Lyssaviruses and Bats: Emergence and Zoonotic Threat
Abstract
:1. Introduction
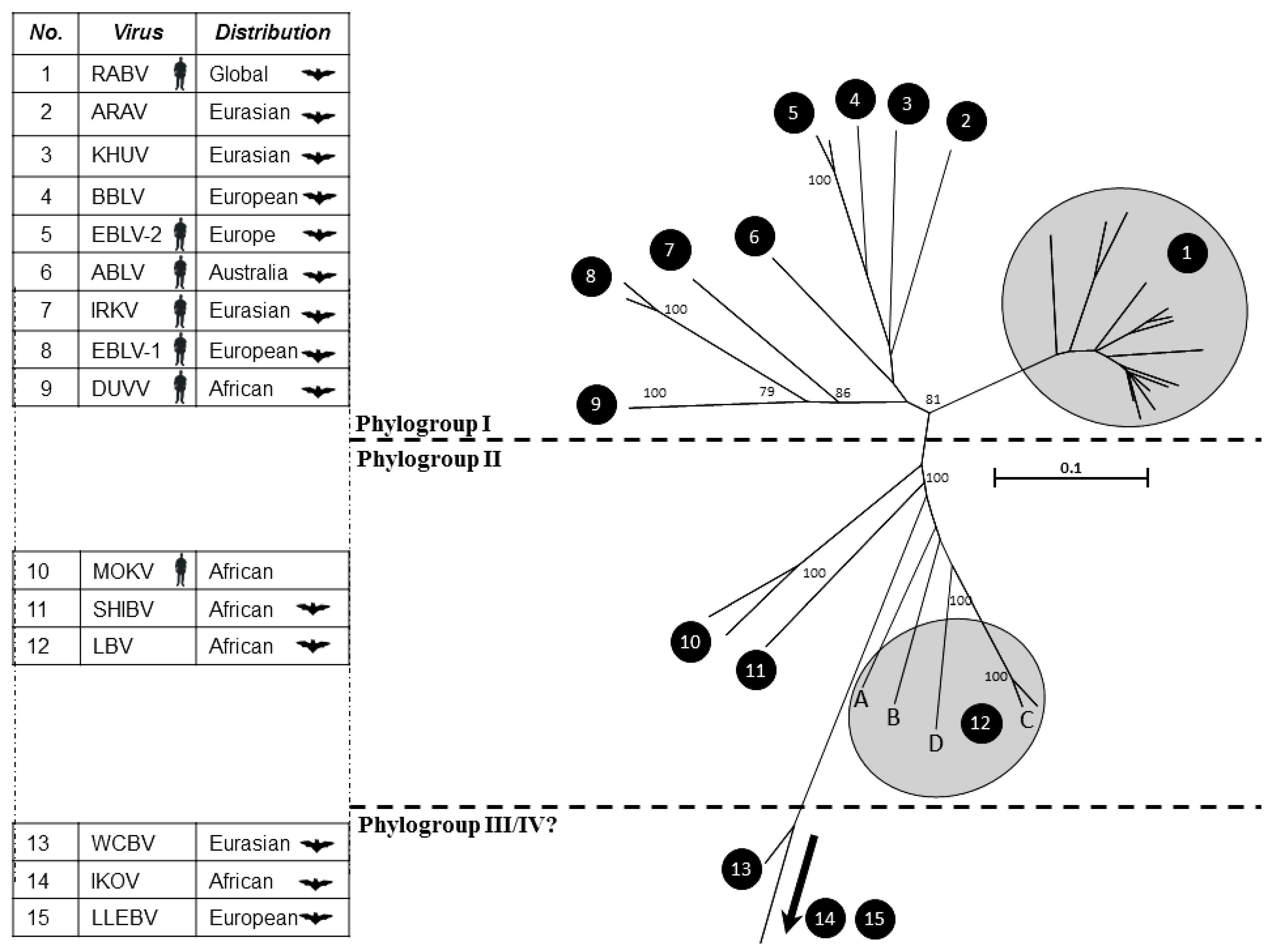
| Geographical distribution | Lyssavirus species | Bat species most commonly associated with lyssavirus infection | Common name | Transmission from bats implicated in human fatalities |
|---|---|---|---|---|
| The Americas | Rabies virus (RABV) | Eptesicus fuscus | Big brown bat | Yes |
| Tadarida brasiliensis | Mexican/Brazilian free-tail bat | Yes | ||
| Lasionycteris noctivagens | Silver-haired bat | Yes | ||
| Perimyotis subflavus | Tri-coloured bat | Yes | ||
| Desmodus rotundus | Vampire bat | Yes | ||
| Africa | Lagos Bat Virus (LBV) | Eidolon helvum | Straw coluored fruit bat | No |
| Rousettus aegyptiacus | Egyptian fruit bat | No | ||
| Epomorphorus wahlbergi | Wahlberg’s epauletted fruit bat | No | ||
| Shimoni Bat Virus (SHIBV) | Hipposideros commersoni | Commerson’s leaf-nosed bat | No | |
| Duvenhage virus (DUVV) | Miniopterus sp? | Undefined | Yes | |
| Nycteris thebaica | Egyptian slit-faced bat | Yes | ||
| Eurasia | European Bat Lyssavirus type 1 (EBLV-1) | Eptesicus serotinus | Serotine bat | Yes |
| European Bat Lyssavirus type 2 (EBLV-2) | Myotis daubentonii | Daubenton’s bat | Yes | |
| Bokeloh Bat Lyssavirus (BBLV) | Myotis nattereri | Natterer’s bat | No | |
| Aravan virus (ARAV) | Myotis blythi | Lesser mouse-eared bat | No | |
| Irkut Virus (IRKV) | Murina leucogaster | Greater tube-nosed bat | Yes | |
| Khujand Virus (KHUV) | Myotis mystacinus | Whiskered bat | No | |
| West Caucasian Bat Virus (WCBV) | Miniopterus schreibersii | Common bent-winged bat | No | |
| Lleida Bat Lyssavirus (LLEBV) * | Miniopterus schreibersii | Common bent-winged bat | No | |
| Australasia | Australian Bat Lyssavirus (ABLV) | Pteropus alecto | Black flying fox and related sp. | Yes |
| Saccolaimus flaviventris | Yellow-bellied sheath-tailed bat | Yes |
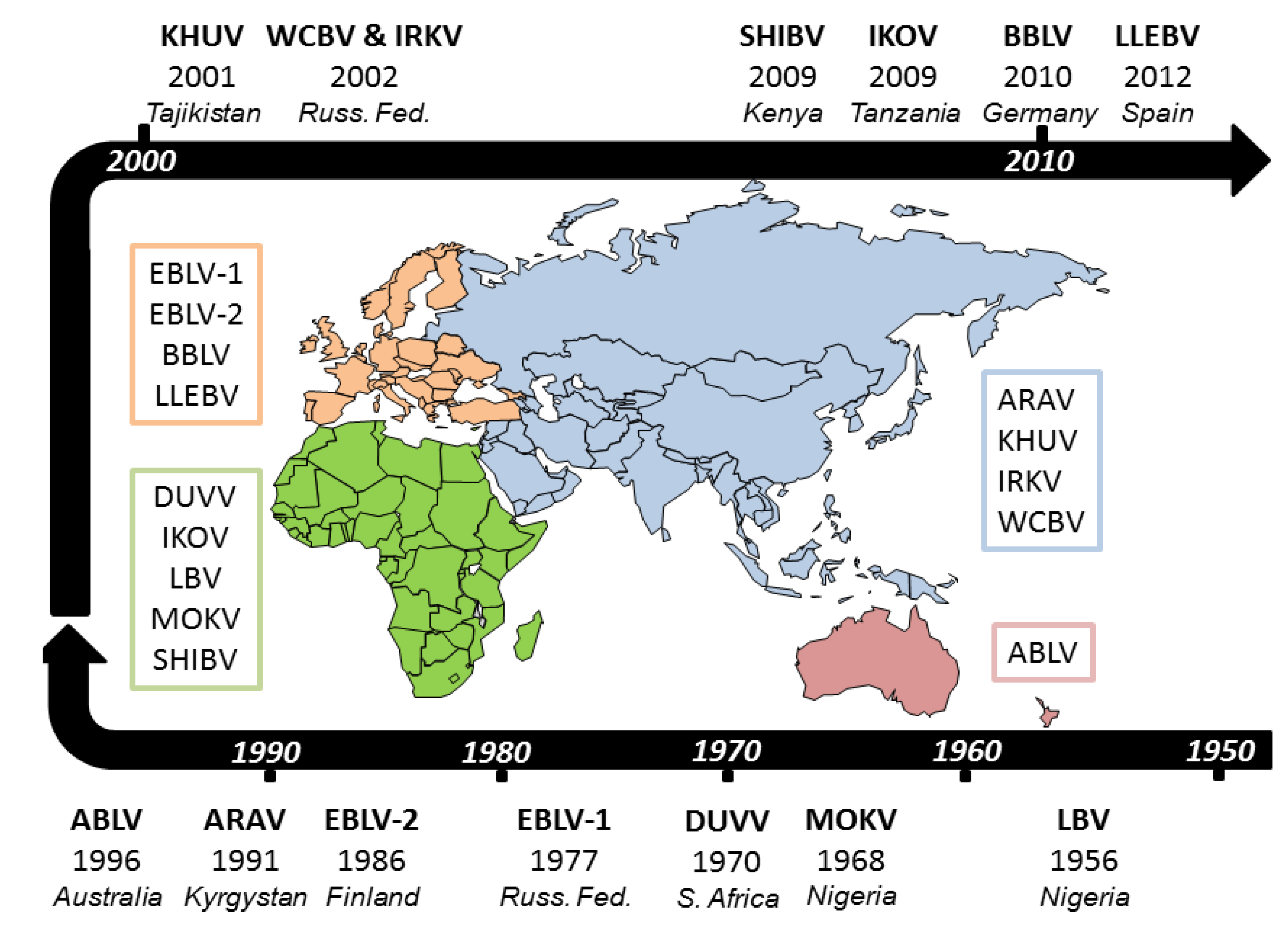
2. Infection of Different Bat Species
3. Transmission and Maintenance
4. Incubation Period
5. Serological Profiles
6. Novel Lyssavirus Detection
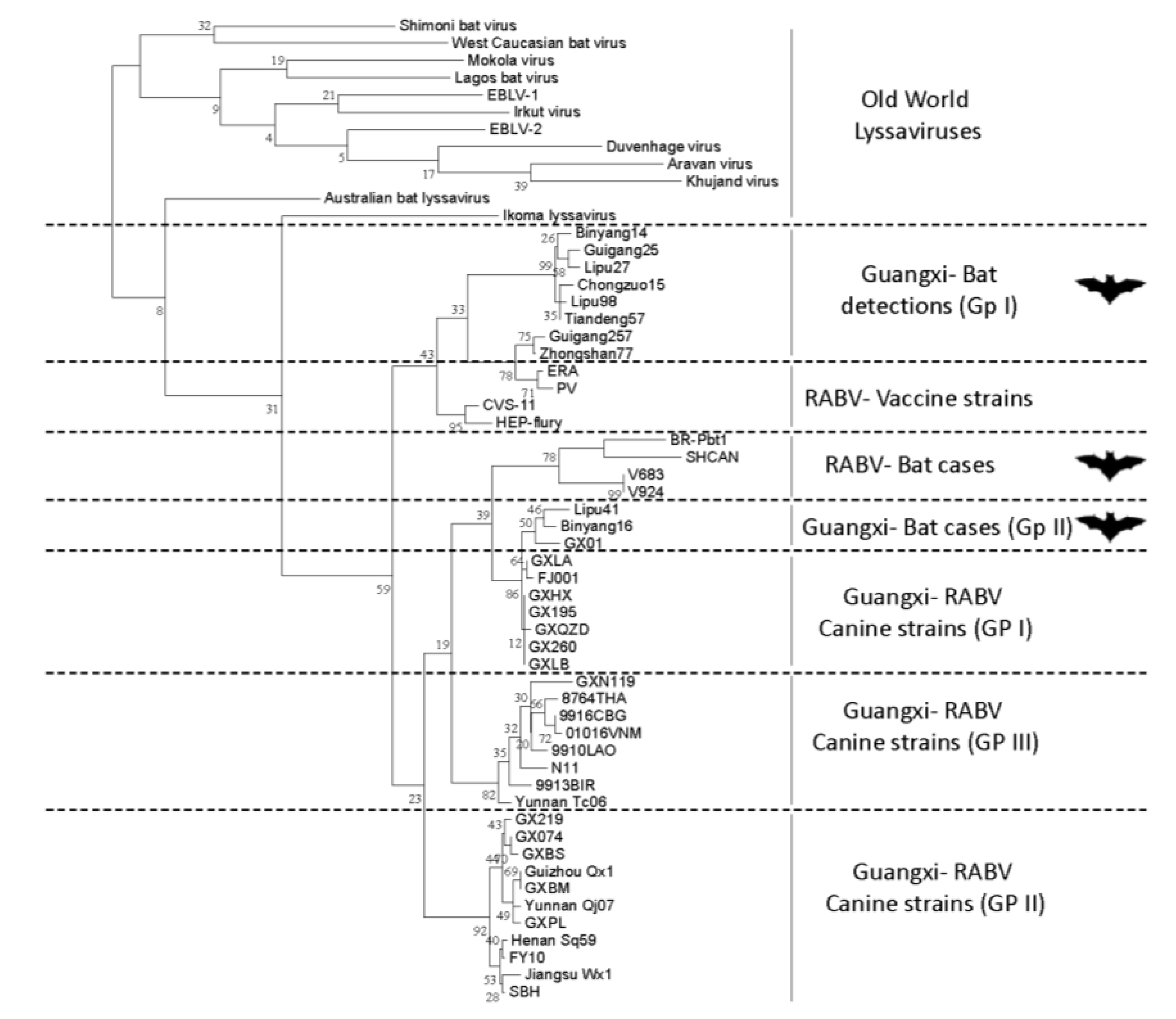
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Fooks, A.R.; Banyard, A.C.; Horton, D.L.; Johnson, N.; McElhinney, L.M.; Jackson, A.C. Current status of rabies and prospects for elimination. Lancet 2014. [Google Scholar] [CrossRef]
- Badrane, H.; Tordo, N. Host switching in lyssavirus history from the chiroptera to the carnivora orders. J. Virol. 2001, 75, 8096–8104. [Google Scholar] [PubMed]
- Streicker, D.G.; Turmelle, A.S.; Vonhof, M.J.; Kuzmin, I.V.; McCracken, G.F.; Rupprecht, C.E. Host phylogeny constrains cross-species emergence and establishment of rabies virus in bats. Science 2010, 329, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Carini, A. About one large epizootie of rabies. Ann. Inst. Pasteur. 1911, 25, 843–846. [Google Scholar]
- Pawan, J.L. Rabies in the vampire bat of trinidad, with special reference to the clinical course and the latency of infection. Ann. Trop. Med. Parasitol. 1936, 30, 410–422. [Google Scholar]
- Sulkin, S.E.; Greve, M.J. Human rabies caused by bat bite. Tex. State J. Med. 1954, 50, 620–621. [Google Scholar] [PubMed]
- Boulger, L.R.; Porterfield, J.S. Isolation of a virus from nigerian fruit bats. Trans. R. Soc. Trop. Med. Hyg. 1958, 52, 421–424. [Google Scholar] [CrossRef] [PubMed]
- Schneider, L.G.; Cox, J.H. Bat lyssaviruses in europe. Curr. Top. Microbiol. Immunol. 1994, 187, 207–218. [Google Scholar] [PubMed]
- Kemp, G.E.; Causey, O.R.; Moore, D.L.; Odelola, A.; Fabiyi, A. Mokola virus. Further studies on iban 27377, a new rabies-related etiologic agent of zoonosis in Nigeria. Am. J. Trop. Med. Hyg. 1997, 21, 356–359. [Google Scholar]
- Shope, R.E.; Murphy, F.A.; Harrison, A.K.; Causey, O.R.; Kemp, G.E.; Simpson, D.I.; Moore, D.L. Two african viruses serologically and morphologically related to rabies virus. J. Virol. 1970, 6, 690–692. [Google Scholar] [PubMed]
- Familusi, J.B.; Moore, D.L. Isolation of a rabies related virus from the cerebrospinal fluid of a child with ‘aseptic meningitis’. Afr. J. Med. Sci. 1972, 3, 93–96. [Google Scholar] [PubMed]
- Sabeta, C.T.; Markotter, W.; Mohale, D.K.; Shumba, W.; Wandeler, A.I.; Nel, L.H. Mokola virus in domestic mammals, South Africa. Emerg. Infect. Dis. 2007, 13, 1371–1373. [Google Scholar] [CrossRef] [PubMed]
- Nel, L.; Jacobs, J.; Jaftha, J.; von Teichman, B.; Bingham, J.; Olivier, M. New cases of mokola virus infection in south africa: A genotypic comparison of southern african virus isolates. Virus Genes 2000, 20, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Kgaladi, J.; Wright, N.; Coertse, J.; Markotter, W.; Marston, D.; Fooks, A.R.; Freuling, C.M.; Muller, T.F.; Sabeta, C.T.; Nel, L.H. Diversity and epidemiology of mokola virus. PLoS Negl. Trop. Dis. 2013, 7, e2511. [Google Scholar] [CrossRef] [PubMed]
- Familusi, J.B.; Osunkoya, B.O.; Moore, D.L.; Kemp, G.E.; Fabiyi, A. A fatal human infection with mokola virus. Am. J. Trop. Med. Hyg. 1972, 21, 959–963. [Google Scholar] [PubMed]
- Le Gonidec, G.; Rickenbach, A.; Robin, Y.; Heme, G. Isolation of a strain of mokola virus in cameroon. Ann. Microbiol. 1978, 129, 245–249. [Google Scholar]
- Meredith, C.D.; Prossouw, A.P.; Koch, H.P. An unusual case of human rabies thought to be of chiropteran origin. S. Afr. Med. J. 1971, 45, 767–769. [Google Scholar] [PubMed]
- King, A.; Crick, J. Rabies-related viruses. In Rabies; Campbell, J.B., Charlton, K.M., Eds.; Kluwer Academic Publishers: Boston, MA, USA, 1988; pp. 177–200. [Google Scholar]
- O’Shea, T.J.; Cryan, P.M.; Cunningham, A.A.; Fooks, A.R.; Hayman, D.T.; Luis, A.D.; Peel, A.J.; Plowright, R.K.; Wood, J.L. Bat flight and zoonotic viruses. Emerg. Infect. Dis. 2014, 20, 741–745. [Google Scholar] [CrossRef] [PubMed]
- Vogel, G. Are bats spreading ebola across Sub-Saharan Africa? Science 2014, 344, 140. [Google Scholar] [CrossRef] [PubMed]
- Olival, K.J.; Hayman, D.T. Filoviruses in bats: Current knowledge and future directions. Viruses 2014, 6, 1759–1788. [Google Scholar] [CrossRef] [PubMed]
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef] [PubMed]
- Hall, R.J.; Wang, J.; Peacey, M.; Moore, N.E.; McInnes, K.; Tompkins, D.M. New alphacoronavirus in mystacina tuberculata bats, New Zealand. Emerg. Infect. Dis. 2014, 20, 697–700. [Google Scholar] [PubMed]
- Banyard, A.C.; Hayman, D.T.; Freuling, C.M.; Mueller, T.; Fooks, A.R.; Johnson, N. Bat rabies. In Rabies, 3rd ed.; Jackson, A.C., Ed.; Elsevier: San Diego, CA, USA, 2013. [Google Scholar]
- Burnett, C.D. Bat rabies in Illinois: 1965 to 1986. J. Wildl. Dis. 1989, 25, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Childs, J.E.; Trimarchi, C.V.; Krebs, J.W. The epidemiology of bat rabies in new york state, 1988–1992. Epidemiol. Infect. 1994, 113, 501–511. [Google Scholar] [CrossRef] [PubMed]
- Crawford-Miksza, L.K.; Wadford, D.A.; Schnurr, D.P. Molecular epidemiology of enzootic rabies in California. J. Clin. Virol. 1999, 14, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Nadin-Davis, S.A.; Huang, W.; Armstrong, J.; Casey, G.A.; Bahloul, C.; Tordo, N.; Wandeler, A.I. Antigenic and genetic divergence of rabies viruses from bat species indigenous to Canada. Virus Res. 2001, 74, 139–156. [Google Scholar] [CrossRef] [PubMed]
- Mondul, A.M.; Krebs, J.W.; Childs, J.E. Trends in national surveillance for rabies among bats in the United States (1993–2000). J. Am. Vet. Med. Assoc. 2003, 222, 633–639. [Google Scholar] [CrossRef] [PubMed]
- Blanton, J.D.; Dyer, J.; McBrayer, J.; Rupprecht, C.E. Rabies surveillance in the united states during 2011. J. Am. Vet. Med. Assoc. 2012, 241, 712–722. [Google Scholar] [CrossRef] [PubMed]
- Sodre, M.M.; da Gama, A.R.; de Almeida, M.F. Updated list of bat species positive for rabies in Brazil. Rev. Inst. Med. Trop. Sao Paulo 2010, 52, 75–81. [Google Scholar] [PubMed]
- Oliveira Rde, N.; de Souza, S.P.; Lobo, R.S.; Castilho, J.G.; Macedo, C.I.; Carnieli, P., Jr.; Fahl, W.O.; Achkar, S.M.; Scheffer, K.C.; Kotait, I. Rabies virus in insectivorous bats: Implications of the diversity of the nucleoprotein and glycoprotein genes for molecular epidemiology. Virology 2010, 405, 352–360. [Google Scholar] [CrossRef] [PubMed]
- McElhinney, L.M.; Marston, D.A.; Leech, S.; Freuling, C.M.; van der Poel, W.H.; Echevarria, J.; Vazquez-Moron, S.; Horton, D.L.; Muller, T.; Fooks, A.R. Molecular epidemiology of bat lyssaviruses in europe. Zoonoses Public Health 2013, 60, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.W. The annual migrations of three species of west african fruit bats (chiroptera: Pteropodidae). Can. J. Zool. 1983, 61, 2266–2272. [Google Scholar] [CrossRef]
- Peel, A.J.; Sargan, D.R.; Baker, K.S.; Hayman, D.T.; Barr, J.A.; Crameri, G.; Suu-Ire, R.; Broder, C.C.; Lembo, T.; Wang, L.F.; et al. Continent-wide panmixia of an african fruit bat facilitates transmission of potentially zoonotic viruses. Nat. Commun. 2013, 4, 2770. [Google Scholar] [CrossRef] [PubMed]
- Luis, A.D.; Hayman, D.T.; O'Shea, T.J.; Cryan, P.M.; Gilbert, A.T.; Pulliam, J.R.; Mills, J.N.; Timonin, M.E.; Willis, C.K.; Cunningham, A.A.; et al. A comparison of bats and rodents as reservoirs of zoonotic viruses: Are bats special? Proc. Biol. Sci./R. Soc. 2013, 280, 20122753. [Google Scholar]
- Constantine, D.G. Rabies transmission by nonbite route. Public Health Rep. 1962, 77, 287–289. [Google Scholar] [CrossRef] [PubMed]
- Serra-Cobo, J.; Lopez-Roig, M.; Segui, M.; Sanchez, L.P.; Nadal, J.; Borras, M.; Lavenir, R.; Bourhy, H. Ecological factors associated with european bat lyssavirus seroprevalence in Spanish bats. PLoS One 2013, 8, e64467. [Google Scholar] [CrossRef] [PubMed]
- Hutson, A.M.; Mickleburgh, S.P.; Racey, P.A. Microchiropteran Bats: Global Status Survey and Conservation Action Plan; Information Press: Oxford, UK, 2001. [Google Scholar]
- Harris, S.L.; Brookes, S.M.; Jones, G.; Hutson, A.M.; Fooks, A.R. Passive surveillance (1987 to 2004) of united kingdom bats for european bat lyssaviruses. Vet. Rec. 2006, 159, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Harris, S.L.; Aegerter, J.N.; Brookes, S.M.; McElhinney, L.M.; Jones, G.; Smith, G.C.; Fooks, A.R. Targeted surveillance for european bat lyssaviruses in english bats (2003–2006). J. Wildl. Dis. 2009, 45, 1030–1041. [Google Scholar] [CrossRef] [PubMed]
- Hughes, G.J.; Kuzmin, I.V.; Schmitz, A.; Blanton, J.; Manangan, J.; Murphy, S.; Rupprecht, C.E. Experimental infection of big brown bats (eptesicus fuscus) with eurasian bat lyssavirus aravan, khujand, and irkut virus. Arch. Virol. 2006, 151, 2021–2035. [Google Scholar] [CrossRef] [PubMed]
- Freuling, C.; Vos, A.; Johnson, N.; Kaipf, I.; Denzinger, A.; Neubert, L.; Mansfield, K.; Hicks, D.; Nunez, A.; Tordo, N. Experimental infection of serotine bats (eptesicus serotinus) with european bat lyssavirus type 1a. J. Gen. Virol. 2009, 90, 2493–2502. [Google Scholar] [CrossRef] [PubMed]
- Johnson, N.; Vos, A.; Neubert, L.; Freuling, C.; Mansfield, K.L.; Kaipf, I.; Denzinger, A.; Hicks, D.; Nunez, A.; Franka, R. Experimental study of european bat lyssavirus type-2 infection in daubenton’s bats (myotis daubentonii). J. Gen. Virol. 2008, 89, 2662–2672. [Google Scholar] [CrossRef] [PubMed]
- Johnson, N.; McKimmie, C.S.; Mansfield, K.L.; Wakeley, P.R.; Brookes, S.M.; Fazakerley, J.K.; Fooks, A.R. Lyssavirus infection activates interferon gene expression in the brain. J. Gen. Virol. 2006, 87, 2663–2667. [Google Scholar] [CrossRef] [PubMed]
- Yamaoka, S.; Ito, N.; Ohka, S.; Kaneda, S.; Nakamura, H.; Agari, T.; Masatani, T.; Nakagawa, K.; Okada, K.; Okadera, K.; et al. Involvement of the rabies virus phosphoprotein gene in neuroinvasiveness. J. Virol. 2013, 87, 12327–12338. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Tang, L.; Tseggai, T.; Guo, Y.; Fu, Z.F. Wild-type rabies virus phosphoprotein is associated with viral sensitivity to type i interferon treatment. Arch. Virol. 2013, 158, 2297–2305. [Google Scholar] [CrossRef] [PubMed]
- Lieu, K.G.; Brice, A.; Wiltzer, L.; Hirst, B.; Jans, D.A.; Blondel, D.; Moseley, G.W. The rabies virus interferon antagonist p protein interacts with activated stat3 and inhibits gp130 receptor signaling. J. Virol. 2013, 87, 8261–8265. [Google Scholar] [CrossRef] [PubMed]
- Wiltzer, L.; Larrous, F.; Oksayan, S.; Ito, N.; Marsh, G.A.; Wang, L.F.; Blondel, D.; Bourhy, H.; Jans, D.A.; Moseley, G.W. Conservation of a unique mechanism of immune evasion across the lyssavirus genus. J. Virol. 2012, 86, 10194–10199. [Google Scholar] [CrossRef] [PubMed]
- Schatz, J.; Fooks, A.R.; McElhinney, L.; Horton, D.; Echevarria, J.; Vazquez-Moron, S.; Kooi, E.A.; Rasmussen, T.B.; Muller, T.; Freuling, C.M. Bat rabies surveillance in Europe. Zoonoses Public Health 2013, 60, 22–34. [Google Scholar] [CrossRef] [PubMed]
- Kgaladi, J.; Nel, L.H.; Markotter, W. Comparison of pathogenic domains of rabies and african rabies-related lyssaviruses and pathogenicity observed in mice. Onderstepoort J. Vet. Res. 2013, 80, 511. [Google Scholar] [CrossRef] [PubMed]
- Banyard, A.C.; Healy, D.M.; Brookes, S.M.; Voller, K.; Hicks, D.J.; Núñez, A.; Fooks, A.R. Lyssavirus infection: 'Low dose, multiple exposure' in the mouse model. Virus Res. 2014, 181, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Paweska, J.T.; Blumberg, L.H.; Liebenberg, C.; Hewlett, R.H.; Grobbelaar, A.A.; Leman, P.A.; Croft, J.E.; Nel, L.H.; Nutt, L.; Swanepoel, R. Fatal human infection with rabies-related duvenhage virus, South Africa. Emerg. Infect. Dis. 2006, 12, 1965–1967. [Google Scholar] [CrossRef] [PubMed]
- Van Thiel, P.P.; van den Hoek, J.A.; Eftimov, F.; Tepaske, R.; Zaaijer, H.J.; Spanjaard, L.; de Boer, H.E.; van Doornum, G.J.; Schutten, M.; Osterhaus, A.; et al. Fatal case of human rabies (duvenhage virus) from a bat in Kenya: The Netherlands, December 2007. Euro Surveill. 2008, 13. [Google Scholar]
- Fekadu, M.; Shaddock, J.H.; Baer, G.M. Intermittent excretion of rabies virus in the saliva of a dog two and six months after it had recovered from experimental rabies. Am. J. Trop. Med. Hyg. 1981, 30, 1113–1115. [Google Scholar] [PubMed]
- Fekadu, M.; Chandler, F.W.; Harrison, A.K. Pathogenesis of rabies in dogs inoculated with an ethiopian rabies virus strain. Immunofluorescence, histologic and ultrastructural studies of the central nervous system. Arch. Virol. 1982, 71, 109–126. [Google Scholar]
- Fekadu, M.; Shaddock, J.H.; Baer, G.M. Excretion of rabies virus in the saliva of dogs. J. Infect. Dis. 1982, 145, 715–719. [Google Scholar] [CrossRef] [PubMed]
- Johnson, N.; Fooks, A.; McColl, K. Human rabies case with long incubation, Australia. Emerg. Infect. Dis. 2008, 14, 1950–1951. [Google Scholar] [CrossRef] [PubMed]
- Shankar, V.; Dietzschold, B.; Koprowski, H. Direct entry of rabies virus into the central nervous system without prior local replication. J. Virol. 1991, 65, 2736–2738. [Google Scholar] [PubMed]
- Pajamo, K.; Harkess, G.; Goddard, T.; Marston, D.; McElhinney, L.; Johnson, N.; Fooks, A.R. Isolation of european bat lyssavirus type 2 (eblv-2) in a daubenton’s bat in the uk with a minimum incubation period of 9 months. Rabies Bulletin Europe 2008, 32, 6–7. [Google Scholar]
- Boots, M.; Sasaki, A. ‘Small worlds’ and the evolution of virulence: Infection occurs locally and at a distance. Proc. Biol. Sci. 1999, 266, 1933–1938. [Google Scholar] [CrossRef] [PubMed]
- Boots, M.; Hudson, P.J.; Sasaki, A. Large shifts in pathogen virulence relate to host population structure. Science 2004, 303, 842–844. [Google Scholar] [CrossRef] [PubMed]
- Ewald, P.W. The evolution of virulence. Sci. Am. 1993, 268, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Keeling, M.J. Modelling the persistence of measles. Trends Microbiol. 1997, 5, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Banyard, A.C.; Hayman, D.T.; Johnson, N.; McElhinney, L.; Fooks, A.R. Bats and lyssaviruses. In Research Advances in Rabies; Jackson, A.C., Ed.; Elsevier/Academic Press: London, UK, 2011; Volume 79, pp. 239–289. [Google Scholar]
- Arguin, P.M.; Murray-Lillibridge, K.; Miranda, M.E.; Smith, J.S.; Calaor, A.B.; Rupprecht, C.E. Serologic evidence of lyssavirus infections among bats, the philippines. Emerg. Infect. Dis. 2002, 8, 258–262. [Google Scholar] [CrossRef] [PubMed]
- Kuzmin, I.V.; Niezgoda, M.; Carroll, D.S.; Keeler, N.; Hossain, M.J.; Breiman, R.F.; Ksiazek, T.G.; Rupprecht, C.E. Lyssavirus surveillance in bats, Bangladesh. Emerg. Infect. Dis. 2006, 12, 486–488. [Google Scholar] [CrossRef] [PubMed]
- Lumlertdacha, B.; Boongird, K.; Wanghongsa, S.; Wacharapluesadee, S.; Chanhome, L.; Khawplod, P.; Hemachudha, T.; Kuzmin, I.; Rupprecht, C.E. Survey for bat lyssaviruses, Thailand. Emerg. Infect. Dis. 2005, 11, 232–236. [Google Scholar] [CrossRef] [PubMed]
- Reynes, J.M.; Molia, S.; Audry, L.; Hout, S.; Ngin, S.; Walston, J.; Bourhy, H. Serologic evidence of lyssavirus infection in bats, Cambodia. Emerg. Infect. Dis. 2004, 10, 2231–2234. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Wang, L.; Lu, Z.; Xuan, H.; Han, X.; Xia, X.; Zhao, F.; Tu, C. Seroprevalence of rabies virus antibodies in bats from Southern China. Vector Borne Zoonotic Dis. 2010, 10, 177–181. [Google Scholar] [CrossRef] [PubMed]
- Turmelle, A.S.; Jackson, F.R.; Green, D.; McCracken, G.F.; Rupprecht, C.E. Host immunity to repeated rabies virus infection in big brown bats. J. Gen. Virol. 2010, 91, 2360–2366. [Google Scholar] [CrossRef] [PubMed]
- Almeida, M.F.; Martorelli, L.F.; Aires, C.C.; Sallum, P.C.; Durigon, E.L.; Massad, E. Experimental rabies infection in haematophagous bats desmodus rotundus. Epidemiol. Infect. 2005, 133, 523–527. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, A.T.; Petersen, B.W.; Recuenco, S.; Niezgoda, M.; Gómez, J.; Laguna-Torres, V.A.; Rupprecht, C. Evidence of rabies virus exposure among humans in the peruvian Amazon. Am. J. Trop. Med. Hyg. 2012, 87, 206–215. [Google Scholar] [CrossRef] [PubMed]
- DeDiaz, A.M.; Fuenzalida, E.; Bell, J.F. Non-fatal rabies in dogs and cats. Ann. Microbiol. (Paris) 1975, 126, 503–509. [Google Scholar]
- Lafon, M. Modulation of the immune response in the nervous system by rabies virus. Curr. Top. Microbiol. Immunol. 2005, 289, 239–258. [Google Scholar] [PubMed]
- Galelli, A.; Baloul, L.; Lafon, M. Abortive rabies virus central nervous infection is controlled by t lymphocyte local recruitment and induction of apoptosis. J. Neurovirol. 2000, 6, 359–372. [Google Scholar] [CrossRef] [PubMed]
- ICTV. Virus Taxonomy: 2013 Release. Available online: http://ictvonline.org/virusTaxonomy.asp/ (accessed on 22 May 2014).
- Marston, D.A.; Horton, D.L.; Ngeleja, C.; Hampson, K.; McElhinney, L.M.; Banyard, A.C.; Haydon, D.; Cleaveland, S.; Rupprecht, C.E.; Bigambo, M.; et al. Ikoma lyssavirus, highly divergent novel lyssavirus in an African civet. Emerg. Infect. Dis. 2012, 18, 664–667. [Google Scholar] [CrossRef] [PubMed]
- Marston, D.A.; Ellis, R.J.; Horton, D.; Kuzmin, I.; Wise, E.L.; McElhinney, L.; Banyard, A.C.; Ngeleja, C.; Keyyu, J.; Cleveland, S.; et al. Complete genomic sequence of ikoma lyssavirus. J. Virol. 2012, 86, 10242–10243. [Google Scholar] [CrossRef] [PubMed]
- Horton, D.L.; Banyard, A.C.; Marston, D.A.; Wise, E.; Selden, D.; Nunez, A.; Hicks, D.; Lembo, T.; Cleaveland, S.; Peel, A.J.; et al. Antigenic and genetic characterization of a divergent african virus, ikoma lyssavirus. J. Gen. Virol. 2014, 95, 1025–1032. [Google Scholar] [CrossRef] [PubMed]
- Schatz, J.; Teifke, J.P.; Mettenleiter, T.C.; Aue, A.; Stiefel, D.; Muller, T.; Freuling, C.M. Lyssavirus distribution in naturally infected bats from Germany. Vet. Microbiol. 2014, 169, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Picard-Meyer, E.; Borel, C.; Moinet, M.; Servat, A.; Rasquin, P.; Cliquet, F. Découverte d’un Vespertilion de Natterer infecté par le Lyssavirus BBLV en Moselle en 2012. Bull. Epidémiol.-Santé Anim. Aliment. 2012, 55, 25. [Google Scholar]
- Picard-Meyer, E.; Servat, A.; Robardet, E.; Moinet, M.; Borel, C.; Cliquet, F. Isolation of bokeloh bat lyssavirus in myotis nattereri in France. Arch. Virol. 2013, 158, 2333–2340. [Google Scholar] [CrossRef] [PubMed]
- Freuling, C.M.; Beer, M.; Conraths, F.J.; Finke, S.; Hoffmann, B.; Keller, B.; Kliemt, J.; Mettenleiter, T.C.; Muhlbach, E.; Teifke, J.P.; et al. Novel lyssavirus in natterer’s bat, Germany. Emerg. Infect. Dis. 2011, 17, 1519–1522. [Google Scholar] [PubMed]
- Freuling, C.M.; Abendroth, B.; Beer, M.; Fischer, M.; Hanke, D.; Hoffmann, B.; Hoper, D.; Just, F.; Mettenleiter, T.C.; Schatz, J.; et al. Molecular diagnostics for the detection of bokeloh bat lyssavirus in a bat from bavaria, Germany. Virus Res. 2013, 177, 201–204. [Google Scholar] [CrossRef] [PubMed]
- Arechiga Ceballos, N.; Vazquez Moron, S.; Berciano, J.M.; Nicolas, O.; Aznar Lopez, C.; Juste, J.; Rodriguez Nevado, C.; Aguilar Setien, A.; Echevarria, J.E. Novel lyssavirus in bat, Spain. Emerg. Infect. Dis. 2013, 19, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.L.; Wang, W.; Yin, W.L.; Tang, H.B.; Pan, Y.; Liang, X.; Liu, Q.; Xiong, Y.; Minamoto, N.; Luo, T.R. Lyssavirus surveillance in bats of Southern China’s Guangxi province. Virus Genes 2013, 46, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, S.; Zhao, J.; Zhang, F.; Hu, R. Isolation of irkut virus from a murina leucogaster bat in China. PLoS Negl. Trop. Dis. 2013, 7, e2097. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, N.; Zhang, S.; Zhang, F.; Lian, H.; Wang, Y.; Zhang, J.; Hu, R. Analysis of the complete genome of the first irkut virus isolate from China: Comparison across the lyssavirus genus. Mol. Biol. Evol. 2013, 69, 687–693. [Google Scholar]
- Leslie, M.J.; Messenger, S.; Rohde, R.E.; Smith, J.; Cheshier, R.; Hanlon, C.; Rupprecht, C.E. Bat-associated rabies virus in skunks. Emerg. Infect. Dis. 2006, 12, 1274–1277. [Google Scholar] [CrossRef] [PubMed]
- Daoust, P.Y.; Wandeler, A.I.; Casey, G.A. Cluster of rabies cases of probable bat origin among red foxes in Prince Edward Island, Canada. J. Wildl. Dis. 1996, 32, 403–406. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Banyard, A.C.; Evans, J.S.; Luo, T.R.; Fooks, A.R. Lyssaviruses and Bats: Emergence and Zoonotic Threat. Viruses 2014, 6, 2974-2990. https://doi.org/10.3390/v6082974
Banyard AC, Evans JS, Luo TR, Fooks AR. Lyssaviruses and Bats: Emergence and Zoonotic Threat. Viruses. 2014; 6(8):2974-2990. https://doi.org/10.3390/v6082974
Chicago/Turabian StyleBanyard, Ashley C., Jennifer S. Evans, Ting Rong Luo, and Anthony R. Fooks. 2014. "Lyssaviruses and Bats: Emergence and Zoonotic Threat" Viruses 6, no. 8: 2974-2990. https://doi.org/10.3390/v6082974
APA StyleBanyard, A. C., Evans, J. S., Luo, T. R., & Fooks, A. R. (2014). Lyssaviruses and Bats: Emergence and Zoonotic Threat. Viruses, 6(8), 2974-2990. https://doi.org/10.3390/v6082974




