Spatial and Temporal Pattern of Norovirus Dispersal in an Oyster Growing Region in the Northeast Pacific
Abstract
:1. Introduction
2. Materials and Methods
2.1. Field Study
2.2. Norovirus Analysis
2.3. Norovirus Confirmation
2.4. Escherichia coli Analysis
2.5. Pathogen Prevalence and Correlation to Environmental Variables
2.6. Spatial and Temporal Analysis
3. Results
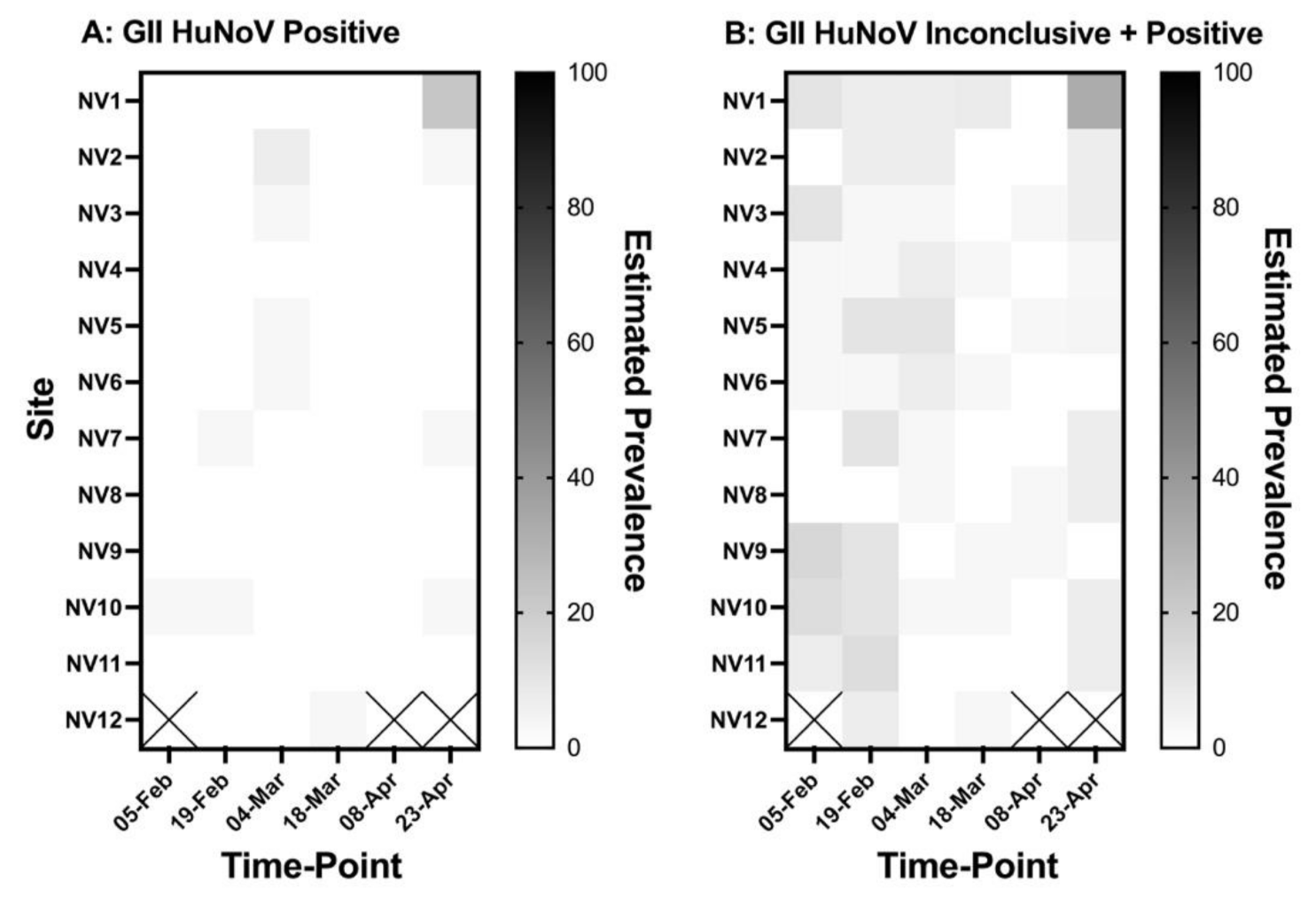
3.1. Description of Field Data
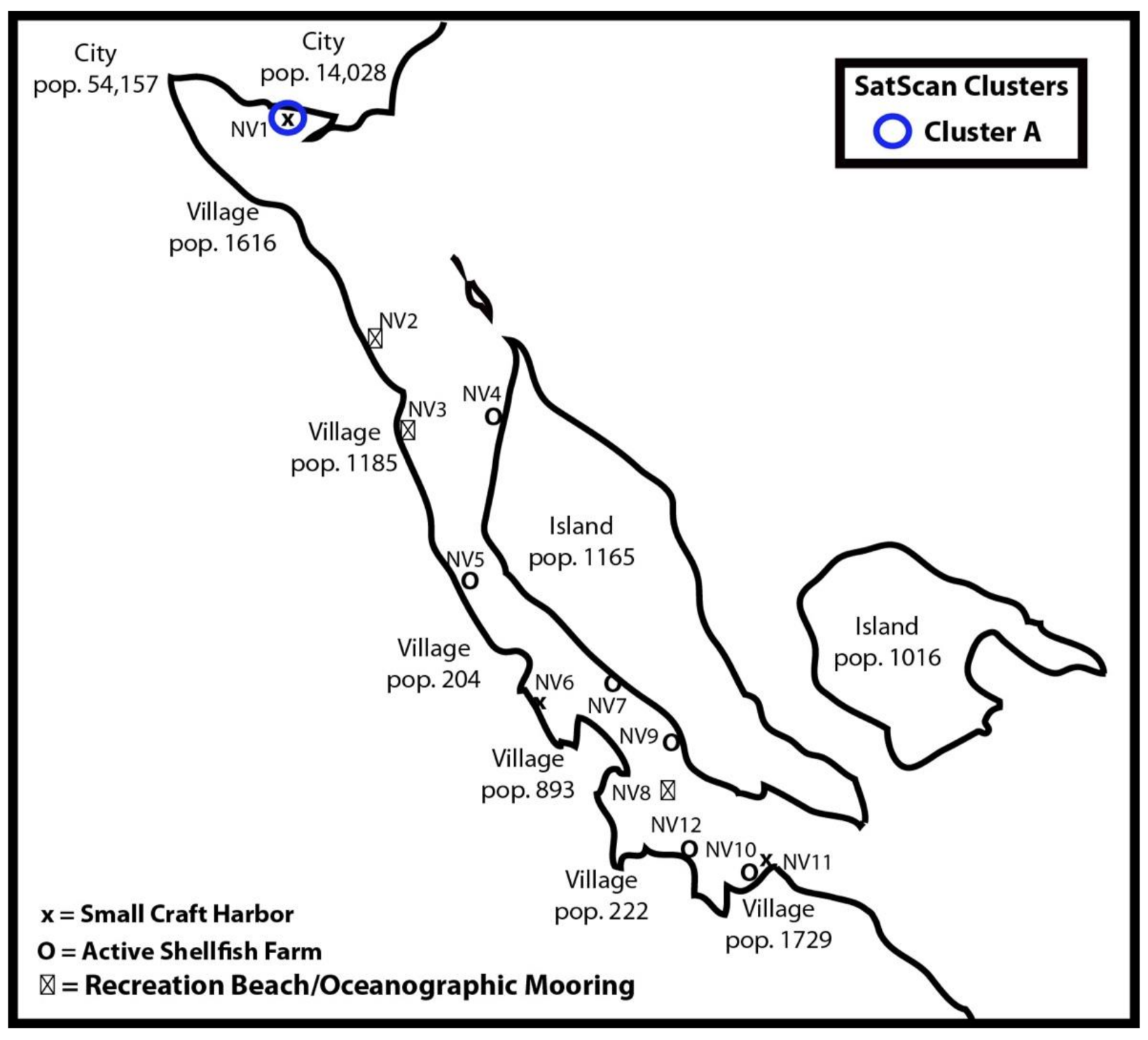
3.2. Cluster Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Belliot, G.; Lopman, B.A.; Ambert-Balay, K.; Pothier, P. The burden of norovirus gastroenteritis: An important foodborne and healthcare-related infection. Clin. Microbiol. Infect. 2014, 20, 724–730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, M.K.; Murray, R.; Flockhart, L.; Pintar, K.; Pollari, F.; Fazil, A.; Nesbitt, A.; Marshall, B. Estimates of the burden of foodbourne illness in Canada for 30 specified pathogens and unspecified agents, circa 2006. Foodborne Pathog. Dis. 2013, 10, 639–648. [Google Scholar] [CrossRef]
- Chhabra, P.; de Graaf, M.; Parra, G.I.; Chan, M.C.-W.; Green, K.; Martella, V.; Wang, Q.; White, P.A.; Katayama, K.; Vennema, H.; et al. Updated classification of norovirus genogroups and genotypes. J. Gen. Virol. 2019, 100, 1393–1406. [Google Scholar] [CrossRef]
- Zheng, D.-P.; Ando, T.; Fankhauser, R.L.; Beard, R.S.; Glass, R.I.; Monroe, S.S. Norovirus classification and proposed strain nomenclature. Virology 2006, 15, 312–323. [Google Scholar] [CrossRef] [Green Version]
- Teunis, P.F.M.; Moe, C.L.; Liu, P.; Miller, S.E.; Lindesmith, L.; Baric, R.S.; Le Pendu, J.; Calderon, R.L. Norwalk virus: How infectious is it? J. Med. Virol. 2008, 80, 1468–1476. [Google Scholar] [CrossRef]
- Karst, S.M. Pathogenesis of noroviruses, emerging RNA viruses. Viruses 2010, 2, 748–781. [Google Scholar] [CrossRef]
- Meghnath, K.; Hasselback, P.; McCormick, R.; Prystajecky, N.; Taylor, M.; McIntyre, L.; Man, S.; Whitfield, Y.; Warshawsky, B.; McKinley, M.; et al. Outbreaks of norovirus and acute gastroenteritis associated with British Columbia oysters, 2016–2017. Food Environ. Virol. 2019, 11, 138–148. [Google Scholar] [CrossRef] [PubMed]
- David, S.T.; McIntyre, L.; MacDougall, L.; Kelly, D.; Liem, S.; Schallie, K.; McNabb, A.; Houde, A.; Mueller, P.; Ward, P.; et al. Consumption of oysters from geographically dispersed harvest sites, British Columbia, Canada, 2004. Foodborne Pathog. Dis. 2007, 4, 349–358. [Google Scholar] [CrossRef]
- Campos, C.J.A.; Lees, D.N. Environmental transmission of human norovirus in shellfish waters. Appl. Environ. Microbiol. 2014, 80, 3552–3561. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burkhardt, W.; Calci, K.R. Selective accumulation may account for shellfish-associated viral illness. Appl. Environ. Microbiol. 2000, 66, 1375–1378. [Google Scholar] [CrossRef] [Green Version]
- Le Guyader, F.S.; Loisy, F.; Atmar, R.L.; Hutson, A.M.; Estes, M.K.; Ruvoen-Clouet, N.; Pommepuy, M.; Le Pendu, J. Norwalk virus-pecific binding to oyster digestive tissues. Emerg. Infect. Dis. 2006, 12, 931–936. [Google Scholar] [CrossRef] [PubMed]
- Pouillot, R.; Smith, M.; Van Doren, J.M.; Catford, A.; Holtzman, J.; Calci, K.R.; Edwards, R.; Goblick, G.; Roberts, C.; Stobo, J.; et al. Risk assessment of norovirus illness from consumption of raw oysters in the United States and in Canada. Risk Anal. 2021, 42, 344–369. [Google Scholar] [CrossRef] [PubMed]
- Hassard, F.; Sharp, J.H.; Taft, H.; LeVay, L.; Harris, J.P.; McDonald, J.E.; Tuson, K.; Wilson, J.; Jones, D.L.; Malham, S.K. Critical review on the public health impact of norovirus contamination in shellfish and the environment: A UK perspective. Food Environ. Virol. 2017, 9, 123–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Canada Government. Canadian Shellfish Sanitation Program Manual. Available online: https://inspection.canada.ca/food-safety-for-industry/food-specific-requirements-and-guidance/fish/canadian-shellfish-sanitation-program/eng/1527251566006/1527251566942?chap=0#s3c2 (accessed on 10 January 2022).
- McIntyre, L.; Galanis, E.; Prystajecky, N.; Kosatsky, T. BC oysters and norovirus: Hundreds of cases in months with an “R”. BC Med. J. 2017, 59, 326–327. [Google Scholar]
- B.C. Seafood Industry. British Columbia Seafood Industry Year in Review 2016. Available online: https://www2.gov.bc.ca/assets/gov/farming-natural-resources-and-industry/agriculture-and-seafood/statistics/industry-and-sector-profiles/year-in-review/bcseafood_yearinreview_2016.pdf (accessed on 8 February 2022).
- Dumbauld, B.R.; Ruesink, J.L.; Rumrill, S.S. The ecological role of bivalve shellfish aquaculture in the estuarine environment: A review with application to oyster and clam culture in West Coast (USA) estuaries. Aquaculture 2009, 290, 196–223. [Google Scholar] [CrossRef]
- Kroneman, A.; Vennema, H.; Deforhe, K.; Avoort, A.v.d.; Penaranda, S.; Oberste, M.S.; Vinje, J.; Koopmans, M. An automated genotyping tool for enteroviruses and noroviruses. J. Clin. Virol. 2011, 51, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Cowling, D.W.; Gardner, A.A.; Johnson, W.O. Comparison of methods for estimation of individual-level prevalence based on pooled samples. Prev. Vet. Med. 1999, 39, 211–225. [Google Scholar] [CrossRef]
- Kulldorff, M. A spatial scan statistic. Commun. Stat. Theory Methods 1997, 26, 1481–1496. [Google Scholar] [CrossRef]
- Lowther, J.A.; Gustar, N.E.; Powell, A.L.; Hartnell, R.E.; Lees, D.N. Two-year systematic study to assess norovirus contamination in oysters from commercial harvesting areas in the United Kingdom. Appl. Environ. Microbiol. 2012, 78, 5812–5817. [Google Scholar] [CrossRef] [Green Version]
- Laurin, E.; Thakur, K.; Mohr, P.G.; Hick, P.; Crane, M.S.J.; Gardner, A.A.; Moody, N.J.; Colling, A.; Ernst, I. To pool or not to pool? Guidelines for pooling samples for use in surveillance testing of infectious diseases in aquatic animals. J. Fish Dis. 2019, 42, 1471–1491. [Google Scholar] [CrossRef]
- Torok, V.; Hodgson, K.; McLeod, C.; Tan, J.; Malhi, N.; Turnbull, A. National survey of foodborne viruses in Australian oysters at production. Food Microbiol. 2018, 69, 196–203. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, J.; Le Saux, J.-C.; Lora, M.; Atmar, R.L.; Le Guyader, F.S. Norovirus contamination on French marketed oysters. Int. J. Food Microbiol. 2013, 166, 244–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DePaola, A.; Jones, J.L.; Woods, J.; Burkhardt III, W.; Calci, K.R.; Krantz, J.A.; Bower, J.C.; Kasturi, K.; Byars, R.H.; Jacobs, E.; et al. Bacterial and viral pathogens in live oysters: 2007 United States market survey. Appl. Environ. Microbiol. 2010, 76, 2754–2768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Costantini, V.; Loisy, F.; Joens, L.; Le Guyader, F.S.; Saif, L.J. Human and animal enteric Caliciviruses in oysters from different coastal regions of the United States. Appl. Environ. Microbiol. 2006, 72, 1800–1809. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Liu, D.; Zhang, Z.; Hewitt, J.; Li, X.; Hou, P.; Wang, D.; Wu, Q. Surveillance of human norovirus in oysters collected from production area in Shandong Province, China during 2017–2018. Food Control 2021, 121, 107649. [Google Scholar] [CrossRef]
- Polo, D.; Varela, M.F.; Romalde, J.L. Detection and quantification of hepatitis A virus and norovirus in Spanish authorized shellfish harvesting areas. Int. J. Food Microbiol. 2015, 193, 43–50. [Google Scholar] [CrossRef]
- Lowther, J.A.; Gustar, N.E.; Hartnell, E.; Lees, D.N. Comparison of norovirus RNA levels in outbreak-related oysters with background environmental levels. J. Food Prot. 2012, 75, 389–393. [Google Scholar] [CrossRef]
- Taylor, S.C.; Nadeau, K.; Abbasi, M.; Lachance, C.; Nguyen, M.; Fenrich, J. The ultimate qPCR experiment: Producing publication quality, reproducible data the first time. Trends Biotechnol. 2019, 37, 761–774. [Google Scholar] [CrossRef] [Green Version]
- Lowther, J.A.; Cross, L.; Stapleton, T.; Gustar, N.E.; Walker, D.I.; Sills, M.; Treagus, S.; Pollington, V.; Lees, D.N. Use of F-specific RNA bacteriophage to estimate infectious norovirus levels in oysters. Food Environ. Virol. 2019, 11, 247–258. [Google Scholar] [CrossRef]
- Brake, F.; Kiermeier, A.; Ross, T.; Holds, G.; Landinez, L.; McLeod, C. Spatial and temporal distribution of norovirus and E. coli in Sydney rock oystes following a sewage overflow into an estuary. Food Environ. Virol. 2018, 10, 7–15. [Google Scholar] [CrossRef]
- Goblick, G.N.; Anbarchian, J.M.; Woods, J.; Burkhardt, W.; Calci, K. Evaluating the dilution of wastewater treatment plant effluent and viral impacts on shellfish growing area in Mobile Bay, Alabama. J. Shellfish Res. 2011, 30, 979–987. [Google Scholar] [CrossRef]
- Kraay, A.N.M.; Han, P.; Kambhampati, A.K.; Wikswo, M.E.; Mirza, S.A.; Lopman, B.A. Impact of nonpharmaceutical interventions for severe acute respiratory syndrome coronaviurs 2 on norovirus outbreaks: An analysis of outbreaks by 9 US States. J. Inf. Dis. 2021, 224, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Desdouits, M.; Piquet, J.C.; Wacrenier, C.; Le Mennec, C.; Parnaudeau, S.; Jousse, S.; Rocq, S.; Bigault, L.; Contrant, M.; Garry, P. Can shellfish be used to monitor SARS-CoV-2 in the coastal environment. Sci. Total Environ. 2021, 778, 146270. [Google Scholar] [CrossRef] [PubMed]
| Target | Forward | Probe | Reverse | Reference |
|---|---|---|---|---|
| HuNoV GI | CGCTGGATGCGNTTCCAT | TGGACAGGAGATCGC | CCTTAGACGCCATCATCATTTAC | ISO 15216-1 |
| HuNoV GII | ATGTTCAGRTGGATGAGRTTCTCWGA | AGCACGTGGGAGGGCGATCG | ISO 15216-1 | |
| MS2 | ATTCCGACTGCGAGCTTATT | ATTCCCTCAGCAATCGCAGCAAACT | TTCGACATGGGTAATCCTCA | 17 |
| E. coli ybbW | TGATTGGCAAAATCTGGCCG | GAAATCGCCCAAATCGCCAT | 18 |
| Cluster | Sites | RT-PCR Confirm | Cluster Size (km) | Time Frame | OvE | p-Value |
|---|---|---|---|---|---|---|
| A | NV1 | GII.2 | 0 | 8 April to 23 April 2020 | 10.44 | 0.006 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Green, T.J.; Yin Walker, C.; Leduc, S.; Michalchuk, T.; McAllister, J.; Roth, M.; Janes, J.K.; Krogh, E.T. Spatial and Temporal Pattern of Norovirus Dispersal in an Oyster Growing Region in the Northeast Pacific. Viruses 2022, 14, 762. https://doi.org/10.3390/v14040762
Green TJ, Yin Walker C, Leduc S, Michalchuk T, McAllister J, Roth M, Janes JK, Krogh ET. Spatial and Temporal Pattern of Norovirus Dispersal in an Oyster Growing Region in the Northeast Pacific. Viruses. 2022; 14(4):762. https://doi.org/10.3390/v14040762
Chicago/Turabian StyleGreen, Timothy J., Chen Yin Walker, Sarah Leduc, Trevor Michalchuk, Joe McAllister, Myron Roth, Jasmine K. Janes, and Erik T. Krogh. 2022. "Spatial and Temporal Pattern of Norovirus Dispersal in an Oyster Growing Region in the Northeast Pacific" Viruses 14, no. 4: 762. https://doi.org/10.3390/v14040762
APA StyleGreen, T. J., Yin Walker, C., Leduc, S., Michalchuk, T., McAllister, J., Roth, M., Janes, J. K., & Krogh, E. T. (2022). Spatial and Temporal Pattern of Norovirus Dispersal in an Oyster Growing Region in the Northeast Pacific. Viruses, 14(4), 762. https://doi.org/10.3390/v14040762






