Redesign and Validation of a Real-Time RT-PCR to Improve Surveillance for Avian Influenza Viruses of the H9 Subtype
Abstract
:1. Introduction
2. Materials and Methods
2.1. In Silico Update
2.2. rRT-PCR Assay Set Up
2.3. Analytical Specificity (Asp)
2.4. Analytical Sensitivity (ASe) and Repeatability
2.5. Diagnostic Sensitivity (DSe) and Specificity (DSp)
2.6. Reproducibility and Robustness
2.7. Procedural Modifications for Pan-H9 rRT-PCR Deployment
2.7.1. Automatic and Manual Nucleic Acids Isolation Kits
2.7.2. One-Step Real-Time RT-PCR Kits
2.8. Comparison with Existing Molecular Diagnostic Methods
2.9. Statistical Analysis
3. Results
3.1. Analytical Performance
3.1.1. Exclusivity, Inclusivity and Selectivity
3.1.2. Analytical Sensitivity and Repeatability
3.2. Diagnostic Performance with Clinical Samples
3.3. Interlaboratory Reproducibility
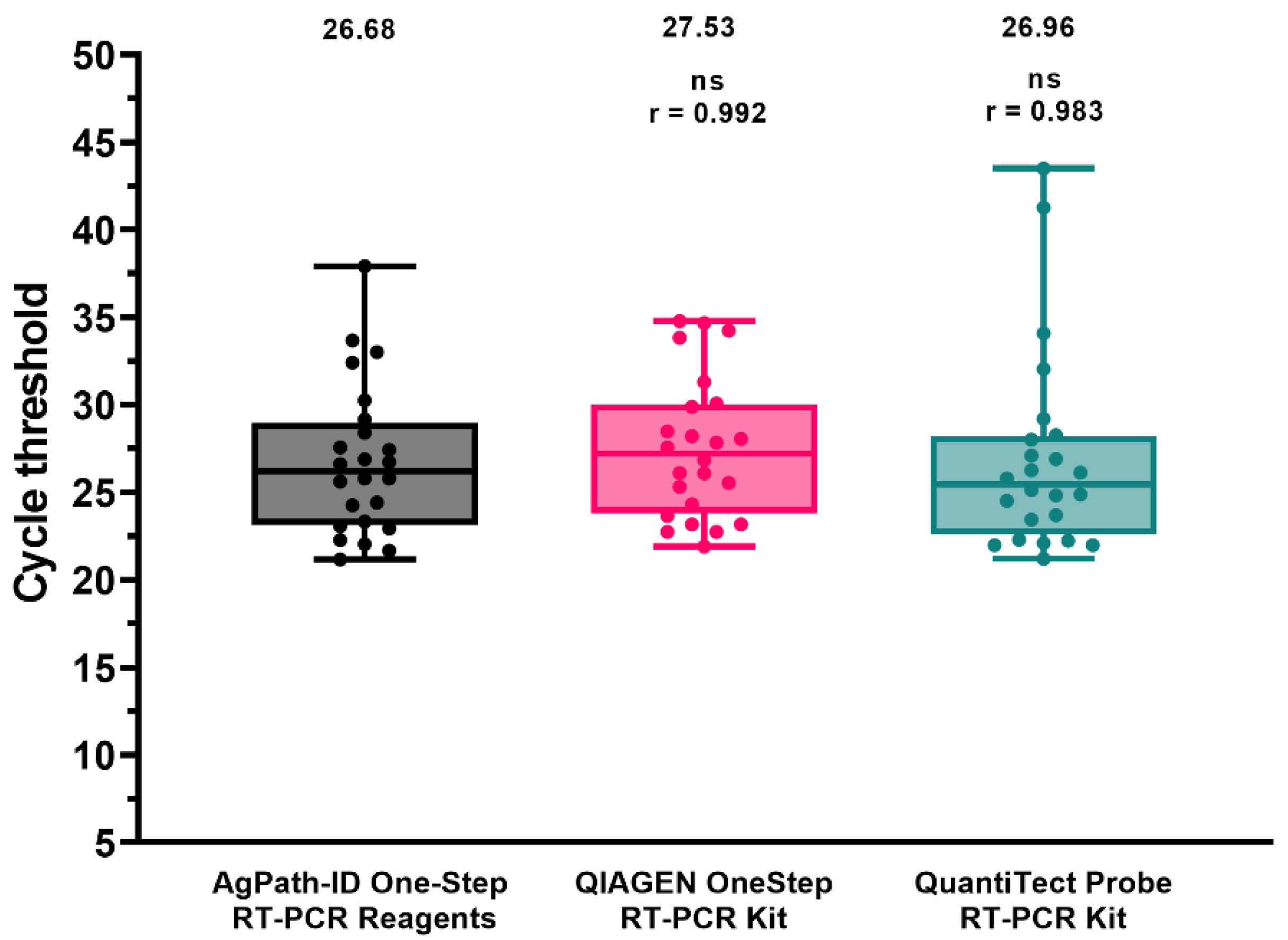
3.4. Potential for Pan-H9 rRT-PCR Deployment
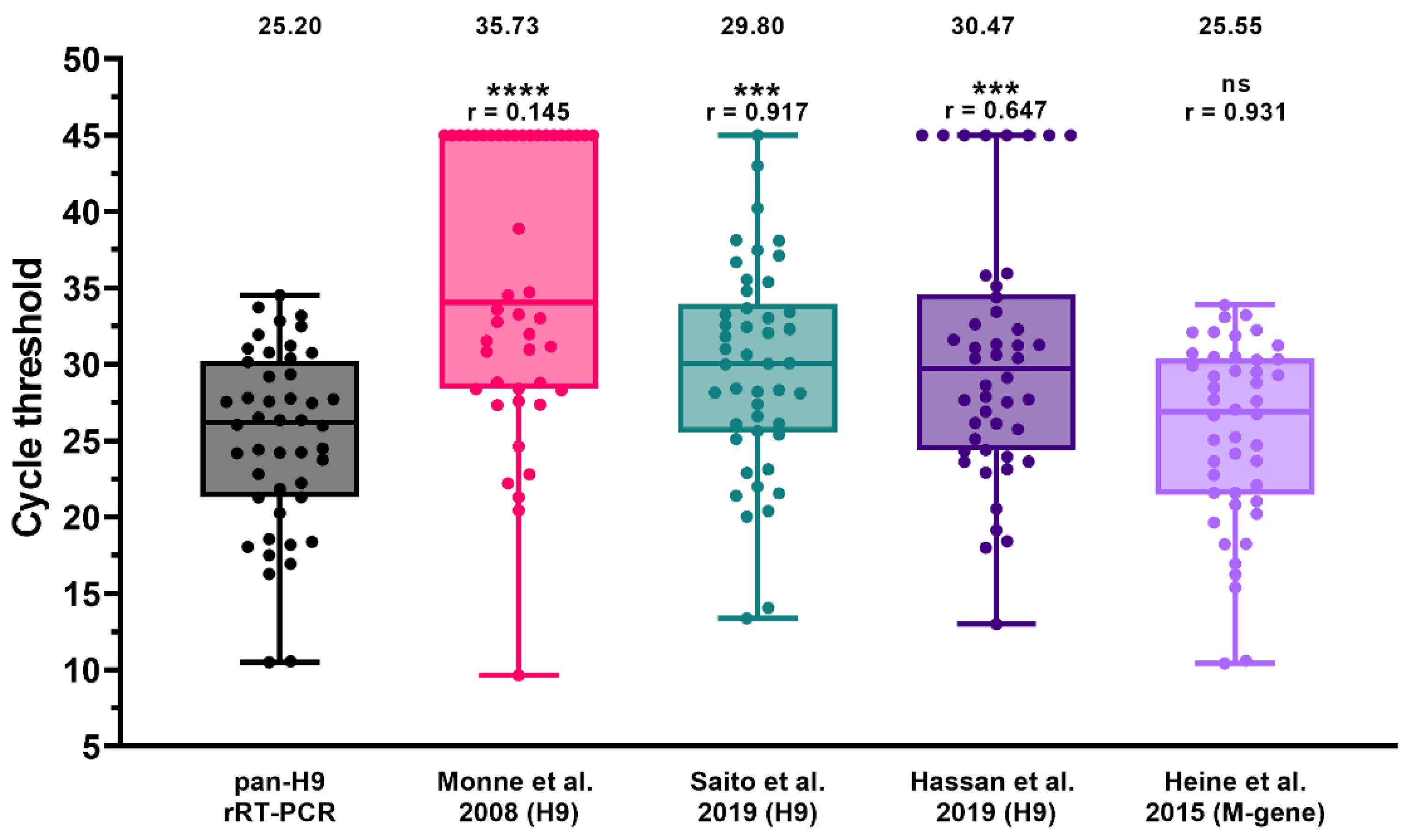
3.5. Comparison of the Pan-H9 rRT-PCRs with Other Assays
4. Discussion
5. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Homme, P.J.; Easterday, B.C. Avian Influenza Virus Infections. I. Characteristics of Influenza A-Turkey-Wisconsin-1966 Virus. Avian Dis. 1970, 14, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Capua, I.; Alexander, D.J. Ecology, Epidemiology and Human Health Implications of Avian Influenza Virus Infections. In Avian Influenza and Newcastle Disease; Springer: Milano, Italy, 2009; pp. 1–18. [Google Scholar] [CrossRef]
- Nagy, A.; Mettenleiter, T.C.; Abdelwhab, E.M. A Brief Summary of the Epidemiology and Genetic Relatedness of Avian Influenza H9N2 Virus in Birds and Mammals in the Middle East and North Africa. Epidemiol. Infect. 2017, 145, 3320–3333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peacock, T.H.P.; James, J.; Sealy, J.E.; Iqbal, M. A Global Perspective on H9N2 Avian Influenza Virus. Viruses 2019, 11, 620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carnaccini, S.; Perez, D.R. H9 Influenza Viruses: An Emerging Challenge. Cold Spring Harb. Perspect. Med. 2020, 10, a038588. [Google Scholar] [CrossRef] [PubMed]
- Channa, A.A.; Tariq, M.; Nizamani, Z.A.; Kalhoro, N.H. Prevalence of Avian Influenza H5, H7 and H9 Viruses in Commercial Layers in Karachi, Pakistan. Iran. J. Vet. Res. 2021, 22, 352–355. [Google Scholar] [CrossRef]
- Gupta, S.D.; Hoque, M.A.; Fournié, G.; Henning, J. Patterns of Avian Influenza A (H5) and A (H9) Virus Infection in Backyard, Commercial Broiler and Layer Chicken Farms in Bangladesh. Transbound. Emerg. Dis. 2021, 68, 137–151. [Google Scholar] [CrossRef]
- Kim, Y.; Biswas, P.K.; Giasuddin, M.; Hasan, M.; Mahmud, R.; Chang, Y.-M.; Essen, S.; Samad, M.A.; Lewis, N.S.; Brown, I.H.; et al. Prevalence of Avian Influenza A (H5) and A (H9) Viruses in Live Bird Markets, Bangladesh. Emerg. Infect. Dis. 2018, 24, 2309–2316. [Google Scholar] [CrossRef]
- Luo, S.; Xie, Z.; Li, M.; Li, D.; Xie, L.; Huang, J.; Zhang, M.; Zeng, T.; Wang, S.; Fan, Q.; et al. Survey of Low Pathogenic Avian Influenza Viruses in Live Poultry Markets in Guangxi Province, Southern China, 2016–2019. Sci. Rep. 2021, 11, 23223. [Google Scholar] [CrossRef]
- Nugroho, C.; Silaen, O.; Kurnia, R.; Soejoedono, R.; Poetri, O.; Soebandrio, A. Isolation and Molecular Characterization of the Hemagglutinin Gene of H9N2 Avian Influenza Viruses from Poultry in Java, Indonesia. J. Adv. Vet. Anim. Res. 2021, 8, 423–434. [Google Scholar] [CrossRef]
- Rahman, M.M.; Nooruzzaman, M.; Kabiraj, C.K.; Mumu, T.T.; Das, P.M.; Chowdhury, E.H.; Islam, M.R. Surveillance on Respiratory Diseases Reveals Enzootic Circulation of Both H5 and H9 Avian Influenza Viruses in Small-scale Commercial Layer Farms of Bangladesh. Zoonoses Public Health 2021, 68, 896–907. [Google Scholar] [CrossRef]
- Shaban, S.; Kyei, F.; Awuni, J.; Danquah, A.; Odoom, T.; Yingar, D.N.Y.T.; Ababio, P.T.; Emikpe, B.O. Dynamics of Influenza A (Avian Influenza) Virus in Poultry in the Greater Accra Region of Ghana amongst the Production Levels. J. Immunoass. Immunochem. 2022, 43, 1952426. [Google Scholar] [CrossRef] [PubMed]
- Nili, H.; Asasi, K. Natural Cases and an Experimental Study of H9N2 Avian Influenza in Commercial Broiler Chickens of Iran. Avian Pathol. 2002, 31, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.A.; Cho, S.H.; Kim, H.S.; Seo, S.H. H9N2 Influenza Viruses Isolated from Poultry in Korean Live Bird Markets Continuously Evolve and Cause the Severe Clinical Signs in Layers. Vet. Microbiol. 2006, 118, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Banet-Noach, C.; Perk, S.; Simanov, L.; Grebenyuk, N.; Rozenblut, E.; Pokamunski, S.; Pirak, M.; Tendler, Y.; Panshin, A. H9N2 influenza viruses from Israeli poultry: A five-year outbreak. Avian Dis. 2007, 51 (Suppl. S1), 290–296. [Google Scholar] [CrossRef] [PubMed]
- Nili, H.; Mohammadi, A.; Habibi, H.; Firouzi, S. Pathogenesis of H9N2 Virus in Chukar Partridges. Avian Pathol. 2013, 42, 230–234. [Google Scholar] [CrossRef]
- Jakhesara, S.J.; Bhatt, V.D.; Patel, N.V.; Prajapati, K.S.; Joshi, C.G. Isolation and Characterization of H9N2 Influenza Virus Isolates from Poultry Respiratory Disease Outbreak. SpringerPlus 2014, 3, 196. [Google Scholar] [CrossRef] [Green Version]
- Bonfante, F.; Mazzetto, E.; Zanardello, C.; Fortin, A.; Gobbo, F.; Maniero, S.; Bigolaro, M.; Davidson, I.; Haddas, R.; Cattoli, G.; et al. A G1-Lineage H9N2 Virus with Oviduct Tropism Causes Chronic Pathological Changes in the Infundibulum and a Long-Lasting Drop in Egg Production. Vet. Res. 2018, 49, 83. [Google Scholar] [CrossRef] [Green Version]
- Chuan, Z.Y.; Bin, Z.; Hui, S.Z.; Jing, W.X.; Hui, F.X.; Xi, G.L.; Ying, L.; Yan, C.X.; Feng, Z.Z. Replication and Pathology of Duck Influenza Virus Subtype H9N2 in Chukar. Biomed. Environ. Sci. 2018, 31, 306–310. [Google Scholar] [CrossRef]
- Awuni, J.A.; Bianco, A.; Dogbey, O.J.; Fusaro, A.; Yingar, D.T.; Salviato, A.; Ababio, P.T.; Milani, A.; Bonfante, F.; Monne, I. Avian Influenza H9N2 Subtype in Ghana: Virus Characterization and Evidence of Co-Infection. Avian Pathol. 2019, 48, 470–476. [Google Scholar] [CrossRef]
- Kariithi, H.M.; Welch, C.N.; Ferreira, H.L.; Pusch, E.A.; Ateya, L.O.; Binepal, Y.S.; Apopo, A.A.; Dulu, T.D.; Afonso, C.L.; Suarez, D.L. Genetic Characterization and Pathogenesis of the First H9N2 Low Pathogenic Avian Influenza Viruses Isolated from Chickens in Kenyan Live Bird Markets. Infect. Genet. Evol. 2020, 78, 104074. [Google Scholar] [CrossRef]
- Kye, S.-J.; Park, M.-J.; Kim, N.-Y.; Lee, Y.-N.; Heo, G.-B.; Baek, Y.-K.; Shin, J.-I.; Lee, M.-H.; Lee, Y.-J. Pathogenicity of H9N2 Low Pathogenic Avian Influenza Viruses of Different Lineages Isolated from Live Bird Markets Tested in Three Animal Models: SPF Chickens, Korean Native Chickens, and Ducks. Poult. Sci. 2021, 100, 101318. [Google Scholar] [CrossRef] [PubMed]
- Sei, S. Natural Co-Infection Caused by Avian Influenza H9 Subtype and Infectious Bronchitis Viruses in Broiler Chicken Farms. Vet. Arhiv 2010, 80, 269–281. Available online: https://hrcak.srce.hr/56716 (accessed on 19 January 2019).
- Pan, Q.; Liu, A.; Zhang, F.; Ling, Y.; Ou, C.; Hou, N.; He, C. Co-Infection of Broilers with Ornithobacterium Rhinotracheale and H9N2 Avian Influenza Virus. BMC Vet. Res. 2012, 8, 104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Śmietanka, K.; Minta, Z.; Świętoń, E.; Olszewska, M.; Jóźwiak, M.; Domańska-Blicharz, K.; Wyrostek, K.; Tomczyk, G.; Pikuła, A. Avian Influenza H9N2 Subtype in Poland—Characterization of the Isolates and Evidence of Concomitant Infections. Avian Pathol. 2014, 43, 427–436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonfante, F.; Cattoli, G.; Leardini, S.; Salomoni, A.; Mazzetto, E.; Davidson, I.; Haddas, R.; Terregino, C. Synergy or Interference of a H9N2 Avian Influenza Virus with a Velogenic Newcastle Disease Virus in Chickens Is Dose Dependent. Avian Pathol. 2017, 46, 488–496. [Google Scholar] [CrossRef] [PubMed]
- Naguib, M.M.; El-Kady, M.F.; Lüschow, D.; Hassan, K.E.; Arafa, A.-S.; El-Zanaty, A.; Hassan, M.K.; Hafez, H.M.; Grund, C.; Harder, T.C. New Real Time and Conventional RT-PCRs for Updated Molecular Diagnosis of Infectious Bronchitis Virus Infection (IBV) in Chickens in Egypt Associated with Frequent Co-Infections with Avian Influenza and Newcastle Disease Viruses. J. Virol. Methods 2017, 245, 19–27. [Google Scholar] [CrossRef]
- Horwood, P.F.; Horm, S.V.; Suttie, A.; Thet, S.; Phalla, Y.; Rith, S.; Sorn, S.; Holl, D.; Tum, S.; Ly, S.; et al. Co-Circulation of Influenza A H5, H7, and H9 Viruses and Co-Infected Poultry in Live Bird Markets, Cambodia. Emerg. Infect. Dis. 2018, 24, 352–355. [Google Scholar] [CrossRef] [Green Version]
- Hassan, K.E.; El-Kady, M.F.; El-Sawah, A.A.A.; Luttermann, C.; Parvin, R.; Shany, S.; Beer, M.; Harder, T. Respiratory Disease Due to Mixed Viral Infections in Poultry Flocks in Egypt between 2017 and 2018: Upsurge of Highly Pathogenic Avian Influenza Virus Subtype H5N8 since 2018. Transbound. Emerg. Dis. 2021, 68, 21–36. [Google Scholar] [CrossRef]
- Pusch, E.; Suarez, D. The Multifaceted Zoonotic Risk of H9N2 Avian Influenza. Vet. Sci. 2018, 5, 82. [Google Scholar] [CrossRef] [Green Version]
- Lam, T.T.-Y.; Wang, J.; Shen, Y.; Zhou, B.; Duan, L.; Cheung, C.-L.; Ma, C.; Lycett, S.J.; Leung, C.Y.-H.; Chen, X.; et al. The Genesis and Source of the H7N9 Influenza Viruses Causing Human Infections in China. Nature 2013, 502, 241–244. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Yuan, H.; Gao, R.; Zhang, J.; Wang, D.; Xiong, Y.; Fan, G.; Yang, F.; Li, X.; Zhou, J.; et al. Clinical and Epidemiological Characteristics of a Fatal Case of Avian Influenza A H10N8 Virus Infection: A Descriptive Study. Lancet 2014, 383, 714–721. [Google Scholar] [CrossRef]
- Shanmuganatham, K.K.; Jones, J.C.; Marathe, B.M.; Feeroz, M.M.; Jones-Engel, L.; Walker, D.; Turner, J.; Rabiul Alam, S.M.; Kamrul Hasan, M.; Akhtar, S.; et al. The Replication of Bangladeshi H9N2 Avian Influenza Viruses Carrying Genes from H7N3 in Mammals. Emerg. Microbes Infect. 2016, 5, e35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, Y.-Y.; Ke, C.-W.; Li, Q.; Yuan, R.-Y.; Xiang, D.; Jia, W.-X.; Yu, Y.-D.; Liu, L.; Huang, C.; Qi, W.-B.; et al. Novel Reassortant Influenza A (H5N6) Viruses in Humans, Guangdong, China, 2015. Emerg. Infect. Dis. 2016, 22, 1507–1509. [Google Scholar] [CrossRef] [PubMed]
- Tosh, C.; Nagarajan, S.; Kumar, M.; Murugkar, H.V.; Venkatesh, G.; Shukla, S.; Mishra, A.; Mishra, P.; Agarwal, S.; Singh, B.; et al. Multiple Introductions of a Reassortant H5N1 Avian Influenza Virus of Clade 2.3.2.1c with PB2 Gene of H9N2 Subtype into Indian Poultry. Infect. Genet. Evol. 2016, 43, 173–178. [Google Scholar] [CrossRef]
- Seiler, P.; Kercher, L.; Feeroz, M.M.; Shanmuganatham, K.; Jones-Engel, L.; Turner, J.; Walker, D.; Alam, S.M.R.; Hasan, M.K.; Akhtar, S.; et al. H9N2 Influenza Viruses from Bangladesh: Transmission in Chicken and New World Quail. Influenza Other Respir. Viruses 2018, 12, 814–817. [Google Scholar] [CrossRef] [PubMed]
- Ge, Z.; Xu, L.; Hu, X.; Zhu, S.; Zhao, Y.; Li, Y.; Liu, K.; Gao, R.; Wang, X.; Hu, J.; et al. Phylogenetic and Phenotypic Characterization of Two Novel Clade 2.3.2.1 H5N2 Subtype Avian Influenza Viruses from Chickens in China. Infect. Genet. Evol. 2022, 98, 105205. [Google Scholar] [CrossRef]
- FAO Global AIV with Zoonotic Potential Situation Update—FAO Emergency Prevention System for Animal Health (EMPRES-AH). Available online: https://www.fao.org/ag/againfo/programmes/en/empres/Global_AIV_Zoonotic_Update/situation_update.html (accessed on 22 February 2022).
- Lee, D.-H.; Song, C.-S. H9N2 Avian Influenza Virus in Korea: Evolution and Vaccination. Clin. Exp. Vaccine Res. 2013, 2, 26–33. [Google Scholar] [CrossRef] [Green Version]
- EL Houadfi, M.; Fellahi, S.; Nassik, S.; Guérin, J.-L.; Ducatez, M.F. First Outbreaks and Phylogenetic Analyses of Avian Influenza H9N2 Viruses Isolated from Poultry Flocks in Morocco. Virol. J. 2016, 13, 140. [Google Scholar] [CrossRef] [Green Version]
- Park, K.J.; Kwon, H.-i.; Song, M.-S.; Pascua, P.N.Q.; Baek, Y.H.; Lee, J.H.; Jang, H.-L.; Lim, J.-Y.; Mo, I.-P.; Moon, H.-J.; et al. Rapid Evolution of Low-Pathogenic H9N2 Avian Influenza Viruses Following Poultry Vaccination Programmes. J. Gen. Virol. 2011, 92 Pt 1, 36–50. [Google Scholar] [CrossRef]
- Bahari, P.; Pourbakhsh, S.A.; Shoushtari, H.; Bahmaninejad, M.A. Molecular Characterization of H9N2 Avian Influenza Viruses Isolated from Vaccinated Broiler Chickens in Northeast Iran. Trop. Anim. Health Prod. 2015, 47, 1195–1201. [Google Scholar] [CrossRef]
- El Mellouli, F.; Mouahid, M.; Fusaro, A.; Zecchin, B.; Zekhnini, H.; El Khantour, A.; Giussani, E.; Palumbo, E.; Rguibi Idrissi, H.; Monne, I.; et al. Spatiotemporal Dynamics, Evolutionary History and Zoonotic Potential of Moroccan H9N2 Avian Influenza Viruses from 2016 to 2021. Viruses 2022, 14, 509. [Google Scholar] [CrossRef] [PubMed]
- Ben Shabat, M.; Meir, R.; Haddas, R.; Lapin, E.; Shkoda, I.; Raibstein, I.; Perk, S.; Davidson, I. Development of a Real-Time TaqMan RT-PCR Assay for the Detection of H9N2 Avian Influenza Viruses. J. Virol. Methods 2010, 168, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Lv, J.; Wei, B.; Chai, T.; Xia, X.; Miao, Z.; Yao, M.; Gao, Y.; Huang, R.; Yang, H.; Roesler, U. Development of a Real-Time RT-PCR Method for Rapid Detection of H9 Avian Influenza Virus in the Air. Arch. Virol. 2011, 156, 1795–1801. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Yao, L.; Zhai, F.; Chen, Y.; Lei, J.; Bi, Z.; Hu, J.; Xiao, Q.; Song, S.; Yan, L.; et al. Development and Application of a Triplex Real-Time PCR Assay for the Simultaneous Detection of Avian Influenza Virus Subtype H5, H7 and H9. J. Virol. Methods 2018, 252, 49–56. [Google Scholar] [CrossRef]
- Mirzaei, S.G.; Shoushtari, A.; Nouri, A. Development and Evaluation of Real-Time RT-PCR Test for Quantitative and Qualitative Recognition of Current H9N2 Subtype Avian Influenza Viruses in Iran. Arch. Razi Inst. 2018, 73, 177–182. [Google Scholar] [CrossRef]
- Monne, I.; Ormelli, S.; Salviato, A.; De Battisti, C.; Bettini, F.; Salomoni, A.; Drago, A.; Zecchin, B.; Capua, I.; Cattoli, G. Development and Validation of a One-Step Real-Time PCR Assay for Simultaneous Detection of Subtype H5, H7, and H9 Avian Influenza Viruses. J. Clin. Microbiol. 2008, 46, 1769–1773. [Google Scholar] [CrossRef] [Green Version]
- Hoffmann, B.; Hoffmann, D.; Henritzi, D.; Beer, M.; Harder, T.C. Riems Influenza a Typing Array (RITA): An RT-QPCR-Based Low Density Array for Subtyping Avian and Mammalian Influenza a Viruses. Sci Rep. 2016, 6, 27211. [Google Scholar] [CrossRef] [Green Version]
- Slomka, M.J.; Hanna, A.; Mahmood, S.; Govil, J.; Krill, D.; Manvell, R.J.; Shell, W.; Arnold, M.E.; Banks, J.; Brown, I.H. Phylogenetic and Molecular Characteristics of Eurasian H9 Avian Influenza Viruses and Their Detection by Two Different H9-Specific RealTime Reverse Transcriptase Polymerase Chain Reaction Tests. Vet. Microbiol. 2013, 162, 530–542. [Google Scholar] [CrossRef] [Green Version]
- Khantour, A.E.; Soulaymani, A.; Salek, M.; Maltouf, A.F.; Darkaoui, S.; Mellouli, F.E.; Ducatez, M.; Fellahi, S. Molecular Characterization of the Hemagglutinin Gene of H9N2 Avian Influenza Viruses Isolated from Broiler Flocks in Morocco from 2016 to 2018. Vet. Arh. 2020, 90, 477. [Google Scholar] [CrossRef]
- Saito, S.; Takayama, I.; Nakauchi, M.; Nagata, S.; Oba, K.; Odagiri, T.; Kageyama, T. Development and Evaluation of a New Real-Time RT-PCR Assay for Detecting the Latest H9N2 Influenza Viruses Capable of Causing Human Infection. Microbiol. Immunol. 2019, 63, 21–31. [Google Scholar] [CrossRef]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data—From vision to reality. Eurosurveillance 2017, 22, 30494. [Google Scholar] [CrossRef] [Green Version]
- Kuraku, S.; Zmasek, C.M.; Nishimura, O.; Katoh, K. aLeaves Facilitates On-Demand Exploration of Metazoan Gene Family Trees on MAFFT Sequence Alignment Server with Enhanced Interactivity. Nucleic Acids Res. 2013, 41, W22–W28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT Online Service: Multiple Sequence Alignment, Interactive Sequence Choice and Visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Geneious Prime 2020.1.2. Available online: https://www.geneious.com (accessed on 19 January 2019).
- Nagy, A.; Jiřinec, T.; Černíková, L.; Jiřincová, H.; Havlíčková, M. Large-scale nucleotide sequence alignment and sequence variability assessment to identify the evolutionarily highly conserved regions for universal screening PCR assay design: An example of influenza A virus. Methods Mol. Biol. 2015, 1275, 57–72. [Google Scholar] [CrossRef] [PubMed]
- Nagy, A.; Jiřinec, T.; Jiřincová, H.; Černíková, L.; Havlíčková, M. In Silico Re-Assessment of a Diagnostic RT-QPCR Assay for Universal Detection of Influenza A Viruses. Sci. Rep. 2019, 9, 1630. [Google Scholar] [CrossRef]
- Integrated DNA Technologies OligoAnalyzer. Available online: https://eu.idtdna.com/calc/analyzer (accessed on 20 May 2021).
- Heine, H.G.; Foord, A.J.; Wang, J.; Valdeter, S.; Walker, S.; Morrissy, C.; Wong, F.Y.K.; Meehan, B. Detection of Highly Pathogenic Zoonotic Influenza Virus H5N6 by Reverse-Transcriptase Quantitative Polymerase Chain Reaction. Virol. J. 2015, 12, 18. [Google Scholar] [CrossRef] [Green Version]
- Laconi, A.; Fortin, A.; Bedendo, G.; Shibata, A.; Sakoda, Y.; Awuni, J.A.; Go-Maro, E.; Arafa, A.; Maken Ali, A.S.; Terregino, C.; et al. Detection of Avian Influenza Virus: A Comparative Study of the In Silico and In Vitro Performances of Current RT-QPCR Assays. Sci. Rep. 2020, 10, 8441. [Google Scholar] [CrossRef]
- OIE. Principles and Methods of Validation of Diagnostic Assays for Infectious Disease (Version Adopted in May 2013). In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals; OIE: Paris, France, 2021; Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/1.01.06_VALIDATION.pdf (accessed on 13 July 2021).
- Waugh, C.M.; Clark, G.A.; Hollier, J.; Watson, J. FAO/OIE Global Network of Expertise on Animal Influenzas (OFFLU); CSIRO-ACDP: East Geelong, Australia, 2022. [Google Scholar]
- Hassan, K.E.; Ahrens, A.K.; Ali, A.; El-Kady, M.F.; Hafez, H.M.; Mettenleiter, T.C.; Beer, M.; Harder, T. Improved Subtyping of Avian Influenza Viruses Using an RT-QPCR-Based Low Density Array: ‘Riems Influenza a Typing Array’, Version 2 (RITA-2). Viruses 2022, 14, 415. [Google Scholar] [CrossRef]
- Suttie, A.; Tok, S.; Yann, S.; Keo, P.; Horm, S.V.; Roe, M.; Kaye, M.; Sorn, S.; Holl, D.; Tum, S.; et al. The Evolution and Genetic Diversity of Avian Influenza A (H9N2) Viruses in Cambodia, 2015–2016. PLoS ONE 2019, 14, e0225428. [Google Scholar] [CrossRef] [Green Version]
- Yan, W.; Cui, H.; Engelsma, M.; Beerens, N.; van Oers, M.M.; de Jong, M.C.M.; Li, X.; Liu, Q.; Yang, J.; Teng, Q.; et al. Molecular and Antigenic Characterization of Avian H9N2 Viruses in Southern China. Microbiol. Spectr. 2022, 10, e00822-21. [Google Scholar] [CrossRef]
- Van Borm, S.; Belák, S.; Freimanis, G.; Fusaro, A.; Granberg, F.; Höper, D.; King, D.P.; Monne, I.; Orton, R.; Rosseel, T. Next-Generation Sequencing in Veterinary Medicine: How Can the Massive Amount of Information Arising from High-Throughput Technologies Improve Diagnosis, Control, and Management of Infectious Diseases? In Veterinary Infection Biology: Molecular Diagnostics and High-Throughput Strategies; Cunha, M.V., Inácio, J., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2015; Volume 1247, pp. 415–436. ISBN 978-1-4939-2003-7. [Google Scholar]
| Oligonucleotide | Sequence 5′ → 3′ | Nt. Position 1 |
|---|---|---|
| Pan-H9 for | ATR GGG TTT GCT GCC | 1615–1629 |
| Pan-H9 rev1 | TCA TAT ACA AAT GTT GCA YCT G | 1662–1683 |
| Pan-H9 rev2 | TTA TAT ACA GAT GTT GCA YCT G | 1662–1683 |
| Pan-H9 probe | TTC TGG GCY ATG TCH AAY GG | 1636–1655 |
| Origin | Collection Year | Matrix | Species | No. | |
|---|---|---|---|---|---|
| H9-confirmed cases | Europe | 2018–2022 | Swabs, stool | Mallard, teal, pheasant, goose, other unspecified avian species | 22 |
| Africa | 2019–2021 | Swabs, organs, FTA | Chicken, cockerel, other unspecified avian species | 21 | |
| Middle East | 2019–2021 | FTA | Chicken | 17 | |
| Asia | 2012–2021 | Organs, FTA | Chicken | 10 | |
| H9-negative samples * | Europe | 2018–2021 | Swabs, organs | Chicken, turkey, mallard, teal, pheasant, goose, quail, magpie, partridge, shoveler, duck, swan, gull | 53 |
| Strain | Tracheal Swab | Cloacal Swab | Oviduct | ||||||
|---|---|---|---|---|---|---|---|---|---|
| LoD (Ct) | E (%) | R2 | LoD (Ct) | E (%) | R2 | LoD (Ct) | E (%) | R2 | |
| A/pheasant/Italy/21VIR2284-22/2021/H9N2 (Y439 clade) | 1.51 (33.85) | 90.5 | 0.995 | 1.51 (33.48) | 95.9 | 0.997 | 1.51 (34.80) | 90.4 | 0.998 |
| A/chicken/Nigeria/19VIR8424-15/2019/H9N2 (G1 clade) | 3.16 (34.06) | 99.9 | 0.996 | 3.16 (35.06) | 94.8 | 0.998 | 3.16 (36.04) | 99.4 | 0.998 |
| A/chicken/Malaysia/2630-8/2012/H9N2 (Y280 clade) | 1.41 (33.78) | 95.6 | 0.998 | 1.41 (33.81) | 98.0 | 0.997 | 14.12 (32.01) | 97.8 | 0.996 |
| A/chicken/Malaysia/2630-8/2012/H9N2 (Y280 clade) with intype IC-RNA * | 1.41 (33.53) | 97.2 | 0.998 | n.t. | n.t. | n.t. | 14.12 (32.13) | 90.9 | 0.985 |
| Synthetic RNA * | No. of Mismatches | Performance | ||||
|---|---|---|---|---|---|---|
| For | Probe | Revs | LoD (Ct) | E (%) | R2 | |
| MN038193 (G1) | 2 | 0 | 0 | 101 (38.98) | 85.6 | 0.998 |
| MK553893 (Y280) | 0 | 1 | 6 | 105 (37.56) | 82.3 | 0.982 |
| MN765147 (Y280) | 0 | 0 | 2 | 102 (34.54) | 91.7 | 0.998 |
| MN765086 (Y280) | 0 | 1 (3′-end) | 5 | 104 (38.98) | 78.5 | 0.973 |
| Positive control § | 0 | 0 | 0 | 101 (37.56) | 86.4 | 0.993 |
| Nucleic Acids’ Purification Systems | Tracheal Swab | Cloacal Swab | Oviduct | ||||||
|---|---|---|---|---|---|---|---|---|---|
| LoD (Ct) | E (%) | R2 | LoD (Ct) | E (%) | R2 | LoD (Ct) | E (%) | R2 | |
| QIAsymphony DSP Virus/Pathogen Midi kit (Qiagen, Hilden, Germany) | 3.16 (34.06) | 99.9 | 0.996 | 3.16 (35.06) | 94.8 | 0.998 | 3.16 (36.04) | 99.4 | 0.998 |
| MagMAX Pathogen RNA/DNA Kit (Applied Biosystems, Waltham, MA, USA) | 3.16 (35.29) | 93.8 | 0.993 | 3.16 (33.97) | 98.7 | 0.999 | 3.16 (34.87) | 98.8 | 0.994 |
| NucleoSpin RNA (Macherey-Nagel, Dueren, Germany) | 31.6 (35.47) | 104.9 | 0.995 | 31.6 (34.38) | 95.4 | 0.997 | 31.6 (34.68) | 94.6 | 0.992 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Panzarin, V.; Marciano, S.; Fortin, A.; Brian, I.; D’Amico, V.; Gobbo, F.; Bonfante, F.; Palumbo, E.; Sakoda, Y.; Le, K.T.; et al. Redesign and Validation of a Real-Time RT-PCR to Improve Surveillance for Avian Influenza Viruses of the H9 Subtype. Viruses 2022, 14, 1263. https://doi.org/10.3390/v14061263
Panzarin V, Marciano S, Fortin A, Brian I, D’Amico V, Gobbo F, Bonfante F, Palumbo E, Sakoda Y, Le KT, et al. Redesign and Validation of a Real-Time RT-PCR to Improve Surveillance for Avian Influenza Viruses of the H9 Subtype. Viruses. 2022; 14(6):1263. https://doi.org/10.3390/v14061263
Chicago/Turabian StylePanzarin, Valentina, Sabrina Marciano, Andrea Fortin, Irene Brian, Valeria D’Amico, Federica Gobbo, Francesco Bonfante, Elisa Palumbo, Yoshihiro Sakoda, Kien Trung Le, and et al. 2022. "Redesign and Validation of a Real-Time RT-PCR to Improve Surveillance for Avian Influenza Viruses of the H9 Subtype" Viruses 14, no. 6: 1263. https://doi.org/10.3390/v14061263
APA StylePanzarin, V., Marciano, S., Fortin, A., Brian, I., D’Amico, V., Gobbo, F., Bonfante, F., Palumbo, E., Sakoda, Y., Le, K. T., Chu, D.-H., Shittu, I., Meseko, C., Haido, A. M., Odoom, T., Diouf, M. N., Djegui, F., Steensels, M., Terregino, C., & Monne, I. (2022). Redesign and Validation of a Real-Time RT-PCR to Improve Surveillance for Avian Influenza Viruses of the H9 Subtype. Viruses, 14(6), 1263. https://doi.org/10.3390/v14061263







