Understanding the Genetic Diversity of Picobirnavirus: A Classification Update Based on Phylogenetic and Pairwise Sequence Comparison Approaches
Abstract
:1. Introduction
2. Materials and Methods
2.1. Retrieval of Picobirnavirus Sequences
2.2. Evaluation of the RdRp-200 Phylogenetic Marker
2.3. Multiple Sequence Alignment (MSA) and Evaluation of the MSA Accuracy
2.4. Phylogenetic Tree Reconstruction
2.5. Assessing the Reliability of the Phylogenetic Trees by the Comparison of Topologies
2.6. Taxonomical Demarcation Analysis Using Pairwise Sequence Comparison (PASC) and Sequence Demarcation Tool (SDT)
2.7. Visualization of Phylogenetic Agreement between RdRp and Capsid and Taxon Sampling Evaluation
3. Results
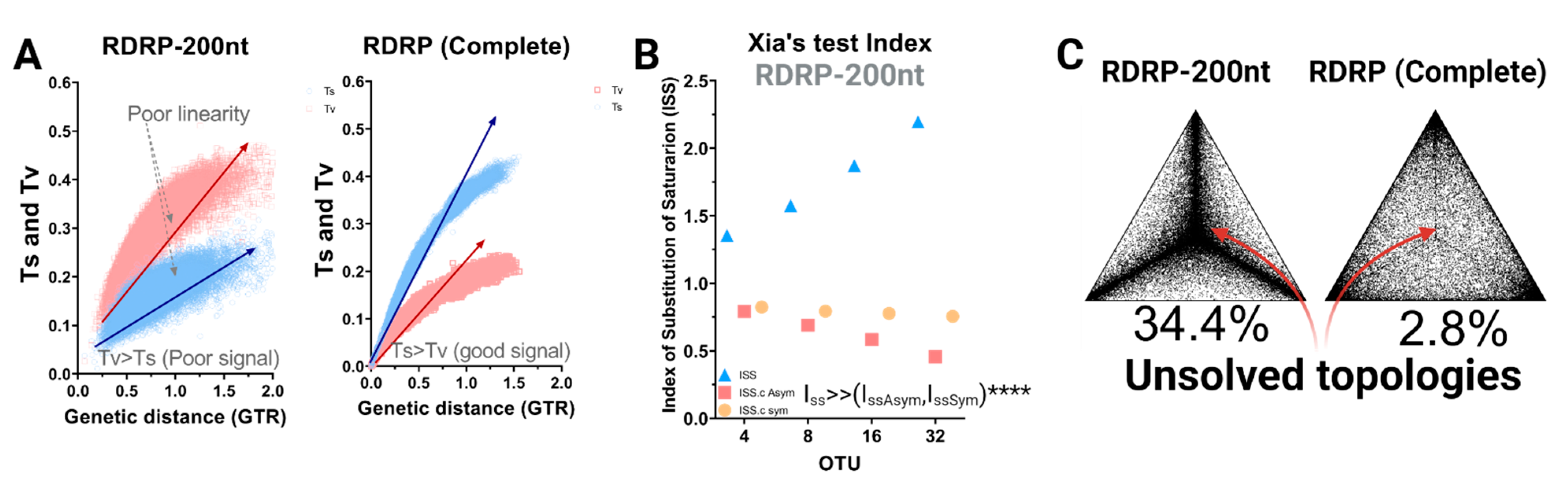
3.1. RdRp-200 Marker Is Saturated and Yields Unresolved Topologies
3.2. Complete Resolution of Taxa Requires Proper Selection of Alignment and Phylogenetic Reconstruction Methods
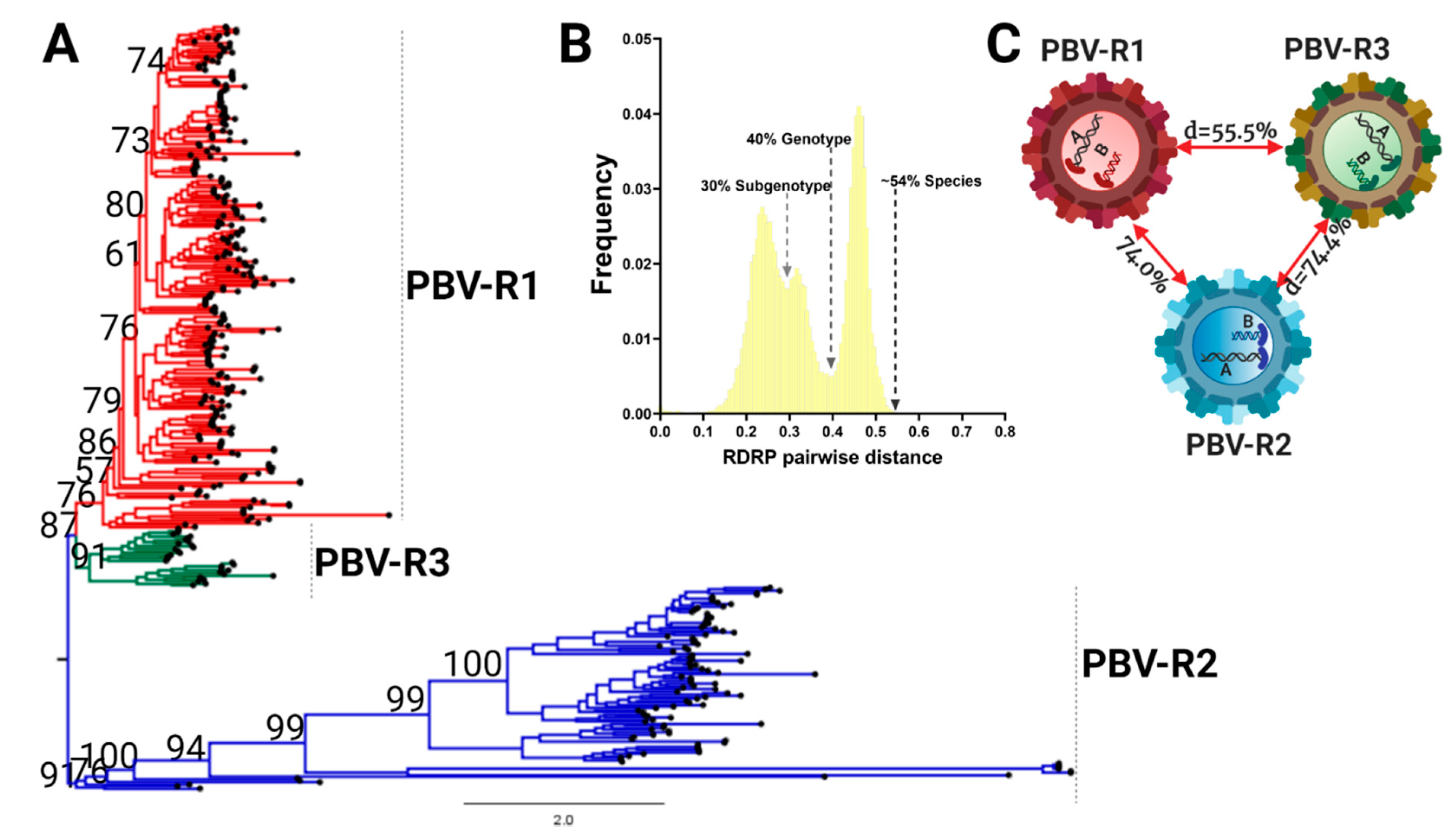
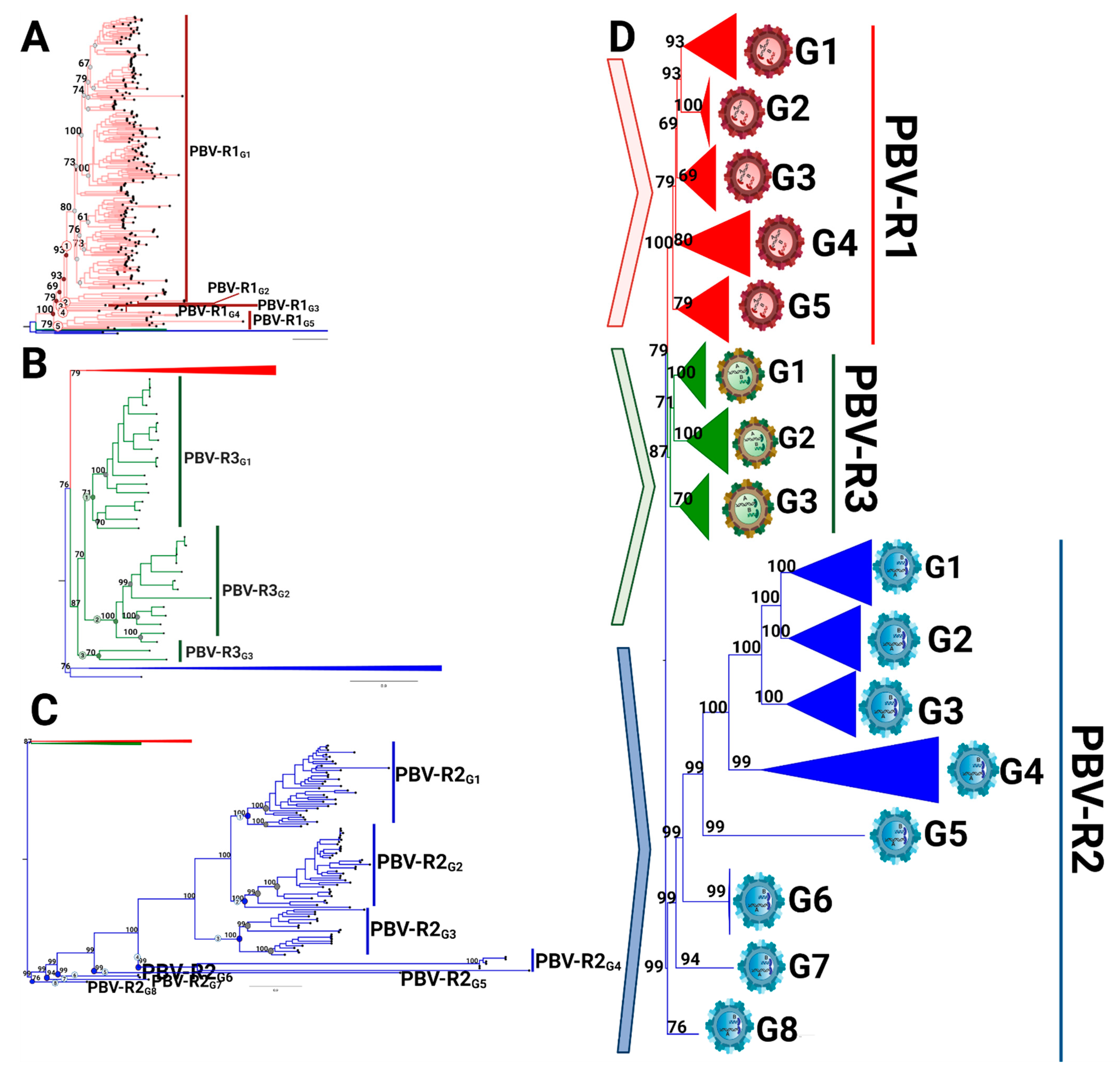
3.3. RdRp Lineages Delimit Three Distinctive Species for the Picobirnavirus Genus
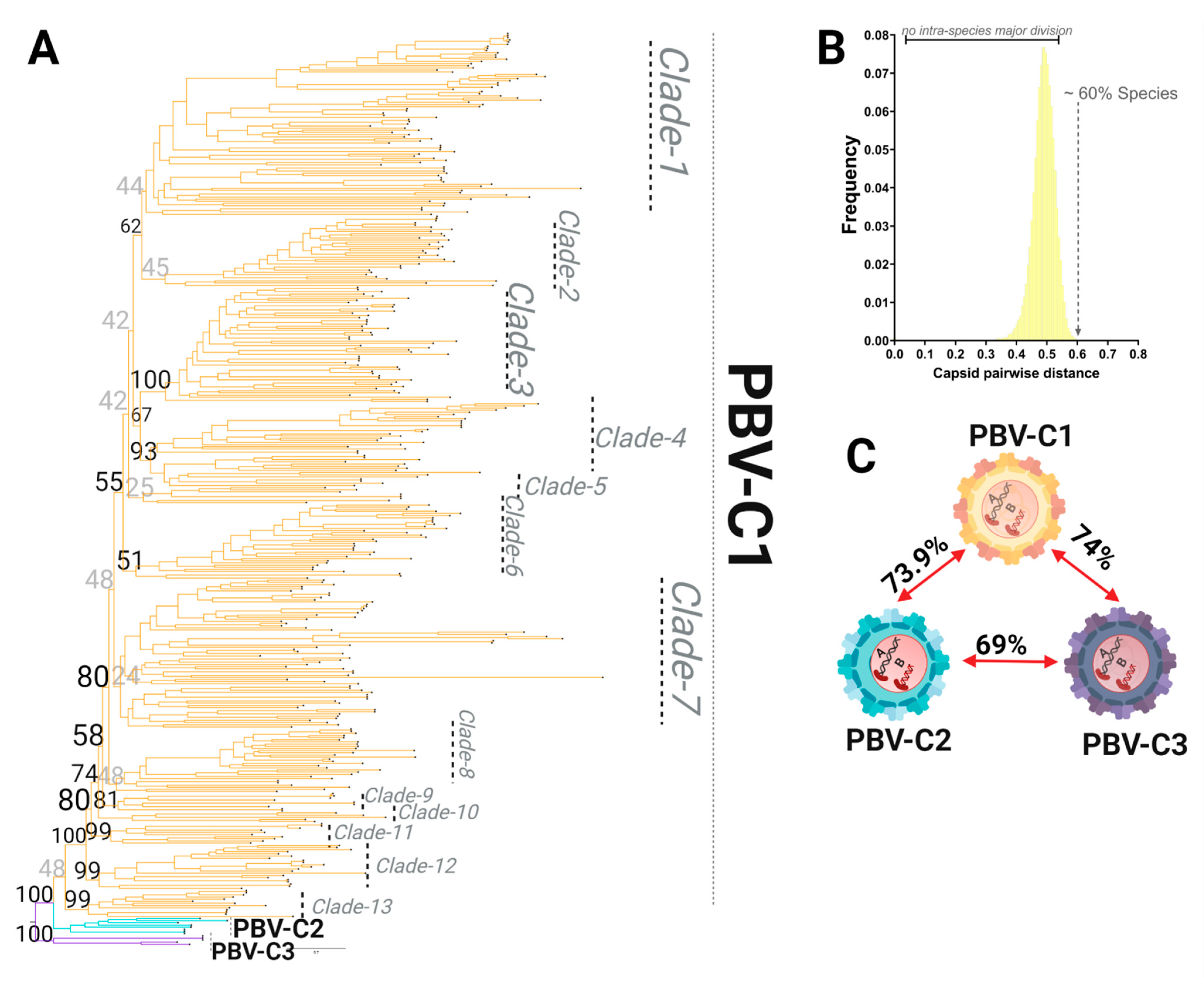
3.4. Capsid Lineages Also Diverged into Three Distinct Species
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Haga, I.R.; Martins, S.S.; Hosomi, S.T.; Vicentini, F.; Tanaka, H.; Gatti, M.S. Identification of a bisegmented double-stranded RNA virus (Picobirnavirus) in faeces of giant anteaters (Myrmecophaga tridactyla). Vet. J. 1999, 158, 234–236. [Google Scholar] [CrossRef]
- Masachessi, G.; Martinez, L.C.; Giordano, M.O.; Barril, P.A.; Isa, B.M.; Ferreyra, L.; Villareal, D.; Carello, M.; Asis, C.; Nates, S.V. Picobirnavirus (PBV) natural hosts in captivity and virus excretion pattern in infected animals. Arch. Virol. 2007, 152, 989–998. [Google Scholar] [CrossRef]
- Woo, P.C.Y.; Teng, J.L.L.; Bai, R.; Tang, Y.; Wong, A.Y.P.; Li, K.S.M.; Lam, C.S.F.; Fan, R.Y.Y.; Lau, S.K.P.; Yuen, K.Y. Novel Picobirnaviruses in Respiratory and Alimentary Tracts of Cattle and Monkeys with Large Intra- and Inter-Host Diversity. Viruses 2019, 11, 574. [Google Scholar] [CrossRef] [Green Version]
- Woo, P.C.; Teng, J.L.; Bai, R.; Wong, A.Y.; Martelli, P.; Hui, S.W.; Tsang, A.K.; Lau, C.C.; Ahmed, S.S.; Yip, C.C.; et al. High Diversity of Genogroup I Picobirnaviruses in Mammals. Front. Microbiol. 2016, 7, 1886. [Google Scholar] [CrossRef] [PubMed]
- Smits, S.L.; van Leeuwen, M.; Schapendonk, C.M.; Schurch, A.C.; Bodewes, R.; Haagmans, B.L.; Osterhaus, A.D. Picobirnaviruses in the human respiratory tract. Emerg. Infect. Dis. 2012, 18, 1539–1540. [Google Scholar] [CrossRef] [PubMed]
- Smits, S.L.; Poon, L.L.; van Leeuwen, M.; Lau, P.N.; Perera, H.K.; Peiris, J.S.; Simon, J.H.; Osterhaus, A.D. Genogroup I and II picobirnaviruses in respiratory tracts of pigs. Emerg. Infect. Dis. 2011, 17, 2328–2330. [Google Scholar] [CrossRef] [PubMed]
- Cummings, M.J.; Tokarz, R.; Bakamutumaho, B.; Kayiwa, J.; Byaruhanga, T.; Owor, N.; Namagambo, B.; Wolf, A.; Mathema, B.; Lutwama, J.J.; et al. Precision surveillance for viral respiratory pathogens: Virome capture sequencing for the detection and genomic characterization of severe acute respiratory infection in Uganda. Clin. Infect. Dis. 2019, 68, 1118–1125. [Google Scholar] [CrossRef] [PubMed]
- Kashnikov, A.Y.; Epifanova, N.V.; Novikova, N.A. Picobirnaviruses: Prevalence, genetic diversity, detection methods. Vavilov J. Genet. Breed. 2020, 24, 661–672. [Google Scholar] [CrossRef]
- Krishnamurthy, S.R.; Wang, D. Extensive conservation of prokaryotic ribosomal binding sites in known and novel picobirnaviruses. Virology 2018, 516, 108–114. [Google Scholar] [CrossRef]
- Kleymann, A.; Becker, A.A.M.J.; Malik, Y.S.; Kobayashi, N.; Ghosh, S. Detection and Molecular Characterization of Picobirnaviruses (PBVs) in the Mongoose: Identification of a Novel PBV Using an Alternative Genetic Code. Viruses 2020, 12, 99. [Google Scholar] [CrossRef] [Green Version]
- Ghosh, S.; Malik, Y.S. The True Host/s of Picobirnaviruses. Front. Vet. Sci. 2021, 7, 615293. [Google Scholar] [CrossRef] [PubMed]
- Fregolente, M.C.; Gatti, M.S. Nomenclature proposal for picobirnavirus. Arch. Virol. 2009, 154, 1953–1954. [Google Scholar] [CrossRef]
- Rosen, B.I.; Fang, Z.Y.; Glass, R.I.; Monroe, S.S. Cloning of human picobirnavirus genomic segments and development of an RT-PCR detection assay. Virology 2000, 277, 316–329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gallagher, C.A.; Navarro, R.; Cruz, K.; Aung, M.S.; Ng, A.; Bajak, E.; Beierschmitt, A.; Lawrence, M.; Dore, K.M.; Ketzis, J.; et al. Detection of picobirnaviruses in vervet monkeys (Chlorocebus sabaeus): Molecular characterization of complete genomic segment-2. Virus Res. 2017, 230, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Malik, Y.S.; Kumar, N.; Sharma, K.; Dhama, K.; Shabbir, M.Z.; Ganesh, B.; Kobayashi, N.; Banyai, K. Epidemiology, phylogeny, and evolution of emerging enteric Picobirnaviruses of animal origin and their relationship to human strains. BioMed Res. Int. 2014, 2014, 780752. [Google Scholar] [CrossRef] [Green Version]
- Knox, M.A.; Gedye, K.R.; Hayman, D.T.S. The Challenges of Analysing Highly Diverse Picobirnavirus Sequence Data. Viruses 2018, 10, 685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pikula, A.; Smietanka, K.; Perez, L.J. Emergence and expansion of novel pathogenic reassortant strains of infectious bursal disease virus causing acute outbreaks of the disease in Europe. Transbound. Emerg. Dis. 2020, 67, 1739–1744. [Google Scholar] [CrossRef]
- De la Cruz, L.; Barrera, M.; Rios, L.; Corona-Gonzalez, B.; Bulnes, C.A.; Diaz-Sanchez, A.A.; Agüero, J.A.; Lobo-Rivero, E.; Perez, L.J. Unraveling the Global Phylodynamic and Phylogeographic Expansion of Mycoplasma gallisepticum: Understanding the Origin and Expansion of This Pathogen in Ecuador. Pathogens 2020, 9, 674. [Google Scholar] [CrossRef]
- Xia, X.; Xie, Z. DAMBE: Software package for data analysis in molecular biology and evolution. J. Hered. 2001, 92, 371–373. [Google Scholar] [CrossRef] [Green Version]
- Xia, X.; Xie, Z.; Salemi, M.; Chen, L.; Wang, Y. An index of substitution saturation and its application. Mol. Phylogenet. Evol. 2003, 26, 1–7. [Google Scholar] [CrossRef]
- Alfonso-Morales, A.; Rios, L.; Martinez-Perez, O.; Dolz, R.; Valle, R.; Perera, C.L.; Bertran, K.; Frias, M.T.; Ganges, L.; Diaz de Arce, H.; et al. Evaluation of a Phylogenetic Marker Based on Genomic Segment B of Infectious Bursal Disease Virus: Facilitating a Feasible Incorporation of this Segment to the Molecular Epidemiology Studies for this Viral Agent. PLoS ONE 2015, 10, e0125853. [Google Scholar] [CrossRef] [PubMed]
- Rios, L.; Coronado, L.; Naranjo-Feliciano, D.; Martinez-Perez, O.; Perera, C.L.; Hernandez-Alvarez, L.; Diaz de Arce, H.; Nunez, J.I.; Ganges, L.; Perez, L.J. Deciphering the emergence, genetic diversity and evolution of classical swine fever virus. Sci. Rep. 2017, 7, 17887. [Google Scholar] [CrossRef] [Green Version]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Mirarab, S.; Warnow, T. FastSP: Linear time calculation of alignment accuracy. Bioinformatics 2011, 27, 3250–3258. [Google Scholar] [CrossRef] [PubMed]
- Nixon, K.C. Phylogeny. In Encyclopedia of Biodiversity, 2nd ed.; Levin, S.A., Ed.; Academic Press: Waltham, MA, USA, 2001; pp. 16–23. [Google Scholar]
- Dhar, A.; Minin, V.N. Maximum Likelihood Phylogenetic Inference. In Encyclopedia of Evolutionary Biology; Kliman, R.M., Ed.; Academic Press: Oxford, UK, 2016; pp. 499–506. [Google Scholar]
- Nascimento, F.F.; Reis, M.D.; Yang, Z. A biologist’s guide to Bayesian phylogenetic analysis. Nat. Ecol. Evol. 2017, 1, 1446–1454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perez, L.J.; de Arce, H.D.; Cortey, M.; Dominguez, P.; Percedo, M.I.; Perera, C.L.; Tarradas, J.; Frias, M.T.; Segales, J.; Ganges, L.; et al. Phylogenetic networks to study the origin and evolution of porcine circovirus type 2 (PCV2) in Cuba. Vet. Microbiol. 2011, 151, 245–254. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kishino, H.; Miyata, T.; Hasegawa, M. Maximum likelihood inference of protein phylogeny and the origin of chloroplasts. J. Mol. Evol. 1990, 31, 151–160. [Google Scholar] [CrossRef]
- Shimodaira, H.; Hasegawa, M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol. Biol. Evol. 1999, 16, 1114–1116. [Google Scholar] [CrossRef] [Green Version]
- Buckley, T.R. Model Misspecification and Probabilistic Tests of Topology: Evidence from Empirical Data Sets. Syst. Biol. 2002, 51, 509–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimodaira, H.; Hasegawa, M. CONSEL: For assessing the confidence of phylogenetic tree selection. Bioinformatics 2001, 17, 1246–1247. [Google Scholar] [CrossRef] [Green Version]
- Strimmer, K.; Rambaut, A. Inferring confidence sets of possibly misspecified gene trees. Proc. R. Soc. Lond. Ser. B Biol. Sci. 2002, 269, 137–142. [Google Scholar] [CrossRef]
- Bao, Y.; Chetvernin, V.; Tatusova, T. Improvements to pairwise sequence comparison (PASC): A genome-based web tool for virus classification. Arch. Virol. 2014, 159, 3293–3304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A Virus Classification Tool Based on Pairwise Sequence Alignment and Identity Calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef]
- Rios, L.; Nunez, J.I.; Diaz de Arce, H.; Ganges, L.; Perez, L.J. Revisiting the genetic diversity of classical swine fever virus: A proposal for new genotyping and subgenotyping schemes of classification. Transbound. Emerg. Dis. 2018, 65, 963–971. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Maust, B.S.; Nickle, D.C.; Learn, G.H.; Liu, Y.; Heath, L.; Pond, S.L.K.; Mullins, J.I. DIVEIN: A web server to analyze phylogenies, sequence divergence, diversity, and informative sites. Biotechniques 2010, 48, 405–408. [Google Scholar] [CrossRef]
- Yu, G.; Smith, D.K.; Zhu, H.; Guan, Y.; Lam, T.T.-Y. ggtree: An r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 2017, 8, 28–36. [Google Scholar] [CrossRef]
- Wickham, H.A.M.; Bryan, J.; Chang, W.; McGowan, L.D.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; Kuhn, M.; et al. Welcome to the tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Xia, X.; Lemey, P. Assessing substitution saturation with DAMBE. In The Phylogenetic Handbook: A Practical Approach to Phylogenetic Analysis and Hypothesis Testing, 2nd ed.; Vandamme, A.-M., Salemi, M., Lemey, P., Eds.; Cambridge University Press: Cambridge, UK, 2009; pp. 615–630. [Google Scholar]
- Wake, D.B. Homoplasy: The Result of Natural Selection, or Evidence of Design Limitations? Am. Nat. 1991, 138, 543–567. [Google Scholar] [CrossRef]
- Hassanin, A.; Lecointre, G.; Tillier, S. The ‘evolutionary signal’ of homoplasy in proteincoding gene sequences and its consequences for a priori weighting in phylogeny. C. R. L’acad. Sci. Ser. III Sci. Vie 1998, 321, 611–620. [Google Scholar] [CrossRef]
- Chowdhury, B.; Garai, G. A review on multiple sequence alignment from the perspective of genetic algorithm. Genomics 2017, 109, 419–431. [Google Scholar] [CrossRef] [PubMed]
- Vialle, R.A.; Tamuri, A.U.; Goldman, N. Alignment Modulates Ancestral Sequence Reconstruction Accuracy. Mol. Biol. Evol. 2018, 35, 1783–1797. [Google Scholar] [CrossRef] [Green Version]
- Yinda, C.K.; Ghogomu, S.M.; Conceição-Neto, N.; Beller, L.; Deboutte, W.; Vanhulle, E.; Maes, P.; Van Ranst, M.; Matthijnssens, J. Cameroonian fruit bats harbor divergent viruses, including rotavirus H, bastroviruses, and picobirnaviruses using an alternative genetic code. Virus Evol. 2018, 4, vey008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kluge, M.; Campos, F.S.; Tavares, M.; de Amorim, D.B.; Valdez, F.P.; Giongo, A.; Roehe, P.M.; Franco, A.C. Metagenomic Survey of Viral Diversity Obtained from Feces of Subantarctic and South American Fur Seals. PLoS ONE 2016, 11, e0151921. [Google Scholar] [CrossRef] [Green Version]
- Ashkenazy, H.; Sela, I.; Levy Karin, E.; Landan, G.; Pupko, T. Multiple Sequence Alignment Averaging Improves Phylogeny Reconstruction. Syst. Biol. 2019, 68, 117–130. [Google Scholar] [CrossRef]
- Brown, J.K.; Zerbini, F.M.; Navas-Castillo, J.; Moriones, E.; Ramos-Sobrinho, R.; Silva, J.C.; Fiallo-Olivé, E.; Briddon, R.W.; Hernández-Zepeda, C.; Idris, A.; et al. Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch. Virol. 2015, 160, 1593–1619. [Google Scholar] [CrossRef]
- Luo, X.-L.; Lu, S.; Jin, D.; Yang, J.; Wu, S.-S.; Xu, J. Marmota himalayana in the Qinghai–Tibetan plateau as a special host for bi-segmented and unsegmented picobirnaviruses. Emerg. Microbes Infect. 2018, 7, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Collier, A.M.; Lyytinen, O.L.; Guo, Y.R.; Toh, Y.; Poranen, M.M.; Tao, Y.J. Initiation of RNA Polymerization and Polymerase Encapsidation by a Small dsRNA Virus. PLoS Pathog. 2016, 12, e1005523. [Google Scholar] [CrossRef] [Green Version]
- Smith, D.B.; Meyers, G.; Bukh, J.; Gould, E.A.; Monath, T.; Scott Muerhoff, A.; Pletnev, A.; Rico-Hesse, R.; Stapleton, J.T.; Simmonds, P.; et al. Proposed revision to the taxonomy of the genus Pestivirus, family Flaviviridae. J. Gen. Virol. 2017, 98, 2106–2112. [Google Scholar] [CrossRef] [PubMed]
- Diaz de Arce, H.; Perez, L.J.; Frias, M.T.; Rosell, R.; Tarradas, J.; Nunez, J.I.; Ganges, L. A multiplex RT-PCR assay for the rapid and differential diagnosis of classical swine fever and other pestivirus infections. Vet. Microbiol. 2009, 139, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Coronado, L.; Perera, C.L.; Rios, L.; Frias, M.T.; Perez, L.J. A Critical Review about Different Vaccines against Classical Swine Fever Virus and Their Repercussions in Endemic Regions. Vaccines 2021, 9, 154. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, L.G.; Mechler-Dreibi, M.L.; Almeida, H.M.S.; Gatto, I.R.H. Bovine Viral Diarrhea Virus: Recent Findings about Its Occurrence in Pigs. Viruses 2020, 12, 600. [Google Scholar] [CrossRef]
- Legoff, J.; Resche-Rigon, M.; Bouquet, J.; Robin, M.; Naccache, S.N.; Mercier-Delarue, S.; Federman, S.; Samayoa, E.; Rousseau, C.; Piron, P.; et al. The eukaryotic gut virome in hematopoietic stem cell transplantation: New clues in enteric graft-versus-host disease. Nat. Med. 2017, 23, 1080–1085. [Google Scholar] [CrossRef] [PubMed]
- Ganesh, B.; Masachessi, G.; Mladenova, Z. Animal picobirnavirus. Virusdisease 2014, 25, 223–238. [Google Scholar] [CrossRef]



| Alignment | Tree | logL | deltaL | bp-RELL | p-KH | p-SH | p-WKH | p-WSH | c-ELW | p-AU |
|---|---|---|---|---|---|---|---|---|---|---|
| Mafft | Clustal_ML | −174,955.2386 | 208.05 | 0.007- | 0.008- | 0.023- | 0.008- | 0.018- | 0.00664- | 0.00499- |
| Muscle_ML | −175,110.1153 | 362.92 | 0- | 0- | 0- | 0- | 0- | 6.55 × 10−24- | 0.000423- | |
| Mafft_ML | −174,747.1905 | 0 | 0.993+ | 0.992+ | 1+ | 0.992+ | 0.999+ | 0.993+ | 0.995+ | |
| Muscle | Clustal_ML | −190,144.7609 | 591.07 | 0- | 0- | 0- | 0- | 0- | 5.17 × 10−97- | 0.00017- |
| Muscle_ML | −189,553.6898 | 0 | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ | 0.999+ | |
| Mafft_ML | −189,976.8135 | 423.12 | 0- | 0- | 0- | 0- | 0- | 9.07 × 10−5- | 0.00115- | |
| Clustal | Clustal_ML | −179,040.1775 | 0 | 1+ | 0.999+ | 1+ | 0.999+ | 1+ | 1+ | 1+ |
| Muscle_ML | −179,564.8764 | 524.7 | 0- | 0- | 0- | 0- | 0- | 5.11 × 10−94- | 0.00129- | |
| Mafft_ML | −179,438.5019 | 398.32 | 0- | 0.001- | 0.001- | 0.001- | 0.002- | 1.44 × 10−12- | 2.68 × 10−5- |
| Alignment | Tree | logL | deltaL | bp-RELL | p-KH | p-SH | p-WKH | p-WSH | c-ELW | p-AU |
|---|---|---|---|---|---|---|---|---|---|---|
| Mafft | NJ | −170,984.6633 | 2041.2 | 0- | 0- | 0- | 0- | 0- | 0- | 1.92 × 10−9- |
| Mafft | ML | −168,943.427 | 0 | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ |
| Mafft | BI | −172,399.3318 | 3455.9 | 0- | 0- | 0- | 0- | 0- | 0- | 3.66 × 10−42- |
| Alignment | Tree | logL | deltaL | bp-RELL | p-KH | p-SH | p-WKH | p-WSH | c-ELW | p-AU |
|---|---|---|---|---|---|---|---|---|---|---|
| Mafft | Clustal_ML | −398,083.4813 | 500.35 | 0- | 0- | 0- | 0- | 0- | 9.57 × 10−10- | 3.6 × 10−5- |
| Mafft_ML | −397,583.133 | 0 | 0.996+ | 0.994+ | 1+ | 0.994+ | 0.999+ | 0.996+ | 0.994+ | |
| Muscle_ML | −397,870.1183 | 286.99 | 0.0039- | 0.0056- | 0.0107- | 0.0056- | 0.0109- | 0.00394- | 0.00696- | |
| Muscle | Clustal_ML | −433,375.5232 | 628.32 | 0- | 0- | 0- | 0- | 0- | 1.49 × 10−89- | 0.00124- |
| Mafft_ML | −433,115.9208 | 368.71 | 0.0002- | 0.0002- | 0.0005- | 0.0002- | 0.0005- | 0.000204- | 0.000896- | |
| Muscle_ML | −432,747.2061 | 0 | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ | 0.999+ | |
| Clustal | Clustal_ML | −411,397.5066 | 0 | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ |
| Mafft_ML | −412,096.1356 | 698.63 | 0- | 0- | 0- | 0- | 0- | 1.9 × 10−101- | 3 × 10−8- | |
| Muscle_ML | −412,145.092 | 747.59 | 0- | 0- | 0- | 0- | 0- | 3.7 × 10−144- | 0.00208- |
| Alignment | Tree | logL | deltaL | bp-RELL | p-KH | p-SH | p-WKH | p-WSH | c-ELW | p-AU |
|---|---|---|---|---|---|---|---|---|---|---|
| Mafft | NJ | −392,286.036 | 768.5 | 0- | 0- | 0- | 0- | 0- | 1.06 × 10−146- | 5.05 × 10−42- |
| Mafft | ML | −389,942.269 | 0 | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ | 1+ |
| Mafft | BI | −392,286.036 | 2343.8 | 0- | 0- | 0- | 0- | 0- | 0- | 1.65 × 10−42- |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perez, L.J.; Cloherty, G.A.; Berg, M.G. Understanding the Genetic Diversity of Picobirnavirus: A Classification Update Based on Phylogenetic and Pairwise Sequence Comparison Approaches. Viruses 2021, 13, 1476. https://doi.org/10.3390/v13081476
Perez LJ, Cloherty GA, Berg MG. Understanding the Genetic Diversity of Picobirnavirus: A Classification Update Based on Phylogenetic and Pairwise Sequence Comparison Approaches. Viruses. 2021; 13(8):1476. https://doi.org/10.3390/v13081476
Chicago/Turabian StylePerez, Lester J., Gavin A. Cloherty, and Michael G. Berg. 2021. "Understanding the Genetic Diversity of Picobirnavirus: A Classification Update Based on Phylogenetic and Pairwise Sequence Comparison Approaches" Viruses 13, no. 8: 1476. https://doi.org/10.3390/v13081476
APA StylePerez, L. J., Cloherty, G. A., & Berg, M. G. (2021). Understanding the Genetic Diversity of Picobirnavirus: A Classification Update Based on Phylogenetic and Pairwise Sequence Comparison Approaches. Viruses, 13(8), 1476. https://doi.org/10.3390/v13081476







