Performance Characteristics of Real-Time PCRs for African Swine Fever Virus Genome Detection—Comparison of Twelve Kits to an OIE-Recommended Method
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples
2.1.1. Field Samples
2.1.2. Animal Trial Samples to Represent Different Genome Loads
2.1.3. African Swine Fever Virus p72 Genotypes Samples
2.2. Viral DNA Extraction
2.3. Molecular Assays
2.4. Data Analysis
3. Results
3.1. Comparison of Kit Handling and Time Requirements
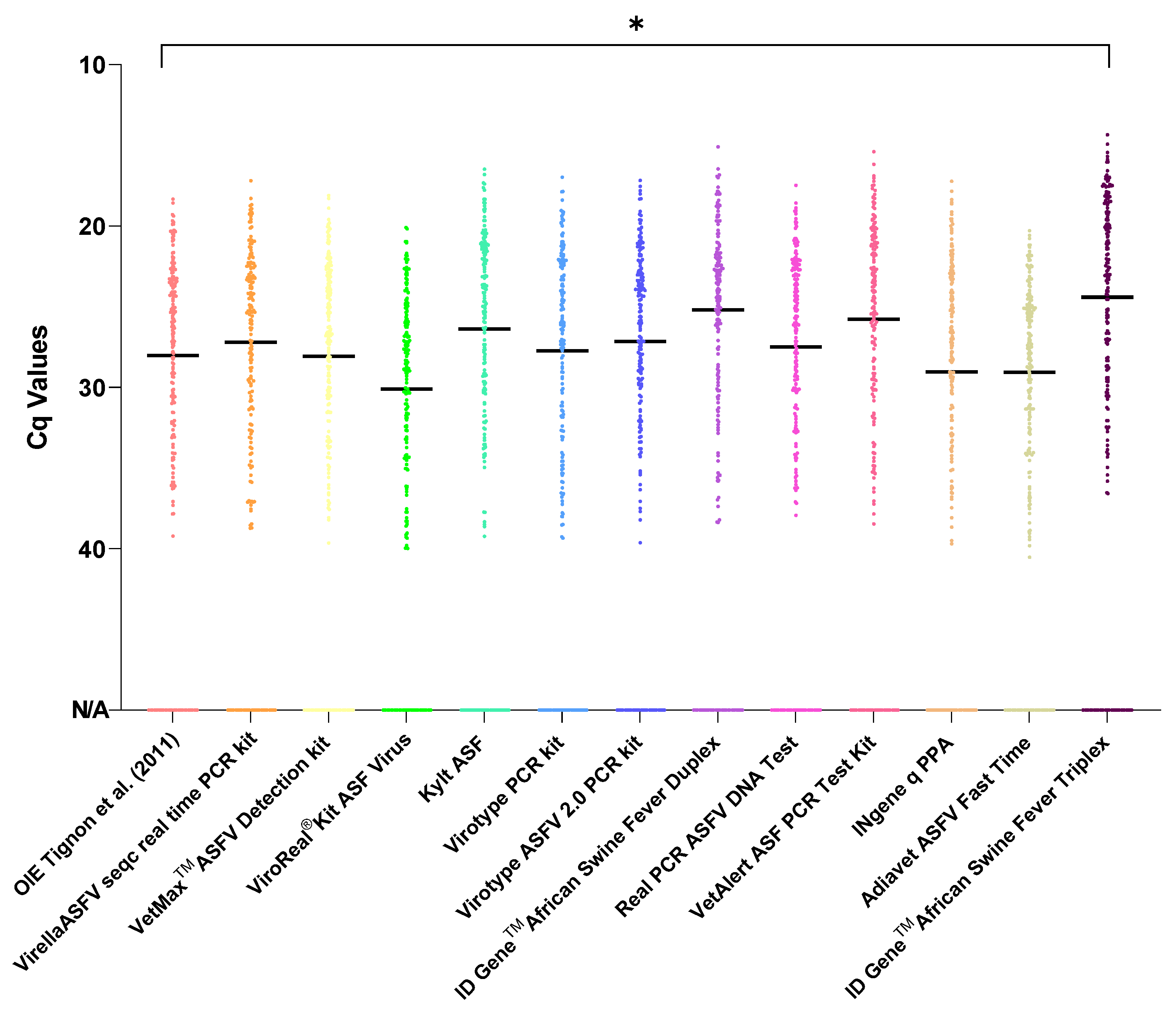
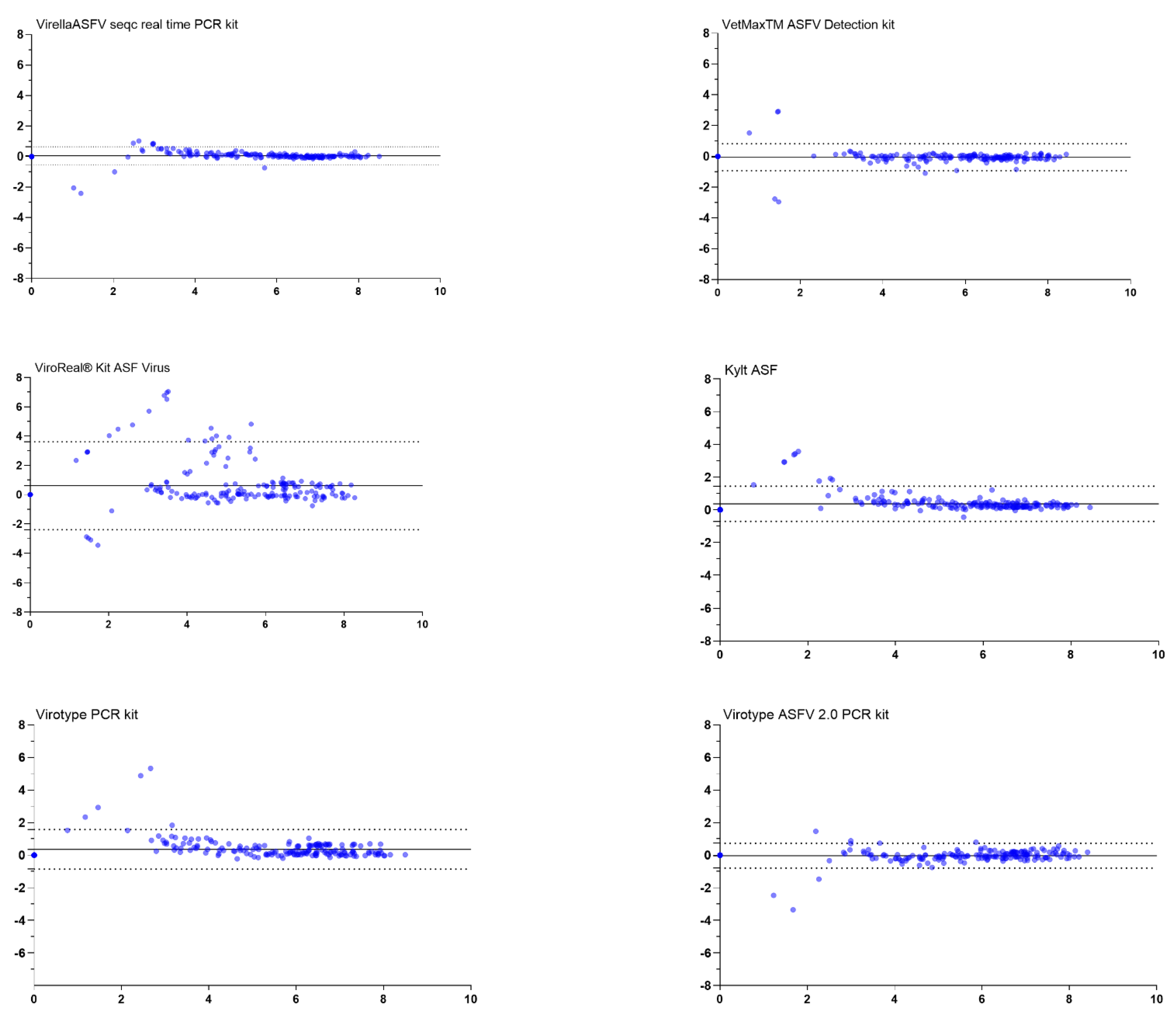
3.2. Evaluation of the Agreement for Viral Genome Detection
3.3. Evaluation of the Internal Control Systems
3.4. Sensitivity, Specificity and Precision
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alejo, A.; Matamoros, T.; Guerra, M.; Andrés, G. A Proteomic Atlas of the African Swine Fever Virus Particle. J. Virol. 2018, 92, e01293-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andrés, G.; Charro, D.; Matamoros, T.; Dillard, R.S.; Abrescia, N.G.A. The cryo-EM structure of African swine fever virus unravels a unique architecture comprising two icosahedral protein capsids and two lipoprotein membranes. J. Biol. Chem. 2020, 295, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gallardo, C.; Sánchez, E.G.; Pérez-Núñez, D.; Nogal, M.; de Leon, P.; Carrascosa, Á.L.; Nieto, R.; Soler, A.; Arias, M.L.; Revilla, Y. African swine fever virus (ASFV) protection mediated by NH/P68 and NH/P68 recombinant live-attenuated viruses. Vaccine 2018, 36, 2694–2704. [Google Scholar] [CrossRef] [PubMed]
- Galindo, I.; Alonso, C. African Swine Fever Virus: A Review. Viruses 2017, 9, 103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Achenbach, J.E.; Gallardo, C.; Pelegrin, E.N.; Rivera-Arroyo, B.; Degefa-Negi, T.; Arias, M.; Jenberie, S.; Mulisa, D.D.; Gizaw, D.; Gelaye, E.; et al. Identification of a New Genotype of African Swine Fever Virus in Domestic Pigs from Ethiopia. Transbound. Emerg. Dis. 2016, 64, 1393–1404. [Google Scholar] [CrossRef] [PubMed]
- Bastos, A.D.S.; Penrith, M.-L.; Crucière, C.; Edrich, J.L.; Hutchings, G.; Roger, F.; Couacy-Hymann, E.; R.Thomson, G. Genotyping field strains of African swine fever virus by partial p72 gene characterisation. Arch. Virol. 2003, 148, 693–706. [Google Scholar] [CrossRef]
- Blome, S.; Franzke, K.; Beer, M. African swine fever-A review of current knowledge. Virus Res. 2020, 287, 198099. [Google Scholar] [CrossRef]
- Boshoff, C.; Bastos, A.; Gerber, L.; Vosloo, W. Genetic characterisation of African swine fever viruses from outbreaks in southern Africa (1973–1999). Vet.-Microbiol. 2007, 121, 45–55. [Google Scholar] [CrossRef] [Green Version]
- Sauter-Louis, C.; Forth, J.H.; Probst, C.; Staubach, C.; Hlinak, A.; Rudovsky, A.; Holland, D.; Schlieben, P.; Göldner, M.; Schatz, J.; et al. Joining the club: First detection of African swine fever in wild boar in Germany. Transbound. Emerg. Dis. 2020, 68, 1744–1752. [Google Scholar] [CrossRef]
- Gallardo, C.; Nieto, R.; Soler, A.; Pelayo, V.; Fernandezpinero, J.; Markowska-Daniel, I.; Pridotkas, G.; Nurmoja, I.; Granta, R.; Simon, A.O.; et al. Assessment of African Swine Fever Diagnostic Techniques as a Response to the Epidemic Outbreaks in Eastern European Union Countries: How To Improve Surveillance and Control Programs. J. Clin. Microbiol. 2015, 53, 2555–2565. [Google Scholar] [CrossRef] [Green Version]
- Tignon, M.; Gallardo, C.; Iscaro, C.; Hutet, E.; Van der Stede, Y.; Kolbasov, D.; De Mia, G.M.; Le Potier, M.-F.; Bishop, R.P.; Arias, M.; et al. Development and inter-laboratory validation study of an improved new real-time PCR assay with internal control for detection and laboratory diagnosis of African swine fever virus. J. Virol. Methods 2011, 178, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Pinero, J.; Gallardo, C.; Elizalde, M.; Robles, A.; Gómez, C.; Bishop, R.; Heath, L.; Couacy-Hymann, E.; Fasina, F.O.; Pelayo, V.; et al. Molecular Diagnosis of African Swine Fever by a New Real-Time PCR Using Universal Probe Library. Transbound. Emerg. Dis. 2012, 60, 48–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- King, D.P.; Reid, S.M.; Hutchings, G.H.; Grierson, S.S.; Wilkinson, P.J.; Dixon, L.K.; Bastos, A.D.; Drew, T.W. Development of a TaqMan® PCR assay with internal amplification control for the detection of African swine fever virus. J. Virol. Methods 2002, 107, 53–61. [Google Scholar] [CrossRef]
- Agüero, M.; Fernández, J.; Romero, L.; Mascaraque, C.S.; Arias, M.; Sánchez-Vizcaíno, J.M. Highly Sensitive PCR Assay for Routine Diagnosis of African Swine Fever Virus in Clinical Samples. J. Clin. Microbiol. 2003, 41, 4431–4434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zani, L.; Forth, J.H.; Forth, L.; Nurmoja, I.; Leidenberger, S.; Henke, J.; Carlson, J.; Breidenstein, C.; Viltrop, A.; Höper, D.; et al. Deletion at the 5′-end of Estonian ASFV strains associated with an attenuated phenotype. Sci. Rep. 2018, 8, 6510. [Google Scholar] [CrossRef]
- Sehl, J.; Pikalo, J.; Schäfer, A.; Franzke, K.; Pannhorst, K.; Elnagar, A.; Blohm, U.; Blome, S.; Breithaupt, A. Comparative Pathology of Domestic Pigs and Wild Boar Infected with the Moderately Virulent African Swine Fever Virus Strain “Estonia 2014”. Pathogens 2020, 9, 662. [Google Scholar] [CrossRef]
- Pietschmann, J.; Guinat, C.; Beer, M.; Pronin, V.; Tauscher, K.; Petrov, A.; Keil, G.; Blome, S. Course and transmission characteristics of oral low-dose infection of domestic pigs and European wild boar with a Caucasian African swine fever virus isolate. Arch. Virol. 2015, 160, 1657–1667. [Google Scholar] [CrossRef]
- Hübner, A.; Petersen, B.; Keil, G.M.; Niemann, H.; Mettenleiter, T.C.; Fuchs, W. Efficient inhibition of African swine fever virus replication by CRISPR/Cas9 targeting of the viral p30 gene (CP204L). Sci. Rep. 2018, 8, 1449. [Google Scholar] [CrossRef] [Green Version]
- Bland, J.M.; Altman, D.G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1986, 1, 307–310. [Google Scholar] [CrossRef]
- Greiner, M.; Gardner, I.A. Epidemiologic issues in the validation of veterinary diagnostic tests. Prev. Vet. Med. 2000, 45, 3–22. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, L.; Noll, L.; Stoy, C.; Porter, E.; Fu, J.; Feng, Y.; Peddireddi, L.; Liu, X.; Dodd, K.A.; et al. Development of a real-time PCR assay for detection of African swine fever virus with an endogenous internal control. Transbound. Emerg. Dis. 2020, 67, 2446–2454. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, B.; Depner, K.; Schirrmeier, H.; Beer, M. A universal heterologous internal control system for duplex real-time RT-PCR assays used in a detection system for pestiviruses. J. Virol. Methods 2006, 136, 200–209. [Google Scholar] [CrossRef] [PubMed]
- Schoder, M.-E.; Tignon, M.; Linden, A.; Vervaeke, M.; Cay, A.B. Evaluation of seven commercial African swine fever virus detection kits and three Taq polymerases on 300 well-characterized field samples. J. Virol. Methods 2020, 280, 113874. [Google Scholar] [CrossRef] [PubMed]


| Protocol/ PCR Kit. | Internal Control | Volume of DNA Eluates per Test (µL) | Cycles | Estimated Run Time (CFX 96) | Pipetting Steps |
|---|---|---|---|---|---|
| OIE Tignon et al. (2011) | Endogenous (β-Actin) | 5 | 45 | 2 h 25 min * | 5 |
| Virella ASFV seqc real-time PCR kit (Gerbion, Kornwestheim, Germany) | Exogenous and endogenous (Succinate-Dehydrogenase) | 6 | 45 | 2 h 18 min | 2 |
| VetMaxTM ASFV Detection kit (Thermo Fisher Scientific, Lissieu, France) | Exogenous | 5 | 45 | 1 h 48 min | 2 |
| ViroReal® Kit ASF Virus (Ingenetix, Vienna, Austria) | Exogenous | 5 (1–8) | 45 | 1 h 31 min | 5 |
| Kylt ASF (AniCon Labor GmbH, Höltinghausen, Germany) | Endogenous (β-Actin) | 4 | 42 | 1 h 40 min | 2 |
| Virotype ASFV PCR kit (Indical, Leipzig, Germany) | Endogenous (β-Actin) | 5 | 40 | 1 h 43 min | 2 |
| Virotype ASFV 2.0 PCR kit (Indical, Leipzig, Germany) | Exogenous and Endogenous (β-Actin) | 5 | 40 | 1 h 2 min | 2 |
| ID GeneTM African Swine Fever Duplex (Innovative Diagnostics, Grabels, France) | Endogenous (housekeeping gene) | 5 | 40 | 1 h 38 min | 2 |
| Real PCR ASFV DNA Test (IDEXX, Hoofddorp, Netherlands) | Endogenous (swine DNA) | 5 | 45 | 1 h 30 min | 4 |
| VetAlert ASF PCR Test Kit (Tetracore, Rockville, USA), not yet approved in Germany | Exogenous | 5 | 45 | 1 h 36 min | 3 |
| INgene q PPA (Ingenasa, Madrid, Spain) | Exogenous | 2 | 45 | 1 h 16 min | 4 |
| Adiavet ASFV Fast Time (Bio-X Diagnostics, Rochefort, Belgium) | Endogenous (RNase P) | 5 | 45 | 1 h 8 min | 2 |
| ID GeneTM African Swine Fever Triplex (Innovative Diagnostics, Grabels, France), not yet approved in Germany | Exogenous and endogenous | 5 | 40 | 1 h 4 min | 2 |
| Kit Name | Se% | 95% CI Min | 95% CI Max | Sp% | 95% CI Min | 95% CI Max |
|---|---|---|---|---|---|---|
| OIE Tignon et al. (2011) | 98.32 | 95.26 | 99.66 | 100 | 87.66 | 100 |
| Virella ASFV seqc real-time PCR kit | 100 | 97.96 | 100 | 100 | 87.66 | 100 |
| VetMaxTM ASFV Detection kit | 98.32 | 95.26 | 99.66 | 100 | 87.66 | 100 |
| ViroReal® Kit ASF Virus | 99.44 | 96.94 | 99.99 | 100 | 87.66 | 100 |
| Kylt ASF | 95.53 | 91.74 | 98.14 | 100 | 87.66 | 100 |
| Virotype ASFV PCR kit | 94.97 | 91.11 | 97.79 | 100 | 87.66 | 100 |
| Virotype ASFV 2.0 PCR kit | 100 | 97.96 | 100 | 100 | 87.66 | 100 |
| ID GeneTM African Swine Fever Duplex | 97.21 | 93.77 | 99.11 | 100 | 87.66 | 100 |
| Real PCR ASFV DNA Test | 98.88 | 96.07 | 99.87 | 100 | 87.66 | 100 |
| VetAlert ASF PCR Test Kit | 100 | 97.96 | 100 | 100 | 87.66 | 100 |
| INgene q PPA | 98.32 | 95.26 | 99.66 | 100 | 87.66 | 100 |
| Adiavet ASFV Fast Time | 100 | 97.96 | 100 | 100 | 87.66 | 100 |
| ID GeneTM African Swine Fever Triplex | 97.77 | 94.5 | 99.4 | 100 | 87.66 | 100 |
| Average | 98.16 | 95.25 | 99.1 | 100 | 87.66 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pikalo, J.; Carrau, T.; Deutschmann, P.; Fischer, M.; Schlottau, K.; Beer, M.; Blome, S. Performance Characteristics of Real-Time PCRs for African Swine Fever Virus Genome Detection—Comparison of Twelve Kits to an OIE-Recommended Method. Viruses 2022, 14, 220. https://doi.org/10.3390/v14020220
Pikalo J, Carrau T, Deutschmann P, Fischer M, Schlottau K, Beer M, Blome S. Performance Characteristics of Real-Time PCRs for African Swine Fever Virus Genome Detection—Comparison of Twelve Kits to an OIE-Recommended Method. Viruses. 2022; 14(2):220. https://doi.org/10.3390/v14020220
Chicago/Turabian StylePikalo, Jutta, Tessa Carrau, Paul Deutschmann, Melina Fischer, Kore Schlottau, Martin Beer, and Sandra Blome. 2022. "Performance Characteristics of Real-Time PCRs for African Swine Fever Virus Genome Detection—Comparison of Twelve Kits to an OIE-Recommended Method" Viruses 14, no. 2: 220. https://doi.org/10.3390/v14020220
APA StylePikalo, J., Carrau, T., Deutschmann, P., Fischer, M., Schlottau, K., Beer, M., & Blome, S. (2022). Performance Characteristics of Real-Time PCRs for African Swine Fever Virus Genome Detection—Comparison of Twelve Kits to an OIE-Recommended Method. Viruses, 14(2), 220. https://doi.org/10.3390/v14020220








