The Origin of the Variola Virus
Abstract
:1. Introduction

2. Smallpox in Ancient Times: Historical Data
3. Dating VARV Evolution
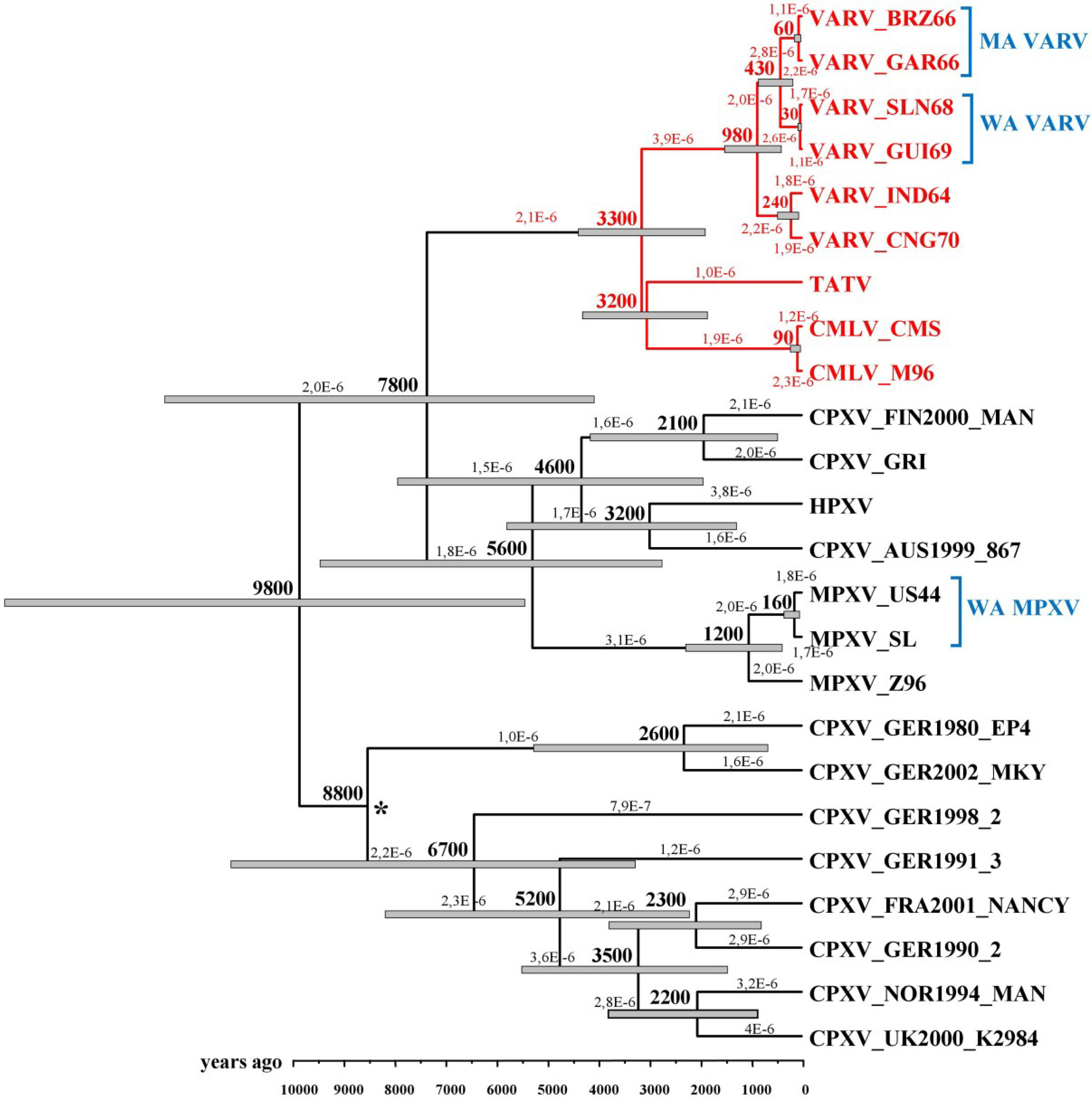
4. VARV Ancestor
5. Where Did VARV Emerge?
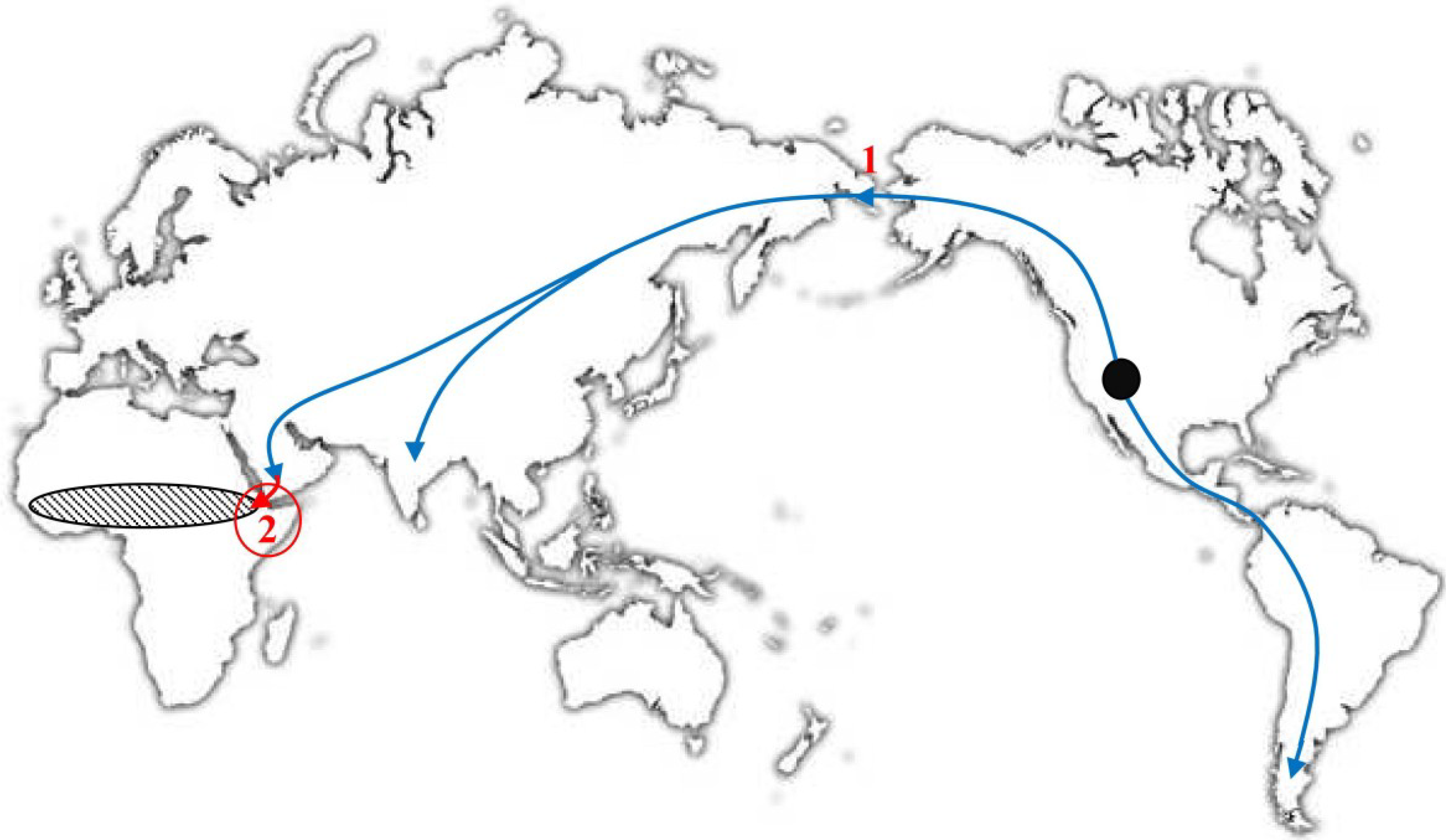
6. The Molecular Evolution of VARV
7. Conclusions
Acknowledgements
Conflicts of Interest
References
- Breman, J.G.; Henderson, D.A. Diagnosis and management of smallpox. N. Engl. J. Med. 2002, 346, 1300–1308. [Google Scholar] [CrossRef] [PubMed]
- Moore, Z.S.; Seward, J.F.; Lane, J.M. Smallpox. Lancet 2006, 367, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Fenner, F.; Henderson, D.A.; Arita, I.; Jezek, Z.; Ladnyi, I.D. Smallpox and Its Eradication; WHO: Geneva, Switzerland, 1988. [Google Scholar]
- Fenner, F. Adventures with poxviruses of vertebrates. FEMS Microbiol. Rev. 2000, 24, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Shchelkunov, S.N.; Marennikova, S.S.; Moyer, R.W. Orthopoxviruses Pathogenic for Humans; Springer-Verlag: Berlin, Germany, 2005. [Google Scholar]
- Moss, B. Poxviridae: The viruses and their replication. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Griffin, D.E., Lamb, R.A., Martin, M.A., Roizman, B., Straus, S.E., Eds.; Lippincott, Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 2905–2946. [Google Scholar]
- Hopkins, D.R. The Greatest Killer: Smallpox in History; University of Chicago Press: Chicago, IL, USA, 2002. [Google Scholar]
- Bray, M.; Buller, M. Looking back at smallpox. Clin. Infect. Dis. 2004, 38, 882–889. [Google Scholar] [CrossRef] [PubMed]
- Moussatché, N.; Damaso, C.R.; McFadden, G. When good vaccines go wild: Feral orthopoxvirus in developing countries and beyond. J. Infect. Dev. Ctries. 2008, 2, 156–173. [Google Scholar] [CrossRef]
- Trindade, G.S.; Emerson, G.L.; Carroll, D.S.; Kroon, E.G.; Damon, I.K. Brazilian vaccinia viruses and their origins. Emerg. Infect. Dis. 2007, 13, 965–972. [Google Scholar] [CrossRef] [PubMed]
- McFadden, G. Poxvirus tropism. Nat. Rev. 2005, 3, 201–213. [Google Scholar]
- Tulman, E.R.; Delhon, G.; Afonso, C.L.; Lu, Z.; Zsak, L.; Sandybaev, N.T.; Kerembekova, U.Z.; Zaitsev, V.L.; Kutish, G.F.; Rock, D.L. Genome of horsepox virus. J. Virol. 2006, 80, 9244–9258. [Google Scholar] [CrossRef] [PubMed]
- Roth, J.R.; Andersson, D.I. Poxvirus use a “gene accordion” to tune out host defenses. Cell 2012, 150, 671–672. [Google Scholar] [CrossRef] [PubMed]
- Duffy, S.; Shackelton, L.A.; Holmes, E.C. Rates of evolutionary change in viruses: Patterns and determinants. Nat. Rev. Genet. 2008, 9, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Esposito, J.J.; Sammons, S.A.; Frace, A.M.; Osborne, J.D.; Olsen-Rasmussen, M.; Zhang, M.; Govil, D.; Damon, I.K.; Kline, R.; Laker, M.; et al. Genome sequence diversity and clues to the evolution of variola smallpox virus. Science 2006, 313, 807–812. [Google Scholar] [CrossRef] [PubMed]
- Esteban, D.J.; Hutchinson, A.P. Genes in the terminal regions of orthopoxvirus genomes experience adaptive molecular evolution. BMC Genomics 2011, 12, e261. [Google Scholar] [CrossRef]
- Gubser, C.; Hue, S.; Kellam, P.; Smith, G.L. Poxvirus genomes: A phylogenetic analysis. J. Gen. Virol. 2004, 85, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Lefkowitz, E.J.; Wang, C.; Upton, C. Poxviruses: Past, present and future. Virus Res. 2006, 117, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Upton, C.; Slack, S.; Hunter, A.L.; Ehlers, A.; Roper, R.L. Poxvirus orthologous clusters: Toward defining the minimum essential poxvirus genome. J. Virol. 2003, 77, 7590–7600. [Google Scholar] [CrossRef] [PubMed]
- Raju, V.K. Susruta of ancient India. Indian J. Ophthalmol. 2003, 51, 119–122. [Google Scholar] [PubMed]
- Littman, R.J. The plague of Athens: Epidemiology and paleopathology. Mt. Sinai J. Med. 2009, 76, 456–467. [Google Scholar] [CrossRef] [PubMed]
- Garnsey, P.; Saller, R. The Roman Empire: Economy, Society, and Culture; University of California Press: Oakland, CA, USA, 1987. [Google Scholar]
- Gubser, C.; Smith, G.L. The sequence of camelpox virus shows it is most closely related to variola virus, the cause of smallpox. J. Gen. Virol. 2002, 83, 855–872. [Google Scholar] [PubMed]
- Domingo, E. Mechanisms of viral emergence. Vet. Res. 2010, 41, e38. [Google Scholar] [CrossRef]
- Wolfe, N.D.; Dunavan, C.P.; Diamond, J. Origins of major human infectious diseases. Nature 2007, 447, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M.E.J.; Gowtage-Sequeria, S. Host range and emerging and reemerging pathogens. Emerg. Infect. Dis. 2005, 11, 1842–1847. [Google Scholar] [CrossRef] [PubMed]
- Kerr, P.J. Myxomatosis in Australia and Europe: A model for emerging infectious diseases. Antivir. Res. 2012, 93, 387–415. [Google Scholar] [CrossRef] [PubMed]
- Kerr, P.J.; Ghedin, E.; DePasse, J.V.; Fitch, A.; Cattadori, I.M.; Hudson, P.J.; Tscharke, D.C.; Read, A.F.; Holmes, E.C. Evolutionary history and attenuation of myxoma virus on two continents. PLoS Pathog. 2012, 8, e1002950. [Google Scholar] [CrossRef] [PubMed]
- Saint, K.M.; French, N.; Kerr, P. Genetic variation in Australian isolates of myxoma virus: An evolutionary and epidemiological study. Arch. Virol. 2001, 146, 1105–1123. [Google Scholar] [CrossRef] [PubMed]
- Babkin, I.V.; Babkina, I.N. A retrospective study of the orthopoxvirus molecular evolution. Infect. Genet. Evol. 2012, 12, 1597–1604. [Google Scholar] [CrossRef] [PubMed]
- Uvarova, E.A.; Shchelkunov, S.N. Species-specific differences in the structure of orthopoxvirus complement-binding protein. Virus Res. 2001, 81, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Belshaw, R.; Sanjuán, R.; Pybus, O.G. Viral mutation and substitution: Units and levels. Curr. Opin. Virol. 2011, 1, 430–435. [Google Scholar] [CrossRef]
- McCollum, A.M.; Li, Y.; Wilkins, K.; Karem, K.L.; Davidson, W.B.; Paddock, C.D.; Reynolds, M.G.; Damon, I.K. Poxvirus viability and signatures in historical relics. Emerg. Infect. Dis. 2014, 20, 177–184. [Google Scholar] [CrossRef]
- Biagini, P.; Theves, C.; Balaresque, P.; Geraut, A.; Cannet, C.; Keyser, C.; Nikolaeva, D.; Gerard, P.; Duchesne, S.; Orlando, L.; et al. Variola virus in a 300-year-old Siberian mummy. N. Engl. J. Med. 2012, 367, 2057–2059. [Google Scholar] [CrossRef] [PubMed]
- Theves, C.; Biagini, P.; Crubezy, E. The rediscovery of smallpox. Clin. Microbiol. Infect. 2014, 20, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Babkina, I.N.; Babkin, I.V.; Le, U.; Ropp, S.; Kline, R.; Damon, I.; Esposito, J.; Sandakhchiev, L.S.; Shchelkunov, S.N. Phylogenetic comparison of the genomes of different strains of variola virus. Dokl. Biochem. Biophys. 2004, 398, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Babkin, I.V.; Shchelkunov, S.N. The time scale in poxvirus evolution. Mol. Biol. Mosk. 2006, 40, 20–24. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Carroll, D.S.; Gardner, S.N.; Walsh, M.C.; Vitalis, E.A.; Damon, I.K. On the origin of smallpox: Correlating variola phylogenics with historical smallpox records. Proc. Natl. Acad. Sci. USA 2007, 104, 15787–15792. [Google Scholar] [CrossRef] [PubMed]
- McLysaght, A.; Baldi, P.F.; Gaut, B.S. Extensive gene gain associated with adaptive evolution of poxviruses. Proc. Natl. Acad. Sci. USA 2003, 100, 15655–15660. [Google Scholar] [CrossRef] [PubMed]
- Emerson, G.L.; Li, Y.; Frace, M.A.; Olsen-Rasmussen, M.A.; Khristova, M.L.; Govil, D.; Sammons, S.A.; Regnery, R.L.; Karem, K.L.; Damon, I.K.; et al. The phylogenetics and ecology of the orthopoxviruses endemic to North America. PLoS One 2009, 4, e7666. [Google Scholar] [CrossRef] [PubMed]
- Babkin, I.V.; Shchelkunov, S.N. Molecular evolution of poxviruses. Genetika 2008, 44, 1029–1044. [Google Scholar] [PubMed]
- Hughes, A.L.; Irausquina, S.; Friedman, R. The evolutionary biology of poxviruses. Infect. Genet. Evol. 2010, 10, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Babkin, I.V.; Babkina, I.N. Molecular dating in the evolution of vertebrate poxviruses. Intervirology 2011, 54, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Firth, C.; Kitchen, A.; Shapiro, B.; Suchard, M.A.; Holmes, E.C.; Rambaut, A. Using time-structured data to estimate evolutionary rates of double-stranded DNA viruses. Mol. Biol. Evol. 2010, 27, 2038–2051. [Google Scholar] [CrossRef] [PubMed]
- Smithson, C.; Purdy, A.; Verster, A.J.; Upton, C. Prediction of steps in the evolution of variola virus host range. PLoS One 2014, 9, e91520. [Google Scholar] [CrossRef] [PubMed]
- Hendrickson, R.C.; Wang, C.; Hatcher, E.L.; Lefkowitz, E.J. Orthopoxvirus genome evolution: The role of gene loss. Viruses 2010, 2, 1933–1967. [Google Scholar] [CrossRef] [PubMed]
- Lourie, B.; Nakano, J.H.; Kemp, G.E.; Setzer, H.W. Isolation of poxvirus from an African rodent. J. Infect. Dis. 1975, 132, 677–681. [Google Scholar] [CrossRef] [PubMed]
- Essbauer, S.; Pfeffer, M.; Meyer, H. Zoonotic poxviruses. Vet. Microbiol. 2010, 140, 229–236. [Google Scholar] [CrossRef] [PubMed]
- Dabrowski, P.W.; Radonić, A.; Kurth, A.; Nitsche, A. Genome-wide comparison of cowpox viruses reveals a new clade related to variola virus. PLoS One 2013, 8, e79953. [Google Scholar] [CrossRef] [PubMed]
- Carroll, D.S.; Emerson, G.L.; Li, Y.; Sammons, S.; Olson, V.; Frace, M.; Nakazawa, Y.; Czerny, C.P.; Tryland, M.; Kolodziejek, J.; et al. Chasing Jenner’s vaccine: Revisiting cowpox virus classification. PLoS One 2011, 6, e23086. [Google Scholar] [CrossRef] [PubMed]
- Shchelkunov, S.N. How long ago did smallpox virus emerge? Arch. Virol. 2009, 154, 1865–1871. [Google Scholar] [CrossRef] [PubMed]
- Babkin, I.V.; Nepomnyashchikh, T.S.; Maksyutov, R.A.; Gutorov, V.V.; Babkina, I.N.; Shchelkunov, S.N. Comparative analysis of variable regions in the variola virus genome. Mol. Biol. Mosk. 2008, 42, 543–553. [Google Scholar] [CrossRef]
- Musser, G.G.; Carleton, M.D. Superfamily Muroidea. In Mammal Species of the World a Taxonomic and Geographic Reference, 3rd ed.; Wilson, D.E., Reeder, D.M., Eds.; Johns Hopkins University Press: Baltimore, MD, USA, 2005; pp. 894–1531. [Google Scholar]
- Piskurek, O.; Okada, N. Poxviruses as possible vectors for horizontal transfer of retroposons from reptiles to mammals. Proc. Natl. Acad. Sci. USA 2007, 104, 12046–12051. [Google Scholar] [CrossRef] [PubMed]
- Bulliet, R. The Camel and the Wheel; Columbia University Press: New York, NY, USA, 1975. [Google Scholar]
- Fagan, B.M.; Beck, C. The Oxford Companion to Archaeology; Oxford University Press: Oxford, UK, 1996. [Google Scholar]
- Pankhurst, R. The Ethiopian Borderlands: Essays in Regional History from Ancient Times to the End of the 18th Century; The Red Sea Press: Trenton, NJ, USA, 1997. [Google Scholar]
- Tillib, S.V. Camel nanoantibody is an efficient tool for research, diagnostics and therapy. Mol. Biol. Mosk. 2011, 45, 77–85. [Google Scholar] [PubMed]
- Muyldermans, S. Nanobodies: Natural single-domain antibodies. Annu. Rev. Biochem. 2013, 82, 775–797. [Google Scholar] [CrossRef] [PubMed]
- Gauthier–Pilters, H.; Dagg, A. The Camel: Its Evolution, Ecology, Behavior, and Relationship to Man; University of Chicago Press: Chicago, IL, USA, 1981. [Google Scholar]
- Cui, P.; Ji, R.; Ding, F.; Qi, D.; Gao, H.; Meng, H.; Yu, J.; Hu, S.; Zhang, H. A complete mitochondrial genome sequence of the wild two-humped camel (Camelus bactrianus ferus): An evolutionary history of camelidae. BMC Genomics 2007, 8, e241. [Google Scholar] [CrossRef]
- Sorci, G.; Cornet, S.; Faivre, B. Immunity and the emergence of virulent pathogens. Infect. Genet. Evol. 2013, 16, 441–446. [Google Scholar] [CrossRef] [PubMed]
- Buckland, P.C.; Dugmore, A.J.; Edwards, K.J. Bronze Age myths? Volcanic activity and human response in the Mediterranean and North Atlantic regions. Antiquity 1997, 71, 581–593. [Google Scholar]
- Bell, M.; Walker, M.J.; Walker, M.J.C. Late Quaternary Environmental Change: Physical and Human Perspectives, 2nd ed.; Prentice Hall: London, UK, 2005. [Google Scholar]
- Bratke, K.A.; McLysaght, A.; Rothenburg, S. A survey of host range genes in poxvirus genomes. Infect. Genet. Evol. 2013, 14, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Haller, S.L.; Peng, C.; McFadden, G.; Rothenburg, S. Poxviruses and the evolution of host range and virulence. Infect. Genet. Evol. 2014, 21, 15–40. [Google Scholar] [CrossRef] [PubMed]
- Yadav, V.N.; Pyaram, K.; Mullick, J.; Sahu, A. Identification of hot spots in the variola virus complement inhibitor SPICE for human complement regulation. J. Virol. 2008, 82, 3283–3294. [Google Scholar] [CrossRef] [PubMed]
- Ciulla, E.; Emery, A.; Konz, D.; Krushkal, J. Evolutionary history of orthopoxvirus proteins similar to human complement regulators. Gene 2005, 355, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Jung, J.U.; Means, R.E. ‘Complementing’ viral infection: Mechanisms for evading innate immunity. Trends Microbiol. 2003, 11, 449–452. [Google Scholar] [CrossRef] [PubMed]
- Rosengard, A.M.; Liu, Y.; Nie, Z.; Jimenez, R. Variola virus immune evasion design: Expression of a highly efficient inhibitor of human complement. Proc. Natl. Acad. Sci. USA 2002, 99, 8808–8813. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M.R.; Rahman, M.M.; Lanchbury, J.S.; Shattuck, D.; Neff, C.; Dufford, M.; van Buuren, N.; Fagan, K.; Barry, M.; Smith, S.; et al. Proteomic screening of variola virus reveals a unique NF–kappaB inhibitor that is highly conserved among pathogenic orthopoxviruses. Proc. Natl. Acad. Sci. USA 2009, 106, 9045–9050. [Google Scholar] [CrossRef] [PubMed]
- Van Vliet, K.; Mohamed, M.R.; Zhang, L.; Villa, N.Y.; Werden, S.J.; Liu, J.; McFadden, G. Poxvirus proteomics and virus–host protein interactions. Microbiol. Mol. Biol. Rev. 2009, 73, 730–749. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Babkin, I.V.; Babkina, I.N. The Origin of the Variola Virus. Viruses 2015, 7, 1100-1112. https://doi.org/10.3390/v7031100
Babkin IV, Babkina IN. The Origin of the Variola Virus. Viruses. 2015; 7(3):1100-1112. https://doi.org/10.3390/v7031100
Chicago/Turabian StyleBabkin, Igor V., and Irina N. Babkina. 2015. "The Origin of the Variola Virus" Viruses 7, no. 3: 1100-1112. https://doi.org/10.3390/v7031100
APA StyleBabkin, I. V., & Babkina, I. N. (2015). The Origin of the Variola Virus. Viruses, 7(3), 1100-1112. https://doi.org/10.3390/v7031100





