Abstract
Tumor suppressor p53 is widely known as ‘the guardian of the genome’ due to its ability to prevent the emergence of transformed cells by the induction of cell cycle arrest and apoptosis. However, recent studies indicate that p53 is also a direct transcriptional target of type I interferons (IFNs) and thus, it is activated by these cytokines upon viral infection. p53 has been shown to contribute to virus-induced apoptosis, therefore dampening the ability of a wide range of viruses to replicate and spread. Interestingly, recent studies also indicate that several IFN-inducible genes such as interferon regulatory factor 9 (IRF9), IRF5, IFN-stimulated gene 15 (ISG15) and toll-like receptor 3 (TLR3) are in fact, p53 direct transcriptional targets. These findings indicate that p53 may play a key role in antiviral innate immunity by both inducing apoptosis in response to viral infection, and enforcing the type I IFN response, and provide a new insight into the evolutionary reasons why many viruses encode p53 antagonistic proteins.
1. Introduction
An important host defense mechanism against virus infection, the type I IFNs, are rapidly induced to activate an antiviral state in infected and neighboring uninfected cells. Type I IFNs bind to the type I IFN receptor formed by IFNAR1 and IFNAR2 subunits [1,2] and trigger the activation of the canonical Jak/signal transducer and activator of transcription (STAT) signaling pathway and the trans-activation of antiviral genes, through a mechanism that depends to a great extent on STAT1, STAT2 and IRF9 [3]. These three transcription factors form the trimeric IFN-stimulated gene factor 3 (ISGF3) complex, which is relocated to the nucleus where it binds to the IFN-stimulated response elements (ISREs) in promoters of target genes to activate their transcription. Since type I IFNs stimulate p53 expression and virus infection activates p53 [4], this tumor suppressor has been recently included as a new component of the cellular antiviral defense mechanisms. Studies by us and others in recent years indicate that p53 is in fact, a key player in innate antiviral immunity by both enforcing the type I IFN response upon viral infection, and inducing apoptosis in infected cells [5-9]. Both actions coordinated by this tumor suppressor gene, help to thwart the replication of a wide variety of viruses both in vitro and in vivo [5-9]. In this review we try to provide a summary of the recent findings that involve p53 in antiviral immunity and that presumably, may help to explain not only why this protein is conserved in invertebrate organisms which do not undergo cancer-related diseases, but also, why this protein is so frequently targeted by viral proteins.
2. Regulation of p53 functions
The potent activity of p53 as inducer of cellular apoptosis and cell cycle arrest demands tight control of its function. In normal cells the level of p53 protein is low due to its short half-life [10] regulated mainly by murine double minute 2 (Mdm2) protein. Mdm2 inhibits p53 transactivation and prompts p53 for proteasomal degradation by promoting its ubiquitylation [11,12]. In contrast, p53 is stabilized and activated when, in response to oncogene expression, another tumor suppressor gene, namely ARF, binds to Mdm2 and blocks its ability to inhibit p53 function [13]. Similarly, the nucleolar protein nucleophosmin (NPM) also interacts with Mdm2 and promotes p53 stability and function [14]. A recently discovered regulator of p53 activity is the type I IFN pathway. IFN-α/β stimulates transcription of the gene encoding p53, through ISRE sites present in its promoter, resulting in an early induction of the protein [4], which is later held in time by the action of ARF [15]. Importantly, type I IFN seems also to induce p53 protein stabilization through posttranslational mechanisms that involve STAT1-dependent transcriptional downregulation of Mdm2 and direct protein interaction with p53, at least upon DNA damage [16]. In addition, although acute IFN treatment does not cause the activation of p53 [4], chronic treatment of primary cells with IFN-β leads to p53 phosphorylation at Ser-15 and engagement of p53 downstream genes, indicative of p53 activation [17]; suggesting that the cellular response to IFN-β is time-dependent. Infection with some viral agents, such as vesicular stomatitis virus (VSV), Newcastle disease virus (NDV) or herpes simplex virus (HSV), probably through the production of interferon, boosts p53 expression, and induces the phosphorylation of p53 at Ser-15 by ataxia telangiectasia mutated (ATM) kinase [4]. Phosphorylation of p53 in other amino acid residues in response to IFN or virus infection cannot be excluded since p53 has been also identified as a substrate for the double-stranded RNA (dsRNA) activated protein kinase R (PKR), an IFN-inducible protein kinase which can induce the phosphorylation of human p53 at Ser-392 in vitro [18,19], an event that has been proposed to be related with the stabilization and activation of p53 [20-22]. In this regard, induction of p53 as a downstream event of tumor necrosis factor alpha (TNF-α)-induced upregulation of PKR has been reported, and impaired p53-mediated responses to doxorubicin in PKR KO cells have been described [17,18,23].
4. p53 direct target genes in the type I interferon pathway
Takaoka and colleagues [4] identified p53 as a type I IFN transcriptional target and demonstrated the existence of a crosstalk between p53 and the IFN pathway both in tumor suppression and antiviral defense [4]. An increasing body of work indicates that in fact, p53 crosstalk takes place both at the level of IFN production and IFN signaling. These findings indicate that p53, through direct transcriptional upregulation of several target genes, influences the type I IFN pathway that further suggests an important role for p53 in innate immunity. Here, we have summarized a list of genes that are transcriptional targets of both type I IFN and p53 and the possible implications of this crosstalk in antiviral immunity.
4.1. IRF9
IRF9 is, along with STAT1 and STAT2 [43], an integral part of the ISGF3 complex, with a critical role in the activation of antiviral genes, as demonstrated by the severely immuno-compromised status of Irf9-/- mice [44]. The ISGF3 complex regulates the expression of antiviral genes harboring ISREs in their promoters, including p53 [4]. However, IRF9 is also a p53 transcriptional target. We have recently shown that p53 induces the transcriptional upregulation of IRF9, which contributes to enhance the IFN signaling pathway leading to upregulation of ISRE-dependent genes such as IRF7, retinoic acid-inducible gene 1 (RIG-I) or MxA [6]. Importantly, and probably due to IRF9-dependent activation of IRF7, p53 also contributed to IFN production in cells infected with viruses such as Sendai (SeV), (Cantell strain) [6]. This link between p53 activity and production of IFN by infected cells has been also described during HIV infection, suggesting that this effect is not dependent on the virus type [45]. Consistent with an important role of p53 at enforcing the type I IFN signaling pathway, mice that conserve wt p53 were more protected from SeV infection than their p53-/- counterparts [6]. Similar to our results showing upregulation of IRF9 by p53 at transcriptional level, Shen and colleagues reported that upregulation of p53 induced in response to influenza virus infection resulted also in the transactivation of IRF9 [46]. Finally, an upregulation of IFN signaling upon Hepatitis C virus (HCV) infection mediated by direct p53-IRF9 protein interaction has been also suggested [47]. Further research is required to determine whether these cellular responses are dependent on the kinetics of infection and/or the multiplicity of infection (MOI) employed in such experiments. The recent findings by Shen and colleagues indicating that p53 levels are upregulated upon infection with influenza A virus, leading to IRF9 and Bax upregulation [46], are consistent with previous studies that demonstrated that p53 is important for influenza-induced apoptosis, and thus, cells retaining wt p53 have an enhanced apoptotic response to influenza A infection, which results in impaired viral replication [7]. These findings indicate that p53 activity contributes to both the enhancement of the type I IFN through IRF9 upregulation as well as virus-induced apoptosis in response to influenza infection. Furthermore, treatment of cancer cells harboring wt p53 with Ribavirin induced mammalian target of rapamycin (mTOR)-dependent activation of p53 that resulted in IRF9 transcriptional upregulation and enhancement of IFN signaling [48]. These studies indicate that p53 directly regulates IRF9 activity and provides a new link between the type I IFN pathway and p53, not only with implications in antiviral defense but also in tumor suppression.
4.2. TLR3
The first member of the TLR protein family was a Drosophila protein implicated in dorsoventral patterning during embryonic development [49]. Further studies indicated that the intracellular domain of Drosophila Toll showed striking similarities to the intracellular domain of the mammalian interleukin-1 (IL-1) receptor [50]. To date different human homologous of Drosophila Toll have been identified and shown to induce activation of several pro-inflammatory cytokines upon pathogen exposure including type I IFNs, TNF-α and IL-8 [51]. TLRs are essential components of innate immunity due to their ability to sense pathogen-associated molecular patterns (PAMPs). Thus, TLRs along with other cellular pathogen sensors such as RIG-I or the NOD-like receptor (NLRs) family of proteins are considered pattern recognition receptors (PRRs) [52]. TLRs are well conserved from mammals to invertebrates underscoring their importance in innate immunity. In mammals, TLRs are composed presumably by 13 family members [53]. Among TLRs, TLR3, TLR7 and TLR9 play a major role in the recognition of virus infection due to their ability to sense double-stranded RNA (dsRNA), single stranded RNA (ssRNA) and CpG DNA motifs respectively [53], which are main subproducts of viral replication [54]. TLR3 is expressed in most cell types even though it might play a major role in recognition of dsRNA by conventional DCs, where it is expressed in cellular endosomes as opposed to fibroblasts in which it is expressed in the cell surface [55]. Upon recognition of dsRNA, TLR3 induces IFN production as well as IL-8, RANTES, and TNF-α, leading to DC maturation which enhances their ability to present virus-derived antigens to naïve T cells by exposure of such antigens by class I and II histocompatibility complex (MHC-I and MHC-II) [56]. Therefore, TLR3 plays a major role at the interface between innate and adaptive immunity.
A recent paper by Taura et al. indicated that TLR3 is a direct p53 downstream target gene [57]. Interestingly, the authors found not only a functional p53 binding site in the TLR3 promoter, but also they showed that TLR3-dependent up-regulation of IL-8 in response to dsRNA, is dependent to a great extent on p53 in the p53 wt HCT116 human colon cancer cell line [57]. Moreover, TLR3 stimulation with synthetic dsRNA (poly I:C) in wt p53 HCT116 cells induced phosphorylation of the TLR3 downstream target IRF3, a key event for the transactivation of the IFN-β promoter, whereas IRF3 phosphorylation was severely impaired in p53-/- cells [57]. In agreement with the author’s findings, unpublished results from our group indicate that cells lacking p53 are in fact, severely impaired in their ability to activate the IFN-β promoter. A still open question is whether p53 affects the function of conventional DCs, which express high levels of TLR3. If so, such findings would imply that p53 functions are important not only for innate antiviral immunity, but also to trigger adaptive immunity. Indeed, recent findings indicating that the type I IFN response is important for DC activation and antigen presentation [58,59] suggest that this may be actually the case. Moreover, this could also explain why p53-/- mice are more sensitive to some viral infections [4-6]. Further research is required to evaluate the possible role of p53 in the interface between innate and adaptive immunity as well as other possible cross-talks between p53 and the TLR family.
4.3. IRF5
IRF family members have been identified as transcription factors that act to prevent different types of cell stress, including viral infection and DNA damage [60]. In fact, the first IRF member discovered, IRF1, was previously shown to cooperate with p53 in the induction of apoptosis in cells undergoing DNA damage, as well as in the induction of cell cycle arrest through p21 [61]. Three members of the IRF family of proteins seem to play a major role as direct activators of the type I IFN response upon viral infection, IRF3, IRF7 and IRF5 [60,62]. These proteins are phosphorylated and translocated to the nucleus upon recognition of viral PAMPs by TLRs where they form part of a multimeric protein complex containing not only the IRFs, but also ATF-2/c-Jun, and NFkappaB, which leads to transactivation of the IFN-β promoter, namely the type I IFN enhanceosome [63].
IRF5 directly interacts with the TLR adaptor protein MyD88 and regulates TLR-dependent expression of pro-inflammatory cytokines such as IL-6 and IL-12 [64]. Interestingly, as opposed to IRF3 and IRF7, IRF5 activity over the type I IFN promoters seems to be to some extent virus-specific. IRF5 activity is upregulated in cells infected with NDV rather than in SeV infected cells [64]. These findings could indicate that IRF5 activity is restricted to viruses that are sensed by TLRs, and not to viruses preferentially recognized by the RIG-I signaling pathway, which seems to be the case of SeV [65]. Another unique feature of IRF5, as opposed to IRF3 and IRF7, is that its expression seems to be restricted to certain types of tissues. Constitutive expression of IRF5 can be detected primarily in lymphoid tissues and peripheral blood leukocytes (PBLs), and low levels of IRF5 can be found in skeletal muscle and prostate [66]. In agreement with the idea that IRF5 function is tissue restricted, MEFs from Irf5-/- mice did not show any defects in their ability to produce type I IFN when infected in vitro with VSV or herpes simplex virus-1 (HSV-1) but these mice were more susceptible to VSV and HSV-1-induced disease in vivo [67].
IRF5 was identified as a direct p53 target gene by Mori and colleagues [68]. In this study the authors noticed an increase in IRF5 levels in cancer cell lines retaining wt p53 exposed to DNA damaging agents, and confirmed that p53 was able to bind to and transactivate the IRF5 promoter [68]. More recent studies by Yanai and colleagues indicate that, in comparison with IRF1 that cooperates with p53 in both the induction of cell cycle arrest and apoptosis in response to DNA damage [61], IRF5 cooperates with p53 only in the induction of apoptosis in response to DNA damage but not in cell cycle arrest [67]. Even though both IRF5 and p53 seem to play an important role in innate immunity, it is currently unknown whether these two proteins cooperate in the induction of type I IFNs and/or antiviral genes.
4.4. ISG15
ISG15 is an ubiquitin homolog that regulates the function of multiple IFN-inducible genes upon activation of the type I IFN pathway [69]. Discovered more than two decades ago, its function as an ubiquitin-like protein modifier (a process that has been termed ISGylation) it’s just starting to be understood [69,70]. ISG15-dependent ISGylated proteins include important antiviral proteins such as PKR, RIG-I, or antiviral effector proteins such as MxA [71]. ISGylation usually drives protein activation since, as opposed to ubiquitylation; ISGylation does not promote protein degradation [69]. Thus, ISG15 impedes virus-dependent degradation of IRF3, prompting IFN-β transactivation [72]. However, the specific mechanisms by which ISG15 promotes the activation of its target genes are in most of the cases largely unknown [69].
The importance of ISG15 in the antiviral immune response is underscored by the fact that is one of the earliest IFN-induced genes [73]. Moreover, ISG15 has been shown to participate in specific aspects of antiviral immunity such as inhibition of EBOV budding and HIV assembly [74,75] and ISGylation by ISG15 has been also shown to be inhibited by influenza B non structural protein 1 (NS1) [76].
ISG15 was identified as a p53 direct target gene by Hummer and colleagues in 2001 [77]. The authors reported the presence of a functional p53 binding site adjacent to the core ISRE site of ISG15 [77]. Intriguingly, this paper suggests that p53 promotes ISG15 upregulation after dsRNA stimulation rather than in response to IFN treatment or virus infection [77]. In view of the recent findings that link p53 and TLR3, it is tempting to speculate that the observed effects on ISG15 could be mediated through p53-dependent upregulation of TLR3 activity. Further investigations are required to decipher whether ISG15 may participate in the p53-dependent antiviral response and whether p53 may induce a different panel of genes in response to virus infection or dsRNA treatment.
6. Summary
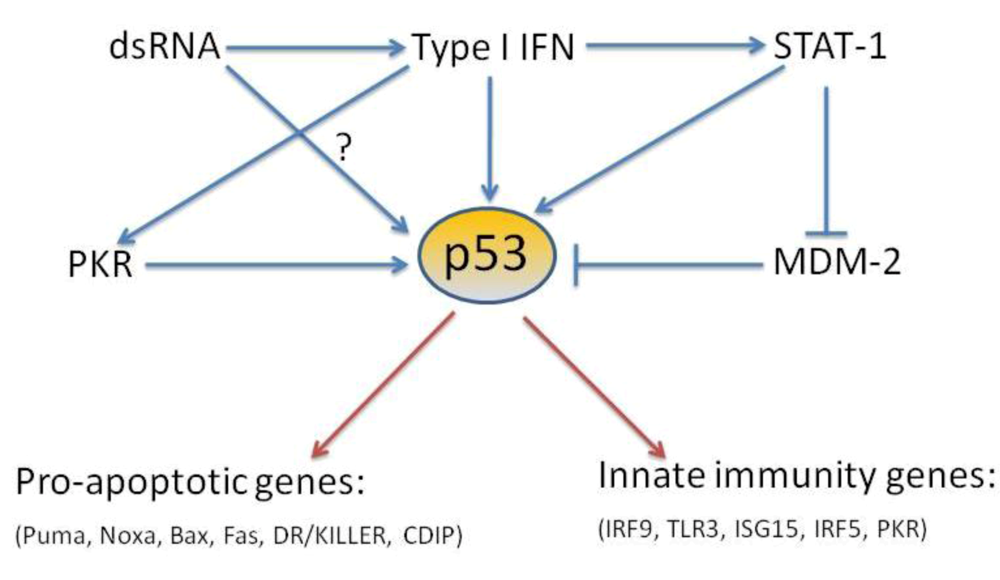
p53 is a tumor suppressor gene which is found mutated or absent in more than 50% of human cancers [109]. In fact, due to the ability of p53 to respond in many ways to DNA damaging insults, this protein has been called ‘the guardian of the genome’ [110]. However, p53 is also present and functional in organisms whose life span precludes them from undergoing cancer-related diseases such as flies and worms [111,112]. Moreover, a recent body of work indicates that besides ‘classic’ p53 inducers such as DNA damage and oncogene expression, its function can be also promoted by hypoxia, nutrient deprivation and viral infection [4,113,114]. These findings, and the recently identified functions of p53 in both virus-induced apoptosis and upregulation of the type I IFN response, are summarized in Figure 1, and suggest, that p53 may act as a virus induced cellular stress sensor. The role of p53 in antiviral immunity may help to explain to some extent why this protein is so highly conserved in evolution.
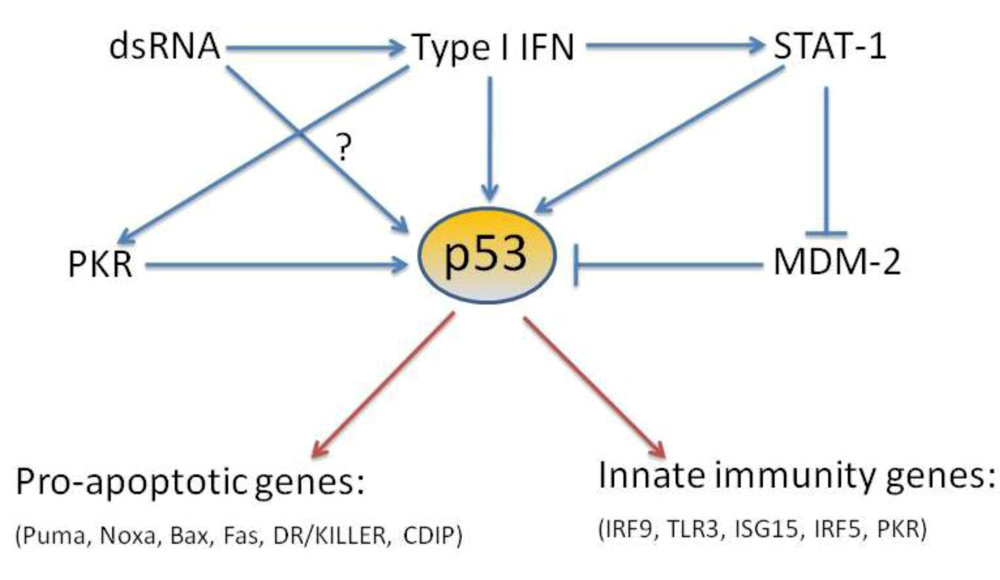
Figure 1.
Schematic of the activators and effectors of p53-dependent response in antiviral immunity.
Figure 1.
Schematic of the activators and effectors of p53-dependent response in antiviral immunity.
Acknowledgments
We thank E. Rodríguez and M. Collado for helpful comments and discussions about the manuscript. C.M.-F. is a recipient of a postdoctoral fellowship from the Spanish Ministry of Education and Science. This work was supported by NIH grants CA80058 and CA85214 to S.A.A. C.R. is funded by Spanish Ministry of Education and Science [BFU2008-03784/BMC] and CSIC [CCG08-CSIC/SAL-3441].
References
- Stark, G.R.; Kerr, I.M.; Williams, B.R.; Silverman, R.H.; Schreiber, R.D. How cells respond to interferons. Annu. Rev. Biochem. 1998, 67, 227–264. [Google Scholar] [CrossRef] [PubMed]
- García-Sastre, A.; Biron, C.A. Type 1 interferons and the virus-host relationship: a lesson in détente. Science 2006, 312, 879–882. [Google Scholar] [CrossRef] [PubMed]
- Platanias, L.C.; Fish, E.N. Signaling pathways activated by interferons. Exp. Hematol. 1999, 27, 1583–1592. [Google Scholar] [CrossRef] [PubMed]
- Takaoka, A.; Hayakawa, S.; Yanai, H.; Stoiber, D.; Negishi, H.; Kikuchi, H.; Sasaki, S.; Imai, K.; Shibue, T.; Honda, K.; Taniguchi, T. Integration of interferon-alpha/beta signalling to p53 responses in tumour suppression and antiviral defence. Nature 2003, 424, 516–523. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Fontela, C.; Garcia, M.A.; Garcia-Cao, I.; Collado, M.; Arroyo, J.; Esteban, M.; Serrano, M.; Rivas, C. Resistance to viral infection of super p53 mice. Oncogene 2005, 24, 3059–3062. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Fontela, C.; Macip, S.; Martínez-Sobrido, L.; Brown, L.; Ashour, J.; García-Sastre, A.; Lee, S.W.; Aaronson, S.A. Transcriptional role of p53 in interferon-mediated antiviral immunity. J. Exp. Med. 2008, 205, 1929–1938. [Google Scholar] [CrossRef] [PubMed]
- Turpin, E.; Luke, K.; Jones, J.; Tumpey, T.; Konan, K.; Schultz-Cherry, S. Influenza virus infection increases p53 activity: role of p53 in cell death and viral replication. J. Virol. 2005, 79, 8802–8811. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, M.L.; Kraft, R.M.; Aubert, M.; Goodwin, E.; DiMaio, D.; Blaho, J.A. p53 and hTERT determine sensitivity to viral apoptosis. J. Virol. 2007, 81, 12985–12995. [Google Scholar] [CrossRef] [PubMed]
- Pampin, M.; Simonin, Y.; Blondel, B.; Percherancier, Y.; Chelbi-Alix, M.K. Cross talk between PML and p53 during poliovirus infection: implications for antiviral defense. J. Virol. 2006, 80, 8582–8592. [Google Scholar] [CrossRef] [PubMed]
- Maki, C.G.; Huibregtse, J.M.; Howley, P.M. In vivo ubiquitination and proteasome-mediated degradation of p53(1). Cancer Res. 1996, 56, 2649–2654. [Google Scholar] [PubMed]
- Haupt, Y.; Maya, R.; Kazaz, A.; Oren, M. Mdm2 promotes the rapid degradation of p53. Nature 1997, 387, 296–269. [Google Scholar] [CrossRef] [PubMed]
- Momand, J.; Wu, H.H.; Dasgupta, G. MDM2-master regulator of the p53 tumor suppressor protein. Gene 2000, 242, 15–29. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, H.; Shimokado, K.; Asahara, T.; Dohi, K.; Niwa, O. Analysis of the c-myc, K-ras and p53 genes in methylcholanthrene-induced mouse sarcomas. Jpn. J. Cancer Res. 1999, 90, 40–47. [Google Scholar] [PubMed]
- Kurki, S.; Peltonen, K.; Latonen, L.; Kiviharju, T.M.; Ojala, P.M.; Meek, D.; Laiho, M. Nucleolar protein NPM interacts with HDM2 and protects tumor suppressor protein p53 from HDM2-mediated degradation. Cancer Cell 2004, 5, 465–475. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, R.; Xue, J.; Pilkinton, M.; Salvi, D.; Kiyokawa, H.; Colamonici, O.R. Different requirements for the cytostatic and apoptotic effects of type I interferons. Induction of apoptosis requires ARF but not p53 in osteosarcoma cell lines. J. Biol. Chem. 2004, 279, 32275–32280. [Google Scholar] [CrossRef] [PubMed]
- Townsend, P.A.; Scarabelli, T.M.; Davidson, S.M.; Knight Jr., R.A.; Latchman, D.S.; Stephanou, A. STAT-1 interacts with p53 to enhance DNA damage-induced apoptosis. J. Biol. Chem. 2004, 279, 5811–5820. [Google Scholar] [CrossRef] [PubMed]
- Moiseeva, O.; Mallette, F.A.; Mukhopadhyay, U.K.; Moores, A.; Ferbeyre, G. DNA damage signaling and p53-dependent senescence after prolonged beta-interferon stimulation. Mol. Biol. Cell 2006, 17, 1583–1592. [Google Scholar] [CrossRef] [PubMed]
- Cuddihy, A.R.; Wong, A.H.; Tam, N.W.; Li, S.; Koromilas, A.E. The double-stranded RNA activated protein kinase PKR physically associates with the tumor suppressor p53 protein and phosphorylates human p53 on serine 392 in vitro. Oncogene 1999, 18, 2690–2702. [Google Scholar] [CrossRef] [PubMed]
- Cuddihy, A.R.; Li, S.; Tam, N.W.; Wong, A.H.; Taya, Y.; Abraham, N.; Bell, J.C.; Koromilas, A.E. Double-stranded-RNA-activated protein kinase PKR enhances transcriptional activation by tumor suppressor p53. Mol. Cell Biol. 1999, 19, 2475–2484. [Google Scholar] [PubMed]
- Ullrich, S.J.; Sakaguchi, K.; Lees-Miller, S.P.; Fiscella, M.; Mercer, W.E.; Anderson, C.W.; Appella, E. Phosphorylation at Ser-15 and Ser-392 in mutant p53 molecules from human tumors is altered compared to wild-type p53. Proc. Natl. Acad. Sci. USA 1993, 90, 5954–5958. [Google Scholar] [CrossRef]
- Sakaguchi, K.; Sakamoto, H.; Lewis, M.S.; Anderson, C.W.; Erickson, J.W.; Appella, E.; Xie, D. Phosphorylation of serine 392 stabilizes the tetramer formation of tumor suppressor protein p53. Biochemistry 1997, 36, 10117–10124. [Google Scholar] [CrossRef] [PubMed]
- Keller, D.M.; Zeng, X.; Wang, Y.; Zhang, Q.H.; Kapoor, M.; Shu, H.; Goodman, R.; Lozano, G.; Zhao, Y.; Lu, H. A DNA damage-induced p53 serine 392 kinase complex contains CK2, hSpt16, and SSRP1. Mol. Cell 2001, 7, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Yeung, M.C.; Lau, A.S. Tumor suppressor p53 as a component of the tumor necrosis factor-induced, protein kinase PKR-mediated apoptotic pathway in human promonocytic U937 cells. J. Biol. Chem. 1998, 273, 25198–25202. [Google Scholar] [CrossRef] [PubMed]
- Son K.N., Pugazhenthi S. Activation of tumor suppressor protein p53 is required for Theiler's murine encephalomyelitis virus-induced apoptosis in M1-D macrophages. J. Virol. 2009, 83, 10770–10777. [Google Scholar] [CrossRef] [PubMed]
- Yoon, C.H.; Lee, E.S.; Lim, D.S.; Bae, Y.S. PKR, a p53 target gene, plays a crucial role in the tumor-suppressor function of p53. Proc. Natl. Acad. Sci. USA 2009, 106, 7852–7857. [Google Scholar] [CrossRef]
- Brown, L.; Ongusaha, P.P.; Kim, H.G.; Nuti, S.; Mandinova, A.; Lee, J.W.; Khosravi-Far, R.; Aaronson, S.A.; Lee, S.W. CDIP, a novel pro-apoptotic gene, regulates TNFalpha-mediated apoptosis in a p53-dependent manner. EMBO J. 2007, 26, 3410–3422. [Google Scholar] [CrossRef] [PubMed]
- Julkunen, I.; Melén, K.; Nyqvist, M.; Pirhonen, J.; Sareneva, T.; Matikainen, S. Inflammatory responses in influenza A virus infection. Vaccine 2000, 19, S32–S37. [Google Scholar] [CrossRef] [PubMed]
- Roux-Lombard, P.; Modoux, C.; Cruchaud, A.; Dayer, J.M. Purified blood monocytes from HIV 1-infected patients produce high levels of TNF alpha and IL-1. Clin. Immunol. Immunopathol. 1989, 50, 374–384. [Google Scholar] [CrossRef] [PubMed]
- Caux, C.; Massacrier, C.; Vanbervliet, B.; Dubois, B.; Van Kooten, C.; Durand, I.; Banchereau, J. Activation of human dendritic cells through CD40 cross-linking. J. Exp. Med. 1994, 180, 1263–1272. [Google Scholar] [CrossRef] [PubMed]
- Bartee, E.; Mohamed, M.R.; McFadden, G. Tumor necrosis factor and interferon: cytokines in harmony. Curr. Opin. Microbiol. 2008, 11, 378–383. [Google Scholar] [CrossRef] [PubMed]
- Belardelli, F. Role of interferons and other cytokines in the regulation of the immune response. APMIS 1995, 103, 161–179. [Google Scholar] [CrossRef] [PubMed]
- Mandron, M.; Martin, H.; Bonjean, B.; Lulé, J.; Tartour, E.; Davrinche, C. Dendritic cell-induced apoptosis of human cytomegalovirus-infected fibroblasts promotes cross-presentation of pp65 to CD8+ T cells. J. Gen. Virol. 2008, 89, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Chattergoon, M.A.; Muthumani, K.; Tamura, Y.; Ramanathan, M.; Shames, J.P.; Saulino, V.; Robinson, T.M.; Montaner, L.J.; Weiner, D.B. DR5 activation of caspase-8 induces DC maturation and immune enhancement in vivo. Mol. Ther. 2008, 16, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Albert, M.L.; Sauter, B.; Bhardwaj, N. Dendritic cells acquire antigen from apoptotic cells and induce class I-restricted CTLs. Nature 1998, 392, 86–89. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Wilder, S.; Bannasch, D.; Israeli, D.; Lehlbach, K.; Li-Weber, M.; Friedman, S.L.; Galle, P.R.; Stremmel, W.; Oren, M.; Krammer, PH. p53 activates the CD95 (APO-1/Fas) gene in response to DNA damage by anticancer drugs. J. Exp. Med. 1998, 188, 2033–2045. [Google Scholar] [CrossRef] [PubMed]
- Burns, T.F.; Bernhard, E.J.; El-Deiry, W.S. Tissue specific expression of p53 target genes suggests a key role for KILLER/DR5 in p53-dependent apoptosis in vivo. Oncogene 2001, 20, 4601–4612. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Qu, L.K.; Koromilas, A.E. Induction of p53-dependent apoptosis in HCT116 tumor cells by RNA viruses and possible implications in virus-mediated oncolysis. Cell Cycle 2004, 3, 1043–1045. [Google Scholar] [PubMed]
- Marques, J.T.; Rebouillat, D.; Ramana, C.V.; Murakami, J.; Hill, J.E.; Gudkov, A.; Silverman, R.H.; Stark, G.R.; Williams, B.R. Down-regulation of p53 by double-stranded RNA modulates the antiviral response. J. Virol. 2005, 79, 11105–11114. [Google Scholar] [CrossRef] [PubMed]
- Casavant, N.C.; Luo, M.H.; Rosenke, K.; Winegardner, T.; Zurawska, A.; Fortunato, E.A. Potential role for p53 in the permissive life cycle of human cytomegalovirus. J. Virol. 2006, 80, 8390–83401. [Google Scholar] [CrossRef] [PubMed]
- Groskreutz, D.J.; Monick, M.M.; Powers, L.S.; Quelle, D.E.; Varga, S.M.; Look, D.C.; Hunninghake, G.W. Respiratory syncytial virus decreases p53 protein to prolong survival of airway epithelial cells. J. Immunol. 2007, 179, 2741–2747. [Google Scholar] [PubMed]
- Royds, J.A.; Hibma, M.; Dix, B.R.; Hananeia, L.; Russell, I.A.; Wiles, A.; Wynford-Thomas, D.; Braithwaite, A.W. p53 promotes adenoviral replication and increases late viral gene expression. Oncogene 2006, 25, 1509–1520. [Google Scholar] [CrossRef] [PubMed]
- Teodoro, J.G.; Branton, P.E. Regulation of apoptosis by viral gene products. J. Virol. 1997, 7, 3620–3627. [Google Scholar]
- Bluyssen, A.R.; Durbin, J.E.; Levy, D.E. ISGF3 gamma p48, a specificity switch for interferon activated transcription factors. Cytokine Growth Factor Rev. 1996, 7, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Kadokawa, Y.; Harada, H.; Matsumoto, M.; Sato, M.; Kashiwazaki, Y.; Tarutani, M.; Tan, R.S.; Takasugi, T.; Matsuyama, T.; Mak, T.W.; Noguchi, S.; Taniguchi, T. Essential and non-redundant roles of p48 (ISGF3 gamma) and IRF-1 in both type I and type II interferon responses, as revealed by gene targeting studies. Genes Cells 1996, 1, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Imbeault, M.; Ouellet, M.; Tremblay, M.J. Microarray study reveals that HIV-1 induces rapid type-I interferon-dependent p53 mRNA up-regulation in human primary CD4+ T cells. Retrovirology 2009, 6. [Google Scholar] [CrossRef]
- Shen, Y.; Wang, X.; Guo, L.; Qiu, Y.; Li, X.; Yu, H.; Xiang, H.; Ma, Z. Influenza A virus induces p53 accumulation in a biphasic pattern. Biochem. Biophys. Res. Commun. 2009, 382, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Dharel, N.; Kato, N.; Muroyama, R.; Taniguchi, H.; Otsuka, M.; Wang, Y.; Jazag, A.; Shao, R.X.; Chang, J.H.; Adler, M.K.; Kawabe, T.; Omata, M. Potential contribution of tumor suppressor p53 in the host defense against hepatitis C virus. Hepatology 2008, 47, 1136–1149. [Google Scholar] [CrossRef] [PubMed]
- Su, W.C.; Liu, W.L.; Cheng, C.W.; Chou, Y.B.; Hung, K.H.; Huang, W.H.; Wu, C.L.; Li, Y.T.; Shiau, A.L.; Lai, M.Y. Ribavirin enhances interferon signaling via stimulation of mTOR and p53 activities. FEBS Lett. 2009, 583, 2793–2798. [Google Scholar] [CrossRef] [PubMed]
- Schneider, D.S.; Hudson, K.L.; Lin, T.Y.; Anderson, K.V. Dominant and recessive mutations define functional domains of Toll, a transmembrane protein required for dorsal-ventral polarity in the Drosophila embryo. Genes Dev. 1991, 5, 797–807. [Google Scholar] [CrossRef] [PubMed]
- Gay, N.J.; Keith, F.J. Drosophila Toll and IL-1 receptor. Nature 1991, 351, 355–356. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Akira, S. Antiviral signaling through pattern recognition receptors. J. Biochem. 2007, 14, 137–145. [Google Scholar]
- Kawai, T.; Akira, S. Innate immune recognition of viral infection. Nat. Immunol. 2006, 7, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Uematsu, S.; Akira, S. Toll-like receptors and innate immunity. J. Mol. Med. 2006, 84, 712–725. [Google Scholar] [CrossRef] [PubMed]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen recognition and innate immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef] [PubMed]
- Muzio, M.; Bosisio, D.; Polentarutti, N.; Stoppacciaro, A.; Mancinelli, R.; van't Veer, C.; Penton-Rol, G.; Ruco, L.P.; Allavena, P.; Mantovani, A. Differential expression and regulation of toll-like receptors (TLR) in human leukocytes: selective expression of TLR3 in dendritic cells. J. Immunol. 2000, 164, 5998–6004. [Google Scholar] [PubMed]
- Kaisho, T.; Akira, S. Regulation of dendritic cell function through Toll-like receptors. Curr. Mol. Med. 2003, 3, 373–385. [Google Scholar] [CrossRef] [PubMed]
- Taura, M.; Eguma, A.; Suico, M.A.; Shuto, T.; Koga, T.; Komune, T.; Sato, T.; Saya, H.; Li, J.D.; Kai, H. p53 regulates Toll-like receptor 3 expression and function in human epithelial cell lines. Mol. Cell. Biol. 2008, 28, 6557–6567. [Google Scholar] [CrossRef] [PubMed]
- Honda, K.; Sakaguchi, S.; Nakajima, C.; Watanabe, A.; Yanai, H.; Matsumoto, M.; Ohteki, T.; Kaisho, T.; Takaoka, A.; Akira, S.; Seya, T.; Taniguchi, T. Selective contribution of IFN-alpha/beta signaling to the maturation of dendritic cells induced by double-stranded RNA or viral infection. Proc. Natl. Acad. Sci. USA 2003, 100, 10872–10877. [Google Scholar] [CrossRef]
- Le Bon, A.; Etchart, N.; Rossmann, C.; Ashton, M.; Hou, S.; Gewert, D.; Borrow, P.; Tough, D.F. Cross-priming of CD8+ T cells stimulated by virus-induced type I interferon. Nat. Immunol. 2003, 4, 1009–1015. [Google Scholar] [CrossRef]
- Honda, K.; Taniguchi, T. IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat. Rev. Immunol. 2006, 6, 644–658. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, N.; Ishihara, M.; Lamphier, M.S.; Nozawa, H.; Matsuyama, T.; Mak, T.W.; Aizawa, S.; Tokino, T.; Oren, M.; Taniguchi, T. Cooperation of the tumour suppressors IRF-1 and p53 in response to DNA damage. Nature 1996, 382, 816–818. [Google Scholar] [CrossRef] [PubMed]
- Barnes, B.; Lubyova, B.; Pitha, P.M. On the role of IRF in host defense. J. Interferon Cytokine Res. 2002, 22, 59–71. [Google Scholar] [PubMed]
- Panne, D.; Maniatis, T.; Harrison, S.C. An atomic model of the interferon-beta enhanceosome. Cell 2007, 129, 1111–1123. [Google Scholar] [CrossRef] [PubMed]
- Takaoka, A.; Yanai, H.; Kondo, S.; Duncan, G.; Negishi, H.; Mizutani, T.; Kano, S.; Honda, K.; Ohba, Y.; Mak, T.W.; Taniguchi, T. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature 2005, 434, 243–249. [Google Scholar] [CrossRef] [PubMed]
- López, C.B.; Moltedo, B.; Alexopoulou, L.; Bonifaz, L.; Flavell, R.A.; Moran; T.M. TLR-independent induction of dendritic cell maturation and adaptive immunity by negative-strand RNA viruses. J. Immunol. 2004, 173, 6882–6889. [Google Scholar] [PubMed]
- Barnes, B.J.; Moore, P.A.; Pitha, P.M. Virus-specific activation of a novel interferon regulatory factor, IRF-5, results in the induction of distinct interferon alpha genes. J. Biol. Chem. 2001, 276, 23382–23390. [Google Scholar] [CrossRef] [PubMed]
- Yanai, H.; Chen, H.M.; Inuzuka, T.; Kondo, S.; Mak, T.W.; Takaoka, A.; Honda, K.; Taniguchi, T. Role of IFN regulatory factor 5 transcription factor in antiviral immunity and tumor suppression. Proc. Natl. Acad. Sci. USA 2007, 104, 3402–3407. [Google Scholar] [CrossRef]
- Mori, T.; Anazawa, Y.; Iiizumi, M.; Fukuda, S.; Nakamura, Y.; Arakawa, H. Identification of the interferon regulatory factor 5 gene (IRF-5) as a direct target for p53. Oncogene 2002, 21, 2914–2918. [Google Scholar] [CrossRef] [PubMed]
- Pitha-Rowe, I.F.; Pitha, P.M. Viral defense, carcinogenesis and ISG15: novel roles for an old ISG. Cytokine Growth Factor Rev. 2007, 18, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Farrell, P.J.; Broeze, R.J.; Lengyel, P. Accumulation of an mRNA and protein in interferon-treated Ehrlich ascites tumour cells. Nature 1979, 279, 523–525. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Denison, C.; Huibregtse, J.M.; Gygi, S.; Krug, R.M. Human ISG15 conjugation targets both IFN-induced and constitutively expressed proteins functioning in diverse cellular pathways. Proc. Natl. Acad. Sci. USA 2005, 102, 10200–10205. [Google Scholar] [CrossRef]
- Lu, G.; Reinert, J.T.; Pitha-Rowe, I.; Okumura, A.; Kellum, M.; Knobeloch, K. P.; Hassel, B.; Pitha, P.M. ISG15 enhances the innate antiviral response by inhibition of IRF-3 degradation. Cell Mol. Biol. (Noisy-le-grand) 2006, 52, 29–41. [Google Scholar] [PubMed]
- Blomstrom, D.C.; Fahey, D.; Kutny, R.; Korant, B.D.; Knight Jr., E. Molecular characterization of the interferon-induced 15-kDa protein. Molecular cloning and nucleotide and amino acid sequence. J. Biol. Chem. 1986, 261, 8811–8816. [Google Scholar] [PubMed]
- Okumura, A.; Pitha, P.M.; Harty, R.N. ISG15 inhibits Ebola VP40 VLP budding in an L-domain-dependent manner by blocking Nedd4 ligase activity. Proc. Natl. Acad. Sci. USA 2008, 105, 3974–3979. [Google Scholar] [CrossRef]
- Okumura, A.; Lu, G.; Pitha-Rowe, I.; Pitha, P.M. Innate antiviral response targets HIV-1 release by the induction of ubiquitin-like protein ISG15. Proc. Natl. Acad. Sci. USA 2006, 103, 1440–1445. [Google Scholar] [CrossRef]
- Yuan, W.; Krug, R.M. Influenza B virus NS1 protein inhibits conjugation of the interferon (IFN)-induced ubiquitin-like ISG15 protein. EMBO J. 2001, 20, 362–371. [Google Scholar] [CrossRef] [PubMed]
- Hummer, B.T.; Li, X.L.; Hassel, B.A. Role for p53 in gene induction by double-stranded RNA. J. Virol. 2001, 75, 7774–7777. [Google Scholar] [CrossRef] [PubMed]
- Levine, A.J. The common mechanisms of transformation by the small DNA tumor viruses: The inactivation of tumor suppressor gene products: p53. Virology 2009, 384, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Klein, G. Perspectives in studies of human tumor viruses. Front Biosci. 2002, 7, 268–274. [Google Scholar] [CrossRef]
- Pan, H.; Yin, C.; Van Dyke, T. Apoptosis and cancer mechanisms. Cancer Surv. 1997, 29, 305–327. [Google Scholar] [PubMed]
- Martin, M.E.; Berk, A.J. Adenovirus E1B 55K represses p53 activation in vitro. J. Virol. 1998, 72, 3146–3154. [Google Scholar] [PubMed]
- Zhang, Q.; Gutsch, D.; Kenney, S. Functional and physical interaction between p53 and BZLF1: implications for Epstein-Barr virus latency. Mol. Cell Biol. 1994, 14, 1929–1938. [Google Scholar] [PubMed]
- Parker, G.A.; Crook, T.; Bain, M.; Sara, E.A.; Farrell, P.J.; Allday, M.J. Epstein-Barr virus nuclear antigen (EBNA)3C is an immortalizing oncoprotein with similar properties to adenovirus E1A and papillomavirus E7. Oncogene 1996, 13, 2541–2549. [Google Scholar] [PubMed]
- Friborg Jr., J.; Kong, W.; Hottiger, M.O.; Nabel, G.J. p53 inhibition by the LANA protein of KSHV protects against cell death. Nature 1999, 402, 889–894. [Google Scholar] [PubMed]
- Rivas, C.; Thlick, A.E.; Parravicini, C.; Moore, P.S.; Chang, Y. Kaposi's sarcoma-associated herpesvirus LANA2 is a B-cell-specific latent viral protein that inhibits p53. J. Virol. 2001, 75, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Damania, B.; Jeong, J.H.; Bowser, B.S.; DeWire, S.M.; Staudt, M.R.; Dittmer, D.P. Comparison of the Rta/Orf50 transactivator proteins of gamma-2-herpesviruses. J. Virol. 2004, 78, 5491–5499. [Google Scholar] [CrossRef] [PubMed]
- Kashanchi, F.; Araujo, J.; Doniger, J.; Muralidhar, S.; Hoch, R.; Khleif, S.; Mendelson, E.; Thompson, J.; Azumi, N.; Brady, J.N.; Luppi, M.; Torelli, G.; Rosenthal, L.J. Human herpesvirus 6 (HHV-6) ORF-1 transactivating gene exhibits malignant transforming activity and its protein binds to p53. Oncogene 1997, 14, 359–367. [Google Scholar] [PubMed]
- Zantema, A.; Fransen, J.A.; Davis-Olivier, A.; Ramaekers, F.C.; Vooijs, G.P.; DeLeys, B.; Van der Eb, A.J. Localization of the E1B proteins of adenovirus 5 in transformed cells, as revealed by interaction with monoclonal antibodies. Virology 1985, 142, 44–58. [Google Scholar] [CrossRef] [PubMed]
- Dobner, T.; Horikoshi, N.; Rubenwolf, S.; Shenk, T. Blockage by adenovirus E4orf6 of transcriptional activation by the p53 tumor suppressor. Science 1996, 272, 1470–1473. [Google Scholar] [PubMed]
- Werness, B.A.; Levine, A.J.; Howley, P.M. Association of human papillomavirus types 16 and 18 E6 proteins with p53. Science 1990, 248, 76–79. [Google Scholar] [PubMed]
- Szekely, L.; Selivanova, G.; Magnusson, K.P.; Klein, G.; Wiman, K.G. EBNA-5, an Epstein-Barr virus-encoded nuclear antigen, binds to the retinoblastoma and p53 proteins. Proc. Natl. Acad. Sci. USA 1993, 90, 5455–5459. [Google Scholar] [CrossRef]
- Lee, H.R.; Toth, Z.; Shin, Y.C.; Lee, J.S.; Chang, H.; Gu, W.; Oh, T.K.; Kim, M.H.; Jung, J.U. Kaposi's sarcoma-associated herpesvirus viral interferon regulatory factor 4 targets MDM2 to deregulate the p53 tumor suppressor pathway. J. Virol. 2009, 83, 6739–6747. [Google Scholar] [CrossRef] [PubMed]
- Santos, C.R.; Vega, F.M.; Blanco, S.; Barcia, R.; Lazo, P.A. The vaccinia virus B1R kinase induces p53 downregulation by an Mdm2-dependent mechanism. Virology 2004, 328, 254–265. [Google Scholar] [CrossRef] [PubMed]
- Massimi, P.; Banks, L. Repression of p53 transcriptional activity by the HPV E7 proteins. Virology 1997, 227, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Seavey, S.E.; Holubar, M.; Saucedo, L.J.; Perry, M.E. The E7 oncoprotein of human papillomavirus type 16 stabilizes p53 through a mechanism independent of p19(ARF). J. Virol. 1999, 73, 7590–7598. [Google Scholar] [PubMed]
- Li, Z; Day, C.P.; Yang, J.Y.; Tsai, W.B.; Lozano, G.; Shih, H.M.; Hung, M.C. Adenoviral E1A targets Mdm4 to stabilize tumor suppressor p53. Cancer Res. 2004, 64, 9080–9085. [Google Scholar] [CrossRef] [PubMed]
- Lowe, S.W; Ruley, H.E. Stabilization of the p53 tumor suppressor is induced by adenovirus 5 E1A and accompanies apoptosis. Genes Dev. 1993, 7, 535–545. [Google Scholar] [CrossRef] [PubMed]
- de Stanchina, E; McCurrach, M.E.; Zindy, F.; Shieh, S.Y.; Ferbeyre, G.; Samuelson, A.V.; Prives, C.; Roussel, M.F.; Sherr, C.J.; Lowe, S.W. E1A signaling to p53 involves the p19(ARF) tumor suppressor. Genes Dev. 1998, 12, 2434–2442. [Google Scholar] [CrossRef] [PubMed]
- Seo, T.; Park, J.; Lee, D.; Hwang, S.G.; Choe, J. Viral interferon regulatory factor 1 of Kaposi's sarcoma-associated herpesvirus binds to p53 and represses p53-dependent transcription and apoptosis. J. Virol. 2001, 75, 6193–6198. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.C.; Nakamura, H.; Liang, X.; Feng, P.; Chang, H.; Kowalik, T.F.; Jung, J.U. Inhibition of the ATM/p53 signal transduction pathway by Kaposi's sarcoma-associated herpesvirus interferon regulatory factor 1. J. Virol. 2006, 80, 2257–2266. [Google Scholar] [CrossRef] [PubMed]
- Kashuba, E.; Mattsson, K.; Pokrovskaja, K.; Kiss, C.; Protopopova, M.; Ehlin-Henriksson, B.; Klein, G.; Szekely, L. EBV-encoded EBNA-5 associates with P14ARF in extranucleolar inclusions and prolongs the survival of P14ARF-expressing cells. Int. J. Cancer 2003, 105, 644–653. [Google Scholar] [CrossRef] [PubMed]
- Moule, M.G.; Collins, C.H.; McCormick, F.; Fried, M. Role for PP2A in ARF signaling to p53. Proc. Natl. Acad. Sci. USA 2004, 101, 14063–14066. [Google Scholar] [CrossRef]
- Feitelson, M.A.; Zhu, M.; Duan, L.X.; London, W.T. Hepatitis B x antigen and p53 are associated in vitro and in liver tissues from patients with primary hepatocellular carcinoma. Oncogene 1993, 8, 1109–1117. [Google Scholar] [PubMed]
- Li, L.; Zhou, S.; Chen, X.; Guo, L.; Li, Z.; Hu, D.; Luo, X.; Ma, X.; Tang, M.; Yi, W.; Tsao, S.W.; Cao, Y. The activation of p53 mediated by Epstein-Barr virus latent membrane protein 1 in SV40 large T-antigen transformed cells. FEBS Lett. 2008, 582, 755–762. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Marker, P.H.; Belcher, J.D.; Wilcken, D.E.; Burns, L.J.; Vercellotti, G.M.; Wang, X.L. Human cytomegalovirus immediate early proteins upregulate endothelial p53 function. FEBS Lett. 2000, 474, 213–216. [Google Scholar] [CrossRef] [PubMed]
- Eckardt-Michel, J.; Lorek, M.; Baxmann, D.; Grunwald, T.; Kei, G.M.; Zimmer, G. The fusion protein of respiratory syncytial virus triggers p53-dependent apoptosis. J. Virol. 2008, 82, 3236–3249. [Google Scholar] [CrossRef] [PubMed]
- Genini, D.; Sheeter, D.; Rought, S.; Zaunders, J.J.; Susin, S.A.; Kroemer, G.; Richman, D.D.; Carson, D.A.; Corbeil, J.; Leoni, L.M. HIV induces lymphocyte apoptosis by a p53-initiated, mitochondrial-mediated mechanism. FASEB J. 2001, 15, 5–6. [Google Scholar] [PubMed]
- Verschuren, E.W.; Klefstrom, J.; Evan, G.I.; Jones, N. The oncogenic potential of Kaposi's sarcoma-associated herpesvirus cyclin is exposed by p53 loss in vitro and in vivo. Cancer Cell 2002, 2, 229–241. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, G.P.; Besaratinia, A. Mutational spectra of human cancer. Hum. Genet. 2009, 125, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.P. Cancer. p53, guardian of the genome. Nature 1992, 58, 15–16. [Google Scholar] [CrossRef]
- Sutcliffe, J.E.; Brehm, A. Of flies and men; p53, a tumour suppressor. FEBS Lett. 2004, 567, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.J.; Abrams, J.M. Lessons from p53 in non-mammalian models. Cell Death Differ. 2006, 13, 909–912. [Google Scholar] [CrossRef] [PubMed]
- Graeber, T.G.; Peterson, J.F.; Tsai, M.; Monica, K.; Fornace Jr., A.J.; Giaccia, A.J. Hypoxia induces accumulation of p53 protein, but activation of a G1-phase checkpoint by low-oxygen conditions is independent of p53 status. Mol. Cell Biol. 1994, 14, 6264–6277. [Google Scholar] [PubMed]
- Yu, X.; Riley, T.; Levine, A.J. The regulation of the endosomal compartment by p53 the tumor suppressor gene. FEBS J. 2009, 276, 2201–2212. [Google Scholar] [CrossRef] [PubMed]
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
