Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus
Abstract
:1. Introduction
2. Material and Methods
2.1. Viral Clones
2.2. Diagnostic Multiplex PCR Analysis for Amplification of SLCMV and ICMV
2.3. Construction of SLMV-[SeM] DNA A and DNA B Partial Dimers
2.4. Construction of SLCMV-[SeT1] DNA A and DNA B Partial Dimers
2.5. Construction of ICMV-[SeT4] DNA A and DNA B Partial Dimers
2.6. Agroinfection of Nicotiana benthamiana
2.7. Southern Blot Analysis
2.8. siRNA Analysis
2.9. Illumina Sequencing and Bioinformatic Analysis of Small RNAs
3. Results and Discussion
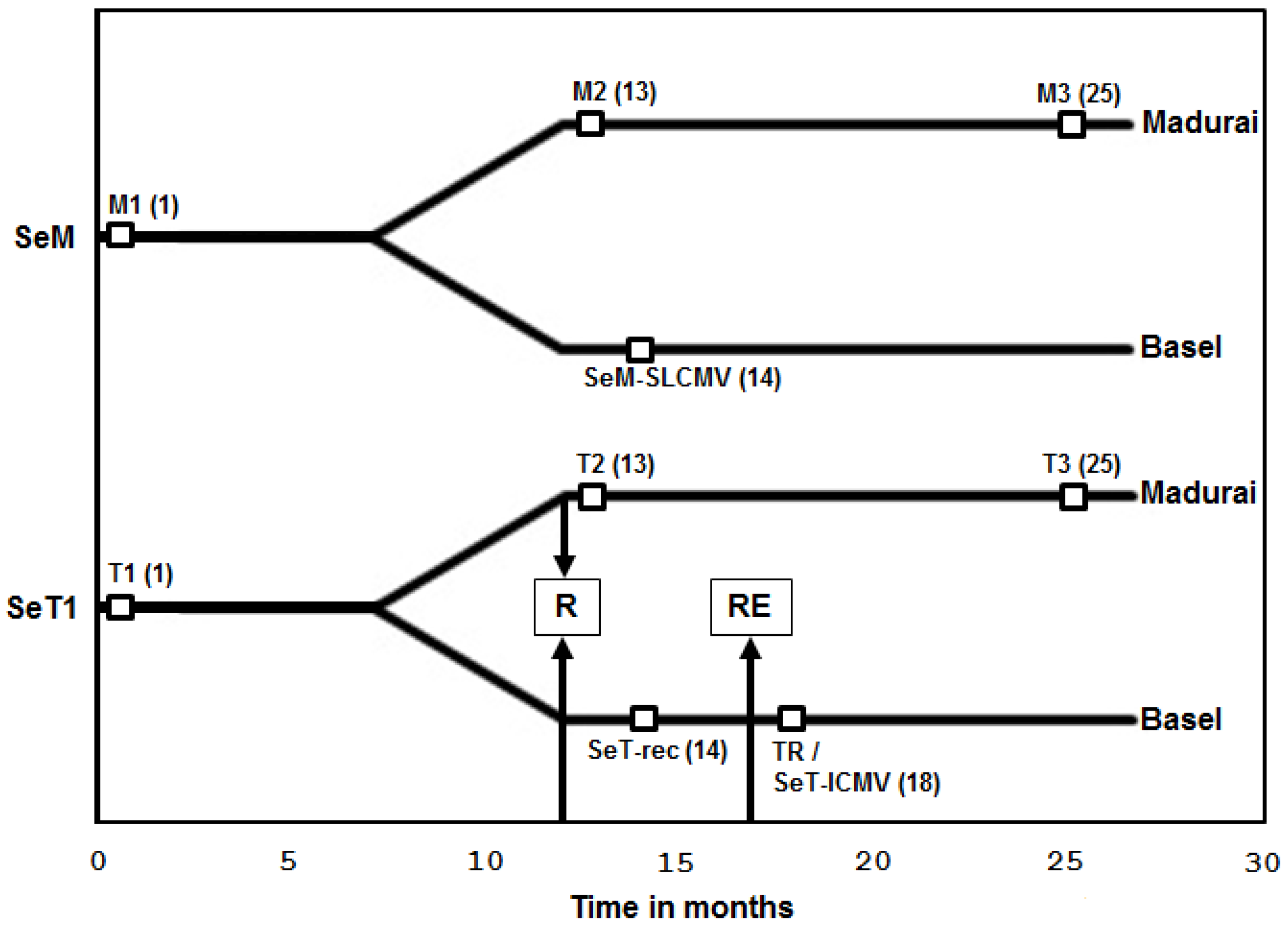
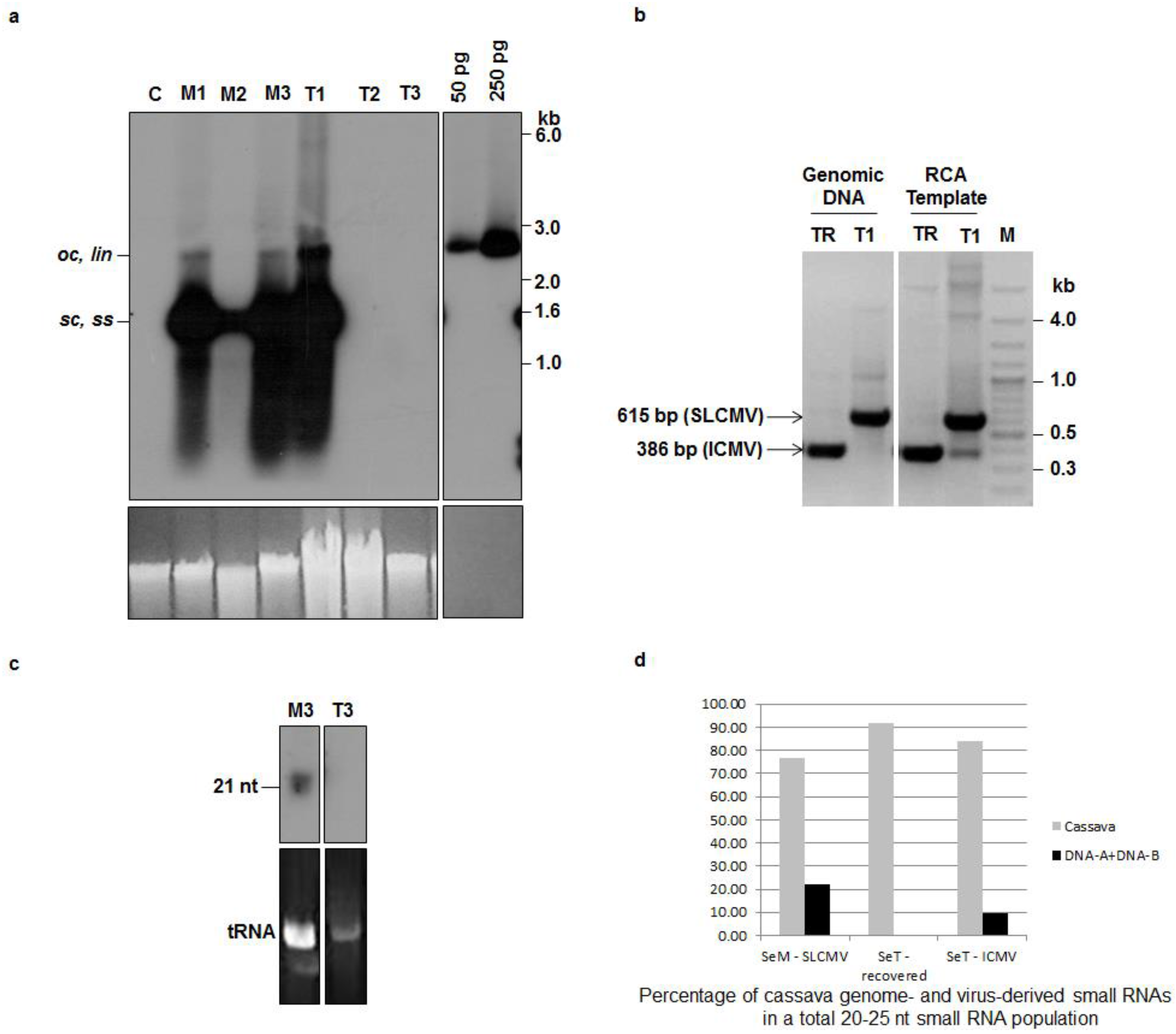
3.1. Recovery of Cassava Plants from SLCMV Infection in the Greenhouse and Re-Emergence of Symptoms due to Latent ICMV
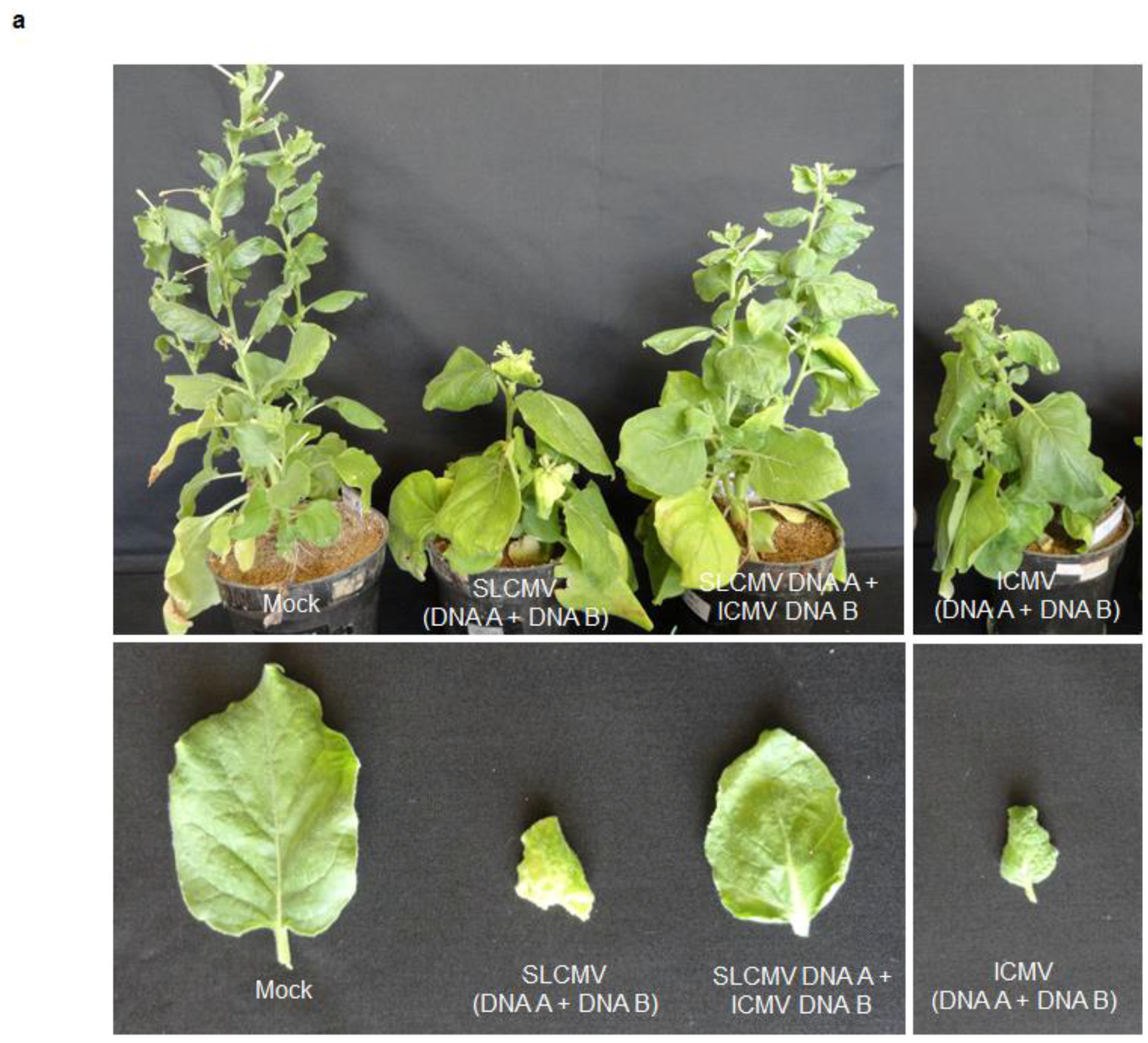
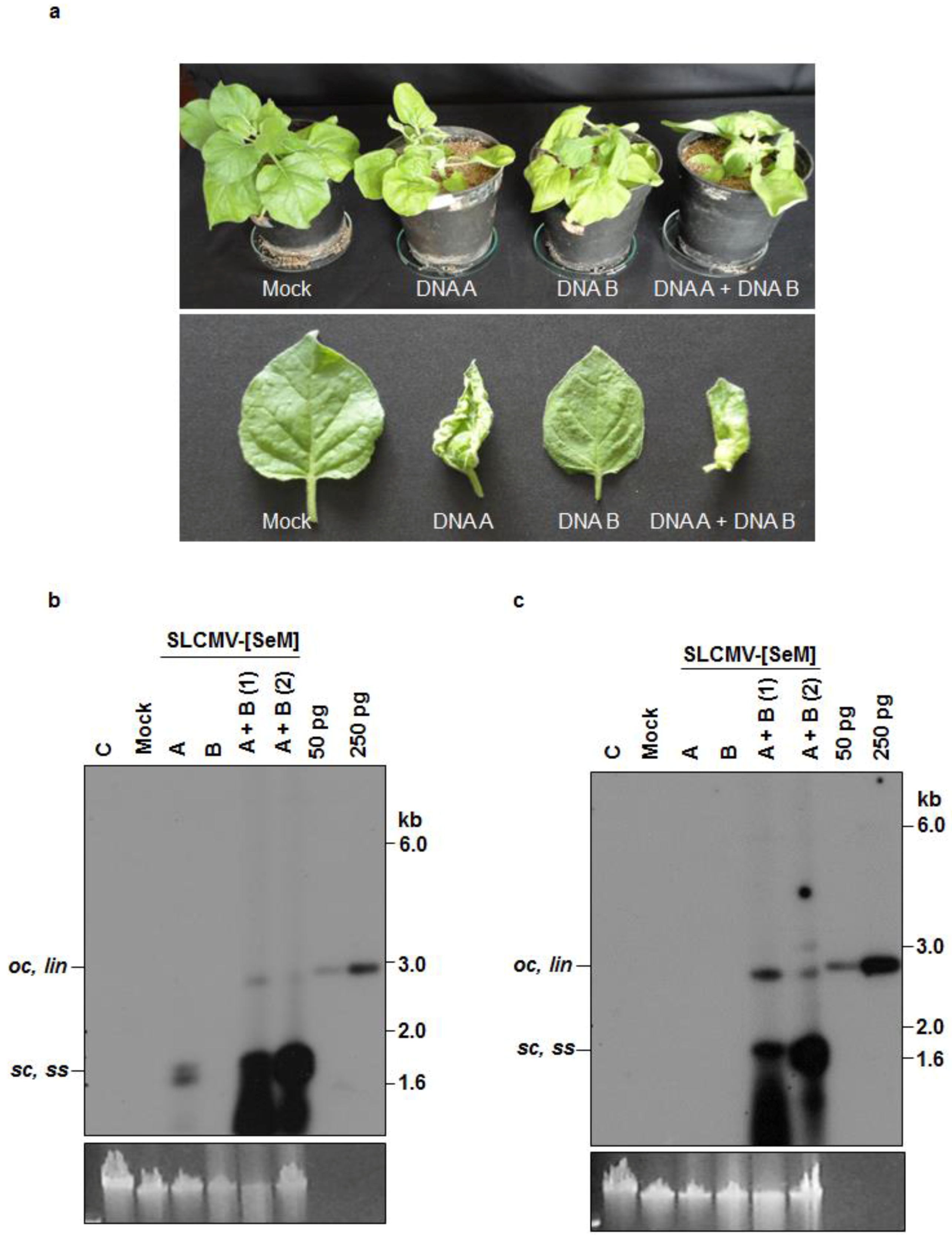
3.2. Infectivity of Cloned SLCMV DNA A and DNA B in Nicotiana benthamiana Plants
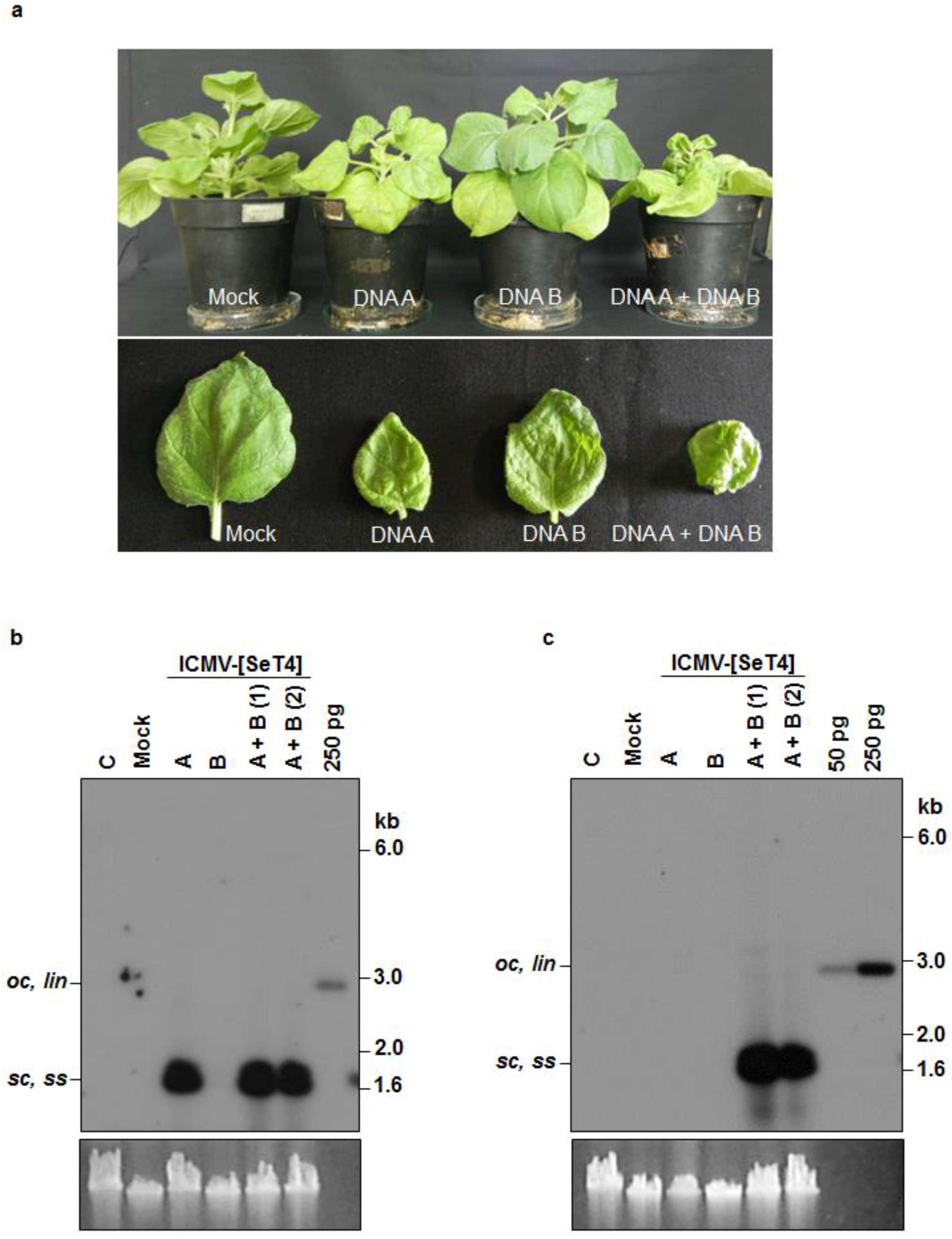
3.3. Infectivity of Cloned ICMV DNA A and DNA B in Nicotiana benthamiana Plants
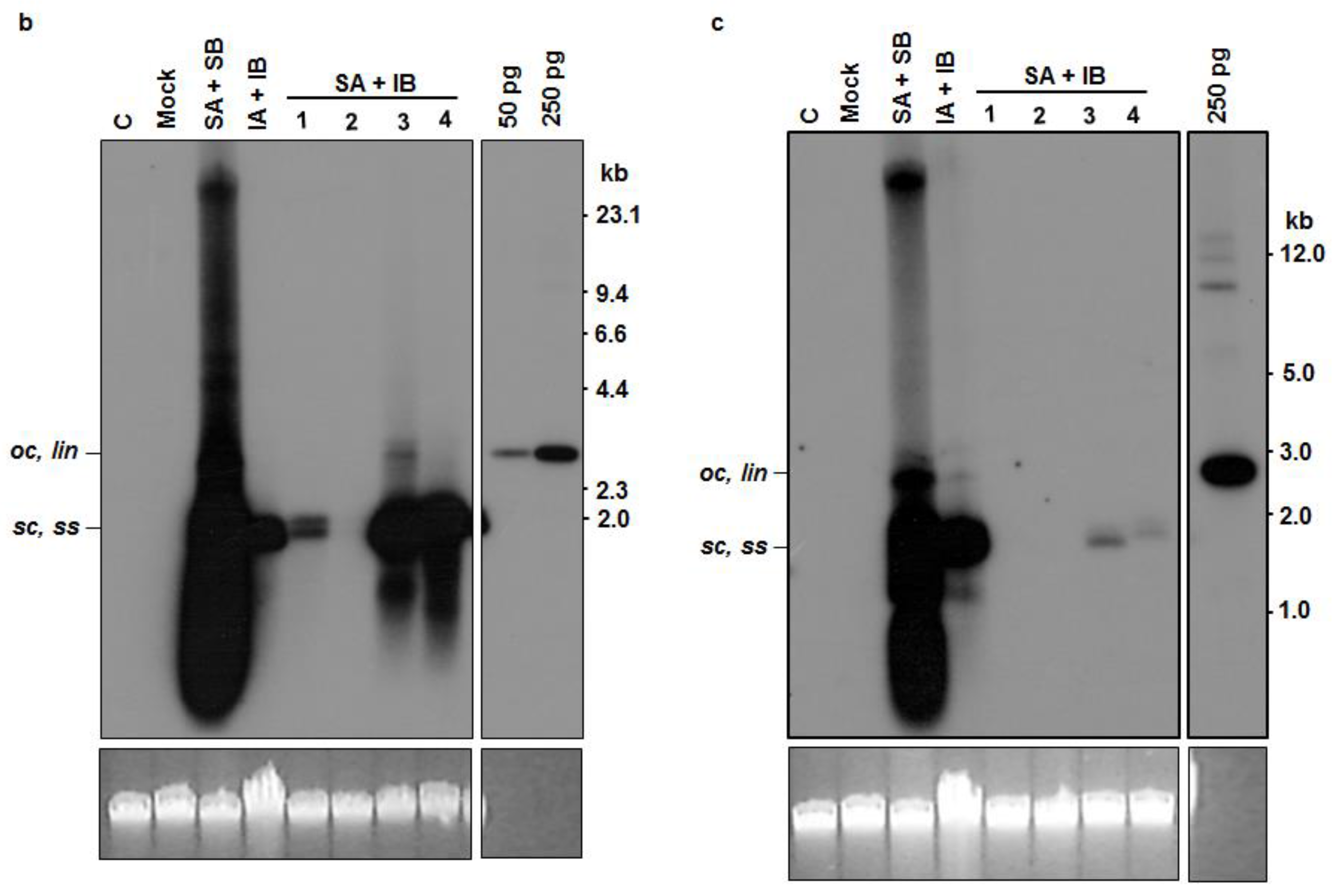
3.4. Analysis of Trans-Replication of ICMV-[TVM4] DNA B by SLCMV-[SeM] DNA A
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fauquet, C.M.; Briddon, R.W.; Brown, J.K.; Moriones, E.; Stanley, J.; Zerbini, M.; Zhou, X. Geminivirus strain demarcation and nomenclature. Arch. Virol. 2008, 153, 783–821. [Google Scholar] [CrossRef] [PubMed]
- Hanley-Bowdoin, L.; Bejarano, E.R.; Robertson, D.; Mansoor, S. Geminiviruses: Masters at redirecting and reprogramming plant processes. Nat. Rev.Microbiol. 2013, 11, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Varsani, A.; Navas-Castillo, J.; Moriones, E.; Hernández-Zepeda, C.; Idris, A.; Brown, J.K.; Zerbini, F.M.; Martin, D.P. Establishment of three new genera in the family Geminiviridae: Becurtovirus, Eragrovirus and Turncurtovirus. Arch. Virol. 2014, 159, 2193–2203. [Google Scholar] [CrossRef] [PubMed]
- Idris, A.M.; Brown, J.K. Molecular analysis of Cotton leaf curl virus-Sudan reveals an evolutionary history of recombination. Virus Genes 2002, 24, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Legg, J.P.; Fauquet, C.M. Cassava mosaic geminiviruses in Africa. Plant Mol. Biol. 2004, 56, 585–599. [Google Scholar] [CrossRef] [PubMed]
- Legg, J.P.; Owor, B.; Sseruwagi, P.; Ndunguru, J. Cassava mosaic virus disease in East and Central Africa: Epidemiology and management of a regional pandemic. Adv. Virus Res. 2006, 67, 355–418. [Google Scholar] [PubMed]
- Abraham, A. Tapioca cultivation in India. In Farm Bulletin No. 17; Indian Council of Agricultural Research: New Delhi, India, 1956; p. 20. [Google Scholar]
- Alagianagalingam, M.N.; Ramakrishnan, K. Cassava mosaic in India. South Indian Hortic. 1966, 14, 441–448. [Google Scholar]
- Saunders, K.; Nazeera, S.; Mali, V.R.; Malathi, V.G.; Briddon, R.W.; Markham, P.G.; Stanley, J. Characterisation of Sri Lankan cassava mosaic virus and Indian cassava mosaic virus: Evidence for acquisition of a DNA B component by a monopartite begomovirus. Virology 2002, 293, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.G.; Robinson, D.J.; Harrison, B.D. Nucleotide sequence evidence for the occurrence of three distinct whitefly transmitted geminiviruses in cassava. J. Gen. Virol. 1993, 74, 2437–2443. [Google Scholar] [CrossRef] [PubMed]
- Patil, B.L.; Rajasubramaniam, S.; Bagchi, C.; Dasgupta, I. Both Indian cassava mosaic virus and Sri Lankan cassava mosaic virusare found in India and exhibit high variability as assessed by PCR-RFLP. Arch. Virol. 2005, 150, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Mittal, D.; Borah, B.K.; Dasgupta, I. Agroinfection of cloned Sri Lankan cassava mosaic virus DNA to Arabidopsis thaliana, Nicotianatabacum and cassava. Arch. Virol. 2008, 153, 2149–2155. [Google Scholar] [CrossRef] [PubMed]
- Chellappan, P.; Vanitharani, R.; Fauquet, C.M. Short interfering RNA accumulation correlates with host recovery in DNA virus infected hosts and gene silencing targets specific viral sequences. J. Virol. 2004, 78, 7465–7477. [Google Scholar] [CrossRef] [PubMed]
- Vanitharani, R.; Chellappan, P.; Fauquet, C.M. Geminiviruses and RNA silencing. Trends Plant Sci. 2005, 10, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Negrete, E.A.; Carrillo-Tripp, J.; Rivera-Bustamante, R.F. RNA silencing against geminivirus: Complementary action of posttranscriptional gene silencing and transcriptional gene silencing in host recovery. J. Virol. 2009, 83, 1332–1340. [Google Scholar] [CrossRef] [PubMed]
- Góngora-Castillo, E.; Ibarra-Laclette, E.; Trejo-Saavedra1, D.L.; Rivera-Bustamante, R.F. Transcriptome analysis of symptomatic and recovered leaves of geminivirus-infected pepper (Capsicum annuum). Virol. J. 2012, 9, 295. [Google Scholar] [CrossRef] [PubMed]
- Hagen, C.; Rojas, M.R.; Kon, T.; Gilbertson, R.L. Recovery from Cucurbit leaf crumple virus (Family Geminiviridae, Genus Begomovirus) infection is an adaptive antiviral response associated with changes in viral small RNAs. Virology 2008, 98, 1029–1037. [Google Scholar]
- Chellappan, P.; Vanitharani, R.; Ogbe, F.; Fauquet, C.M. Effect of temperature on geminivirus-induced RNA silencing in plants. Plant Physiol. 2005, 138, 1828–1841. [Google Scholar] [CrossRef] [PubMed]
- Patil, B.L.; Fauquet, C.M. Studies on differential behavior of cassava mosaic geminivirus DNA components, symptom recovery patterns, and their siRNA profiles. Virus Genes 2015, 50, 474–486. [Google Scholar] [CrossRef] [PubMed]
- Gilbertson, R.L.; Hidayat, S.H.; Paplomatas, E.J.; Rojas, M.R.; Hou, Y.; Maxwell, D.P. Pseudo-recombination between infectious cloned DNA components of tomato mottle and bean dwarf mosaic geminiviruses. J. Gen. Virol. 1993, 74, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Unseld, S.; Ringel, M.; Höfer, P.; Höhnle, M.; Jeske, H.; Bedford, I.D.; Markham, P.G.; Frischmuth, T. Host range and symptom variation of pseudorecombinant virus produced by two distinct bipartite geminiviruses. Arch. Virol. 2000, 145, 1449–1454. [Google Scholar] [CrossRef] [PubMed]
- Unseld, S.; Ringel, M.; Konrad, A.; Lauster, S.; Frischmuth, T. Virus-specific adaptations for the production of a pseudorecombinant virus formed by two distinct bipartite geminiviruses from Central America. Virology 2000, 274, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Fondong, V.N. Geminivirus protein structure and function. Mol. Plant Pathol. 2013, 14, 635–649. [Google Scholar] [CrossRef] [PubMed]
- Argüello-Astorga, G.R.; Guevara-González, R.G.; Herrera-Estrella, L.R.; Rivera-Bustamante, R.F. Geminivirus replication origins have a group-specific organization of iterative elements: A model for replication. Virology 1994, 203, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Fujii, R.; Kitaoka, M.; Hayashi, K. Error-prone rolling circle amplification: The simplest random mutagenesis protocol. Nat. Protoc. 2006, 1, 2493–2497. [Google Scholar] [CrossRef] [PubMed]
- Hajdukiewicz, P.; Svab, Z.; Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol. Biol. 1994, 25, 989–994. [Google Scholar] [CrossRef] [PubMed]
- Ditta, G.; Stanfield, S.; Corbin, D.; Helinski, D.R. Broad host-range DNA cloning system for Gram-negative bacteria: Construction of a gene bank of Rhizobium meliloti. Proc. Natl. Acad. Sci. USA 1980, 77, 7347–7351. [Google Scholar] [CrossRef] [PubMed]
- Jacob, S.S.; Vanitharani, R.; Karthikeyan, A.S.; Chinchore, Y.; Thillaichidambaram, P.; Veluthambi, K. Mungbean yellow mosaic virus-Vi agroinfection by codelivery of DNA A and DNA B from one Agrobacterium strain. Plant Dis. 2003, 87, 247–251. [Google Scholar] [CrossRef]
- Chilton, M.; Currier, T.C.; Farrand, S.K.; Bendich, A.J.; Gordon, M.P.; Nester, E.W. Agrobacterium tumefaciens DNA and PS8 bacteriophage DNA not detected in crown gall tumours. Proc. Natl. Acad. Sci. USA 1974, 71, 3672–3676. [Google Scholar] [CrossRef] [PubMed]
- Grimsley, N.; Hohn, B.; Hohn, T.; Walden, R. “Agroinfection,” an alternative route for viral infection of plants by using the Ti plasmid. Proc. Natl. Acad. Sci. USA 1986, 83, 3282–3286. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, N.; Parameswari, C.; Veluthambi, K. Severe stunting in blackgram caused by the Mungbean yellow mosaic virus (MYMV) KA27 DNA B component is ameliorated by co-infection or post-infection with the KA22 DNA B: MYMV nuclear shuttle protein is the symptom determinant. Virus Res. 2011, 157, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Rogers, S.O.; Bendich, A.J. Extraction of total cellular DNA from plants, algae and fungi. In Plant Molecular Biology Manual, 2nd ed.; Gelvin, S.B., Schilperoort, R.A., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1994; pp. 1–8. [Google Scholar]
- Hong, Y.; Stanley, J. Virus resistance in Nicotiana benthamiana conferred by African cassava mosaic virus replication associated protein (AC1) transgene. Mol. Plant-Microbe Interact. 1996, 9, 219–225. [Google Scholar] [CrossRef]
- Southern, E.M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 1975, 98, 503–517. [Google Scholar] [CrossRef]
- Balaji, V.; Vanitharani, R.; Karthikeyan, A.S.; Anbalagan, S.; Veluthambi, K. Infectivity analysis of two variable DNA B components of Mungbean yellow mosaic virus-Vigna in Vigna mungo and Vignaradiata. J. Biosci. 2004, 29, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Sunitha, S.; Shanmugapriya, G.; Balamani, V.; Veluthambi, K. Mungbean yellow mosaic virus (MYMV) AC4 suppresses post-transcriptional gene silencing and an AC4 hairpin RNA gene reduces MYMV DNA accumulation in transgenic tobacco. Virus Genes 2013, 46, 496–504. [Google Scholar] [CrossRef] [PubMed]
- Fasteris—DNA Sequencing Service. Available online: https://www.fasteris.com/dna/ (accessed on 23 September 2016).
- Phytozome v11.0. Manihot esculenta v6.1 (Cassava). Available online: https://phytozome.jgi.doe.gov/pz/portal.html#!info?alias=Org_Mesculenta (accessed on 23 September 2016).
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler Transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Rothenstein, D.; Haible, D.; Dasgupta, I.; Dutt, N.; Patil, B.L.; Jeske, H. Biodiversity and recombination of cassava-infecting begomoviruses from southern India. Arch. Virol. 2006, 151, 55–69. [Google Scholar] [CrossRef] [PubMed]
- Von Arnim, A.; Stanley, J. Determinants of Tomato golden mosaic virus symptom development located on DNA B. Virology 1992, 186, 286–293. [Google Scholar] [CrossRef]
- Dry, I.B.; Krake, L.R.; Rigden, J.E.; Rezaian, M.A. A novel subviral agent associated with a geminivirus: The first report of a DNA satellite. Proc. Natl. Acad. Sci. USA 1997, 94, 7088–7093. [Google Scholar] [CrossRef] [PubMed]
- Fontes, E.P.; Gladfelter, H.J.; Schaffer, R.L.; Petty, I.T.; Hanley-Bowdoin, L. Geminivirus replication origins have a modular organization. Plant Cell 1994, 6, 405–416. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Vanitharani, R.; Chattopadhyay, B.; Fauquet, C.M. Supervirulent pseudo-recombination and asymmetric synergism between genomic components of two distinct species of begomovirus associated with severe tomato leaf curl disease in India. J. Gen. Virol. 2008, 89, 818–828. [Google Scholar] [CrossRef] [PubMed]

| Geographical Region | Cassava Cultivar | Virus | Isolate | NCBI Accession No. | Length(nt) | Identity |
|---|---|---|---|---|---|---|
| Malappuram | Sengutchi | SLCMV | SeM-DNA-A | KR611577 | 2756 | 98% to Kerala 17 |
| 96% to SeT1 | ||||||
| SLCMV | SeM-DNA-B | KR611578 | 2737 | 97% to Kerala 4 | ||
| 98% to SeT1 | ||||||
| Thiruvananthapuram | Sengutchi | SLCMV | SeT1-DNA-A | KR611579 | 2726 | 99% to Kerala 17 |
| 96% to SeM | ||||||
| SLCMV | SeT1-DNA-B | KR611580 | 2739 | 97% to Kerala 4 | ||
| 98% to SeM | ||||||
| Re-emergent in Basel | Sengutchi | ICMV | SeT4-DNA-A | KU308385 | 2735 | 94% to Kerala 2 |
| ICMV | SeT4-DNA-B | KU308386 | 2716 | 98% to Kerala 6 |
| Component A | Component B | Symptoms (at 14 days post-inoculation (dpi)) | Levels of virus DNA |
|---|---|---|---|
| SLCMV-[SeM] | Upward leaf rolling | Low DNA A | |
| SLCMV-[SeM] | No symptoms | None | |
| SLCMV-[SeM] | SLCMV-[SeM] | Stunting, chlorosis, downward leaf curling | High DNA A and DNA B |
| SLCMV-[SeT1] | Upward leaf rolling | Low DNA A | |
| SLCMV-[SeT1] | No symptoms | None | |
| SLCMV-[SeT1] | SLCMV-[SeT1] | Stunting, chlorosis, downward leaf curling | High DNA A and DNA B |
| ICMV-[SeT4] | Very mild leaf blade curling | High DNA A | |
| ICMV-[SeT4] | No symptoms | None | |
| ICMV-[SeT4] | ICMV-[SeT4] | Stunting and mild to moderate leaf blade curling | High DNA A and DNA B |
| SLCMV-[SeM] | ICMV-[SeT4] | Mild leaf blade curling | High DNA A and low DNA B |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karthikeyan, C.; Patil, B.L.; Borah, B.K.; Resmi, T.R.; Turco, S.; Pooggin, M.M.; Hohn, T.; Veluthambi, K. Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus. Viruses 2016, 8, 264. https://doi.org/10.3390/v8100264
Karthikeyan C, Patil BL, Borah BK, Resmi TR, Turco S, Pooggin MM, Hohn T, Veluthambi K. Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus. Viruses. 2016; 8(10):264. https://doi.org/10.3390/v8100264
Chicago/Turabian StyleKarthikeyan, Chockalingam, Basavaprabhu L. Patil, Basanta K. Borah, Thulasi R. Resmi, Silvia Turco, Mikhail M. Pooggin, Thomas Hohn, and Karuppannan Veluthambi. 2016. "Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus" Viruses 8, no. 10: 264. https://doi.org/10.3390/v8100264
APA StyleKarthikeyan, C., Patil, B. L., Borah, B. K., Resmi, T. R., Turco, S., Pooggin, M. M., Hohn, T., & Veluthambi, K. (2016). Emergence of a Latent Indian Cassava Mosaic Virus from Cassava Which Recovered from Infection by a Non-Persistent Sri Lankan Cassava Mosaic Virus. Viruses, 8(10), 264. https://doi.org/10.3390/v8100264






