Structural and Functional Properties of the Hepatitis C Virus p7 Viroporin
Abstract
:1. Introduction
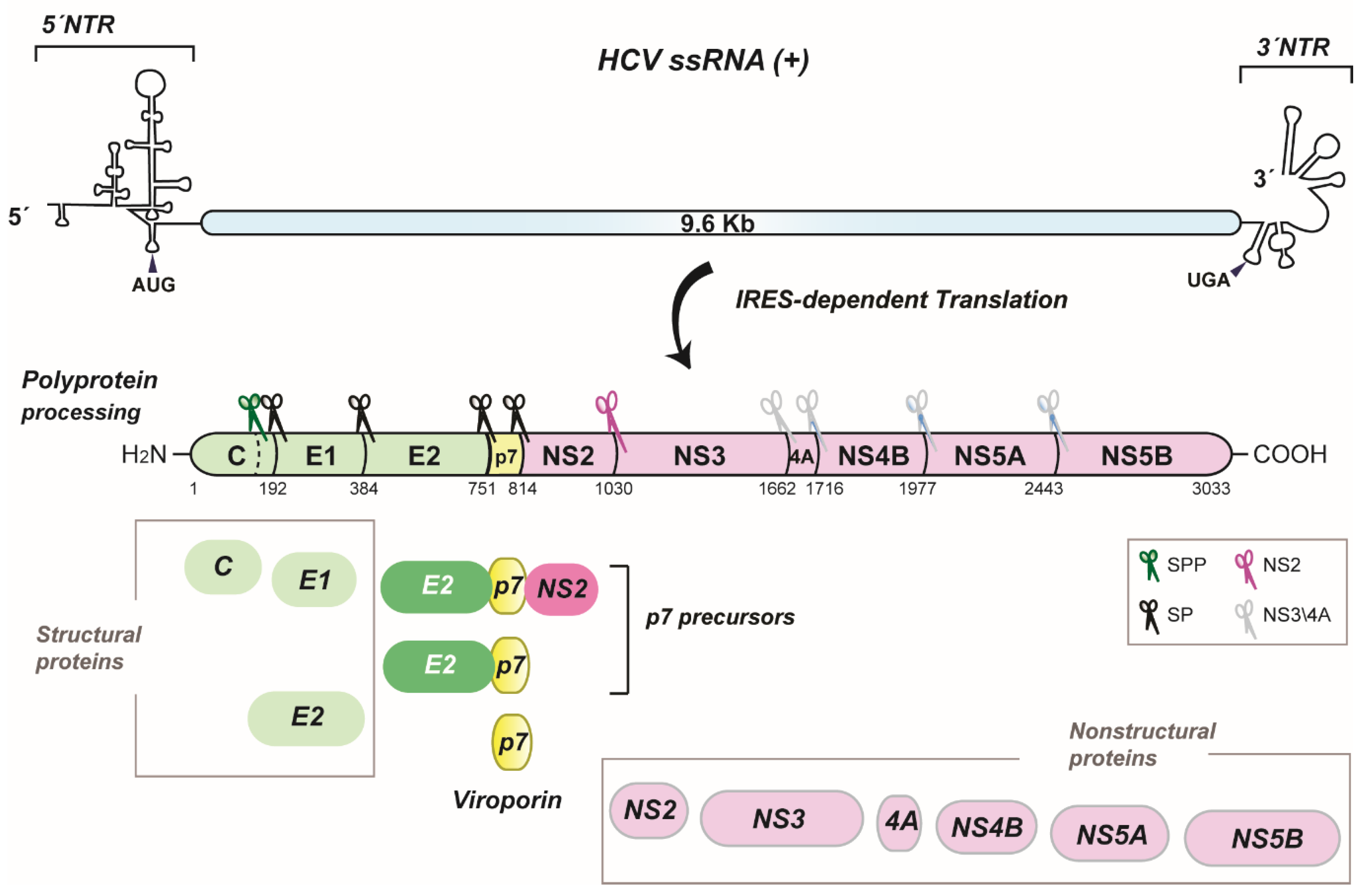
2. HCV p7 Topology and Structure
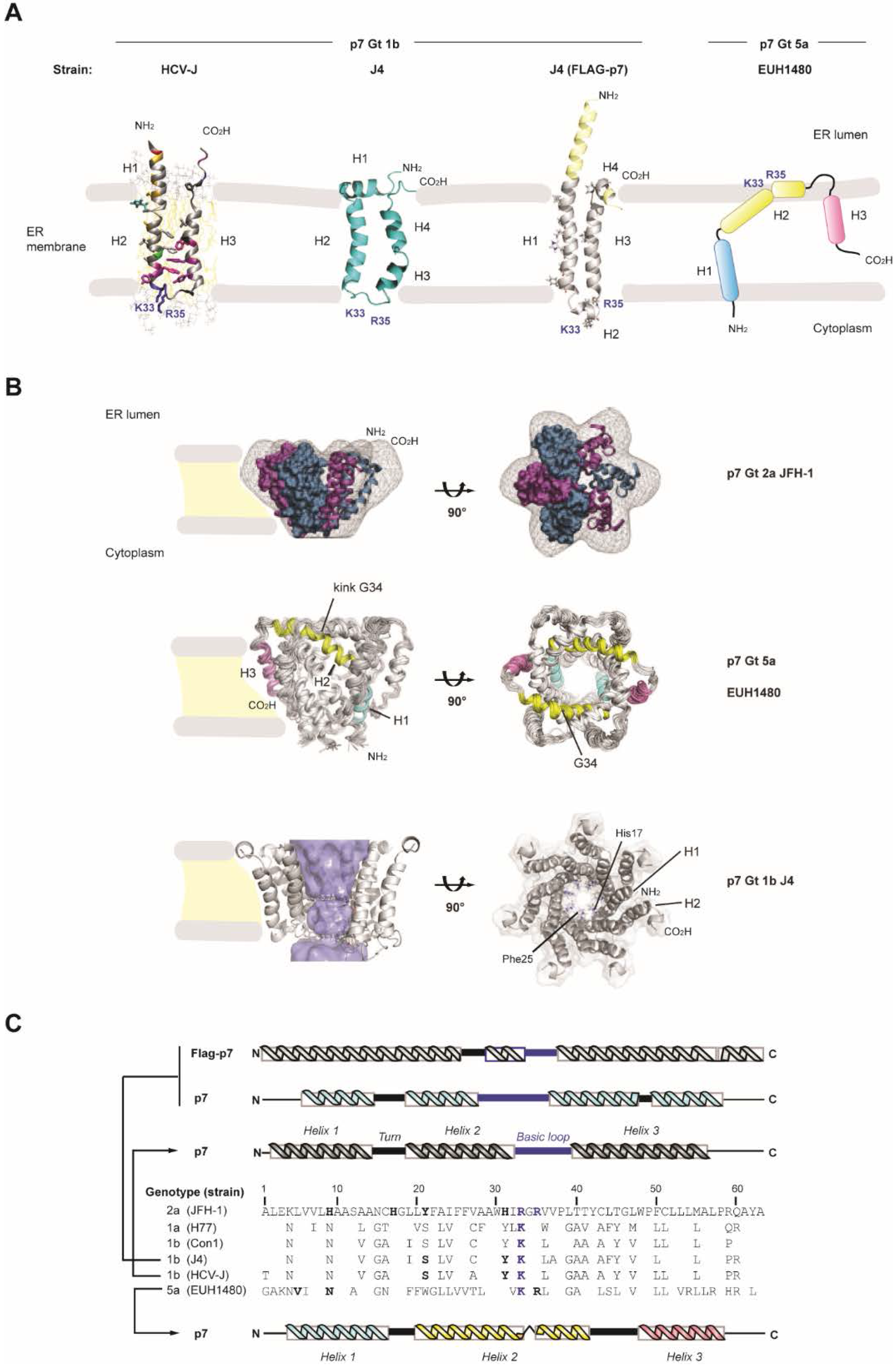
3. HCV p7 Ion Channel and Pore-Like Function
4. Role of p7 Channel Activity in Production and Release of Infectious HCV Particles
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Simmonds, P. The origin of hepatitis C virus. Curr. Top. Microbiol. Immunol. 2013, 369, 1–15. [Google Scholar] [PubMed]
- Niepmann, M. Hepatitis C virus RNA translation. Curr. Top. Microbiol. Immunol. 2013, 369, 143–166. [Google Scholar] [PubMed]
- Moradpour, D.; Penin, F. Hepatitis C virus proteins: From structure to function. Curr. Top. Microbiol. Immunol. 2013, 369, 113–142. [Google Scholar] [PubMed]
- Lohmann, V. Hepatitis C virus RNA replication. Curr. Top. Microbiol. Immunol. 2013, 369, 167–198. [Google Scholar] [PubMed]
- Lin, C.; Lindenbach, B.D.; Pragai, B.M.; McCourt, D.W.; Rice, C.M. Processing in the hepatitis C virus E2-NS2 region: Identification of p7 and two distinct E2-specific products with different C termini. J. Virol. 1994, 68, 5063–5073. [Google Scholar] [PubMed]
- Mizushima, H.; Hijikata, M.; Asabe, S.; Hirota, M.; Kimura, K.; Shimotohno, K. Two hepatitis C virus glycoprotein E2 products with different C termini. J. Virol. 1994, 68, 6215–6222. [Google Scholar] [PubMed]
- Carrere-Kremer, S.; Montpellier, C.; Lorenzo, L.; Brulin, B.; Cocquerel, L.; Belouzard, S.; Penin, F.; Dubuisson, J. Regulation of hepatitis C virus polyprotein processing by signal peptidase involves structural determinants at the p7 sequence junctions. J. Biol. Chem. 2004, 279, 41384–41392. [Google Scholar] [CrossRef] [PubMed]
- Lohmann, V.; Korner, F.; Koch, J.; Herian, U.; Theilmann, L.; Bartenschlager, R. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 1999, 285, 110–113. [Google Scholar] [CrossRef] [PubMed]
- Carrere-Kremer, S.; Montpellier-Pala, C.; Cocquerel, L.; Wychowski, C.; Penin, F.; Dubuisson, J. Subcellular localization and topology of the p7 polypeptide of hepatitis C virus. J. Virol. 2002, 76, 3720–3730. [Google Scholar] [CrossRef] [PubMed]
- Harada, T.; Tautz, N.; Thiel, H.J. E2-p7 region of the bovine viral diarrhea virus polyprotein: Processing and functional studies. J. Virol. 2000, 74, 9498–9506. [Google Scholar] [CrossRef] [PubMed]
- Sanz, M.A.; Madan, V.; Nieva, J.L; Carrasco, L. The Alphavirus 6 K Protein. In Viral Membrane Proteins: Structure, Function, and Drug Design; Fischer, W.B., Ed.; Kluwer Academic/Plenum Publishers: New York, NY, USA, 2005; Volume 1, pp. 233–242. [Google Scholar]
- Griffin, S.D.; Beales, L.P.; Clarke, D.S.; Worsfold, O.; Evans, S.D.; Jaeger, J.; Harris, M.P.; Rowlands, D.J. The p7 protein of hepatitis C virus forms an ion channel that is blocked by the antiviral drug, Amantadine. FEBS Lett. 2003, 535, 34–38. [Google Scholar] [CrossRef]
- Pavlovic, D.; Neville, D.C.; Argaud, O.; Blumberg, B.; Dwek, R.A.; Fischer, W.B.; Zitzmann, N. The hepatitis C virus p7 protein forms an ion channel that is inhibited by long-alkyl-chain iminosugar derivatives. Proc. Natl. Acad. Sci. USA 2003, 100, 6104–6108. [Google Scholar] [CrossRef] [PubMed]
- Premkumar, A.; Wilson, L.; Ewart, G.D.; Gage, P.W. Cation-selective ion channels formed by p7 of hepatitis C virus are blocked by hexamethylene amiloride. FEBS Lett. 2004, 557, 99–103. [Google Scholar] [CrossRef]
- Clarke, D.; Griffin, S.; Beales, L.; Gelais, C.S.; Burgess, S.; Harris, M.; Rowlands, D. Evidence for the formation of a heptameric ion channel complex by the hepatitis C virus p7 protein in vitro. J. Biol. Chem. 2006, 281, 37057–37068. [Google Scholar] [CrossRef] [PubMed]
- Luik, P.; Chew, C.; Aittoniemi, J.; Chang, J.; Wentworth, P., Jr.; Dwek, R.A.; Biggin, P.C.; Venien-Bryan, C.; Zitzmann, N. The 3-dimensional structure of a hepatitis C virus p7 ion channel by electron microscopy. Proc. Natl. Acad. Sci. USA 2009, 106, 12712–12716. [Google Scholar] [CrossRef] [PubMed]
- Montserret, R.; Saint, N.; Vanbelle, C.; Salvay, A.G.; Simorre, J.P.; Ebel, C.; Sapay, N.; Renisio, J.G.; Bockmann, A.; Steinmann, E.; et al. NMR structure and ion channel activity of the p7 protein from hepatitis C virus. J. Biol. Chem. 2010, 285, 31446–31461. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Date, T.; Miyamoto, M.; Furusaka, A.; Tokushige, K.; Mizokami, M.; Wakita, T. Efficient replication of the genotype 2a hepatitis C virus subgenomic replicon. Gastroenterology 2003, 125, 1808–1817. [Google Scholar] [CrossRef] [PubMed]
- Wakita, T.; Pietschmann, T.; Kato, T.; Date, T.; Miyamoto, M.; Zhao, Z.; Murthy, K.; Habermann, A.; Krausslich, H.G.; Mizokami, M.; et al. Production of infectious hepatitis C virus in tissue culture from a cloned viral genome. Nat. Med. 2005, 11, 791–796. [Google Scholar] [CrossRef] [PubMed]
- Steinmann, E.; Penin, F.; Kallis, S.; Patel, A.H.; Bartenschlager, R.; Pietschmann, T. Hepatitis C virus p7 protein is crucial for assembly and release of infectious virions. PLoS Pathog. 2007, 3, e103. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.T.; Murray, C.L.; Eastman, D.K.; Tassello, J.; Rice, C.M. Hepatitis C virus p7 and NS2 proteins are essential for production of infectious virus. J. Virol. 2007, 81, 8374–8383. [Google Scholar] [CrossRef] [PubMed]
- Sakai, A.; Claire, M.S.; Faulk, K.; Govindarajan, S.; Emerson, S.U.; Purcell, R.H.; Bukh, J. The p7 polypeptide of hepatitis C virus is critical for infectivity and contains functionally important genotype-specific sequences. Proc. Natl. Acad. Sci. USA 2003, 100, 11646–11651. [Google Scholar] [CrossRef] [PubMed]
- Saunier, B.; Triyatni, M.; Ulianich, L.; Maruvada, P.; Yen, P.; Kohn, L.D. Role of the asialoglycoprotein receptor in binding and entry of hepatitis C virus structural proteins in cultured human hepatocytes. J. Virol. 2003, 77, 546–559. [Google Scholar] [CrossRef] [PubMed]
- Griffin, S.; Stgelais, C.; Owsianka, A.M.; Patel, A.H.; Rowlands, D.; Harris, M. Genotype-dependent sensitivity of hepatitis C virus to inhibitors of the p7 ion channel. Hepatology 2008, 48, 1779–1790. [Google Scholar] [CrossRef] [PubMed]
- Vieyres, G.; Brohm, C.; Friesland, M.; Gentzsch, J.; Wolk, B.; Roingeard, P.; Steinmann, E.; Pietschmann, T. Subcellular localization and function of an epitope-tagged p7 viroporin in hepatitis C virus-producing cells. J. Virol. 2013, 87, 1664–1678. [Google Scholar] [CrossRef] [PubMed]
- Nieva, J.L.; Madan, V.; Carrasco, L. Viroporins: Structure and biological functions. Nat. Rev. Microbiol. 2012, 10, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Scott, C.; Griffin, S.D. Viroporins: Structure, function and potential as antiviral targets. J. Gen. Virol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Madan, V.; Castello, A.; Carrasco, L. Viroporins from RNA viruses induce caspase-dependent apoptosis. Cell Microbiol. 2008, 10, 437–451. [Google Scholar] [CrossRef] [PubMed]
- Klimkait, T.; Strebel, K.; Hoggan, M.D.; Martin, M.A.; Orenstein, J.M. The human immunodeficiency virus type 1-specific protein vpu is required for efficient virus maturation and release. J. Virol. 1990, 64, 621–629. [Google Scholar] [PubMed]
- Sanz, M.A.; Carrasco, L. Sindbis virus variant with a deletion in the 6K gene shows defects in glycoprotein processing and trafficking: Lack of complementation by a wild-type 6K gene in trans. J. Virol. 2001, 75, 7778–7784. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Watanabe, S.; Kim, J.H.; Hatta, M.; Kawaoka, Y. Novel approach to the development of effective H5N1 influenza A virus vaccines: Use of M2 cytoplasmic tail mutants. J. Virol. 2008, 82, 2486–2492. [Google Scholar] [CrossRef] [PubMed]
- Regla-Nava, J.A.; Nieto-Torres, J.L.; Jimenez-Guardeno, J.M.; Fernandez-Delgado, R.; Fett, C.; Castano-Rodriguez, C.; Perlman, S.; Enjuanes, L.; DeDiego, M.L. Severe acute respiratory syndrome coronaviruses with mutations in the E protein are attenuated and promising vaccine candidates. J. Virol. 2015, 89, 3870–3887. [Google Scholar] [CrossRef] [PubMed]
- Atoom, A.M.; Jones, D.M.; Russell, R.S. Evidence suggesting that HCV p7 protects E2 glycoprotein from premature degradation during virus production. Virus Res. 2013, 176, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Isherwood, B.J.; Patel, A.H. Analysis of the processing and transmembrane topology of the E2p7 protein of hepatitis C virus. J. Gen. Virol. 2005, 86, 667–676. [Google Scholar] [CrossRef] [PubMed]
- OuYang, B.; Xie, S.; Berardi, M.J.; Zhao, X.; Dev, J.; Yu, W.; Sun, B.; Chou, J.J. Unusual architecture of the p7 channel from hepatitis C virus. Nature 2013, 498, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Patargias, G.; Zitzmann, N.; Dwek, R.; Fischer, W.B. Protein-protein interactions: Modeling the hepatitis C virus ion channel p7. J. Med. Chem. 2006, 49, 648–655. [Google Scholar] [CrossRef] [PubMed]
- Chandler, D.E.; Penin, F.; Schulten, K.; Chipot, C. The p7 protein of hepatitis C virus forms structurally plastic, minimalist ion channels. PLoS. Comput. Biol. 2012, 8, e1002702. [Google Scholar] [CrossRef] [PubMed]
- Kalita, M.M.; Griffin, S.; Chou, J.J.; Fischer, W.B. Genotype-specific differences in structural features of hepatitis C virus (HCV) p7 membrane protein. Biochim. Biophys. Acta 2015, 1848, 1383–1392. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.L.; Thompson, G.S.; Kalverda, A.P.; Kankanala, J.; Bentham, M.; Wetherill, L.F.; Thompson, J.; Barker, A.M.; Clarke, D.; Noerenberg, M.; et al. Structure-guided design affirms inhibitors of hepatitis C virus p7 as a viable class of antivirals targeting virion release. Hepatology 2014, 59, 408–422. [Google Scholar] [CrossRef] [PubMed]
- Cook, G.A.; Dawson, L.A.; Tian, Y.; Opella, S.J. Three-dimensional structure and interaction studies of hepatitis C virus p7 in 1,2-dihexanoyl-sn-glycero-3-phosphocholine by solution nuclear magnetic resonance. Biochemistry 2013, 52, 5295–5303. [Google Scholar] [CrossRef] [PubMed]
- Radoicic, J.; Lu, G.J.; Opella, S.J. NMR structures of membrane proteins in phospholipid bilayers. Q. Rev. Biophys. 2014, 47, 249–283. [Google Scholar] [CrossRef] [PubMed]
- Griffin, S.D.; Harvey, R.; Clarke, D.S.; Barclay, W.S.; Harris, M.; Rowlands, D.J. A conserved basic loop in hepatitis C virus p7 protein is required for amantadine-sensitive ion channel activity in mammalian cells but is dispensable for localization to mitochondria. J. Gen. Virol. 2004, 85, 451–461. [Google Scholar] [CrossRef] [PubMed]
- Brass, V.; Berke, J.M.; Montserret, R.; Blum, H.E.; Penin, F.; Moradpour, D. Structural determinants for membrane association and dynamic organization of the hepatitis C virus NS3–4A complex. Proc. Natl. Acad. Sci. USA 2008, 105, 14545–14550. [Google Scholar] [CrossRef] [PubMed]
- Cross, T.A.; Murray, D.T.; Watts, A. Helical membrane protein conformations and their environment. Eur. Biophys. J. 2013, 42, 731–755. [Google Scholar] [CrossRef] [PubMed]
- Guinea, R.; Carrasco, L. Influenza virus M2 protein modifies membrane permeability in E. coli cells. FEBS Lett. 1994, 343, 242–246. [Google Scholar] [CrossRef]
- Sanz, M.A.; Perez, L.; Carrasco, L. Semliki Forest virus 6K protein modifies membrane permeability after inducible expression in Escherichia coli cells. J. Biol. Chem. 1994, 269, 12106–12110. [Google Scholar] [PubMed]
- Perez, M.; Garcia-Barreno, B.; Melero, J.A.; Carrasco, L.; Guinea, R. Membrane permeability changes induced in Escherichia coli by the SH protein of human respiratory syncytial virus. Virology 1997, 235, 342–351. [Google Scholar] [CrossRef] [PubMed]
- Aldabe, R.; Barco, A.; Carrasco, L. Membrane permeabilization by poliovirus proteins 2B and 2BC. J. Biol. Chem. 1996, 271, 23134–23137. [Google Scholar] [PubMed]
- Gonzalez, M.E.; Carrasco, L. The human immunodeficiency virus type 1 Vpu protein enhances membrane permeability. Biochemistry 1998, 37, 13710–13719. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.S.; Liao, C.L.; Tsao, C.H.; Chen, M.C.; Liu, C.I.; Chen, L.K.; Lin, Y.L. Membrane permeabilization by small hydrophobic nonstructural proteins of Japanese encephalitis virus. J. Virol. 1999, 73, 6257–6264. [Google Scholar] [PubMed]
- Sanz, M.A.; Madan, V.; Carrasco, L.; Nieva, J.L. Interfacial domains in Sindbis virus 6K protein. Detection and functional characterization. J. Biol. Chem. 2003, 278, 2051–2057. [Google Scholar] [CrossRef] [PubMed]
- Han, Z.; Harty, R.N. The NS3 protein of bluetongue virus exhibits viroporin-like properties. J. Biol. Chem. 2004, 279, 43092–43097. [Google Scholar] [CrossRef] [PubMed]
- Madan, V.; Garcia, M.J.; Sanz, M.A.; Carrasco, L. Viroporin activity of murine hepatitis virus E protein. FEBS Lett. 2005, 579, 3607–3612. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Orba, Y.; Okada, Y.; Sunden, Y.; Kimura, T.; Tanaka, S.; Nagashima, K.; Hall, W.W.; Sawa, H. The human polyoma JC virus agnoprotein acts as a viroporin. PLoS Pathog. 2010, 6, e1000801. [Google Scholar] [CrossRef] [PubMed]
- Agirre, A.; Barco, A.; Carrasco, L.; Nieva, J.L. Viroporin-mediated membrane permeabilization. Pore formation by nonstructural poliovirus 2B protein. J. Biol. Chem. 2002, 277, 40434–40441. [Google Scholar] [CrossRef] [PubMed]
- Gladue, D.P.; Holinka, L.G.; Largo, E.; Fernandez, S.I.; Carrillo, C.; O’Donnell, V.; Baker-Branstetter, R.; Lu, Z.; Ambroggio, X.; Risatti, G.R.; et al. Classical swine fever virus p7 protein is a viroporin involved in virulence in swine. J. Virol. 2012, 86, 6778–6791. [Google Scholar] [CrossRef] [PubMed]
- Wetherill, L.F.; Holmes, K.K.; Verow, M.; Muller, M.; Howell, G.; Harris, M.; Fishwick, C.; Stonehouse, N.; Foster, R.; Blair, G.E.; et al. High-risk human papillomavirus E5 oncoprotein displays channel-forming activity sensitive to small-molecule inhibitors. J. Virol. 2012, 86, 5341–5351. [Google Scholar] [CrossRef] [PubMed]
- Wozniak, A.L.; Griffin, S.; Rowlands, D.; Harris, M.; Yi, M.; Lemon, S.M.; Weinman, S.A. Intracellular proton conductance of the hepatitis C virus p7 protein and its contribution to infectious virus production. PLoS Pathog. 2010, 6, e1001087. [Google Scholar] [CrossRef] [PubMed]
- Stgelais, C.; Tuthill, T.J.; Clarke, D.S.; Rowlands, D.J.; Harris, M.; Griffin, S. Inhibition of hepatitis C virus p7 membrane channels in a liposome-based assay system. Antivir. Res. 2007, 76, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Gervais, C.; Do, F.; Cantin, A.; Kukolj, G.; White, P.W.; Gauthier, A.; Vaillancourt, F.H. Development and validation of a high-throughput screening assay for the hepatitis C virus p7 viroporin. J. Biomol. Screen. 2011, 16, 363–369. [Google Scholar] [CrossRef] [PubMed]
- Pinto, L.H.; Holsinger, L.J.; Lamb, R.A. Influenza virus M2 protein has ion channel activity. Cell 1992, 69, 517–528. [Google Scholar] [CrossRef]
- Plugge, B.; Gazzarrini, S.; Nelson, M.; Cerana, R.; van Etten, J.L.; Derst, C.; DiFrancesco, D.; Moroni, A.; Thiel, G. A potassium channel protein encoded by chlorella virus PBCV-1. Science 2000, 287, 1641–1644. [Google Scholar] [CrossRef] [PubMed]
- Bentham, M.J.; Foster, T.L.; McCormick, C.; Griffin, S. Mutations in hepatitis C virus p7 reduce both the egress and infectivity of assembled particles via impaired proton channel function. J. Gen. Virol. 2013, 94, 2236–2248. [Google Scholar] [CrossRef] [PubMed]
- Hong, M.; DeGrado, W.F. Structural basis for proton conduction and inhibition by the influenza M2 protein. Protein Sci. 2012, 21, 1620–1633. [Google Scholar] [CrossRef] [PubMed]
- Chew, C.F.; Vijayan, R.; Chang, J.; Zitzmann, N.; Biggin, P.C. Determination of pore-lining residues in the hepatitis C virus p7 protein. Biophys. J. 2009, 96, L10–L12. [Google Scholar] [CrossRef] [PubMed]
- Meshkat, Z.; Audsley, M.; Beyer, C.; Gowans, E.J.; Haqshenas, G. Reverse genetic analysis of a putative, influenza virus M2 HXXXW-like motif in the p7 protein of hepatitis C virus. J. Viral Hepat. 2009, 16, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Brohm, C.; Steinmann, E.; Friesland, M.; Lorenz, I.C.; Patel, A.; Penin, F.; Bartenschlager, R.; Pietschmann, T. Characterization of determinants important for hepatitis C virus p7 function in morphogenesis by using trans-complementation. J. Virol. 2009, 83, 11682–11693. [Google Scholar] [CrossRef] [PubMed]
- Stgelais, C.; Foster, T.L.; Verow, M.; Atkins, E.; Fishwick, C.W.; Rowlands, D.; Harris, M.; Griffin, S. Determinants of hepatitis C virus p7 ion channel function and drug sensitivity identified in vitro. J. Virol. 2009, 83, 7970–7981. [Google Scholar] [CrossRef] [PubMed]
- Atkins, E.; Tatineni, R.; Li, H.; Gretch, D.; Harris, M.; Griffin, S. The stability of secreted, acid-labile H77/JFH-1 hepatitis C virus (HCV) particles is altered by patient isolate genotype 1a p7 sequences. Virology 2014, 448, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Atkins, E.; Bruckner, J.; McArdle, S.; Qiu, W.C.; Thomassen, L.V.; Scott, J.; Shuhart, M.C.; Livingston, S.; Townshend-Bulson, L.; et al. Genetic and functional heterogeneity of the hepatitis C virus p7 ion channel during natural chronic infection. Virology 2012, 423, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.L.; Verow, M.; Wozniak, A.L.; Bentham, M.J.; Thompson, J.; Atkins, E.; Weinman, S.A.; Fishwick, C.; Foster, R.; Harris, M.; et al. Resistance mutations define specific antiviral effects for inhibitors of the hepatitis C virus p7 ion channel. Hepatology 2011, 54, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, T.; Miles, A.J.; Scheinost, J.C.; Offer, J.; Wentworth, P., Jr.; Dwek, R.A.; Wallace, B.A.; Biggin, P.C.; Zitzmann, N. The influence of different lipid environments on the structure and function of the hepatitis C virus p7 ion channel protein. Mol. Membr. Biol. 2011, 28, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Gan, S.W.; Surya, W.; Vararattanavech, A.; Torres, J. Two different conformations in hepatitis C virus p7 protein account for proton transport and dye release. PLoS ONE 2014, 9, e78494. [Google Scholar] [CrossRef] [PubMed]
- Perez-Berna, A.J.; Guillen, J.; Moreno, M.R.; Bernabeu, A.; Pabst, G.; Laggner, P.; Villalain, J. Identification of the membrane-active regions of hepatitis C virus p7 protein: Biophysical characterization of the loop region. J. Biol. Chem. 2008, 283, 8089–8101. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, T.; Leser, G.P.; Lamb, R.A. The ion channel activity of the influenza virus M2 protein affects transport through the Golgi apparatus. J. Cell Biol. 1996, 133, 733–747. [Google Scholar] [CrossRef] [PubMed]
- Merz, A.; Long, G.; Hiet, M.S.; Brugger, B.; Chlanda, P.; Andre, P.; Wieland, F.; Krijnse-Locker, J.; Bartenschlager, R. Biochemical and morphological properties of hepatitis C virus particles and determination of their lipidome. J. Biol. Chem. 2011, 286, 3018–3032. [Google Scholar] [CrossRef] [PubMed]
- Catanese, M.T.; Uryu, K.; Kopp, M.; Edwards, T.J.; Andrus, L.; Rice, W.J.; Silvestry, M.; Kuhn, R.J.; Rice, C.M. Ultrastructural analysis of hepatitis C virus particles. Proc. Natl. Acad. Sci. USA 2013, 110, 9505–9510. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Acosta, E.G.; Stoeck, I.K.; Long, G.; Hiet, M.S.; Mueller, B.; Fackler, O.T.; Kallis, S.; Bartenschlager, R. Apolipoprotein E likely contributes to a maturation step of infectious hepatitis C virus particles and interacts with viral envelope glycoproteins. J. Virol. 2014, 88, 12422–12437. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.; Dumans, A.; Beaumont, E.; Etienne, L.; Roingeard, P.; Meunier, J.C. The association of hepatitis C virus glycoproteins with apolipoproteins E and B early in assembly is conserved in lipoviral particles. J. Biol. Chem. 2014, 289, 18904–18913. [Google Scholar] [CrossRef] [PubMed]
- Jirasko, V.; Montserret, R.; Appel, N.; Janvier, A.; Eustachi, L.; Brohm, C.; Steinmann, E.; Pietschmann, T.; Penin, F.; Bartenschlager, R. Structural and functional characterization of nonstructural protein 2 for its role in hepatitis C virus assembly. J. Biol. Chem. 2008, 283, 28546–28562. [Google Scholar] [CrossRef] [PubMed]
- Jirasko, V.; Montserret, R.; Lee, J.Y.; Gouttenoire, J.; Moradpour, D.; Penin, F.; Bartenschlager, R. Structural and functional studies of nonstructural protein 2 of the hepatitis C virus reveal its key role as organizer of virion assembly. PLoS Pathog. 2010, 6, e1001233. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Anantpadma, M.; Timpe, J.M.; Shanmugam, S.; Singh, S.M.; Lemon, S.M.; Yi, M. Hepatitis C virus NS2 protein serves as a scaffold for virus assembly by interacting with both structural and nonstructural proteins. J. Virol. 2011, 85, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Popescu, C.I.; Callens, N.; Trinel, D.; Roingeard, P.; Moradpour, D.; Descamps, V.; Duverlie, G.; Penin, F.; Heliot, L.; Rouille, Y.; et al. NS2 protein of hepatitis C virus interacts with structural and non-structural proteins towards virus assembly. PLoS Pathog. 2011, 7, e1001278. [Google Scholar] [CrossRef] [PubMed]
- Stapleford, K.A.; Lindenbach, B.D. Hepatitis C virus NS2 coordinates virus particle assembly through physical interactions with the E1-E2 glycoprotein and NS3-NS4A enzyme complexes. J. Virol. 2011, 85, 1706–1717. [Google Scholar] [CrossRef] [PubMed]
- Tedbury, P.; Welbourn, S.; Pause, A.; King, B.; Griffin, S.; Harris, M. The subcellular localization of the hepatitis C virus non-structural protein NS2 is regulated by an ion channel-independent function of the p7 protein. J. Gen. Virol. 2011, 92, 819–830. [Google Scholar] [CrossRef] [PubMed]
- Boson, B.; Granio, O.; Bartenschlager, R.; Cosset, F.L. A concerted action of hepatitis C virus p7 and nonstructural protein 2 regulates core localization at the endoplasmic reticulum and virus assembly. PLoS Pathog. 2011, 7, e1002144. [Google Scholar] [CrossRef] [PubMed]
- Hagen, N.; Bayer, K.; Rosch, K.; Schindler, M. The intraviral protein interaction network of hepatitis C virus. Mol. Cell Proteom. 2014, 13, 1676–1689. [Google Scholar] [CrossRef] [PubMed]
- Scheel, T.K.; Prentoe, J.; Carlsen, T.H.; Mikkelsen, L.S.; Gottwein, J.M.; Bukh, J. Analysis of functional differences between hepatitis C virus NS5A of genotypes 1–7 in infectious cell culture systems. PLoS Pathog. 2012, 8, e1002696. [Google Scholar] [CrossRef] [PubMed]
- Griffin, S.; Clarke, D.; McCormick, C.; Rowlands, D.; Harris, M. Signal peptide cleavage and internal targeting signals direct the hepatitis C virus p7 protein to distinct intracellular membranes. J. Virol. 2005, 79, 15525–15536. [Google Scholar] [CrossRef] [PubMed]
- Haqshenas, G.; Mackenzie, J.M.; Dong, X.; Gowans, E.J. Hepatitis C virus p7 protein is localized in the endoplasmic reticulum when it is encoded by a replication-competent genome. J. Gen. Virol. 2007, 88, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Pietschmann, T.; Kaul, A.; Koutsoudakis, G.; Shavinskaya, A.; Kallis, S.; Steinmann, E.; Abid, K.; Negro, F.; Dreux, M.; Cosset, F.L.; et al. Construction and characterization of infectious intragenotypic and intergenotypic hepatitis C virus chimeras. Proc. Natl. Acad. Sci. USA 2006, 103, 7408–7413. [Google Scholar] [CrossRef] [PubMed]
- Gentzsch, J.; Brohm, C.; Steinmann, E.; Friesland, M.; Menzel, N.; Vieyres, G.; Perin, P.M.; Frentzen, A.; Kaderali, L.; Pietschmann, T. Hepatitis c Virus p7 is critical for capsid assembly and envelopment. PLoS Pathog. 2013, 9, e1003355. [Google Scholar] [CrossRef] [PubMed]
- Paul, D.; Madan, V.; Bartenschlager, R. Hepatitis C virus RNA replication and assembly: Living on the fat of the land. Cell Host Microbe 2014, 16, 569–579. [Google Scholar] [CrossRef] [PubMed]
- Pavlovic, D.; Fischer, W.; Hussey, M.; Durantel, D.; Durantel, S.; Branza-Nichita, N.; Woodhouse, S.; Dwek, R.A.; Zitzmann, N. Long alkylchain iminosugars block the HCV p7 ion channel. Adv. Exp. Med. Biol. 2005, 564, 3–4. [Google Scholar] [PubMed]
- Gottwein, J.M.; Jensen, T.B.; Mathiesen, C.K.; Meuleman, P.; Serre, S.B.; Lademann, J.B.; Ghanem, L.; Scheel, T.K.; Leroux-Roels, G.; Bukh, J. Development and application of hepatitis C reporter viruses with genotype 1 to 7 core-nonstructural protein 2 (NS2) expressing fluorescent proteins or luciferase in modified JFH1 NS5A. J. Virol. 2011, 85, 8913–8928. [Google Scholar] [CrossRef] [PubMed]
- Meredith, L.W.; Zitzmann, N.; McKeating, J.A. Differential effect of p7 inhibitors on hepatitis C virus cell-to-cell transmission. Antivir. Res. 2013, 100, 636–639. [Google Scholar] [CrossRef] [PubMed]
- Mihm, U.; Grigorian, N.; Welsch, C.; Herrmann, E.; Kronenberger, B.; Teuber, G.; von Wagner, M.; Hofmann, W.P.; Albrecht, M.; Lengauer, T.; et al. Amino acid variations in hepatitis C virus p7 and sensitivity to antiviral combination therapy with amantadine in chronic hepatitis C. Antivir. Ther. 2006, 11, 507–519. [Google Scholar] [PubMed]
- Castelain, S.; Bonte, D.; Penin, F.; Francois, C.; Capron, D.; Dedeurwaerder, S.; Zawadzki, P.; Morel, V.; Wychowski, C.; Duverlie, G. Hepatitis C Virus p7 membrane protein quasispecies variability in chronically infected patients treated with interferon and ribavirin, with or without amantadine. J. Med. Virol. 2007, 79, 144–154. [Google Scholar] [CrossRef] [PubMed]
- Khoury, G.; Ewart, G.; Luscombe, C.; Miller, M.; Wilkinson, J. Antiviral efficacy of the novel compound BIT225 against HIV-1 release from human macrophages. Antimicrob. Agents Chemother. 2010, 54, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Luscombe, C.A.; Huang, Z.; Murray, M.G.; Miller, M.; Wilkinson, J.; Ewart, G.D. A novel Hepatitis C virus p7 ion channel inhibitor, BIT225, inhibits bovine viral diarrhea virus in vitro and shows synergism with recombinant interferon-alpha-2b and nucleoside analogues. Antiviral Res. 2010, 86, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Bichmann, L.; Wang, Y.T.; Fischer, W.B. Docking assay of small molecule antivirals to p7 of HCV. Comput. Biol. Chem. 2014, 53, 308–317. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madan, V.; Bartenschlager, R. Structural and Functional Properties of the Hepatitis C Virus p7 Viroporin. Viruses 2015, 7, 4461-4481. https://doi.org/10.3390/v7082826
Madan V, Bartenschlager R. Structural and Functional Properties of the Hepatitis C Virus p7 Viroporin. Viruses. 2015; 7(8):4461-4481. https://doi.org/10.3390/v7082826
Chicago/Turabian StyleMadan, Vanesa, and Ralf Bartenschlager. 2015. "Structural and Functional Properties of the Hepatitis C Virus p7 Viroporin" Viruses 7, no. 8: 4461-4481. https://doi.org/10.3390/v7082826
APA StyleMadan, V., & Bartenschlager, R. (2015). Structural and Functional Properties of the Hepatitis C Virus p7 Viroporin. Viruses, 7(8), 4461-4481. https://doi.org/10.3390/v7082826




