3.1. Cloning, Sequencing and Sequence Analysis of Begomoviruses
Leaf samples of field infected weeds
Ageratum conyzoides (isolate ACL) and
Sonchus oleraceous (SOL), showing yellow vein symptoms (
Figure 1, panels A and B), were collected from Lahore, Pakistan, in 2006 and 2005, respectively. A leaf sample of turnip (
Brassica rapa var. rapa), exhibiting foliar vein yellowing, was collected from Faisalabad, Pakistan during 2006 (isolate ABF). Additionally a leaf sample originating from an
A. conyzoides plant, collected in Nepal during the 1990s (isolate ACN), showing a novel symptom was included. This plant appeared superficially healthy (
Figure 1, panel C). However, closer inspection of leaves under transmitted light showed vein darkening (
Figure 1, panel D) and, on the lower leaf surface, showed raised veins and structures on veins resembling enations (
Figure 1, panel E) which on the upper leaf surface were depressions (
Figure 1, panel F)—we shall refer to this novel symptom as “dimples”.
The complete nucleotide sequences of single begomovirus clones from each of the four isolates were determined. The sequences were determined to be 2750 bp, 2749 bp, 2747 bp and 2746 bp, respectively, in length and are available in the databases under accession numbers given in
Table 1. Analyses of the sequences shows them, in all respects, to be typical of the genomes of monopartite and DNA A components of bipartite begomoviruses encoding two genes (for the CP and V2 protein) in the virion-sense and four genes (for the Rep, TrAP, REn and C4 protein) in the complementary-sense. The positions and coding capacity of the predicted genes are given in
Table 1. Between the virion- and complementary-sense genes lies a non-coding intergenic region which contains a predicted hairpin structure with, within the loop, the conserved (between all geminiviruses) nonanucleotide sequence (TAATATTAC) which forms part of the origin or virion-strand DNA replication.
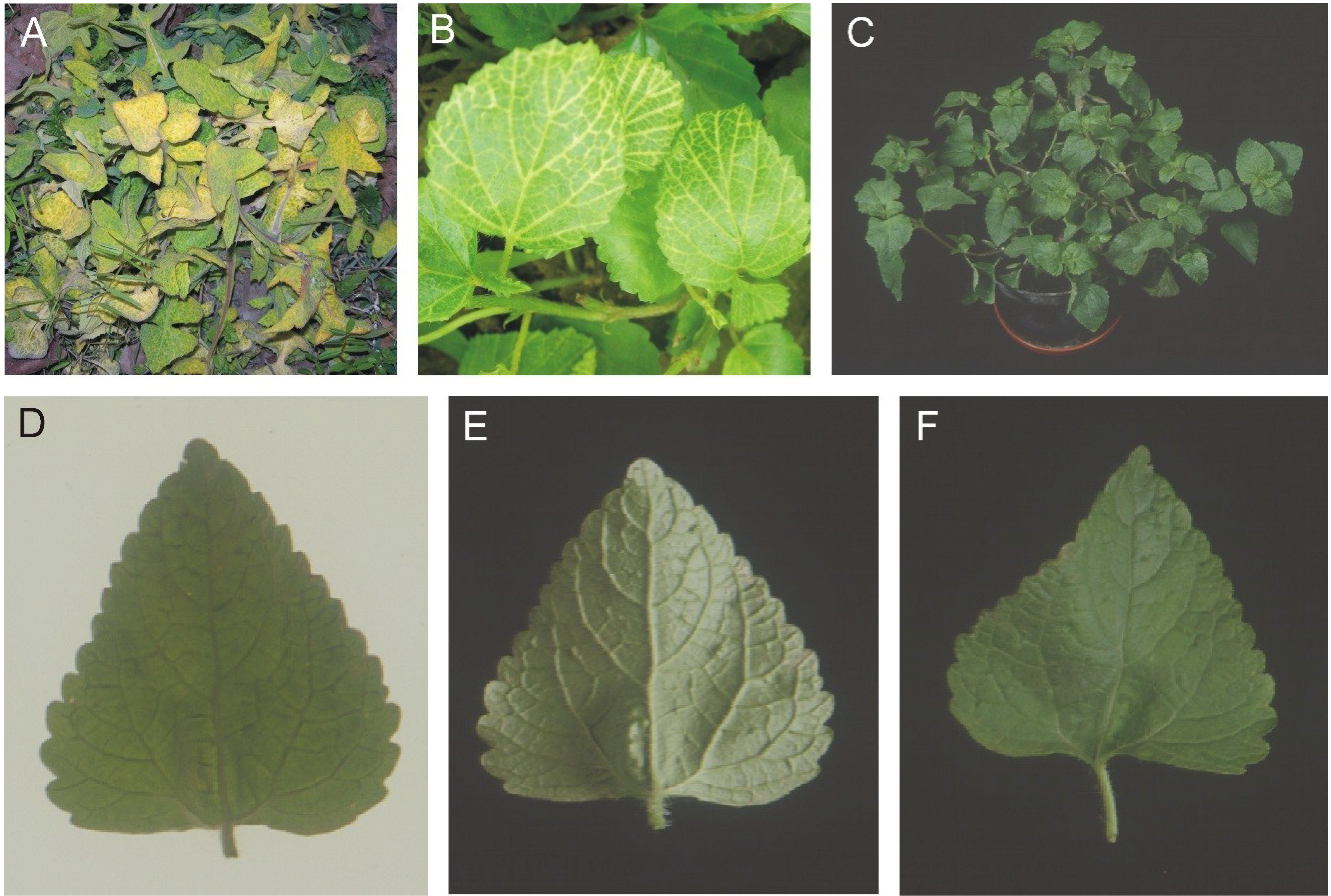
Figure 1.
Symptoms exhibited by the plants from which AEV clones were obtained. Sonchus oleraceous with yellow veins (A) Ageratum conyzoides with yellow veins (B), A. conyzoides from Nepal (C). A leaf from the A. conyzoides plant from Nepal is photographed under transmitted light to highlight the vein darkening (D) and under reflected light on the underside (E) and upper side (F) to highlight the dimple structures.
Figure 1.
Symptoms exhibited by the plants from which AEV clones were obtained. Sonchus oleraceous with yellow veins (A) Ageratum conyzoides with yellow veins (B), A. conyzoides from Nepal (C). A leaf from the A. conyzoides plant from Nepal is photographed under transmitted light to highlight the vein darkening (D) and under reflected light on the underside (E) and upper side (F) to highlight the dimple structures.
Table 1.
Origins of the virus isolates and features of the begomovirus and betasatellite clones obtained.
Table 1.
Origins of the virus isolates and features of the begomovirus and betasatellite clones obtained.
| Isolate | Origin (year) | Plant Species (Symptoms *) | Component | Accession No. | Size (bp) | ORF | Start/Stop Codon (Nucleotide Coordinates) | Predicted Size (no. of Amino Acids) | Predicted Molecular Weight (kDa) |
|---|
| SOL | Lahore, Pakistan (2005) | Sonchus oleraceus (YV) | begomovirus | AM261836 | 2750 | V2 | 135/482 | 115 | 13.3 |
| | | | CP | 295/1065 | 256 | 29.7 |
| | | | Rep | 2599/1514 | 361 | 40.7 |
| | | | TrAP | 1611/1180 | 143 | 16.2 |
| | | | REn | 1466/1062 | 134 | 15.9 |
| | | | C4 | 2442/2185 | 85 | 9.3 |
| betasatellite | AM412239 | 1368 | βC1 | 605/189 | 138 | 16.1 |
| ACL | Lahore, Pakistan (2006) | Ageratum conyzoides (YV) | begomovirus | AM698011 | 2749 | V2 | 134/481 | 115 | 13.3 |
| | | | CP | 294/1064 | 256 | 29.6 |
| | | | Rep | 2598/1513 | 361 | 40.8 |
| | | | TrAP | 1610/1206 | 134 | 15.0 |
| | | | REn | 1465/1061 | 134 | 15.9 |
| | | | C4 | 2441/2184 | 85 | 9.4 |
| betasatellite | AM698010 | 1355 | βC1 | 596/180 | 138 | 16.1 |
| ACN | Nepal (2001) | Ageratum conyzoides (LC, EN) | begomovirus | AJ437618 | 2746 | V2 | 134/481 | 115 | 13.3 |
| | | | CP | 294/1064 | 256 | 29.5 |
| | | | Rep | 2597/1512 | 361 | 40.6 |
| | | | TrAP | 1610/1206 | 134 | 15.1 |
| | | | REn | 1465/1061 | 134 | 15.9 |
| | | | C4 | 2441/2184 | 85 | 9.2 |
| betasatellite | - | - | - | - | - | - |
| ABF | Faisalabad, Pakistan (2006) | Brassica rapa var. rapa (YV) | begomovirus | AM701770 | 2747 | V2 | 134/481 | 115 | 13.3 |
| | | | CP | 294/1064 | 256 | 29.6 |
| | | | Rep | 2598/1513 | 361 | 40.8 |
| | | | TrAP | 1610/1206 | 134 | 15.2 |
| | | | REn | 1465/1061 | 134 | 15.8 |
| | | | C4 | 2441/2184 | 85 | 9.4 |
| betasatellite | AM701771 | 1359 | βC1 | 596/180 | 138 | 16.1 |
Despite being isolated from widely different regions (Nepal and Pakistan) and with a 10 year gap, the four sequences show high levels of sequence conservation (between 95.2%–98.7% nucleotide sequence identity). Based on the 89% nucleotide sequence identity demarcation threshold for identification of begomovirus species, this indicates that all four clones are isolates of a single species [
27,
30].
3.2. Comparison of the Sequences Obtained Here with Sequences Available in the Databases
The AEV clone obtained from Nepal was obtained some time ago and was at the time recognized as a distinct species within the genus
Begomovirus and given the name
Ageratum enation virus [
30]. The sequence of AEV-Nepal thus represents the type isolate of this begomovirus species. The other three sequences, with greater than 89% nucleotide sequence identity to AEV-Nepal, thus are variants of AEV. A Blast search of the GenBank nucleotide sequence database with the four AEV sequences presented here identified 26 additional sequences originating from India with high sequence identity (>89%): GQ268327 isolated from
Trichosanthes dioica (unpublished), EU867513 isolated from
Amaranthus cruentus (unpublished), KC795968 and KC818421 isolated from tomato (unpublished), JX436472 isolated from tomato (unpublished), HE861940 isolated from soybean (unpublished), JQ911765 and HM149260 isolated from
Papaver somniferum (unpublished), JX436473 isolated from Fenugreek (unpublished), JQ911767, KJ488990 and KJ488991 isolated from
Ageratum sp. (unpublished), JF728865 and JF728867 (isolated from carrot [
31]), JF728860-JF728864 and JF728866 isolated from
A. conyzoides [
31], JF682242 isolated from
Amaranthus (unpublished), FN794201 isolated from
Crassocephalum crepidioides [
32], FN543099 isolated from
Zinnia elegans [
33], FJ177031 isolated from
Cleome gynandra [
34] and FN794198 isolated from
A. conyzoides [
32]. Some of these sequences have previously been reported as isolates of AEV. Overall these 30 sequences (including the 4 identified here) show between 88.9% and 100% nucleotide sequence identity (
Table S1), with the highest levels of identity to isolates of
Tobacco curly shoot virus (TbCSV). This confirms the 30 sequences as isolates of a distinct begomovirus species, which has previously been named AEV [
30].
A closer analysis of the AEV sequences shows them to fall into three groups. The first group consists of all the AEV sequences characterized here, as well as GQ268327, EU867513, KC818421, KC795968, and FJ177031. These nine sequences show between 93.9% and 100% nucleotide sequence identity (
Table S1). The sequences in the second group, consisting of FN543099, FN794198, FN794201, JF728860-64, JF728866, JX436472, JX436473, JQ911765, JF682242, JQ911767, HM149260, KJ488991, KJ488990 and HE861940 show between 94.6% and 100% identity. Between the two groups the identity levels vary between 90.6%. and 92.9%. Based on the presently applicable demarcation threshold for strains within a species (93%; [
27]), this would indicate that the two groups represent strains of AEV, for which we propose the names “Nepal” and “India”, respectively. The third group of sequences (JF728865 and JF728867) show relatively high sequence identities to both the Nepal (group 1—91.8% to 98.1% identity) and the India (group 2—94.3% to 99.2% identity) strain AEV sequences, making it difficult to assign them to a strain.
Interestingly the four AEV sequences have Rep IRD sequences predicted to be either FKIN (AM261836, AM698011, AM701770) or LKIN (AJ437618). The other three Nepal strain isolates also have LKIN. In contrast, all India strain isolates have an IRD with the predicted sequence FQIY. The corresponding iteron sequences are GGT/AGT for all Nepal strain isolates and either GGTG/AC/A, or possibly GTACT, for all India strain isolates and both problematic isolates. The alignment of all AEV sequences suggested that the differences between the Nepal and India strain isolates may be due to recombination across the origin of replication, possibly with Papaya leaf crumple virus (PaLCrV) as the donor of the ori for the India strain (results not shown). PaLCrV occurs in India and has IRD and iteron sequences identical to those of India strain AEV sequences.
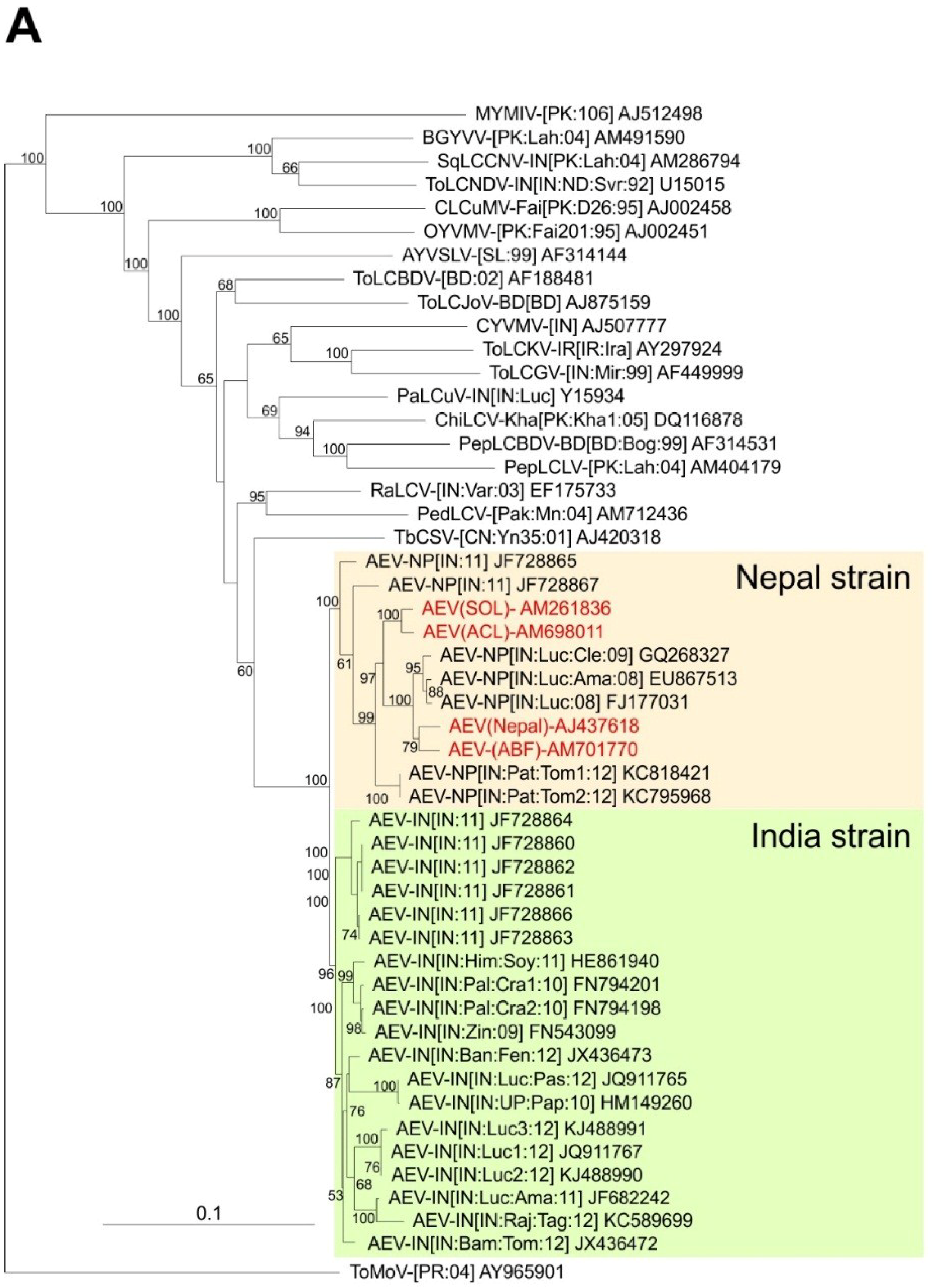
A phylogenetic dendrogram based upon an alignment of the four sequences obtained here, the 26 other AEV sequences available in the databases and selected other begomovirus sequences available in the databases is shown in
Figure 2A. This shows the 30 AEV sequences to form a group distinct from all other sequences in the tree but to be most closely related to TbCSV. The division of AEV sequences into two distinct groups, for which we have proposed the names Nepal and India strains, are well supported by bootstrapping. The two problematic AEV isolates (JF728865 and JF728867) group with, but are distinct from and basal to the Nepal strain isolates, despite having overall more sequence identity to isolates of the India strain. JF728867 has a Nepal strain IRD (FKIN), whereas JF728865 has an Indian strain IRD (FQIY). It is for this reason that we proposed, at least provisionally, to include these two isolates under the Nepal strain, awaiting the availability of further sequences and analyses.
Figure 2.
Phylogenetic analyses of begomovirus and betasatellite sequences. Phylogenetic dendrograms based upon alignments of the complete nucleotide sequences of the genomes (or DNA A genomic components of) begomoviruses (
A) and betasatellites (
B) identified here with selected sequences available in the databases. The Neighbour-joining method was used for construction of the phylogenetic dendrograms. Horizontal distances are proportional to mutation distances whereas vertical distances are arbitrary. The numbers at each branch indicate percentage bootstrap confidence scores (1000 replicates). The begomovirus acronyms used are
Ageratum enation virus (AEV),
Ageratum yellow vein Sri Lanka virus (AYVSLV),
Bitter gourd yellow vein virus (BGYVV),
Chili leaf curl virus (ChiLCV),
Cotton leaf curl Multan virus (CLCuMV),
Croton yellow vein mosaic virus (CYVMV),
Okra yellow vein mosaic virus (OYVMV),
Mungbean yellow mosaic India virus (MYMIV),
Papaya leaf curl virus (PaLCuV),
Pedilanthus leaf curl virus (PedLCV),
Pepper leaf curl Bangladesh virus (PepLCBDV),
Pepper leaf curl Lahore virus (PepLCLV),
Radish leaf curl virus (RaLCV),
Squash leaf curl China virus (SLCCNV),
Tobacco curly shoot virus (TbCSV),
Tomato leaf curl Bangladesh virus (ToLCBDV),
Tomato leaf curl Gujarat virus (ToLCGV),
Tomato leaf curl Joydebpur virus (ToLCJoV),
Tomato leaf curl Karnataka virus (ToLCKV) and
Tomato leaf curl New Delhi virus (ToLCNDV). Isolate descriptors are as given in [
27]. The tree was rooted on the sequence of the DNA A component of
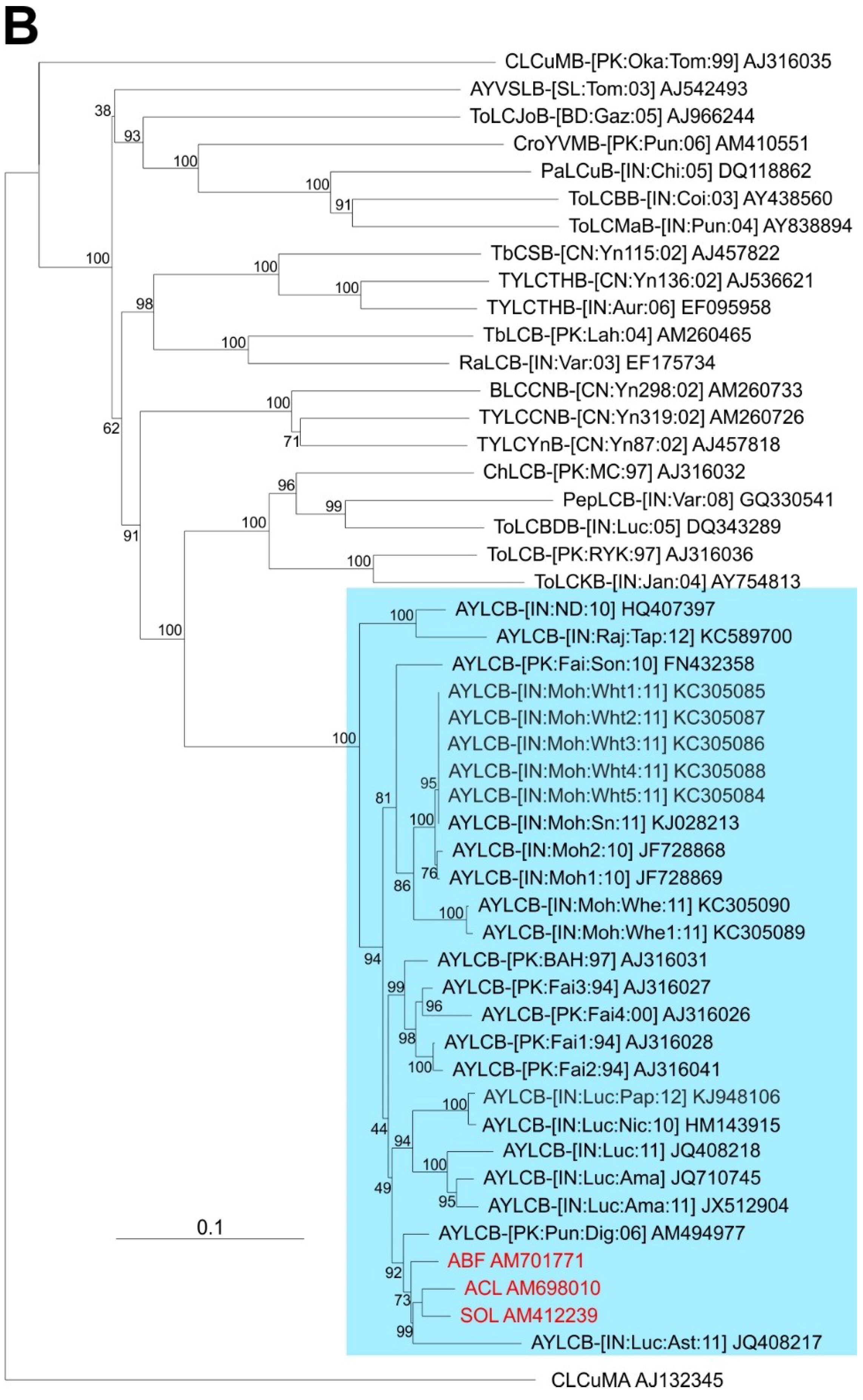
Tomato mottle virus (ToMoV) as an outgroup. The AEV isolates and the two strains of AEV (India and Nepal) are indicated on the right of the tree. The betasatellite acronyms used are Ageratum yellow leaf curl betasatellite (AYLCB), Ageratum yellow vein Sri Lanka betasatellite (AYVSLB), Bean leaf curl China betasatellite (BLCCNB), Chili leaf curl betasatellite (ChLCB), Cotton leaf curl Multan betasatellite (CLCuMB), Tomato leaf curl Joydebpur betasatellite (ToLCJoB), Croton yellow vein mosaic betasatellite (CroYVMB), Papaya leaf curl betasatellite (PaLCuB), Pepper leaf curl betasatellite (PepLCB), Radish leaf curl betasatellite (RaLCB), Tobacco curly shoot betasatellite (TbCSB), Tobacco leaf curl betasatellite (TbLCB), Tomato leaf curl Bangalore betasatellite (ToLCBB), Tomato leaf curl Bangladesh betasatellite (ToLCBDB), Tomato leaf curl Karnataka betasatellite (ToLCKB), Tomato leaf curl Maharashtra betasatellite (ToLCMaB), Tomato leaf curl betasatellite (ToLCB), Tomato yellow leaf curl China betasatellite (TYLCCNB), Tomato yellow leaf curl Thailand betasatellite (TYLCTHB) and Tomato yellow leaf curl Yunan betasatellite (TYLCYnB). Isolate descriptors are as given in [
8]. The tree was rooted on the sequence of Cotton leaf curl Multan alphasatellite (CLCuMA) as outgroup. For both trees the sequences obtained here are highlighted in red text and the database accession numbers of isolates are given.
Figure 2.
Phylogenetic analyses of begomovirus and betasatellite sequences. Phylogenetic dendrograms based upon alignments of the complete nucleotide sequences of the genomes (or DNA A genomic components of) begomoviruses (
A) and betasatellites (
B) identified here with selected sequences available in the databases. The Neighbour-joining method was used for construction of the phylogenetic dendrograms. Horizontal distances are proportional to mutation distances whereas vertical distances are arbitrary. The numbers at each branch indicate percentage bootstrap confidence scores (1000 replicates). The begomovirus acronyms used are
Ageratum enation virus (AEV),
Ageratum yellow vein Sri Lanka virus (AYVSLV),
Bitter gourd yellow vein virus (BGYVV),
Chili leaf curl virus (ChiLCV),
Cotton leaf curl Multan virus (CLCuMV),
Croton yellow vein mosaic virus (CYVMV),
Okra yellow vein mosaic virus (OYVMV),
Mungbean yellow mosaic India virus (MYMIV),
Papaya leaf curl virus (PaLCuV),
Pedilanthus leaf curl virus (PedLCV),
Pepper leaf curl Bangladesh virus (PepLCBDV),
Pepper leaf curl Lahore virus (PepLCLV),
Radish leaf curl virus (RaLCV),
Squash leaf curl China virus (SLCCNV),
Tobacco curly shoot virus (TbCSV),
Tomato leaf curl Bangladesh virus (ToLCBDV),
Tomato leaf curl Gujarat virus (ToLCGV),
Tomato leaf curl Joydebpur virus (ToLCJoV),
Tomato leaf curl Karnataka virus (ToLCKV) and
Tomato leaf curl New Delhi virus (ToLCNDV). Isolate descriptors are as given in [
27]. The tree was rooted on the sequence of the DNA A component of
Tomato mottle virus (ToMoV) as an outgroup. The AEV isolates and the two strains of AEV (India and Nepal) are indicated on the right of the tree. The betasatellite acronyms used are Ageratum yellow leaf curl betasatellite (AYLCB), Ageratum yellow vein Sri Lanka betasatellite (AYVSLB), Bean leaf curl China betasatellite (BLCCNB), Chili leaf curl betasatellite (ChLCB), Cotton leaf curl Multan betasatellite (CLCuMB), Tomato leaf curl Joydebpur betasatellite (ToLCJoB), Croton yellow vein mosaic betasatellite (CroYVMB), Papaya leaf curl betasatellite (PaLCuB), Pepper leaf curl betasatellite (PepLCB), Radish leaf curl betasatellite (RaLCB), Tobacco curly shoot betasatellite (TbCSB), Tobacco leaf curl betasatellite (TbLCB), Tomato leaf curl Bangalore betasatellite (ToLCBB), Tomato leaf curl Bangladesh betasatellite (ToLCBDB), Tomato leaf curl Karnataka betasatellite (ToLCKB), Tomato leaf curl Maharashtra betasatellite (ToLCMaB), Tomato leaf curl betasatellite (ToLCB), Tomato yellow leaf curl China betasatellite (TYLCCNB), Tomato yellow leaf curl Thailand betasatellite (TYLCTHB) and Tomato yellow leaf curl Yunan betasatellite (TYLCYnB). Isolate descriptors are as given in [
8]. The tree was rooted on the sequence of Cotton leaf curl Multan alphasatellite (CLCuMA) as outgroup. For both trees the sequences obtained here are highlighted in red text and the database accession numbers of isolates are given.
![]()
![]()
3.3. Cloning, Sequencing and Sequence Analysis of Betasatellites
Three full-length betasatellite clones were obtained. Unfortunately the satellite for isolate ACN was not obtained for comparison, although the presence of a betasatellite with this isolate has been shown previously [
35]. The three sequences have a length typical of betasatellites (between 1355 to 1368 bp,
Table 1), being approximately half the size of their helper begomoviruses. Analysis of the sequences show them to have an arrangement typical of this class of satellites consisting of a single open reading frame in the complementary-sense encoding the βC1 protein (
Table 1), a region of sequence rich in adenine and a sequence highly conserved between all betasatellites—the satellite conserved region [
35].
Comparison of the three betasatellite sequences obtained here to sequences available in the databases shows them to be most similar to Ageratum yellow leaf curl betasatellite (AYLCB; with between 90.6% and 99.8% nucleotide sequence identity to the 18 other sequences available in the databases). Six defective AYLCB are available in the databases (JQ408217, KC305086-90) ranging in size from 891 bp to 1270 bp, which are included in the analysis but not included in the calculated identity values. To all other betasatellite sequences available in the databases the percentage nucleotide sequence identity was less than 62% (not shown) with the highest sequence identity levels (57.5%–61.3%) to an isolate of Tomato leaf curl Bangladesh betasatellite (AJ542489).
A phylogenetic dendrogram, based upon an alignment of the full-length sequences of the 3 betasatellites obtained here, all available AYLCB sequences in the databases and selected other betasatellite sequences from the databases is shown in
Figure 2B. This shows AYLCB to form a clade with Chili leaf curl betasatellite, Pepper leaf curl betasatellite, Tomato leaf curl Bangladesh betasatellite, Tomato leaf curl betasatellite and Tomato leaf curl Karnataka betasatellite—a grouping well supported by bootstrapping. It is interesting to note that all AYLCB isolates from Pakistan, with the exception of FN432358, are distinct from those originating from India. This suggests that AYLCB in these two areas are evolving independently. Possibly FN432358, an “Indian” isolate of AYLCB occurring in Pakistan, indicates that infrequent exchanges between India and Pakistan take place. Also, the Pakistan AYLCB isolates characterized prior to 2000 are distinct from those characterized more recently, which may indicate temporal changes that have occurred in this betasatellite species.
3.4. Analysis of the Infectivity of AEV and Betasatellite Clones
Partial repeat constructs of the AEV and betasatellite clones isolated from
S. oleraceus (isolate SOL) and
A. conyzoides (isolate ACL) were produced for
Agrobacterium-mediated inoculation of plants. The results of the infectivity studies are summarized in
Table 2. Inoculation of
N. benthamiana with the AEV and AYLCB clones isolated from
S. oleraceus (AEV
SOL and AYLCB
SOL) led to the appearance of symptoms consisting of vein thickening and dimples (structures that, on the underside of the leaf, resembled enations but on the upper leaf surface were depressions;
Figure 3, panels D and E) on veins at 12 days post inoculation (dpi) of leaves developing subsequent to inoculation. At approximately 18 to 20 dpi the dimples were more pronounced and were associated with severe leaf curling, followed by foliar chlorosis at 30 dpi with the veins being raised on the lower surface and depressed on the upper surface (
Figure 3, panels F and G). Older plants were stunted, in comparison to healthy
N. benthamiana plants, with leaves showing a distinct downward curling and chlorosis (panel H).
For
N. tabacum, inoculation with AEV
SOL and AYLCB
SOL initially resulted in mild vein yellowing (at approx. 15 dpi;
Figure 3, panel A) followed by sunken veins (on the upper leaf surface; appearing as swollen vein on the lower surface)) and dimple structures, similar to those seen in
N. benthamiana, at 20 dpi (
Figure 3, panels B and C). However,
N. tabacum did not exhibit the leaf curling and chlorosis evident in
N. benthamiana.A. conyzoides plants infected with AEV
SOL/AYLCB
SOL initially showed mild leaf crumpling at 12 dpi. At approx. 18 dpi the leaves developing subsequent to inoculation showed crumpling and some leaf curling (
Figure 3, panel I).
For infected
S. lycopersicon plants inoculated with AEV
SOL and AYLCB
SOL the leaves developing subsequent to inoculation initially showed vein yellowing at 15 dpi. By 21 dpi leaves were narrow, rolled at the edges and showed a patchy necrosis developing from the veins (
Figure 3, panel J).
Inoculation of
N. benthamiana with the clones obtained from isolate ACL (AEV
ACL and AYLCB
ACL) led to the first symptoms appearing within 12 dpi, consisting of narrow leaves with a pointed apex with some foliar chlorosis for leaves developing subsequent to inoculation. As plants developed, the leaves remained small and chlorotic with some twisting (
Figure 4, panels B and C). Inoculated
A. conyzoides plants similarly showed the first symptoms within 12 dpi consisting of crumpling in leaves developing after inoculation. Subsequently plants remained severely stunted, with severe crumpling and deformation of leaves (
Figure 4, panel A). For tomato (
S. lycopersicon) plants inoculated with AEV
ACL and AYLCB
ACL the symptoms were not very distinct with older plants showing stunting, reduced leaf size and a mild foliar chlorosis (
Figure 4, panel D). On some older leaves a mild veinal necrosis developed, but this was not as pronounced as for tomato plants inoculated with AEV
SOL/AYLCB
SOL.
Table 2.
Infectivity of AEV, isolates SOL and ACL, with their AYLCB by Agrobacterium-mediated inoculation.
Table 2.
Infectivity of AEV, isolates SOL and ACL, with their AYLCB by Agrobacterium-mediated inoculation.
| Inoculum | Plant Species | Infectivity (Plants Infected/Plants Inoculated) | Latent Period # (days) | PCR Detection | Symptoms * |
|---|
| Virus | Betasatellite |
|---|
| AEVSOL and AYLCBSOL | N. benthamiana | 60/60 | 12–18 | + | + | D, LC, C |
| N. tabacum | 20/20 | 12–18 | + | + | VY, D |
| S. lycopersicon | 20/20 | 15–21 | + | + | C, N |
| A. conyzoides | 2/2 | 15–18 | + | + | CR, LC |
| S. oleraceus | 0/5 | - | - | - | - |
| AEVSOL | N. benthamiana | 22/30 | 12–18 | + | - | - |
| AYLCBSOL | N. benthamiana | 00/30 | - | - | - | - |
| pGreen0029 | N. benthamiana | 00/20 | - | - | - | - |
| AEVACL and AYLCBACL | N. benthamiana | 10/10 | 12–18 | + | + | R, LC, C |
| S. lycopersicon | 10/10 | 15–21 | + | + | SG, C, R, N |
| A. conyzoides | 2/2 | 15–18 | + | + | CR, LC |
| S. oleraceus | 0/5 | - | - | - | - |
| none | N. benthamiana | 00/20 | - | - | - | - |
| pGreen0029 | N. benthamiana | 00/20 | - | - | - | - |
The inoculations of AEV
SOL alone, AYLCB
SOL alone, and pGreen vector alone, in
N. benthamina did not show any apparent symptom (
Table 2). None of the symptoms was observed in healthy control
N. benthamiana plants. The
N. benthamiana plants, inoculated with AEV
SOL and AYLCB
SOL, produced dimples, leaf curl and chlorosis symptoms. A fragment of the expected size was obtained from
N. benthamiana plants inoculated with AEV
SOL alone in a PCR. The PCR product was sequenced and showed 100% nucleotide sequence identity to the inoculated AEV. The results showed that AEV
SOL alone is infectious but does not induce symptoms.
All attempts to infect S. oleraceus with AEV and AYLCB were unsuccessful. This was likely due to the plants exuding a white latex following inoculation by the pin-prick method used here, meaning that the Agrobacterium inoculum was likely not retained by the plant long enough for T-DNA transfer and establishment of an infection.
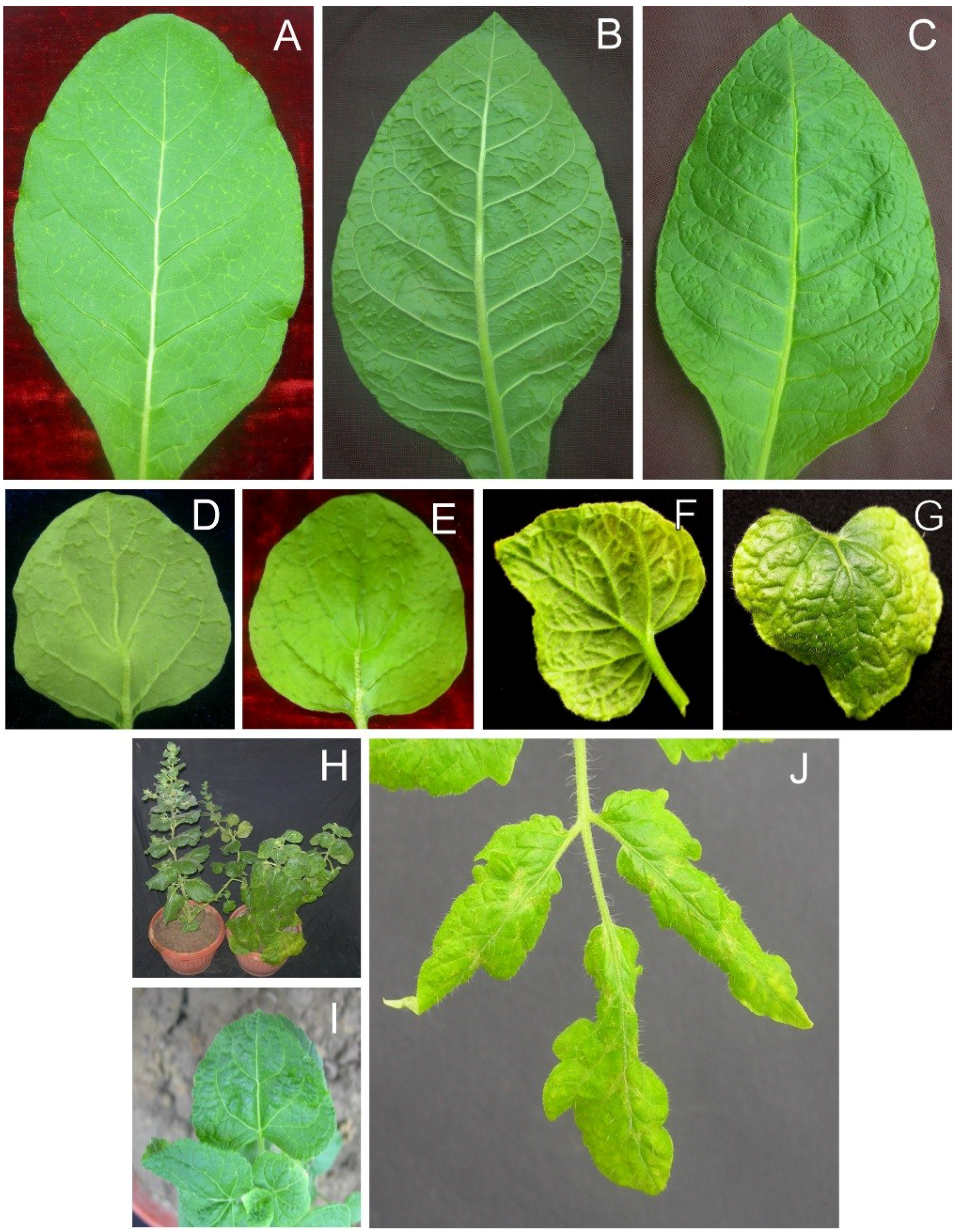
Figure 3.
Symptoms of plants infected with AEV and AYLCB clones obtained from isolate SOL. Nicotiana tabacum plants infected with AEVSOL/AYLCBSOL initially (15 dpi) showed mild vein yellowing on leaves developing subsequent to inoculation (A) which developed into what looked, on the underside of the leaf, like vein swelling (B) but on the upper leaf surface was evidently depression of the veins (C) at 20 dpi. Infected N. benthamiana initially (12 dpi) exhibited enations on the veins on the undersides of leaves (D) that on the upper leaf surface were seen to be depressions (dimples; E). In older N. benthamiana infections (30 dpi) leaves showed extensive chlorosis with raised veins on the lower surface (F) and sunken veins on the upper surface (G) giving a crumpled appearance. A comparison of a healthy, non-inoculated N. benthamiana plant (left) and an AEVSOL/AYLCBSOL infected plant at 30 dpi is shown in panel (H). The infected plant showed some chlorosis and overall downward curved leaves. The height of the plant was not greatly affected. Ageratum conyzoides plants infected with AEVSOL/AYLCBS°L showed mild leaf crumpling but no vein yellowing (I). Infected Solanum lycopersicon plants showed chlorosis of leaves developing after inoculation and vein necrosis (J).
Figure 3.
Symptoms of plants infected with AEV and AYLCB clones obtained from isolate SOL. Nicotiana tabacum plants infected with AEVSOL/AYLCBSOL initially (15 dpi) showed mild vein yellowing on leaves developing subsequent to inoculation (A) which developed into what looked, on the underside of the leaf, like vein swelling (B) but on the upper leaf surface was evidently depression of the veins (C) at 20 dpi. Infected N. benthamiana initially (12 dpi) exhibited enations on the veins on the undersides of leaves (D) that on the upper leaf surface were seen to be depressions (dimples; E). In older N. benthamiana infections (30 dpi) leaves showed extensive chlorosis with raised veins on the lower surface (F) and sunken veins on the upper surface (G) giving a crumpled appearance. A comparison of a healthy, non-inoculated N. benthamiana plant (left) and an AEVSOL/AYLCBSOL infected plant at 30 dpi is shown in panel (H). The infected plant showed some chlorosis and overall downward curved leaves. The height of the plant was not greatly affected. Ageratum conyzoides plants infected with AEVSOL/AYLCBS°L showed mild leaf crumpling but no vein yellowing (I). Infected Solanum lycopersicon plants showed chlorosis of leaves developing after inoculation and vein necrosis (J).
![]()
Figure 4.
Symptoms of plants infected with AEV and AYLCB clones obtained from isolate ACL. Ageratum conyzoides plants infected with AEVACL/AYLCBACL with severely distorted leaves but no evidence of vein yellowing (A). The leaves of N. benthamiana plants developing subsequent to inoculation were very narrow with sharp, curled tips and some chlorosis ((B) and some lateral curling (C)). Symptoms in Solanum lycopersicon were mild with a reduction in leaf size, mild chlorosis and plan stunting ((D), right) in comparisons to a healthy, non-inoculated plant (left). Plants were photographed at approx. 30 dpi.
Figure 4.
Symptoms of plants infected with AEV and AYLCB clones obtained from isolate ACL. Ageratum conyzoides plants infected with AEVACL/AYLCBACL with severely distorted leaves but no evidence of vein yellowing (A). The leaves of N. benthamiana plants developing subsequent to inoculation were very narrow with sharp, curled tips and some chlorosis ((B) and some lateral curling (C)). Symptoms in Solanum lycopersicon were mild with a reduction in leaf size, mild chlorosis and plan stunting ((D), right) in comparisons to a healthy, non-inoculated plant (left). Plants were photographed at approx. 30 dpi.