Clinical Tick-Borne Encephalitis in a Roe Deer (Capreolus capreolus L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Case History, Clinical Signs and Autopsy Findings
2.2. Virologic and Molecular Testing
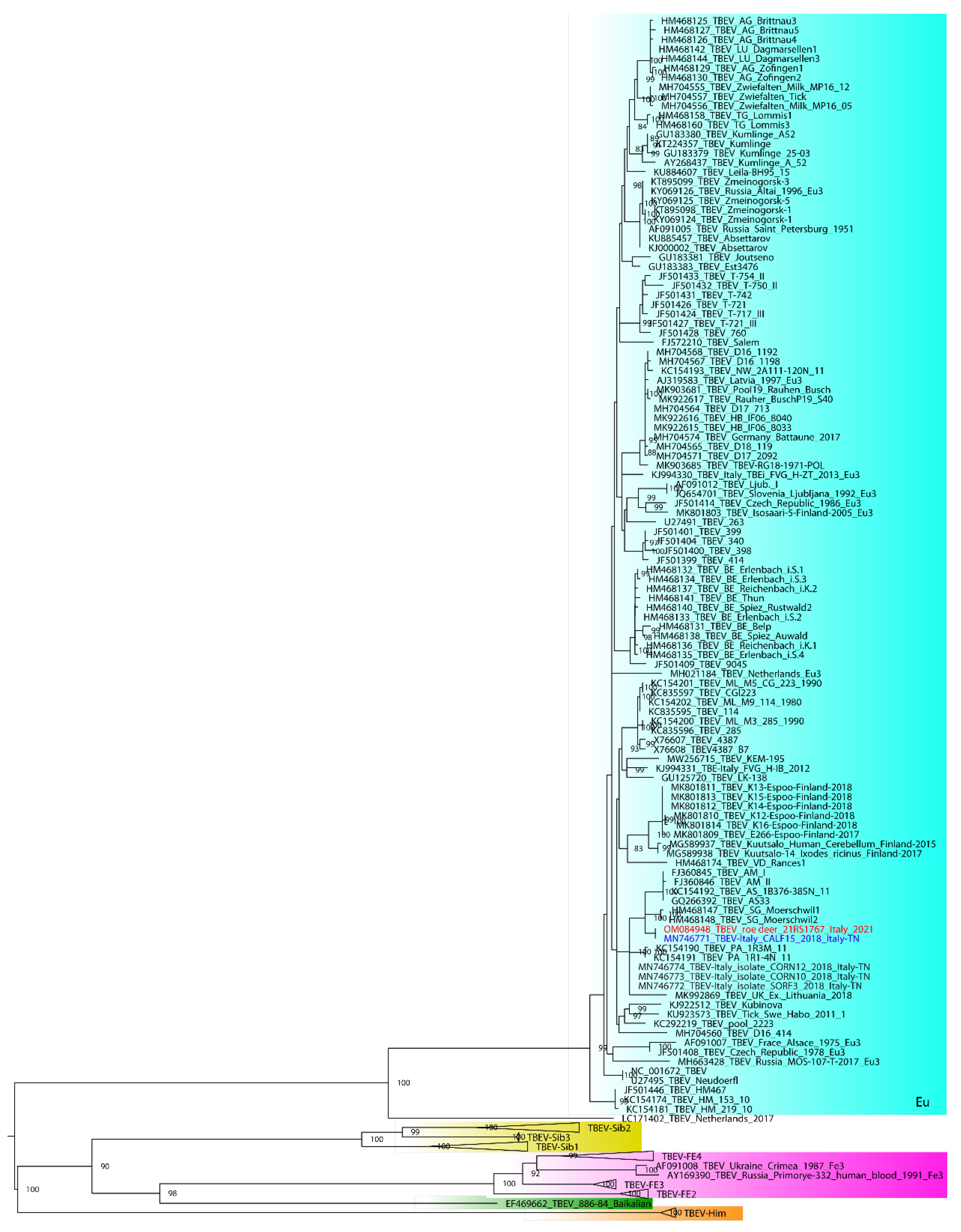
2.3. Illumina MiSeq Sequencing, Bioinformatics Analysis and Phylogeny
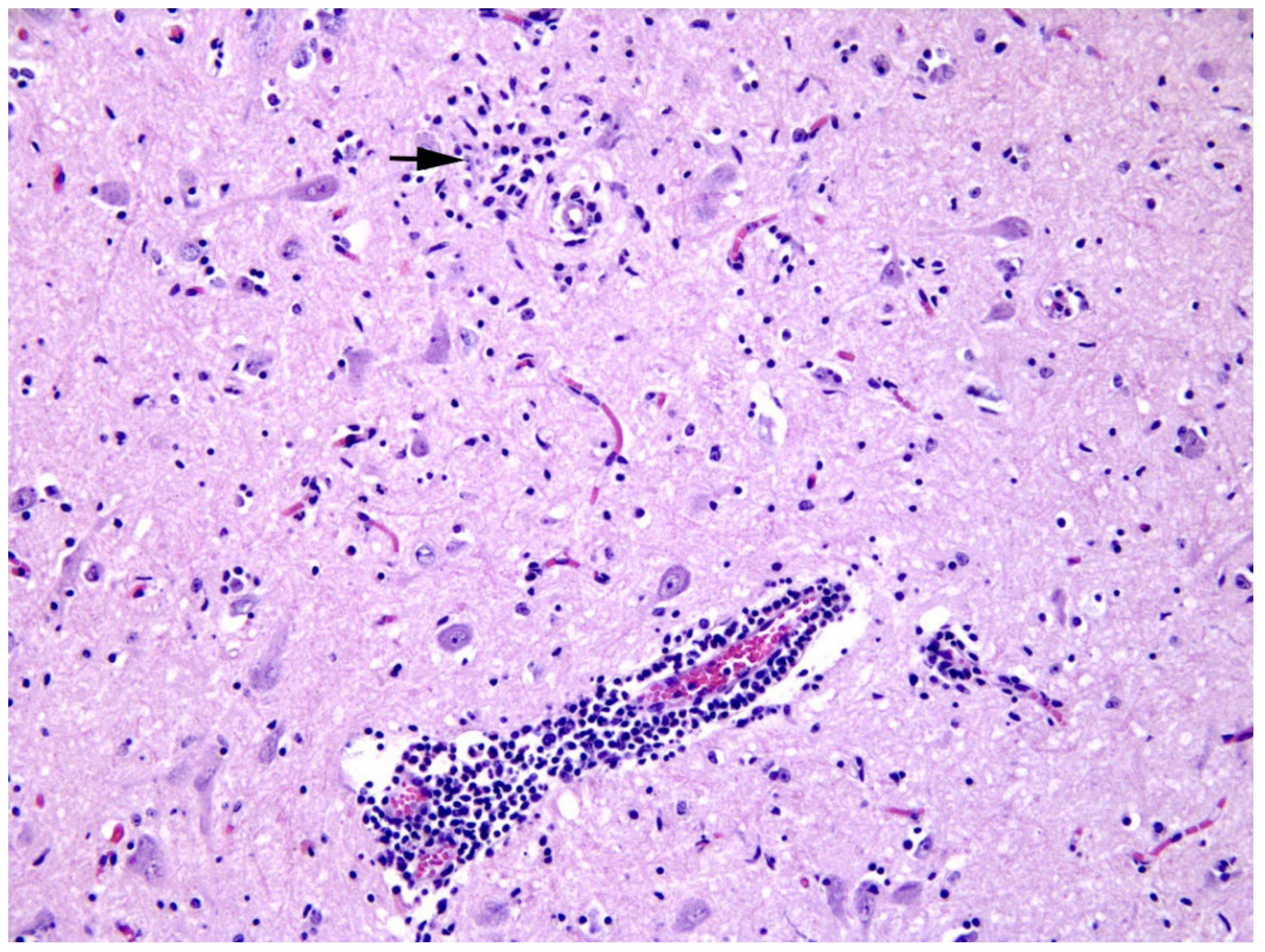
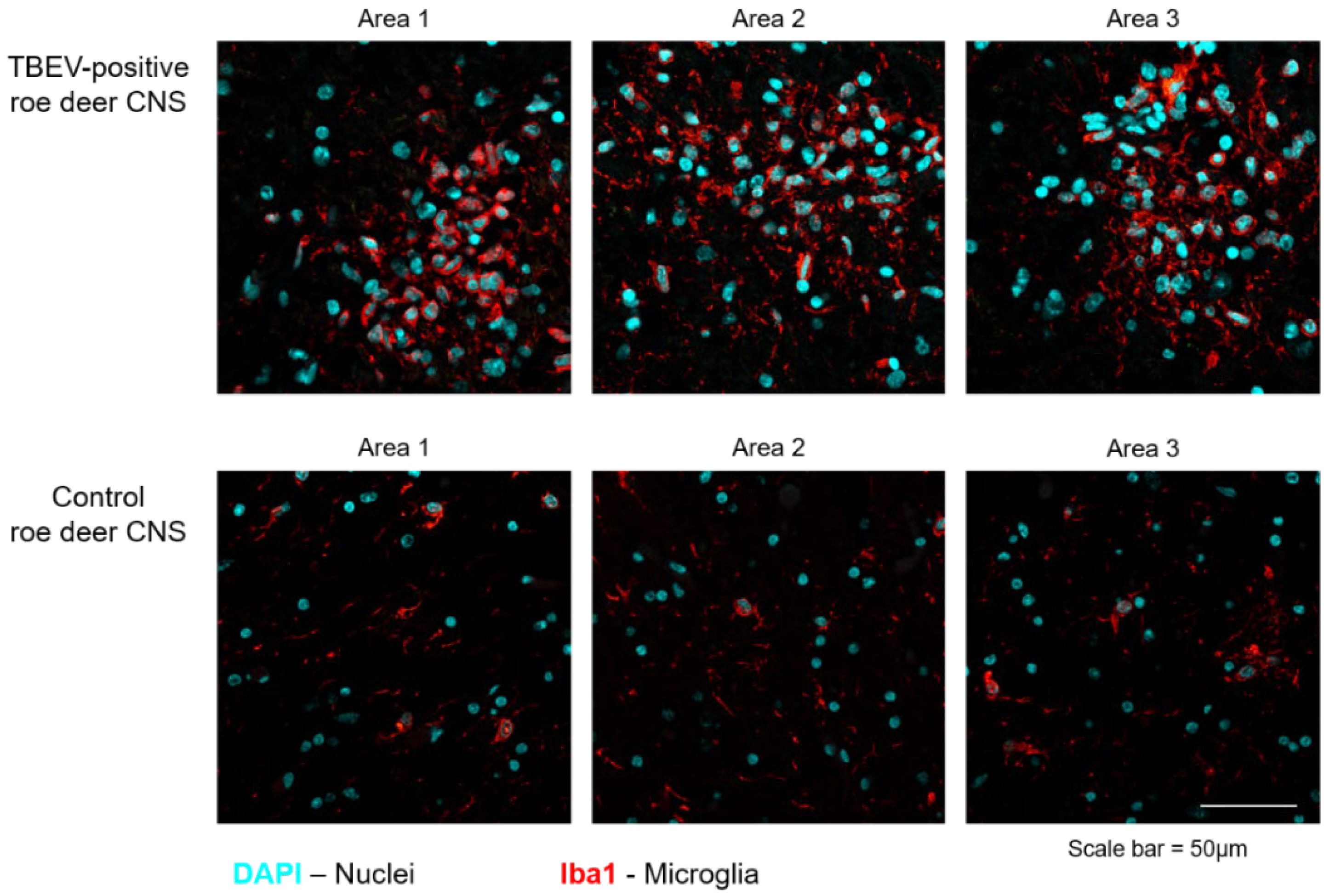
2.4. Histological Examination, Immunohistochemistry and Immunofluorescence
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
References
- Lindquist, L.; Vapalahti, O. Tick-borne encephalitis. Lancet 2008, 371, 1861–1871. [Google Scholar] [CrossRef]
- Kovalev, S.Y.; Mukhacheva, T.A. Reconsidering the classification of tick-borne encephalitis virus within the Siberian subtype gives new insights into its evolutionary history. Infect. Genet. Evol. 2017, 55, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Kellman, E.M.; Offerdahl, D.K.; Melik, W.; Bloom, M.E. Viral Determinants of Virulence in Tick-Borne Flaviviruses. Viruses 2018, 10, 329. [Google Scholar] [CrossRef] [PubMed]
- Gritsun, T.S.; Lashkevich, V.A.; Gould, E.A. Tick-borne encephalitis. Antivir. Res. 2003, 57, 129–146. [Google Scholar] [CrossRef]
- Ruzek, D.; Avšič Županc, T.; Borde, J.; Chrdle, A.; Eyer, L.; Karganova, G.; Kholodilov, I.; Knap, N.; Kozlovskaya, L.; Matveev, A.; et al. Tick-borne encephalitis in Europe and Russia: Review of pathogenesis, clinical features, therapy, and vaccines. Antivir. Res. 2019, 164, 23–51. [Google Scholar] [CrossRef]
- Süss, J. Epidemiology and ecology of TBE relevant to the production of effective vaccines. Vaccine 2003, 21, 19–35. [Google Scholar] [CrossRef]
- Cisak, E.; Wójcik-Fatla, A.; Zając, V.; Sroka, J.; Buczek, A.; Dutkiewicz, J. Prevalence of tick-borne encephalitis virus (TBEV) in samples of raw milk taken randomly from cows, goats and sheep in eastern Poland. Ann. Agric. Environ. Med. 2010, 17, 283–286. [Google Scholar]
- Nuttall, P.A.; Labuda, M. Dynamics of infection in tick vectors and at the tick-host interface. Adv. virus Res. 2003, 60, 232–272. [Google Scholar] [CrossRef]
- Labuda, M.; Kozuch, O.; Zuffová, E.; Elecková, E.; Hails, R.S.; Nuttall, P.A. Tick-Borne Encephalitis Virus transmission between Ticks Cofeeding on Specific Immune Natural Rodent Hosts. Virology 1997, 235, 138–143. [Google Scholar] [CrossRef]
- Labuda, M.; Randolph, S.E. Survival strategy of tick-borne encephalitis virus: Cellular basis and environmental determinants. Zent. Für Bakteriol. 1999, 289, 513–524. [Google Scholar] [CrossRef]
- Randolph, S.E.; Gern, L.; Nuttall, P.A. Co-feeding ticks: Epidemiological significance for tick-borne pathogen transmission. Parasitol. Today 1996, 12, 472–479. [Google Scholar] [CrossRef]
- Slovák, M.; Kazimírováa, M.; Siebenstichová, M.; Ustaníková, K.; Klempa, B.; Gritsun, T.; Gould, E.A.; Nuttall, P.A. Survival dynamics of tick-borne encephalitis virus in Ixodes ricinus ticks. Ticks Tick-Borne Dis. 2014, 5, 962–969. [Google Scholar] [CrossRef] [PubMed]
- Jaenson, T.G.T.; Petersson, E.H.; Jaenson, D.; Kindberg, J.; Pettersson, J.H.; Hjertqvist, M.; Medlock, J.M.; Bengtsson, H. The importance of wildlife in the ecology and epidemiology of the TBE virus in Sweden: Incidence of human TBE correlates with abundance of deer and hares. Parasites Vectors 2018, 11, 477. [Google Scholar] [CrossRef] [PubMed]
- Randolph, S.E.; Green, R.M.; Peacey, M.F.; Rogers, D.J. Seasonal synchrony: The key to tick-borne encephalitis foci identified by satellite data. Parasitology 2000, 121, 15–23. [Google Scholar] [CrossRef]
- Rosà, R.; Tagliapietra, V.; Manica, M.; Arnoldi, D.; Hauffe, H.C.; Rossi, C.; Rosso, F.; Henttonen, H.; Rizzoli, A. Changes in host densities and co-feeding pattern efficiently predict tick-borne encephalitis hazard in an endemic focus in northern Italy. Int. J. Parasitol. 2019, 49, 779–787. [Google Scholar] [CrossRef]
- Hekrlová, A.; Kubíček, O.; Lány, P.; Rosenbergová, K.; Schánilec, P. Tick-borne encephalitis in dogs: Application of "nested real-time RT-PCR" for intravital virus detection. Berl. Munch. Tierarztl. Wochenschr. 2015, 128, 397–401. [Google Scholar]
- Beckmann, K.; Oevermann, A.; Golini, L.; Steffen, F.; Kircher, P.R.; Carrera, I. MRI findings in a case of canine tick born meningoencephalomyelitis. Schweiz. Arch. Tierheilkd. 2014, 156, 395–399. [Google Scholar] [CrossRef]
- Pfeffer, M.; Dobler, G. Tick-borne encephalitis virus in dogs—Is this an issue? Parasit. Vectors 2011, 4, 59. [Google Scholar] [CrossRef]
- Leschnik, M.W.; Kirtz, G.C.; Thalhammer, J.G. Tick-borne encephalitis (TBE) in dogs. Int. J. Med. Microbiol. 2002, 291, 66–69. [Google Scholar] [CrossRef]
- Andersson, E.; Kendall, A.; Url, A.; Auer, A.; Leschnik, M. The first RT-qPCR confirmed case of tick-borne encephalitis in a dog in Scandinavia. Acta Vet. Scand. 2020, 62, 51. [Google Scholar] [CrossRef] [PubMed]
- Klaus, C.; Hörügel, U.; Hoffmann, B.; Beer, M. Tick-borne encephalitis virus (TBEV) infection in horses: Clinical and laboratory findings and epidemiological investigations. Vet. Microbiol. 2013, 163, 368–372. [Google Scholar] [CrossRef] [PubMed]
- Fouché, N.; Oesch, S.; Ziegler, U.; Gerber, V. Clinical presentation and laboratory diagnostic work-up of a horse with tick-borne encephalitis in Switzerland. Viruses 2021, 13, 1474. [Google Scholar] [CrossRef] [PubMed]
- Süss, J.; Gelpi, E.; Klaus, C.; Bagon, A.; Liebler-Tenorio, E.M.; Budka, H.; Stark, B.; Müller, W.; Hotzel, H. Tickborne encephalitis in naturally exposed monkey (Macaca sylvanus). Emerg. Infect. Dis. 2007, 13, 905–907. [Google Scholar] [CrossRef] [PubMed]
- Klaus, C.; Hoffmann, B.; Beer, M.; Müller, W.; Stark, B.; Bader, W.; Stiasny, K.; Heinz, F.X.; Süss, J. Seroprevalence of tick-borne encephalitis (TBE) in naturally exposed monkeys (Macaca sylvanus) and sheep and prevalence of TBE virus in ticks in a TBE endemic area in Germany. Ticks Tick-Borne Dis. 2010, 1, 141–144. [Google Scholar] [CrossRef] [PubMed]
- Böhm, B.; Schade, B.; Bauer, B.; Hoffmann, B.; Hoffmann, D.; Ziegler, U.; Beer, M.; Klaus, C.; Weissenböck, H.; Böttcher, J. Tick-borne encephalitis in a naturally infected sheep. BMC Vet. Res. 2017, 13, 267. [Google Scholar] [CrossRef] [PubMed]
- Zindel, W.; Wyler, R. Tick-borne encephalitis in a goat in lower Prätigau. Schweiz. Arch. Tierheilkd. 1983, 125, 383–386. [Google Scholar] [PubMed]
- Bagó, Z.; Bauder, B.; Kolodziejek, J.; Nowotny, N.; Weissenböck, H. Tickborne encephalitis in a mouflon (Ovis ammon musimon). Vet. Rec. 2002, 150, 218–220. [Google Scholar] [CrossRef] [PubMed]
- Salat, J.; Ruzek, D. Tick-borne encephalitis in domestic animals. Acta Virol. 2020, 64, 226–232. [Google Scholar] [CrossRef]
- Kiffner, C.; Lödige, C.; Alings, M.; Vor, T.; Rühe, F. Abundance estimation of Ixodes ticks (Acari: Ixodidae) on roe deer (Capreolus capreolus). Exp. Appl. Acarol. 2010, 52, 73–84. [Google Scholar] [CrossRef]
- Kriz, B.; Daniel, M.; Benes, C.; Maly, M. The role of game (wild boar and roe deer) in the spread of tick-borne encephalitis in the Czech Republic. Vector Borne Zoonotic Dis. 2014, 14, 801–807. [Google Scholar] [CrossRef]
- Gerth, H.J.; Grimshandl, D.; Stage, B.; Döller, G.; Kunz, C. Roe deer as sentinels for endemicity of tick-borne encephalitis virus. Epidemiol. Infect. 1995, 115, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Carpi, G.; Cagnacci, F.; Neteler, M.; Rizzoli, A. Tick infestation on roe deer in relation to geographic and remotely sensed climatic variables in a tick-borne encephalitis endemic area. Epidemiol. Infect. 2008, 136, 1416–1424. [Google Scholar] [CrossRef] [PubMed]
- Knap, N.; Avšič-Županc, T. Correlation of TBE incidence with red deer and roe deer abundance in Slovenia. PLoS ONE 2013, 8, 66380. [Google Scholar] [CrossRef] [PubMed]
- Pugliese, A.; Rosà, R. Effect of host populations on the intensity of ticks and the prevalence of tick-borne pathogens: How to interpret the results of deer exclosure experiments. Parasitology 2008, 135, 1531–1544. [Google Scholar] [CrossRef] [PubMed]
- Rizzoli, A.; Hauffe, H.C.; Tagliapietra, V.; Neteler, M.; Rosà, R. Forest structure and roe deer abundance predict tick-borne encephalitis risk in Italy. PLoS ONE 2009, 4, e4336. [Google Scholar] [CrossRef] [PubMed]
- Król, N.; Chitimia-Dobler, L.; Dobler, G.; Karliuk, Y.; Birka, S.; Obiegala, A.; Pfeffer, M. Tick burden on European roe deer (Capreolus capreolus) from Saxony, Germany, and detection of tick-borne encephalitis virus in attached ticks. Parasitol. Res. 2020, 119, 1387–1392. [Google Scholar] [CrossRef] [PubMed]
- De Benedictis, P.; De Battisti, C.; Dacheux, L.; Marciano, S.; Ormelli, S.; Salomoni, A.; Caenazzo, S.T.; Lepelletier, A.; Bourhy, H.; Capua, I.; et al. Lyssavirus detection and typing using pyrosequencing. J. Clin. Microbiol. 2011, 49, 1932–1938. [Google Scholar] [CrossRef]
- OIE manual rabies chapter 3.1.17. 2018. Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.01.17_RABIES.pdf (accessed on 7 November 2021).
- Schwaiger, M.; Cassinotti, P. Development of a quantitative real-time RT-PCR assay with internal control for the laboratory detection of tick-borne encephalitis virus (TBEV) RNA. J. Clin. Virol. 2003, 27, 136–145. [Google Scholar] [CrossRef]
- Da Rold, G.; Ravagnan, S.; Soppelsa, F.; Porcellato, E.; Soppelsa, M.; Obber, F.; Citterio, C.V.; Carlin, S.; Danesi, P.; Montarsi, F.; et al. Ticks are more suitable than red foxes for monitoring zoonotic tick-borne pathogens in northeastern Italy. Parasit. Vectors. 2018, 11, 137. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Dai, X.; Shang, G.; Lu, S.; Yang, J.; Xu, J. A new subtype of eastern tick-borne encephalitis virus discovered in Qinghai-Tibet Plateau, China. Emerg. Microbes Infect. 2018, 25, 74. [Google Scholar] [CrossRef] [PubMed]
- Ecker, M.; Allison, S.L.; Meixner, T.; Heinz, F.X. Sequence analysis and genetic classification of tick-borne encephalitis viruses from Europe and Asia. J. Gen. Virol. 1999, 80, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Kupča, A.M.; Essbauer, S.; Zoeller, G.; de Mendonça, P.G.; Brey, R.; Rinder, M.; Pfister, K.; Spiegel, M.; Doerrbecker, B.; Pfeffer, M.; et al. Isolation and molecular characterization of a tick-borne encephalitis virus strain from a new tick-borne encephalitis focus with severe cases in Bavaria, Germany. Ticks Tick-Borne Dis. 2010, 1, 44–51. [Google Scholar] [CrossRef]
- Rey, F.A.; Heinz, F.X.; Mandl, C.; Kunz, C.; Harrison, S.C. The envelope glycoprotein from tick-borne encephalitis virus at 2 A resolution. Nature 1995, 375, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Alfano, N.; Tagliapietra, V.; Rosso, F.; Ziegler, U.; Arnoldi, D.; Rizzoli, A. Tick-borne encephalitis foci in northeast Italy revealed by combined virus detection in ticks, serosurvey on goats and human cases. Emerg. microbes Infect. 2020, 9, 474–484. [Google Scholar] [CrossRef]
- Weissenböck, H.; Suchy, A.; Holzmann, H. Tick-borne encephalitis in dogs: Neuropathological findings and distribution of antigen. Acta Neuropathol. 1998, 95, 361–366. [Google Scholar] [CrossRef]
- Pulkkinen, L.; Butcher, S.J.; Anastasina, M. Tick-Borne Encephalitis Virus: A Structural View. Viruses 2018, 10, 350. [Google Scholar] [CrossRef]
- Rezza, G.; Farchi, F.; Pezzotti, P.; Pezzotti, P.; Ruscio, M.; Lo Presti, A.; Ciccozzi, M.; Mondardini, V.; Paternoster, C.; Bassetti, M.; et al. TBE Virology Group Tick-borne encephalitis in north-east Italy: A 14-year retrospective study, January 2000 to December 2013. EuroSurveillance 2015, 20, 4. [Google Scholar] [CrossRef]
- Jaenson, T.G.; Hjertqvist, M.; Bergström, T.; Lundkvist, A. Why is tick-borne encephalitis increasing? A review of the key factors causing the increasing incidence of human TBE in Sweden. Parasit. Vectors 2012, 5, 184. [Google Scholar] [CrossRef]
- Paulsen, K.M.; Das Neves, C.G.; Granquist, E.G.; Madslien, K.; Stuen, S.; Pedersen, B.N.; Vikse, R.; Rocchi, M.; Laming, E.; Stiasny, K.; et al. Cervids as sentinel-species for tick-borne encephalitis virus in Norway—A serological study. Zoonoses Public Health 2020, 67, 342–351. [Google Scholar] [CrossRef]
- Opalińska, P.; Wierzbicka, A.; Asman, M.; Rączka, G.; Dyderski, M.K.; Nowak-Chmura, M. Fivefold higher abundance of ticks (Acari: Ixodida) on the European roe deer (Capreolus capreolus L.) forest than field ecotypes. Sci. Rep. 2021, 11, 10649. [Google Scholar] [CrossRef] [PubMed]
- Pautienius, A.; Armonaite, A.; Simkute, E.; Zagrabskaite, R.; Buitkuviene, J.; Alpizar-Jara, R.; Grigas, J.; Zakiene, I.; Zienius, D.; Salomskas, A.; et al. Cross-Sectional Study on the Prevalence and Factors Influencing Occurrence of Tick-Borne Encephalitis in Horses in Lithuania. Pathogens 2021, 10, 140. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Beier, S.; Flade, I.; Górska, A.; El-Hadidi, M.; Mitra, S.; Ruscheweyh, H.J.; Tappu, R. MEGAN Community Edition—Interactive Exploration and Analysis of Large-Scale Microbiome Sequencing Data. PLoS Comput. Biol. 2016, 12, e1004957. [Google Scholar] [CrossRef]
- Peng, Y.; Leung, H.C.; Yiu, S.M.; Chin, F.Y. IDBA-UD: A de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 2012, 28, 1420–1428. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Wilm, A.; Aw, P.P.; Bertrand, D.; Yeo, G.H.; Ong, S.H.; Wong, C.H.; Khor, C.C.; Petric, R.; Hibberd, M.L.; Nagarajan, N. LoFreq: A sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic Acids Res. 2012, 40, 11189–11201. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Milne, I.; Bayer, M.; Cardle, L.; Shaw, P.; Stephen, G.; Wright, F.; Marshall, D. Tablet—Next generation sequence assembly visualization. Bioinformatics 2010, 26, 401–402. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Da Rold, G.; Obber, F.; Monne, I.; Milani, A.; Ravagnan, S.; Toniolo, F.; Sgubin, S.; Zamperin, G.; Foiani, G.; Vascellari, M.; et al. Clinical Tick-Borne Encephalitis in a Roe Deer (Capreolus capreolus L.). Viruses 2022, 14, 300. https://doi.org/10.3390/v14020300
Da Rold G, Obber F, Monne I, Milani A, Ravagnan S, Toniolo F, Sgubin S, Zamperin G, Foiani G, Vascellari M, et al. Clinical Tick-Borne Encephalitis in a Roe Deer (Capreolus capreolus L.). Viruses. 2022; 14(2):300. https://doi.org/10.3390/v14020300
Chicago/Turabian StyleDa Rold, Graziana, Federica Obber, Isabella Monne, Adelaide Milani, Silvia Ravagnan, Federica Toniolo, Sofia Sgubin, Gianpiero Zamperin, Greta Foiani, Marta Vascellari, and et al. 2022. "Clinical Tick-Borne Encephalitis in a Roe Deer (Capreolus capreolus L.)" Viruses 14, no. 2: 300. https://doi.org/10.3390/v14020300
APA StyleDa Rold, G., Obber, F., Monne, I., Milani, A., Ravagnan, S., Toniolo, F., Sgubin, S., Zamperin, G., Foiani, G., Vascellari, M., Drzewniokova, P., Castellan, M., De Benedictis, P., & Citterio, C. V. (2022). Clinical Tick-Borne Encephalitis in a Roe Deer (Capreolus capreolus L.). Viruses, 14(2), 300. https://doi.org/10.3390/v14020300







