Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics
Abstract
:1. Introduction
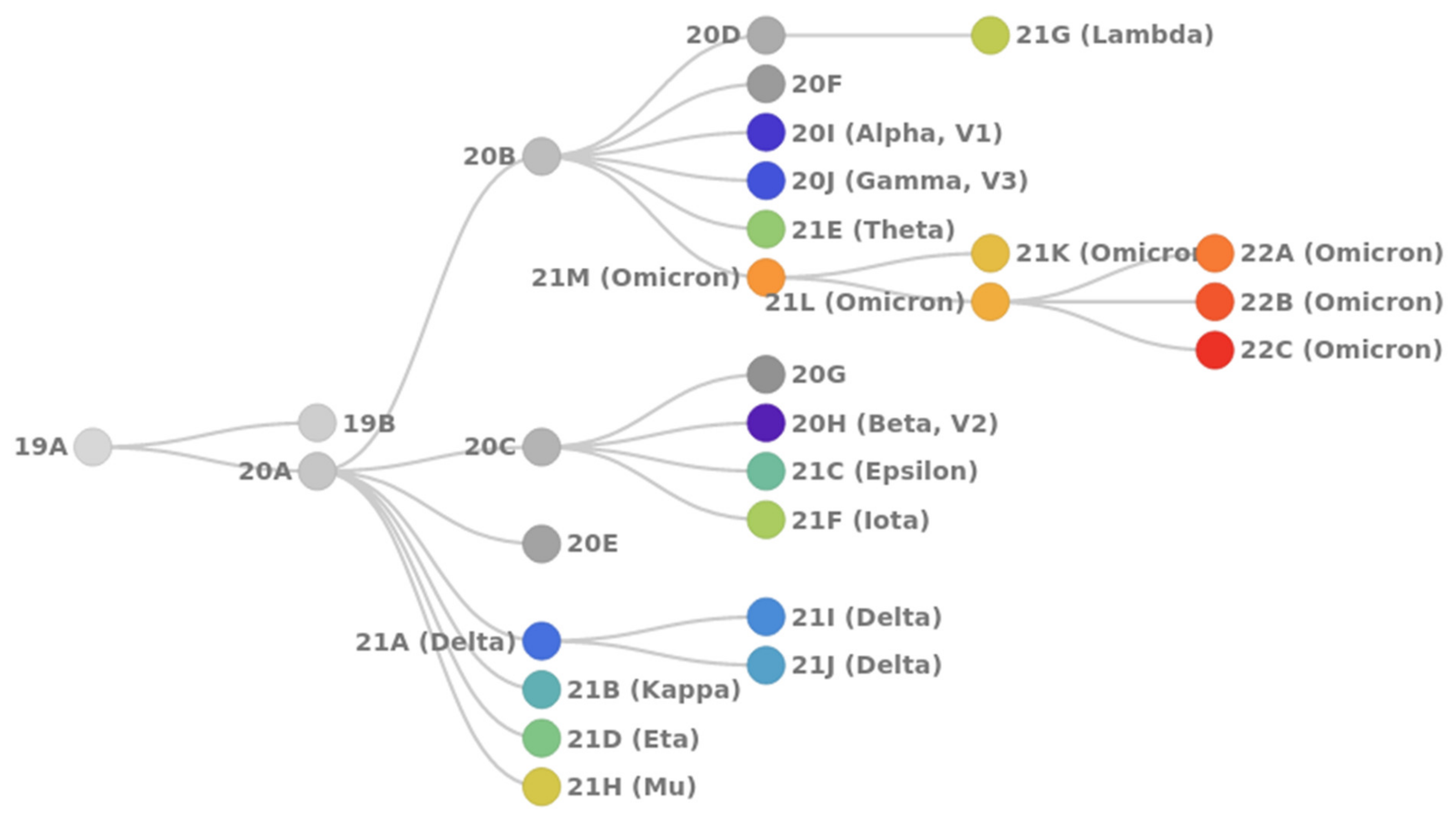
- 20I Alpha (B.1.1.7, V1): first VoC described in the United Kingdom in late December 2020;
- 20H Beta (B.1.351, V2): first VoC reported in South Africa in December 2020;
- 20J Gamma (P.1, V3): first reported in Brazil in early January 2021;
- 21A Delta (B.1.617.2): first reported in India in December 2020; includes sublineages 21I and 21J;
- 21M Omicron (B.1.1.529): first reported in South Africa in November 2021; includes two sublineages: 21K (BA.1) and 21L (BA.2).
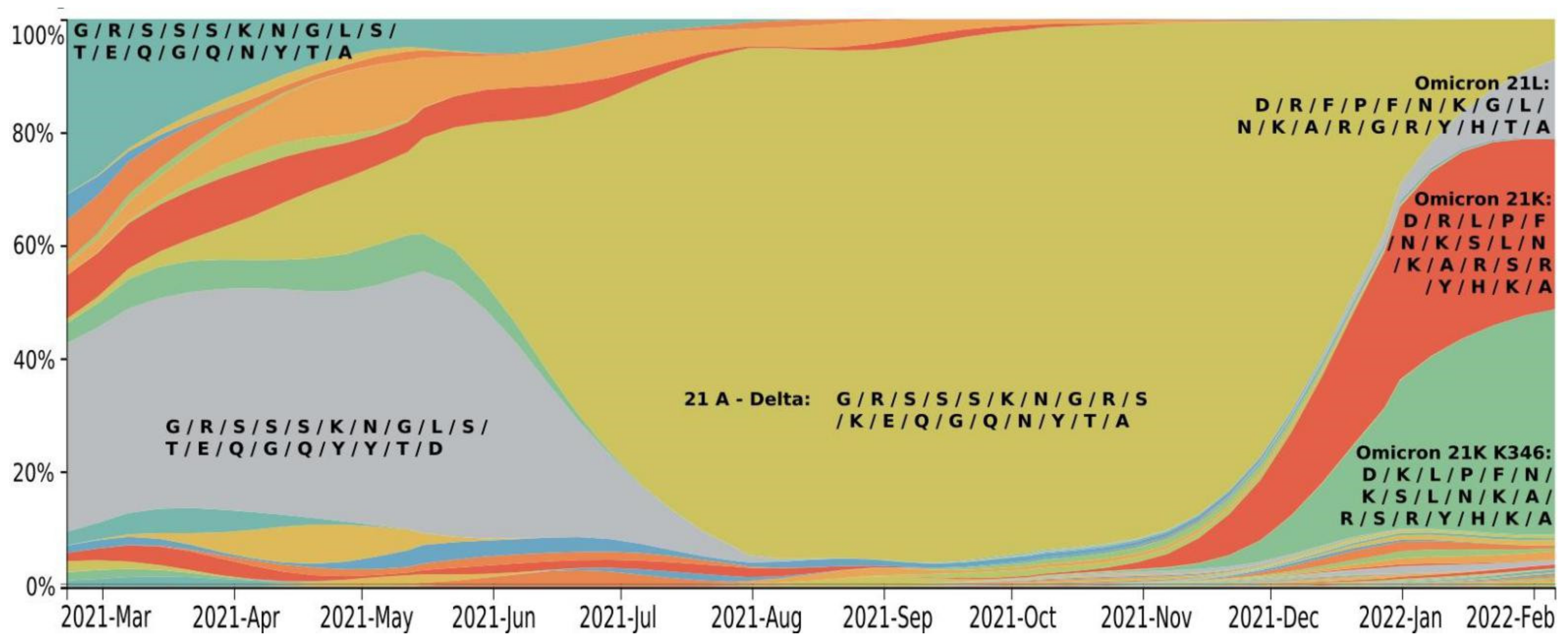
2. Mutation Dynamic Is Associated with Omicron Evolution
3. Omicron Mutations Are Significant for SARS-CoV-2 Evolution
4. Omicron Is Characterized by High Immune Evasion
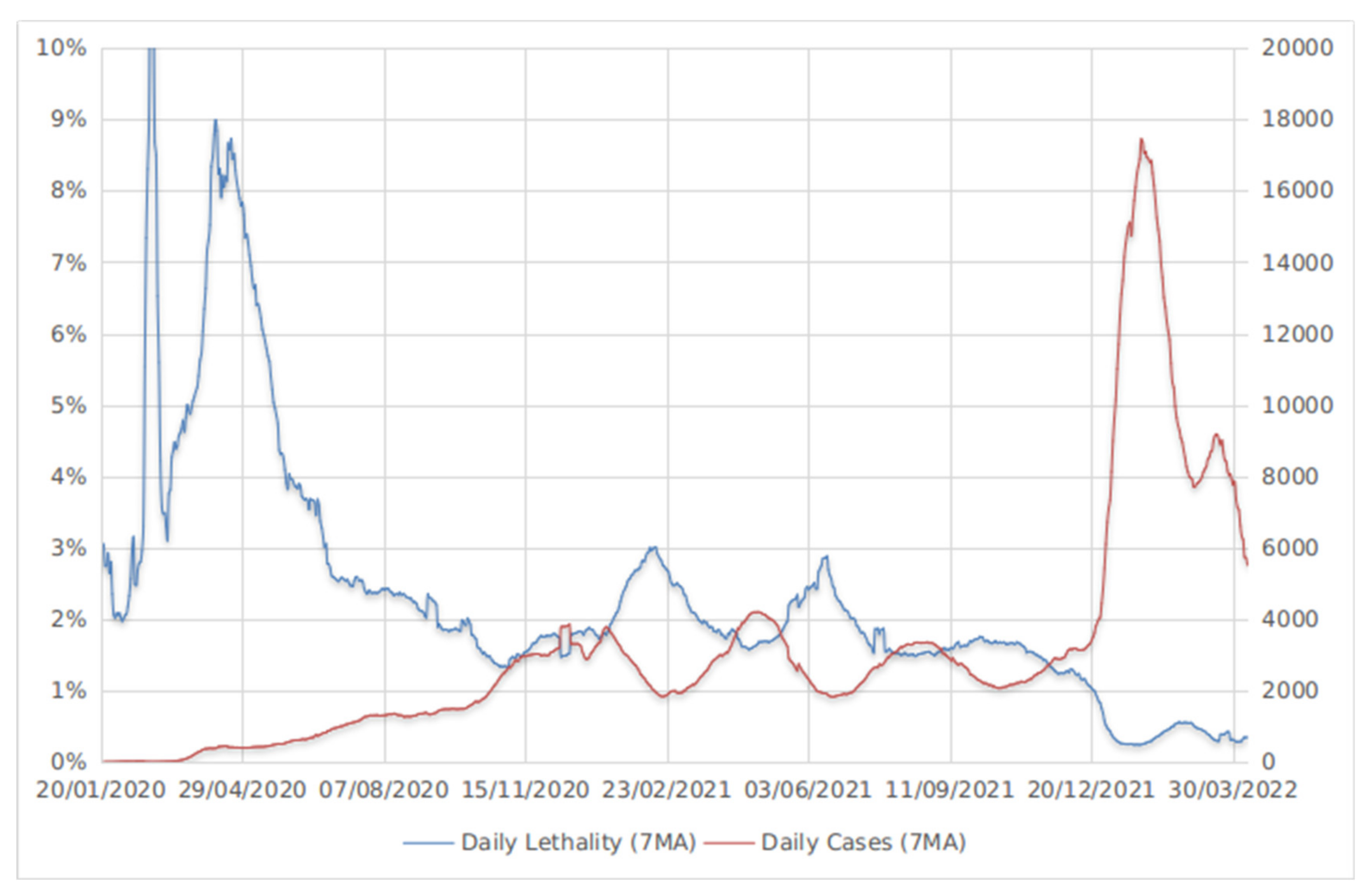
5. Omicron Has Achieved Balance between High Transmissibility and Low Mortality
6. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 14 May 2022).
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Guo, F.; Liu, K.; Wang, H.; Rao, S.; Yang, P.; Jiang, C. Endocytosis of the receptor-binding domain of SARS-CoV spike protein together with virus receptor ACE2. Virus Res. 2008, 136, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Belouzard, S.; Chu, V.C.; Whittaker, G.R. Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites. Proc. Natl. Acad. Sci. USA 2009, 106, 5871–5876. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, W.; Gui, M.; Wang, X.; Xiang, Y. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog. 2018, 14, e1007236. [Google Scholar] [CrossRef]
- Walls, A.C.; Tortorici, M.A.; Snijder, J.; Xiong, X.; Bosch, B.-J.; Rey, F.A.; Veesler, D. Tectonic conformational changes of a coronavirus spike glycoprotein promote membrane fusion. Proc. Natl. Acad. Sci. USA 2017, 114, 11157–11162. [Google Scholar] [CrossRef] [Green Version]
- Li, F. Structure, function, and evolution of coronavirus spike proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [Green Version]
- Clade Naming & Definitions-SARS-CoV-2 Workflow Documentation. Available online: https://docs.nextstrain.org/projects/ncov/en/latest/reference/naming_clades.html (accessed on 13 May 2022).
- Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/activities/tracking-SARS-CoV-2-variants (accessed on 13 May 2022).
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef]
- Sagulenko, P.; Puller, V.; Neher, R.A. TreeTime: Maximum-likelihood phylodynamic analysis. Virus Evol. 2018, 4, vex042. [Google Scholar] [CrossRef]
- Leung, K.; Shum, M.H.; Leung, G.M.; Lam, T.T.; Wu, J.T. Early transmissibility assessment of the N501Y mutant strains of SARS-CoV-2 in the United Kingdom, October to November 2020. Eurosurveillance 2021, 26, 2002106. [Google Scholar] [CrossRef]
- Boehm, E.; Kronig, I.; Neher, R.A.; Eckerle, I.; Vetter, P.; Kaiser, L. Novel SARS-CoV-2 variants: The pandemics within the pandemic. Clin. Microbiol. Infect. 2021, 27, 1109–1117. [Google Scholar] [CrossRef]
- “RCoV19 Version 4.0 2019新型冠状病毒信息库.” RCoV19—2019新型冠状病毒信息库. Available online: https://ngdc.cncb.ac.cn/ncov/ (accessed on 13 May 2022).
- Zhang, X.; Wu, S.; Wu, B.; Yang, Q.; Chen, A.; Li, Y.; Zhang, Y.; Pan, T.; Zhang, H.; He, X. SARS-CoV-2 Omicron strain exhibits potent capabilities for immune evasion and viral entrance. Signal Transduct. Target. Ther. 2021, 6, 430. [Google Scholar] [CrossRef] [PubMed]
- Casalino, L.; Gaieb, Z.; Goldsmith, J.A.; Hjorth, C.K.; Dommer, A.C.; Harbison, A.M.; Fogarty, C.A.; Barros, E.P.; Taylor, B.C.; McLellan, J.S.; et al. Beyond Shielding: The Roles of Glycans in the SARS-CoV-2 Spike Protein. ACS Cent. Sci. 2020, 6, 1722–1734. [Google Scholar] [CrossRef] [PubMed]
- Nextstrain. Available online: https://nextstrain.org/ (accessed on 13 May 2022).
- Kandeel, M.; Mohamed, M.E.M.; Abd El-Lateef, H.M.; Venugopala, K.N.; El-Beltagi, H.S. Omicron Variant Genome Evolution and Phylogenetics. J. Med. Virol. 2021, 94, 1627–1632. [Google Scholar] [CrossRef] [PubMed]
- Berkhout, B.; Herrera-Carrillo, E. SARS-CoV-2 Evolution: On the Sudden Appearance of the Omicron Variant. J. Virol. 2022, 96, e00090-22. [Google Scholar] [CrossRef]
- Wei, C.; Shan, K.J.; Wang, W.; Zhang, S.; Huan, Q.; Qian, W. Evidence for a Mouse Origin of the SARS-CoV-2 Omicron Variant. J. Genet. Genom. 2021, 48, 1111–1121. [Google Scholar] [CrossRef]
- Xu, Y.; Wu, C.; Cao, X.; Gu, C.; Liu, H.; Jiang, M.; Wang, X.; Yuan, Q.; Wu, K.; Liu, J.; et al. Structural and biochemical mechanism for increased infectivity and immune evasion of Omicron BA.2 variant compared to BA.1 and their possible mouse origins. Cell Res. 2022, 32, 609–620. [Google Scholar] [CrossRef]
- Tarcsai, K.R.; Corolciuc, O.; Tordai, A.; Ongrádi, J. SARS-CoV-2 infection in HIV-infected patients: Potential role in the high mutational load of the Omicron variant emerging in South Africa. GeroScience 2022, 1–9. [Google Scholar] [CrossRef]
- Hoffman, S.A.; Costales, C.; Sahoo, M.K.; Palanisamy, S.; Yamamoto, F.; Huang, C.; Verghese, M.; Solis, D.A.; Sibai, M.; Subramanian, A.; et al. SARS-CoV-2 Neutralization Resistance Mutations in Patient with HIV/AIDS, California, USA. Emerg. Infect. Dis. 2021, 27, 2720–2723. [Google Scholar] [CrossRef]
- Abdelnabi, R.; Foo, C.S.; Zhang, X.; Lemmens, V.; Maes, P.; Slechten, B.; Raymenants, J.; André, E.; Weynand, B.; Dallmeier, K.; et al. The omicron (B.1.1.529) SARS-CoV-2 variant of concern does not readily infect Syrian hamsters. Antivir. Res. 2022, 198, 105253. [Google Scholar] [CrossRef]
- Frost, S.D.W.; Magalis, B.R.; Kosakovsky Pond, S.L. Neutral Theory and Rapidly Evolving Viral Pathogens. Mol. Biol. Evol. 2018, 35, 1348–1354. [Google Scholar] [CrossRef]
- Zhou, B.; Thao, T.T.N.; Hoffmann, D.; Taddeo, A.; Ebert, N.; Labroussaa, F.; Pohlmann, A.; King, J.; Steiner, S.; Kelly, J.N.; et al. SARS-CoV-2 spike D614G change enhances replication and transmission. Nature 2021, 592, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.J.; Chiba, S.; Halfmann, P.; Ehre, C.; Kuroda, M.; Dinnon, K.H.; Leist, S.R.; Schäfer, A.; Nakajima, N.; Takahashi, K.; et al. SARS-CoV-2 D614G Variant Exhibits Efficient Replication Ex Vivo and Transmission in Vivo. Science 2020, 370, 1464–1468. [Google Scholar] [CrossRef] [PubMed]
- Yurkovetskiy, L.; Wang, X.; Pascal, K.E.; Tomkins-Tinch, C.; Nyalile, T.P.; Wang, Y.; Baum, A.; Diehl, W.E.; Dauphin, A.; Carbone, C.; et al. Structural and Functional Analysis of the D614G SARS-CoV-2 Spike Protein Variant. Cell 2020, 183, 739–751. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.; Hill, V.; McCrone, J.T.; Price, A.; Jorgensen, D.; O’Toole, Á.; Southgate, J.; Johnson, R.; Jackson, B.; Nascimento, F.F.; et al. Evaluating the Effects of SARS-CoV-2 Spike Mutation D614G on Transmissibility and Pathogenicity. Cell 2021, 184, 64–75. [Google Scholar] [CrossRef]
- Zhan, X.-Y.; Zhang, Y.; Zhou, X.; Huang, K.; Qian, Y.; Leng, Y.; Yan, L.; Huang, B.; He, Y. Molecular Evolution of SARS-CoV-2 Structural Genes: Evidence of Positive Selection in Spike Glycoprotein. bioRxiv 2020. [Google Scholar] [CrossRef]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence That D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827. [Google Scholar] [CrossRef]
- Ozono, S.; Zhang, Y.; Ode, H.; Sano, K.; Tan, T.S.; Imai, K.; Miyoshi, K.; Kishigami, S.; Ueno, T.; Iwatani, Y.; et al. SARS-CoV-2 D614G Spike Mutation Increases Entry Efficiency with Enhanced ACE2-Binding Affinity. Nat. Commun. 2021, 12, 848. [Google Scholar] [CrossRef]
- Plante, J.A.; Liu, Y.; Liu, J.; Xia, H.; Johnson, B.A.; Lokugamage, K.G.; Zhang, X.; Muruato, A.E.; Zou, J.; Fontes-Garfias, C.R.; et al. Spike Mutation D614G Alters SARS-CoV-2 Fitness. Nature 2020, 592, 116–121. [Google Scholar] [CrossRef]
- Gobeil, S.M.; Janowska, K.; McDowell, S.; Mansouri, K.; Parks, R.; Manne, K.; Stalls, V.; Kopp, M.F.; Henderson, R.; Edwards, R.J.; et al. D614G Mutation Alters SARS-CoV-2 Spike Conformation and Enhances Protease Cleavage at the S1/S2 Junction. Cell Rep. 2021, 34, 108630. [Google Scholar] [CrossRef]
- Benton, D.J.; Wrobel, A.G.; Roustan, C.; Borg, A.; Xu, P.; Martin, S.R.; Rosenthal, P.B.; Skehel, J.J.; Gamblin, S.J. The Effect of the D614G Substitution on the Structure of the Spike Glycoprotein of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2021, 118, e2022586118. [Google Scholar] [CrossRef]
- Verkhivker, G.M.; Agajanian, S.; Oztas, D.; Gupta, G. Computational Analysis of Protein Stability and Allosteric Interaction Networks in Distinct Conformational Forms of the SARS-CoV-2 Spike D614G Mutant: Reconciling Functional Mechanisms through Allosteric Model of Spike Regulation. J. Biomol. Struct. Dyn. 2021, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Saito, A.; Irie, T.; Suzuki, R.; Maemura, T.; Nasser, H.; Uriu, K.; Kosugi, Y.; Shirakawa, K.; Sadamasu, K.; Kimura, I.; et al. SARS-CoV-2 Spike P681R Mutation Enhances and Accelerates Viral Fusion. bioRxiv 2021. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, M.; Gao, J. Enhanced Receptor Binding of SARS-CoV-2 through Networks of Hydrogen-Bonding and Hydrophobic Interactions. Proc. Natl. Acad. Sci. USA 2020, 117, 13967–13974. [Google Scholar] [CrossRef]
- Zhu, X.; Mannar, D.; Srivastava, S.S.; Berezuk, A.M.; Demers, J.P.; Saville, J.W.; Leopold, K.; Li, W.; Dimitrov, D.S.; Tuttle, K.S.; et al. Cryo-Electron Microscopy Structures of the N501Y SARS-CoV-2 Spike Protein in Complex with ACE2 and 2 Potent Neutralizing Antibodies. PLoS Biol. 2021, 19, e3001237. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; VanBlargan, L.A.; Bloyet, L.M.; Rothlauf, P.W.; Chen, R.E.; Stumpf, S.; Zhao, H.; Errico, J.M.; Theel, E.S.; Liebeskind, M.J.; et al. Identification of SARS-CoV-2 Spike Mutations That Attenuate Monoclonal and Serum Antibody Neutralization. Cell Host Microbe 2021, 29, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Greaney, A.J.; Loes, A.N.; Crawford, K.H.D.; Starr, T.N.; Malone, K.D.; Chu, H.Y.; Bloom, J.D. Comprehensive Mapping of Mutations to the SARS-CoV-2 Receptor-Binding Domain That Affect Recognition by Polyclonal Human Serum Antibodies. Cell Host Microbe 2021, 29, 463–476. [Google Scholar] [CrossRef] [PubMed]
- Tchesnokova, V.; Kulakesara, H.; Larson, L.; Bowers, V.; Rechkina, E.; Kisiela, D.; Sledneva, Y.; Choudhury, D.; Maslova, I.; Deng, K.; et al. Acquisition of the L452R mutation in the ACE2-binding interface of Spike protein triggers recent massive expansion of SARS-Cov-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- Weisblum, Y.; Schmidt, F.; Zhang, F.; DaSilva, J.; Poston, D.; Lorenzi, J.C.; Muecksch, F.; Rutkowska, M.; Hoffmann, H.H.; Michailidis, E.; et al. Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. Elife 2020, 9, e61312. [Google Scholar] [CrossRef]
- Andreano, E.; Piccini, G.; Licastro, D.; Casalino, L.; Johnson, N.V.; Paciello, I.; Dal Monego, S.; Pantano, E.; Manganaro, N.; Manenti, A.; et al. SARS-CoV-2 escape in vitro from a highly neutralizing COVID-19 convalescent plasma. bioRxiv 2020. [Google Scholar] [CrossRef]
- Wall, E.C.; Wu, M.; Harvey, R.; Kelly, G.; Warchal, S.; Sawyer, C.; Daniels, R.; Hobson, P.; Hatipoglu, E.; Ngai, Y.; et al. Neutralising antibody activity against SARS-CoV-2 VOCs B.1.617.2 and B.1.351 by BNT162b2 vaccination. Lancet 2021, 397, 2331–2333. [Google Scholar] [CrossRef]
- Goher, S.S.; Ali, F.; Amin, M. The Delta Variant Mutations in the Receptor Binding Domain of SARS-CoV-2 Show Enhanced Electrostatic Interactions with the ACE2. Med. Drug Discov. 2021, 13, 100114. [Google Scholar] [CrossRef] [PubMed]
- Nelson, G.; Buzko, O.; Spilman, P.; Niazi, K.; Rabizadeh, S.; Soon-Shiong, P. Molecular Dynamic Simulation Reveals E484K Mutation Enhances Spike RBD-ACE2 Affinity and the Combination of E484K, K417N and N501Y Mutations (501Y.V2 Variant) Induces Conformational Change Greater than N501Y Mutant Alone, Potentially Resulting in an Escape Mutant. bioRxiv 2021. [Google Scholar] [CrossRef]
- McCallum, M.; Czudnochowski, N.; Rosen, L.E.; Zepeda, S.K.; Bowen, J.E.; Walls, A.C.; Hauser, K.; Joshi, A.; Stewart, C.; Dillen, J.R.; et al. Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement. Science 2022, 375, 864–868. [Google Scholar] [CrossRef] [PubMed]
- Nabel, K.G.; Clark, S.A.; Shankar, S.; Pan, J.; Clark, L.E.; Yang, P.; Coscia, A.; McKay, L.G.A.; Varnum, H.H.; Brusic, V.; et al. Structural basis for continued antibody evasion by the SARS-CoV-2 receptor binding domain. Science 2022, 375, eabl6251. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Lou, F.; Fan, H. SARS-CoV-2 variant Omicron: Currently the most complete “escapee” from neutralization by antibodies and vaccines. Signal Transduct. Target. Ther. 2022, 7, 28. [Google Scholar] [CrossRef]
- Cameroni, E.; Bowen, J.E.; Rosen, L.E.; Saliba, C.; Zepeda, S.K.; Culap, K.; Pinto, D.; VanBlargan, L.A.; De Marco, A.; di Iulio, J.; et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 2022, 602, 664–670. [Google Scholar] [CrossRef]
- Muik, A.; Lui, B.G.; Wallisch, A.K.; Bacher, M.; Mühl, J.; Reinholz, J.; Ozhelvaci, O.; Beckmann, N.; Güimil Garcia, R.C.; Poran, A.; et al. Neutralization of SARS-CoV-2 Omicron by BNT162b2 mRNA vaccine-elicited human sera. Science 2022, 375, 678–680. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Shen, X.; Schmidt, S.D.; O’Dell, S.; McDanal, C.; Feng, W.; Tong, J.; Eaton, A.; Maglinao, M.; Tang, H.; et al. Booster of mRNA-1273 Strengthens SARS-CoV-2 Omicron Neutralization. medRxiv 2021. [Google Scholar] [CrossRef]
- Ai, J.; Zhang, H.; Zhang, Y.; Lin, K.; Zhang, Y.; Wu, J.; Wan, Y.; Huang, Y.; Song, J.; Fu, Z.; et al. Omicron variant showed lower neutralizing sensitivity than other SARS-CoV-2 variants to immune sera elicited by vaccines after boost. Emerg. Microbes Infect. 2022, 11, 337–343. [Google Scholar] [CrossRef]
- Schmidt, F.; Muecksch, F.; Weisblum, Y.; Da Silva, J.; Bednarski, E.; Cho, A.; Wang, Z.; Gaebler, C.; Caskey, M.; Nussenzweig, M.C.; et al. Plasma Neutralization of the SARS-CoV-2 Omicron Variant. N. Engl. J. Med. 2022, 386, 599–601. [Google Scholar] [CrossRef]
- Lu, L.; Mok, B.W.; Chen, L.L.; Chan, J.M.; Tsang, O.T.; Lam, B.H.; Chuang, V.W.; Chu, A.W.; Chan, W.M.; Ip, J.D.; et al. Neutralization of SARS-CoV-2 Omicron variant by sera from BNT162b2 or Coronavac vaccine recipients. Clin. Infect. Dis. 2021, ciab1041. [Google Scholar] [CrossRef] [PubMed]
- Cele, S.; Jackson, L.; Khoury, D.S.; Khan, K.; Moyo-Gwete, T.; Tegally, H.; San, J.E.; Cromer, D.; Scheepers, C.; Amoako, D.G.; et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature 2022, 602, 654–656. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Nair, M.S.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 2021, 593, 130–135. [Google Scholar] [CrossRef] [PubMed]
- Emary, K.R.W.; Golubchik, T.; Aley, P.K.; Ariani, C.V.; Angus, B.; Bibi, S.; Blane, B.; Bonsall, D.; Cicconi, P.; Charlton, S.; et al. Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 variant of concern 202012/01 (B.1.1.7): An exploratory analysis of a randomised controlled trial. Lancet 2021, 397, 1351–1362. [Google Scholar] [CrossRef]
- Wang, G.L.; Wang, Z.Y.; Duan, L.J.; Meng, Q.C.; Jiang, M.D.; Cao, J.; Yao, L.; Zhu, K.L.; Cao, W.C.; Ma, M.J. Susceptibility of Circulating SARS-CoV-2 Variants to Neutralization. N. Engl. J. Med. 2021, 384, 2354–2356. [Google Scholar] [CrossRef]
- Sapkal, G.N.; Yadav, P.D.; Ella, R.; Deshpande, G.R.; Sahay, R.R.; Gupta, N.; Mohan, V.K.; Abraham, P.; Panda, S.; Bhargava, B. Neutralization of UK-Variant VUI-202012/01 with COVAXIN Vaccinated Human Serum. bioRxiv 2021. [Google Scholar] [CrossRef]
- Garcia-Beltran, W.F.; Lam, E.C.; St Denis, K.; Nitido, A.D.; Garcia, Z.H.; Hauser, B.M.; Feldman, J.; Pavlovic, M.N.; Gregory, D.J.; Poznansky, M.C.; et al. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 2021, 184, 2372–2383.e9. [Google Scholar] [CrossRef]
- Hu, J.; Wei, X.Y.; Xiang, J.; Peng, P.; Xu, F.L.; Wu, K.; Luo, F.Y.; Jin, A.S.; Fang, L.; Liu, B.Z.; et al. Reduced neutralization of SARS-CoV-2 B.1.617 variant by convalescent and vaccinated sera. Genes Dis. 2021, 9, 51–61. [Google Scholar] [CrossRef]
- Liu, J.; Liu, Y.; Xia, H.; Zou, J.; Weaver, S.C.; Swanson, K.A.; Cai, H.; Cutler, M.; Cooper, D.; Muik, A.; et al. BNT162b2-elicited neutralization of B.1.617 and other SARS-CoV-2 variants. Nature 2021, 596, 273–275. [Google Scholar] [CrossRef]
- Wibmer, C.K.; Ayres, F.; Hermanus, T.; Madzivhandila, M.; Kgagudi, P.; Oosthuysen, B.; Lambson, B.E.; de Oliveira, T.; Vermeulen, M.; van der Berg, K.; et al. SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. Nat. Med. 2021, 27, 622–625. [Google Scholar] [CrossRef]
- Li, Q.; Nie, J.; Wu, J.; Zhang, L.; Ding, R.; Wang, H.; Zhang, Y.; Li, T.; Liu, S.; Zhang, M.; et al. SARS-CoV-2 501Y.V2 variants lack higher infectivity but do have immune escape. Cell 2021, 184, 2362–2371.e9. [Google Scholar] [CrossRef] [PubMed]
- Cele, S.; Gazy, I.; Jackson, L.; Hwa, S.H.; Tegally, H.; Lustig, G.; Giandhari, J.; Pillay, S.; Wilkinson, E.; Naidoo, Y.; et al. Escape of SARS-CoV-2 501Y.V2 from neutralization by convalescent plasma. Nature 2021, 593, 142–146. [Google Scholar] [CrossRef]
- Madhi, S.A.; Baillie, V.; Cutland, C.L.; Voysey, M.; Koen, A.L.; Fairlie, L.; Padayachee, S.D.; Dheda, K.; Barnabas, S.L.; Bhorat, Q.E.; et al. Efficacy of the ChAdOx1 nCoV-19 Covid-19 Vaccine against the B.1.351 Variant. N. Engl. J. Med. 2021, 384, 1885–1898. [Google Scholar] [CrossRef] [PubMed]
- Mahase, E. Covid-19: Novavax Vaccine Efficacy Is 86% against UK Variant and 60% against South African Variant. BMJ 2021, 372, n296. [Google Scholar] [CrossRef]
- Shinde, V.; Bhikha, S.; Hoosain, Z.; Archary, M.; Bhorat, Q.; Fairlie, L.; Lalloo, U.; Masilela, M.; Moodley, D.; Hanley, S.; et al. Efficacy of NVX-CoV2373 Covid-19 Vaccine against the B.1.351 Variant. N. Engl. J. Med. 2021, 384, 1899–1909. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Zhou, D.; Supasa, P.; Liu, C.; Mentzer, A.J.; Ginn, H.M.; Zhao, Y.; Duyvesteyn, H.M.E.; Tuekprakhon, A.; Nutalai, R.; et al. Antibody evasion by the P.1 strain of SARS-CoV-2. Cell 2021, 184, 2939–2954.e9. [Google Scholar] [CrossRef]
- Straten, K.; Guerra, D.; van Gils, M.J.; Bontjer, I.; Caniels, T.G.; van Willigen, H.D.G.; Wynberg, E.; Poniman, M.; Burger, J.A.; Bouhuijs, J.H.; et al. Mapping the Antigenic Diversification of SARS-CoV-2. medRxiv 2022. [Google Scholar] [CrossRef]
- Suryawanshi, R.K.; Chen, I.P.; Ma, T.; Syed, A.M.; Brazer, N.; Saldhi, P.; Simoneau, C.R.; Ciling, A.; Khalid, M.M.; Sreekumar, B.; et al. Limited Cross-Variant Immunity after Infection with the SARS-CoV-2 Omicron Variant Without Vaccination. medRxiv 2022. [Google Scholar] [CrossRef]
- Nyberg, T.; Ferguson, N.M.; Nash, S.G.; Webster, H.H.; Flaxman, S.; Andrews, N.; Hinsley, W.; Bernal, J.L.; Kall, M.; Bhatt, S.; et al. Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: A cohort study. Lancet 2022, 399, 1303–1312. [Google Scholar] [CrossRef]
- Simmonds, P.; Aiewsakun, P.; Katzouraki, A. Prisoners of War—Host Adaptation and Its Constraints on Virus Evolution. Nat. Rev. Microbiol. 2018, 17, 321–328. [Google Scholar] [CrossRef]
- Geoghegan, J.L.; Holmes, E.C. The Phylogenomics of Evolving Virus Virulence. Nat. Rev. Genet. 2018, 19, 756–769. [Google Scholar] [CrossRef] [PubMed]
- Bull, J.J.; Lauring, A.S. Theory and Empiricism in Virulence Evolution. PLoS Pathog. 2014, 10, e1004387. [Google Scholar] [CrossRef] [PubMed]
- GitHub, CSSEGISandData. Available online: https://github.com/CSSEGISandData/COVID-19/tree/master/csse_covid_19_data (accessed on 13 May 2022).
- Bernhauerová, V. Adapting to Vaccination. Nat. Ecol. Evol. 2022, 6, 673–674. [Google Scholar] [CrossRef]
- Paules, C.I.; Marston, H.D.; Fauci, A.S. Coronavirus Infections-More Than Just the Common Cold. JAMA 2020, 323, 707–708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, X.; Wang, T.; Cai, D.; Hu, Z.; Chen, J.; Liao, H.; Zhi, L.; Wei, H.; Zhang, Z.; Qiu, Y.; et al. Cytokine storm intervention in the early stages of COVID-19 pneumonia. Cytokine Growth Factor Rev. 2020, 53, 38–42. [Google Scholar] [CrossRef]
- Kovacech, B.; Fialova, L.; Filipcik, P.; Skrabana, R.; Zilkova, M.; Paulenka-Ivanovova, N.; Kovac, A.; Palova, D.; Rolkova, G.P.; Tomkova, K.; et al. Monoclonal antibodies targeting two immunodominant epitopes on the Spike protein neutralize emerging SARS-CoV-2 variants of concern. EBioMedicine 2022, 76, 103818. [Google Scholar] [CrossRef]
- Markov, P.V.; Katzourakis, A.; Stilianakis, N.I. Antigenic evolution will lead to new SARS-CoV-2 variants with unpredictable severity. Nat. Rev. Microbiol. 2022, 20, 251–252. [Google Scholar] [CrossRef]
- Bull, J.J.; Rustom, A. Which ‘Imperfect Vaccines’ Encourage the Evolution of Higher Virulence? Evol. Med. Public Health 2022, 10, 202–213. [Google Scholar] [CrossRef]
- He, C.; He, X.; Yang, J.; Lei, H.; Hong, W.; Song, X.; Yang, L.; Li, J.; Wang, W.; Shen, G.; et al. Spike Protein of SARS-CoV-2 Omicron (B.1.1.529) Variant Have a Reduced Ability to Induce the Immune Response. Signal Transduct. Target. Ther. 2022, 7, 119. [Google Scholar] [CrossRef]
- Rössler, A.; Riepler, L.; Bante, D.; von Laer, D.; Kimpel, J. SARS-CoV-2 Omicron Variant Neutralization in Serum from Vaccinated and Convalescent Persons. N. Engl. J. Med. 2022, 386, 698–700. [Google Scholar] [CrossRef]
- Mallory, R.; Formica, N.; Pfeiffer, S.; Wilkinson, B.; Marcheschi, A.; Albert, G.; McFall, H.; Robinson, M.; Plested, J.S.; Zhu, M.; et al. Immunogenicity and Safety Following a Homologous Booster Dose of a SARS-CoV-2 Recombinant Spike Protein Vaccine (NVX-CoV2373): A Phase 2 Randomized Placebo-Controlled Trial. medRxiv 2021. [Google Scholar] [CrossRef]
- Yu, X.; Wei, D.; Xu, W.; Li, Y.; Li, X.; Zhang, X.; Qu, J.; Yang, Z.; Chen, E. Reduced sensitivity of SARS-CoV-2 Omicron variant to antibody neutralization elicited by booster vaccination. Cell Discov. 2022, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Haveri, A.; Solastie, A.; Ekström, N.; Österlund, P.; Nohynek, H.; Nieminen, T.; Palmu, A.A.; Melin, M. Neutralizing antibodies to SARS-CoV-2 Omicron variant after third mRNA vaccination in health care workers and elderly subjects. Eur. J. Immunol. 2022, 52, 816–824. [Google Scholar] [CrossRef] [PubMed]
- Gruell, H.; Vanshylla, K.; Tober-Lau, P.; Hillus, D.; Schommers, P.; Lehmann, C.; Kurth, F.; Sander, L.E.; Klein, F. MRNA Booster Immunization Elicits Potent Neutralizing Serum Activity against the SARS-CoV-2 Omicron Variant. Nat. Med. 2022, 28, 477–480. [Google Scholar] [CrossRef]
- Nemet, I.; Kliker, L.; Lustig, Y.; Zuckerman, N.; Erster, O.; Cohen, C.; Kreiss, Y.; Alroy-Preis, S.; Regev-Yochay, G.; Mendelson, E.; et al. Third BNT162b2 Vaccination Neutralization of SARS-CoV-2 Omicron Infection. N. Engl. J. Med. 2022, 386, 492–494. [Google Scholar] [CrossRef]
- Bar-On, Y.M.; Goldberg, Y.; Mandel, M.; Bodenheimer, O.; Amir, O.; Freedman, L.; Alroy-Preis, S.; Ash, N.; Huppert, A.; Milo, R. Protection by a Fourth Dose of BNT162b2 against Omicron in Israel. N. Engl. J. Med. 2022, 386, 1712–1720. [Google Scholar] [CrossRef]
- Lauring, A.S.; Tenforde, M.W.; Chappell, J.D.; Gaglani, M.; Ginde, A.A.; McNeal, T.; Ghamande, S.; Douin, D.J.; Talbot, H.K.; Casey, J.D.; et al. Clinical severity of, and effectiveness of mRNA vaccines against, covid-19 from omicron, delta, and alpha SARS-CoV-2 variants in the United States: Prospective observational study. BMJ 2022, 376, e069761. [Google Scholar] [CrossRef]
- Choi, A.; Koch, M.; Wu, K.; Chu, L.; Ma, L.; Hill, A.; Nunna, N.; Huang, W.; Oestreicher, J.; Colpitts, T.; et al. Safety and immunogenicity of SARS-CoV-2 variant mRNA vaccine boosters in healthy adults: An interim analysis. Nat. Med. 2021, 27, 2025–2031. [Google Scholar] [CrossRef]
- Massare, M.J.; Patel, N.; Zhou, B.; Maciejewski, S.; Flores, R.; Guebre-Xabier, M.; Tian, J.; Portnoff, A.D.; Fries, L.; Shinde, V.; et al. Combination Respiratory Vaccine Containing Recombinant SARS-CoV-2 Spike and Quadrivalent Seasonal Influenza Hemagglutinin Nanoparticles with Matrix-M Adjuvant. bioRxiv 2021. [Google Scholar] [CrossRef]
| 21L BA.2 | 21K K346 | Title 3 BA.1 | 21A (Delta) | ||
|---|---|---|---|---|---|
| Prevalence | 1 January 2022 | 3% | 26% | 31% | 29% |
| 6 February 2022 | 14% | 40% | 30% | 7% | |
| RBD mutations | 339 | D | D | D | G |
| 346 | R | K | R | R | |
| 371 | F | L | L | S | |
| 373 | P | P | P | S | |
| 375 | F | F | F | S | |
| 417 | N | N | N | K | |
| 440 | K | K | K | N | |
| 446 | G | S | S | G | |
| 452 | L | L | L | R | |
| 477 | N | N | N | S | |
| 478 | K | K | K | K | |
| 484 | A | A | A | E | |
| 493 | R | R | R | Q | |
| 496 | G | S | S | G | |
| 498 | R | R | R | Q | |
| 501 | Y | Y | Y | N | |
| 505 | H | H | H | Y | |
| 547 | T | K | K | T |
| VoC | K417N | E484K | N501Y | L452R | T478K | Immune Escape |
|---|---|---|---|---|---|---|
| Alpha | + | + | ||||
| Beta | + | + | + | +++++ | ||
| Gamma | + (K417T) | + | + | +++ | ||
| Delta | + (E484Q) | + | + | ++ | ||
| Omicron | + | + (E484A) | + | + | +++++++ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kudriavtsev, A.V.; Vakhrusheva, A.V.; Novoseletsky, V.N.; Bozdaganyan, M.E.; Shaitan, K.V.; Kirpichnikov, M.P.; Sokolova, O.S. Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics. Viruses 2022, 14, 1603. https://doi.org/10.3390/v14081603
Kudriavtsev AV, Vakhrusheva AV, Novoseletsky VN, Bozdaganyan ME, Shaitan KV, Kirpichnikov MP, Sokolova OS. Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics. Viruses. 2022; 14(8):1603. https://doi.org/10.3390/v14081603
Chicago/Turabian StyleKudriavtsev, Aleksandr V., Anna V. Vakhrusheva, Valery N. Novoseletsky, Marine E. Bozdaganyan, Konstantin V. Shaitan, Mikhail P. Kirpichnikov, and Olga S. Sokolova. 2022. "Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics" Viruses 14, no. 8: 1603. https://doi.org/10.3390/v14081603
APA StyleKudriavtsev, A. V., Vakhrusheva, A. V., Novoseletsky, V. N., Bozdaganyan, M. E., Shaitan, K. V., Kirpichnikov, M. P., & Sokolova, O. S. (2022). Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics. Viruses, 14(8), 1603. https://doi.org/10.3390/v14081603






