Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasma Cytokine Analysis
2.2. Machine Learning for Modeling
2.3. Receiver Operating Characteristic Curve
2.4. Single-Cell RNA Sequencing Data Analysis
2.5. RNA-Sequencing
2.6. Statistics
3. Results
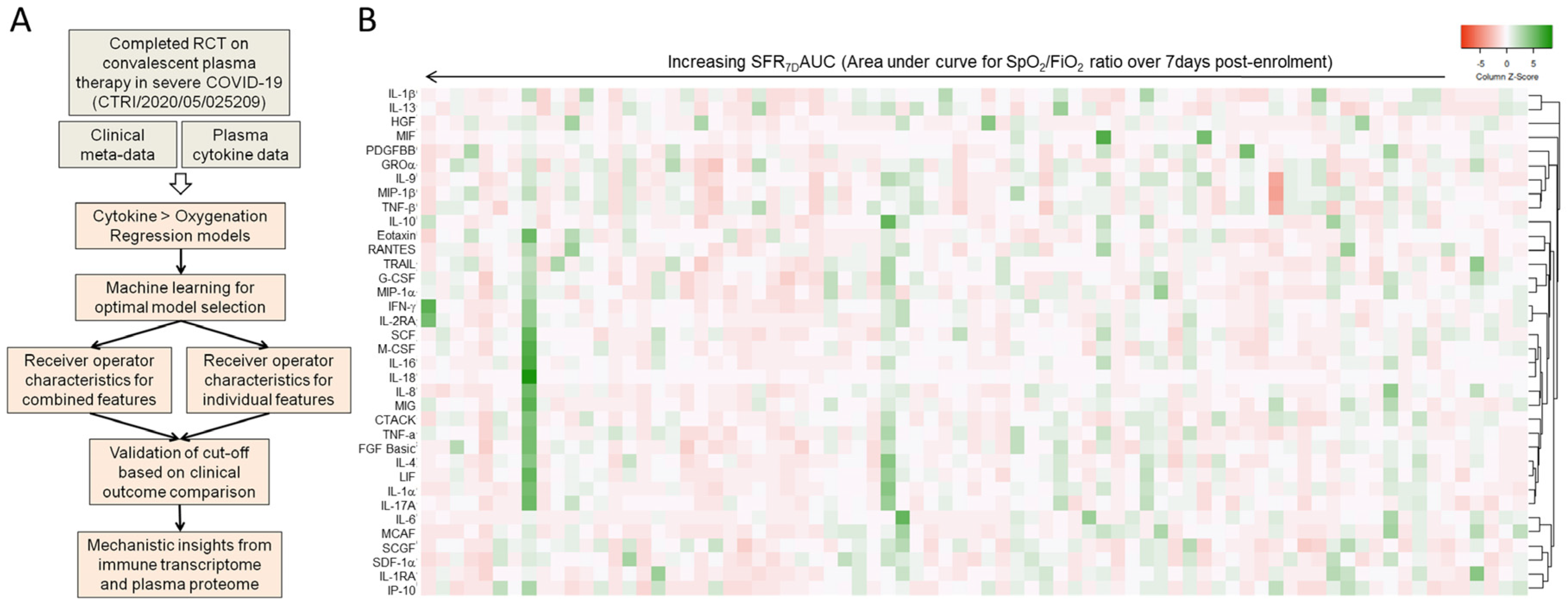
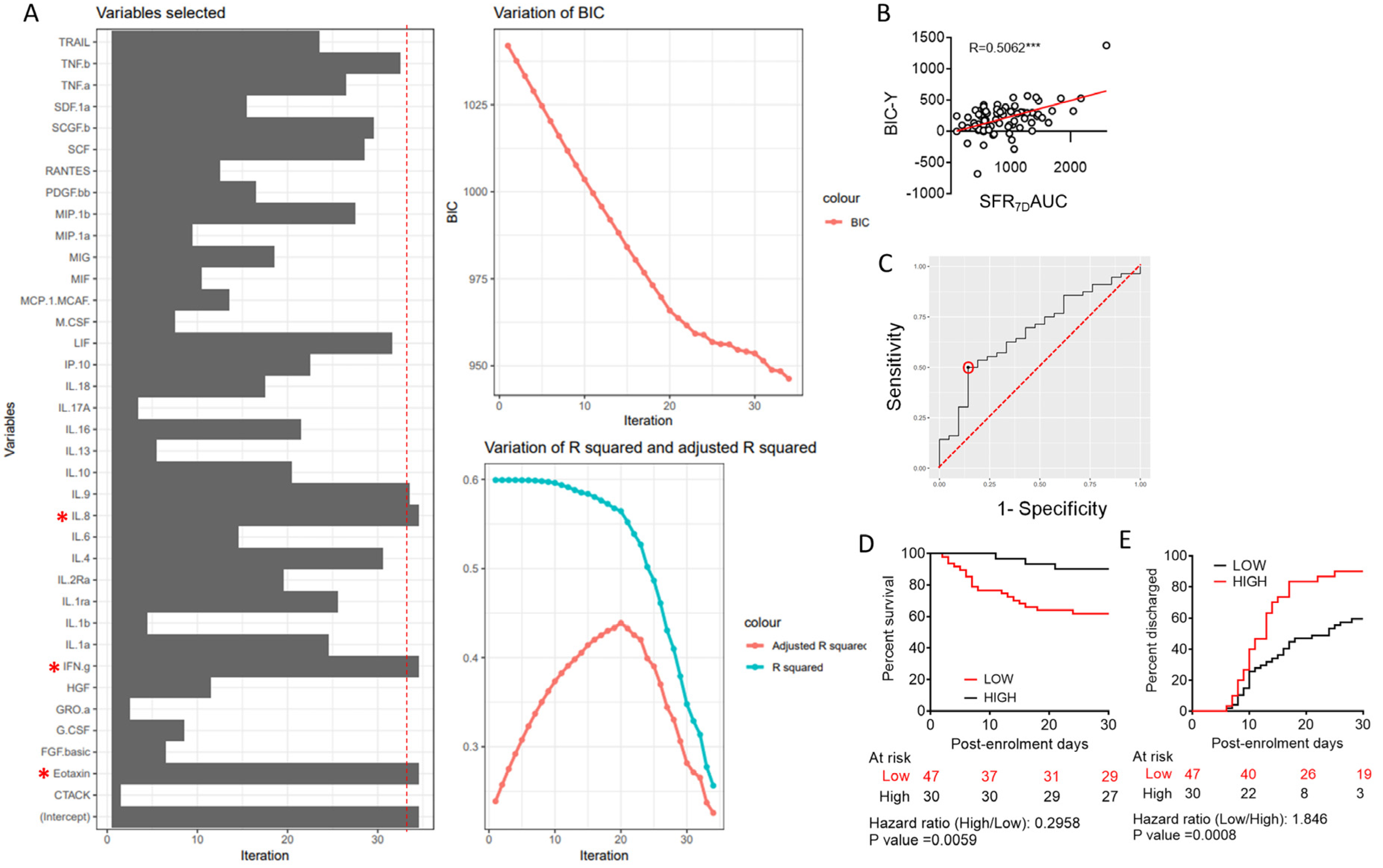
3.1. Machine Learning to Derive a Minimal Model Based on Plasma Cytokine Levels to Predict Clinical Outcome in Severe COVID-19
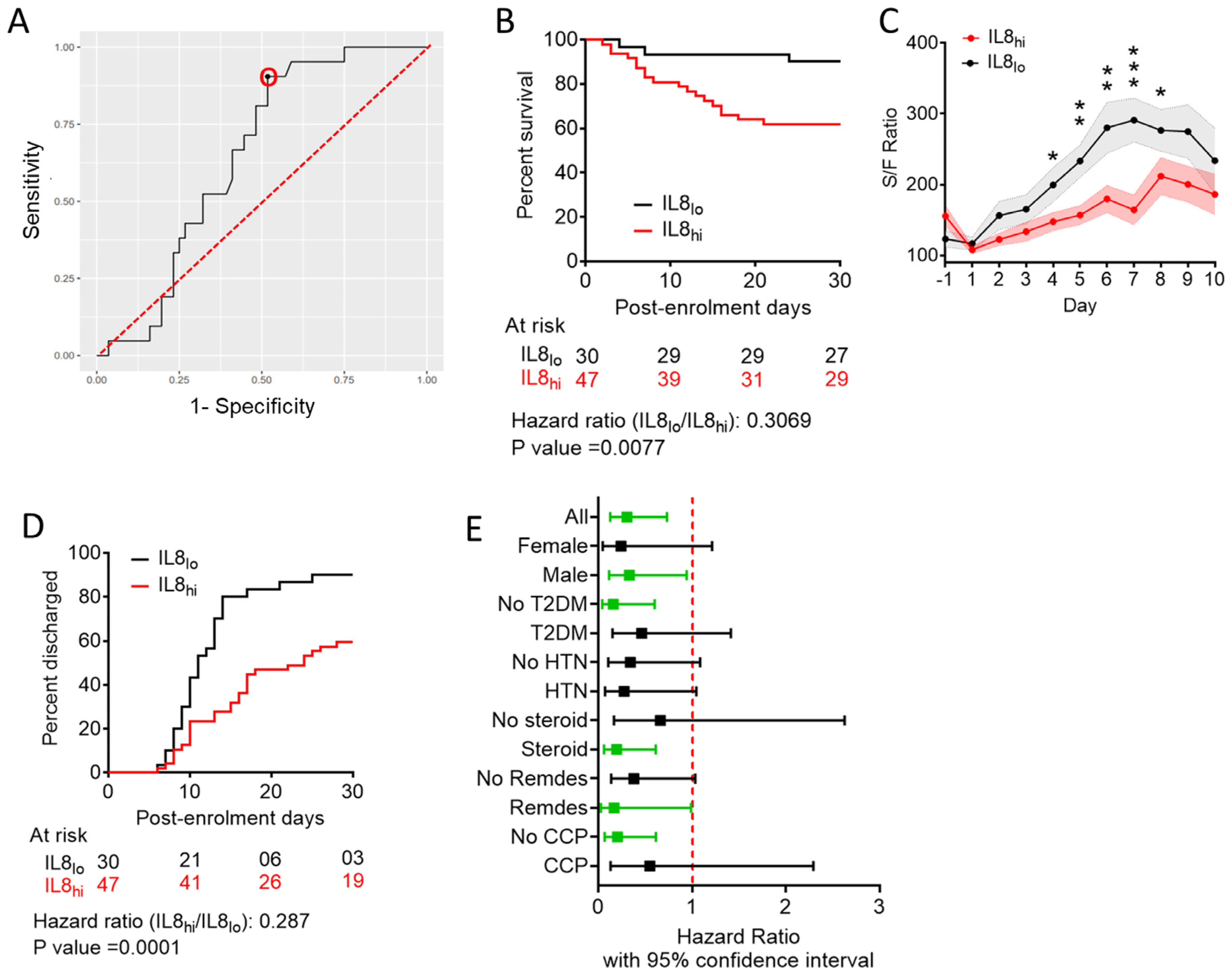
3.2. Predictive Value of Individual Cytokines Featured in BIC-Derived Regression Model and Identification of IL-8 as an Independent Predictor
3.3. Exploring Cellular Sources of IL-8 and IL-8-Responder Cells in Peripheral Blood and Bronchoalveolar Lumen
3.4. Systemic Mechanistic Insight from Transcriptome Studies on Circulating Immune Cells Compared between IL8hi and IL8lo Patients at Different Time-Points
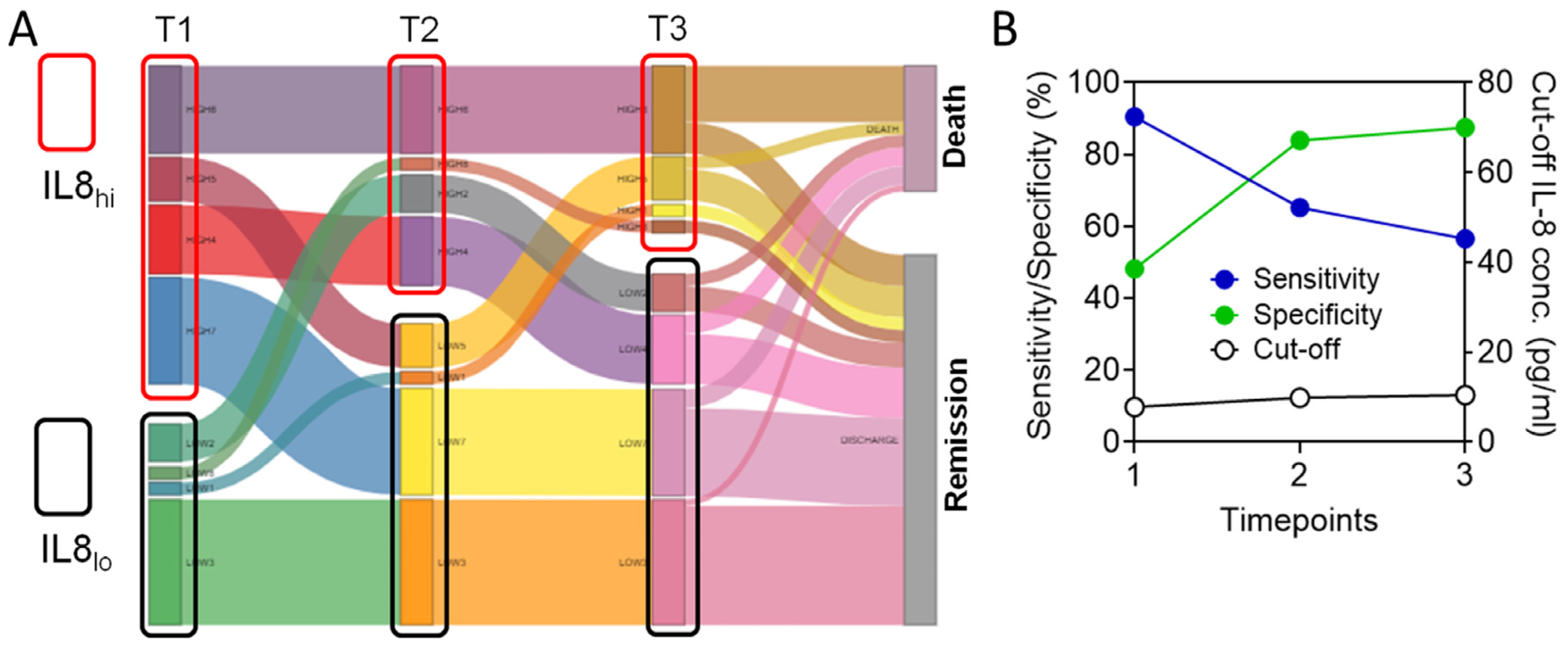
3.5. Circulating IL-8 Dynamics Parallels Disease Course
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- WHO Working Group on the Clinical Characterisation and Management of COVID-19 Infection. A minimal common outcome measure set for COVID-19 clinical research. Lancet Infect. Dis. 2020, 20, e192–e197. [Google Scholar] [CrossRef] [PubMed]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Qu, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef] [PubMed]
- Bergwerk, M.; Gonen, T.; Lustig, Y.; Amit, S.; Lipsitch, M.; Cohen, C.; Mandelboim, M.; Levin, E.G.; Rubin, C.; Indenbaum, V.; et al. COVID-19 Breakthrough Infections in Vaccinated Health Care Workers. N. Engl. J. Med. 2021, 385, 1474–1484. [Google Scholar] [CrossRef]
- Edara, V.V.; Pinsky, B.A.; Suthar, M.S.; Lai, L.; Davis-Gardner, M.E.; Floyd, K.; Flowers, M.W.; Wrammert, J.; Hussaini, L.; Ciric, C.R.; et al. Infection and Vaccine-Induced Neutralizing-Antibody Responses to the SARS-CoV-2 B.1.617 Variants. N. Engl. J. Med. 2021, 385, 664–666. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J.; HLH Across Speciality Collaboration, UK. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Bandopadhyay, P.; D’Rozario, R.; Lahiri, A.; Sarif, J.; Ray, Y.; Paul, S.R.; Roy, R.; Maiti, R.; Chaudhuri, K.; Bagchi, S.; et al. Nature and dimensions of the systemic hyper-inflammation and its attenuation by convalescent plasma in severe COVID-19. J. Infect. Dis. 2021, 224, 565–574. [Google Scholar] [CrossRef]
- Li, H.; Liu, L.; Zhang, D.; Xu, J.; Dai, H.; Tang, N.; Su, X.; Cao, B. SARS-CoV-2 and viral sepsis: Observations and hypotheses. Lancet 2020, 395, 1517–1520. [Google Scholar] [CrossRef]
- Schulte-Schrepping, J.; Reusch, N.; Paclik, D.; Baßler, K.; Schlickeiser, S.; Zhang, B.; Krämer, B.; Krammer, T.; Brumhard, S.; Bonaguro, L.; et al. Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment. Cell 2020, 182, 1419–1440.e23. [Google Scholar] [CrossRef]
- Silvin, A.; Chapuis, N.; Dunsmore, G.; Goubet, A.-G.; Dubuisson, A.; Derosa, L.; Almire, C.; Hénon, C.; Kosmider, O.; Droin, N.; et al. Elevated Calprotectin and Abnormal Myeloid Cell Subsets Discriminate Severe from Mild COVID-19. Cell 2020, 182, 1401–1418.e18. [Google Scholar] [CrossRef]
- Combes, A.J.; Courau, T.; Kuhn, N.F.; Hu, K.H.; Ray, A.; Chen, W.S.; Chew, N.W.; Cleary, S.J.; Kushnoor, D.; Reeder, G.C.; et al. Global absence and targeting of protective immune states in severe COVID-19. Nature 2021, 591, 124–130. [Google Scholar] [CrossRef]
- Sarif, J.; Raychaudhuri, D.; D’Rozario, R.; Bandopadhyay, P.; Singh, P.; Mehta, P.; Hoque, M.A.; Sinha, B.P.; Kushwaha, M.; Sahni, S.; et al. Plasma Gradient of Soluble Urokinase-Type Plasminogen Activator Receptor Is Linked to Pathogenic Plasma Proteome and Immune Transcriptome and Straifies Outcomes in Severe COVID-19. Front. Immunol. 2021, 12, 738093. [Google Scholar] [CrossRef]
- RECOVERY Collaborative Group; Horby, P.; Lim, W.S.; Emberson, J.R.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Brightling, C.; Ustianowski, A.; et al. Dexamethasone in Hospitalized Patients with COVID-19. N. Engl. J. Med. 2021, 384, 693–704. [Google Scholar]
- Dequin, P.F.; Heming, N.; Meziani, F.; Plantefève, G.; Voiriot, G.; Badié, J.; François, B.; Aubron, C.; Ricard, J.D.; Ehrmann, S.; et al. Effect of Hydrocortisone on 21-Day Mortality or Respiratory Support Among Critically Ill Patients With COVID-19: A Randomized Clinical Trial. JAMA 2020, 324, 1298–1306. [Google Scholar] [CrossRef] [PubMed]
- REMAP-CAP Investigators; Gordon, A.C.; Mouncey, P.R.; Al-Beidh, F.; Rowan, K.M.; Nichol, A.D.; Arabi, Y.M.; Annane, D.; Beane, A.; van Bentum-Puijk, W.; et al. Interleukin-6 Receptor Antagonists in Critically Ill Patients with COVID-19. N. Engl. J. Med. 2021, 384, 1491–1502. [Google Scholar]
- Salvarani, C.; Dolci, G.; Massari, M.; Merlo, D.F.; Cavuto, S.; Savoldi, L.; Bruzzi, P.; Boni, F.; Braglia, L.; Turrà, C.; et al. Effect of Tocilizumab vs Standard Care on Clinical Worsening in Patients Hospitalized With COVID-19 Pneumonia: A Randomized Clinical Trial. JAMA Intern. Med. 2021, 181, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Rosas, I.O.; Bräu, N.; Waters, M.; Go, R.C.; Hunter, B.D.; Bhagani, S.; Skiest, D.; Aziz, M.S.; Cooper, N.; Douglas, I.S.; et al. Tocilizumab in Hospitalized Patients with Severe COVID-19 Pneumonia. N. Engl. J. Med. 2021, 384, 1503–1516. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir-Ritonavir in Adults Hospitalized with Severe COVID-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of COVID-19—Final Report. N. Engl. J. Med. 2020, 383, 1813–1826. [Google Scholar] [CrossRef]
- Goldman, J.D.; Lye, D.C.; Hui, D.S.; Marks, K.M.; Bruno, R.; Montejano, R.; Spinner, C.D.; Galli, M.; Ahn, M.-Y.; Nahass, R.G.; et al. Remdesivir for 5 or 10 Days in Patients with Severe COVID-19. N. Engl. J. Med. 2020, 383, 1827–1837. [Google Scholar] [CrossRef]
- Dougan, M.; Nirula, A.; Azizad, M.; Mocherla, B.; Gottlieb, R.L.; Chen, P.; Hebert, C.; Perry, R.; Boscia, J.; Heller, B.; et al. Bamlanivimab plus Etesevimab in Mild or Moderate COVID-19. N. Engl. J. Med. 2021, 385, 1382–1392. [Google Scholar] [CrossRef] [PubMed]
- Ray, Y.; Paul, S.R.; Bandopadhyay, P.; D’Rozario, R.; Sarif, J.; Raychaudhuri, D.; Bhowmik, D.; Lahiri, A.; Vasudevan, J.S.; Maurya, R.; et al. A phase 2 single center open label randomised control trial for convalescent plasma therapy in patients with severe COVID-19. Nat. Commun. 2022, 13, 383. [Google Scholar] [CrossRef] [PubMed]
- Johnson, W.; Li, C.; Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 2007, 8, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Zindler, T.; Frieling, H.; Neyazi, A.; Bleich, S.; Friedel, E. Simulating ComBat: How batch correction can lead to the systematic introduction of false positive results in DNA methylation microarray studies. BMC Bioinform. 2020, 21, 271. [Google Scholar] [CrossRef] [PubMed]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef] [PubMed]
- Howe, E.A.; Sinha, R.; Schlauch, D.; Quackenbush, J. RNA-Seq analysis in MeV. Bioinformatics 2011, 27, 3209–3210. [Google Scholar] [CrossRef]
- Wettenhall, J.M.; Smyth, G.K. Limma GUI: A graphical user interface for linear modeling of microarray data. Bioinformatics 2004, 20, 3705–3706. [Google Scholar] [CrossRef]
- Zhou, G.; Soufan, O.; Ewald, J.; Hancock, R.E.; Basu, N.; Xia, J. NetworkAnalyst 3.0: A visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 2019, 47, W234–W241. [Google Scholar] [CrossRef]
- Liao, M.; Liu, Y.; Yuan, J.; Wen, Y.; Xu, G.; Zhao, J.; Cheng, L.; Li, J.; Wang, X.; Wang, F.; et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 2020, 26, 842–844. [Google Scholar] [CrossRef]
- Park, J.H.; Lee, H.K. Re-analysis of Single Cell Transcriptome Reveals That the NR3C1-CXCL8-Neutrophil Axis Determines the Severity of COVID-19. Front. Immunol. 2020, 11, 2145. [Google Scholar] [CrossRef]
- Osuchowski, M.F.; Winkler, M.S.; Skirecki, T.; Cajander, S.; Shankar-Hari, M.; Lachmann, G.; Monneret, G.; Venet, F.; Bauer, M.; Brunkhorst, F.M.; et al. The COVID-19 puzzle: Deciphering pathophysiology and phenotypes of a new disease entity. Lancet Respir. Med. 2021, 9, 622–642. [Google Scholar] [CrossRef] [PubMed]
- McGonagle, D.; Bridgewood, C.; Meaney, J.F.M. A tricompartmental model of lung oxygenation disruption to explain pulmonary and systemic pathology in severe COVID-19. Lancet Respir. Med. 2021, 9, 665–672. [Google Scholar] [CrossRef] [PubMed]
- Nicolai, L.; Leunig, A.; Brambs, S.; Kaiser, R.; Weinberger, T.; Weigand, M.; Muenchhoff, M.; Hellmuth, J.C.; Ledderose, S.; Schulz, H.; et al. Immunothrombotic Dysregulation in COVID-19 Pneumonia Is Associated With Respiratory Failure and Coagulopathy. Circulation 2020, 142, 1176–1189. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhao, K.; Wei, H.; Chen, W.; Wang, W.; Jia, L.; Liu, Q.; Zhang, J.; Shan, T.; Peng, Z.; et al. Dynamic relationship between D-dimer and COVID-19 severity. Br. J. Haematol. 2020, 190, e24–e27. [Google Scholar] [CrossRef] [PubMed]
- Pons, M.J.; Ymaña, B.; Mayanga-Herrera, A.; Sáenz, Y.; Alvarez-Erviti, L.; Tapia-Rojas, S.; Gamarra, R.; Blanco, A.B.; Moncunill, G.; Ugarte-Gil, M.F. Cytokine Profiles Associated With Worse Prognosis in a Hospitalized Peruvian COVID-19 Cohort. Front. Immunol. 2021, 12, 700921. [Google Scholar] [CrossRef] [PubMed]
- Alosaimi, B.; Mubarak, A.; Hamed, M.E.; Almutairi, A.Z.; Alrashed, A.A.; AlJuryyan, A.; Enani, M.; Alenzi, F.Q.; Alturaiki, W. Complement Anaphylatoxins and Inflammatory Cytokines as Prognostic Markers for COVID-19 Severity and In-Hospital Mortality. Front. Immunol. 2021, 12, 668725. [Google Scholar] [CrossRef]
- Del Valle, D.M.; Kim-Schulze, S.; Huang, H.H.; Beckmann, N.D.; Nirenberg, S.; Wang, B.; Lavin, Y.; Swartz, T.H.; Madduri, D.; Stock, A.; et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat. Med. 2020, 26, 1636–1643. [Google Scholar] [CrossRef]
- McElvaney, O.J.; McEvoy, N.L.; McElvaney, O.F.; Carroll, T.P.; Murphy, M.P.; Dunlea, D.M.; Choileáin, O.N.; Clarke, J.; O’Connor, E.; Hogan, G.; et al. Characterization of the Inflammatory Response to Severe COVID-19 Illness. Am. J. Respir. Crit. Care Med. 2020, 202, 812–821. [Google Scholar] [CrossRef]
- Melero, I.; Villalba-Esparza, M.; Recalde-Zamacona, B.; Jiménez-Sánchez, D.; Teijeira, Á.; Argueta, A.; García-Tobar, L.; Álvarez-Gigli, L.; Sainz, C.; Garcia-Ros, D.; et al. Neutrophil Extracellular Traps, Local IL-8 Expression, and Cytotoxic T-Lymphocyte Response in the Lungs of Patients With Fatal COVID-19. Chest 2022, 162, 1006–1016. [Google Scholar] [CrossRef]
- Bordoni, V.; Mariotti, D.; Matusali, G.; Colavita, F.; Cimini, E.; Ippolito, G.; Agrati, C. SARS-CoV-2 Infection of Airway Epithelium Triggers Pulmonary Endothelial Cell Activation and Senescence Associated with Type I IFN Production. Cells 2022, 11, 2912. [Google Scholar] [CrossRef]
- Bergantini, L.; d’Alessandro, M.; Cameli, P.; Otranto, A.; Luzzi, S.; Bianchi, F.; Bargagli, E. Cytokine profiles in the detection of severe lung involvement in hospitalized patients with COVID-19: The IL-8/IL-32 axis. Cytokine 2022, 151, 155804. [Google Scholar] [CrossRef] [PubMed]
- Tamayo-Velasco, Á.; Martínez-Paz, P.; Peñarrubia-Ponce, M.J.; de la Fuente, I.; Pérez-González, S.; Fernández, I.; Dueñas, C.; Gómez-Sánchez, E.; Lorenzo-López, M.; Gómez-Pesquera, E.; et al. HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients. J. Clin. Med. 2021, 10, 2017. [Google Scholar] [CrossRef] [PubMed]
- Gideon, S.E. Estimating the dimension of a model. Ann. Stat. 1978, 6, 461–464. [Google Scholar]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
D’Rozario, R.; Raychaudhuri, D.; Bandopadhyay, P.; Sarif, J.; Mehta, P.; Liu, C.S.C.; Sinha, B.P.; Roy, J.; Bhaduri, R.; Das, M.; et al. Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19. Viruses 2023, 15, 549. https://doi.org/10.3390/v15020549
D’Rozario R, Raychaudhuri D, Bandopadhyay P, Sarif J, Mehta P, Liu CSC, Sinha BP, Roy J, Bhaduri R, Das M, et al. Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19. Viruses. 2023; 15(2):549. https://doi.org/10.3390/v15020549
Chicago/Turabian StyleD’Rozario, Ranit, Deblina Raychaudhuri, Purbita Bandopadhyay, Jafar Sarif, Priyanka Mehta, Chinky Shiu Chen Liu, Bishnu Prasad Sinha, Jayasree Roy, Ritwik Bhaduri, Monidipa Das, and et al. 2023. "Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19" Viruses 15, no. 2: 549. https://doi.org/10.3390/v15020549
APA StyleD’Rozario, R., Raychaudhuri, D., Bandopadhyay, P., Sarif, J., Mehta, P., Liu, C. S. C., Sinha, B. P., Roy, J., Bhaduri, R., Das, M., Bandyopadhyay, S., Paul, S. R., Chatterjee, S., Pandey, R., Ray, Y., & Ganguly, D. (2023). Circulating Interleukin-8 Dynamics Parallels Disease Course and Is Linked to Clinical Outcomes in Severe COVID-19. Viruses, 15(2), 549. https://doi.org/10.3390/v15020549







