/v9080196
APA StyleOh, W.-T., Kim, R.-Y., Nguyen, V.-G., Chung, H.-C., & Park, B.-K. (2017). Perspectives on the Evolution of Porcine Parvovirus. Viruses, 9(8), 196. https://doi.org/10.3390/v9080196





