SARS-CoV-2 Reinfection in a Healthcare Worker Despite the Presence of Detectable Neutralizing Antibodies
Abstract
1. Introduction
2. Materials and Methods
2.1. Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
2.2. Cell Culture and Virus Isolation
2.3. Viral Whole Genome Sequence Analysis
2.4. Analysis of Humoral Immune Response
2.5. Virus Neutralization Assay (NT)
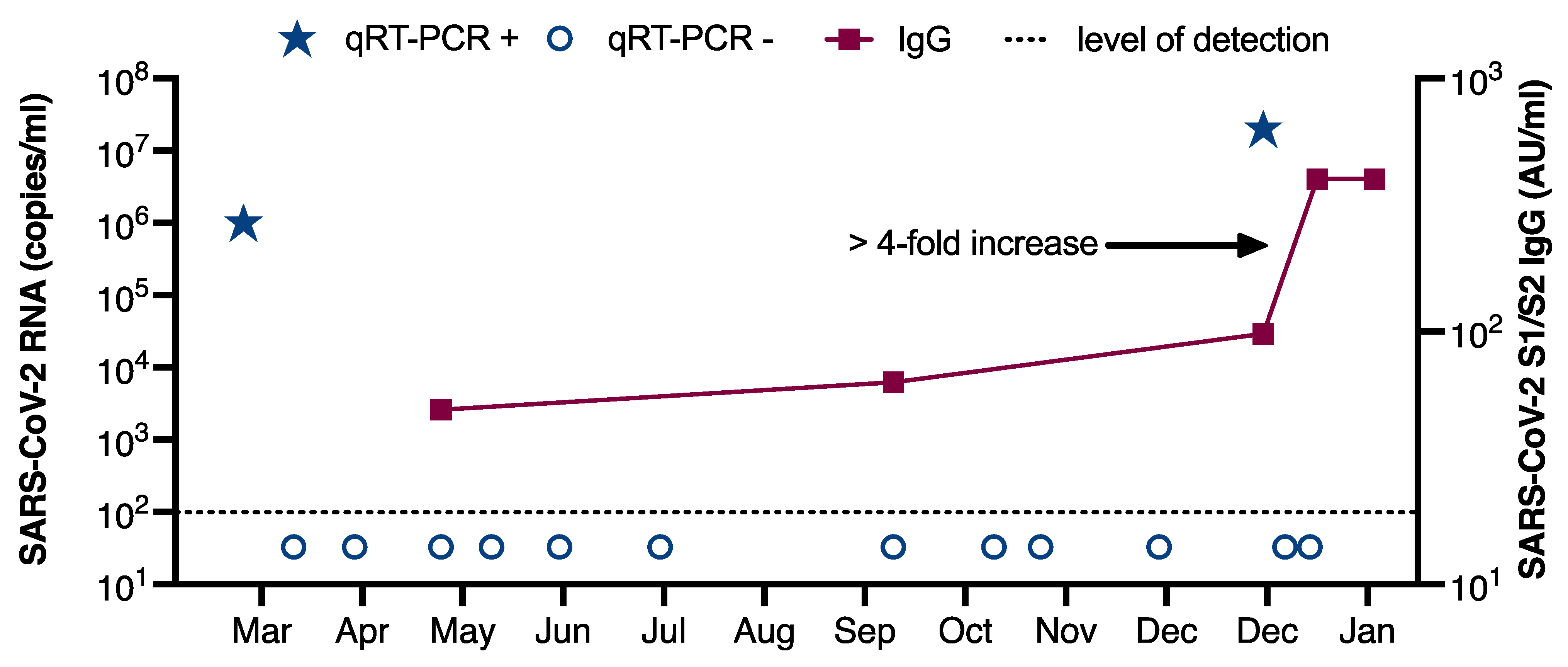
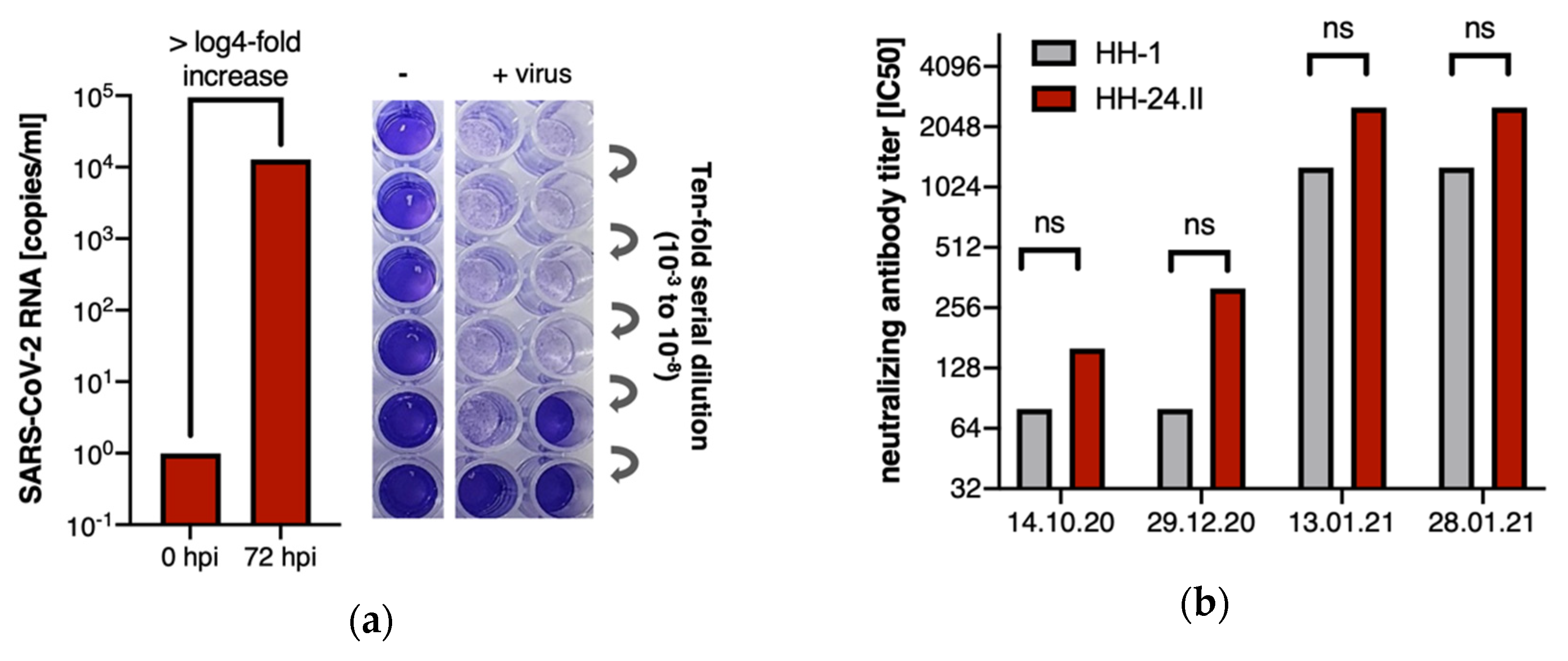
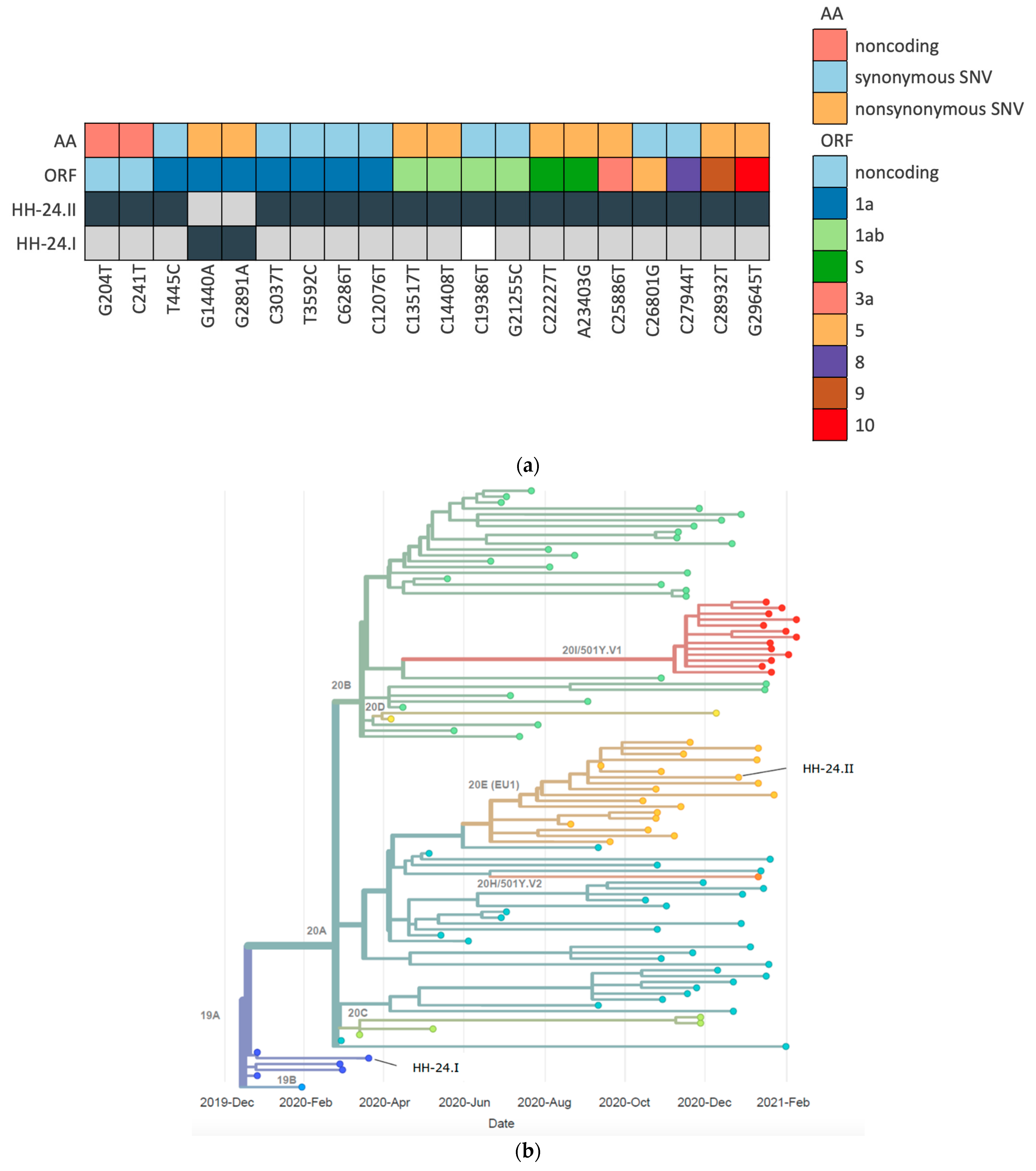
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Coronavirus Disease (COVID-19) Pandemic. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019 (accessed on 7 March 2021).
- Gaebler, C.; Wang, Z.; Lorenzi, J.C.C.; Muecksch, F.; Finkin, S.; Tokuyama, M.; Cho, A.; Jankovic, M.; Schaefer-Babajew, D.; Oliveira, T.Y.; et al. Evolution of antibody immunity to SARS-CoV-2. Nature 2021, 591, 639–644. [Google Scholar] [CrossRef]
- To, K.K.; Hung, I.F.; Ip, J.D.; Chu, A.W.; Chan, W.M.; Tam, A.R.; Fong, C.H.; Yuan, S.; Tsoi, H.W.; Ng, A.C.; et al. COVID-19 re-infection by a phylogenetically distinct SARS-coronavirus-2 strain confirmed by whole genome sequencing. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Tillett, R.L.; Sevinsky, J.R.; Hartley, P.D.; Kerwin, H.; Crawford, N.; Gorzalski, A.; Laverdure, C.; Verma, S.C.; Rossetto, C.C.; Jackson, D.; et al. Genomic evidence for reinfection with SARS-CoV-2: A case study. Lancet Infect. Dis. 2021, 21, 52–58. [Google Scholar] [CrossRef]
- Van Elslande, J.; Vermeersch, P.; Vandervoort, K.; Wawina-Bokalanga, T.; Vanmechelen, B.; Wollants, E.; Laenen, L.; André, E.; Van Ranst, M.; Lagrou, K.; et al. Symptomatic SARS-CoV-2 reinfection by a phylogenetically distinct strain. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Harrington, D.; Kele, B.; Pereira, S.; Couto-Parada, X.; Riddell, A.; Forbes, S.; Dobbie, H.; Cutino-Moguel, T. Confirmed Reinfection with SARS-CoV-2 Variant VOC-202012/01. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef] [PubMed]
- Selhorst, P.; Van Ierssel, S.; Michiels, J.; Mariën, J.; Bartholomeeusen, K.; Dirinck, E.; Vandamme, S.; Jansens, H.; Ariën, K.K. Symptomatic SARS-CoV-2 reinfection of a health care worker in a Belgian nosocomial outbreak despite primary neutralizing antibody response. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Prado-Vivar, B.; Becerra-Wong, M.; Guadalupe, J.J.; Márquez, S.; Gutierrez, B.; Rojas-Silva, P.; Grunauer, M.; Trueba, G.; Barragán, V.; Cárdenas, P. A case of SARS-CoV-2 reinfection in Ecuador. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Gupta, V.; Bhoyar, R.C.; Jain, A.; Srivastava, S.; Upadhayay, R.; Imran, M.; Jolly, B.; Divakar, M.K.; Sharma, D.; Sehgal, P.; et al. Asymptomatic reinfection in two healthcare workers from India with genetically distinct SARS-CoV-2. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Babiker, A.; Marvil, C.; Waggoner, J.J.; Collins, M.; Piantadosi, A. The Importance and Challenges of Identifying SARS-CoV-2 Reinfections. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef]
- Nörz, D.; Frontzek, A.; Eigner, U.; Oestereich, L.; Wichmann, D.; Kluge, S.; Fischer, N.; Aepfelbacher, M.; Pfefferle, S.; Lütgehetmann, M. Pushing beyond specifications: Evaluation of linearity and clinical performance of the cobas 6800/8800 SARS-CoV-2 RT-PCR assay for reliable quantification in blood and other materials outside recommendations. J. Clin. Virol. 2020, 132, 104650. [Google Scholar] [CrossRef]
- Pfefferle, S.; Huang, J.; Nörz, D.; Indenbirken, D.; Lütgehetmann, M.; Oestereich, L.; Günther, T.; Grundhoff, A.; Aepfelbacher, M.; Fischer, N. Complete Genome Sequence of a SARS-CoV-2 Strain Isolated in Northern Germany. Microbiol. Resour. Announc. 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Pfefferle, S.; Günther, T.; Kobbe, R.; Czech-Sioli, M.; Nörz, D.; Santer, R.; Oh, J.; Kluge, S.; Oestereich, L.; Peldschus, K.; et al. SARS Coronavirus-2 variant tracing within the first Coronavirus Disease 19 clusters in northern Germany. Clin. Microbiol. Infect. 2021, 27, 130.e135–130.e138. [Google Scholar] [CrossRef]
- Günther, T.; Czech-Sioli, M.; Indenbirken, D.; Robitaille, A.; Tenhaken, P.; Exner, M.; Ottinger, M.; Fischer, N.; Grundhoff, A.; Brinkmann, M.M. SARS-CoV-2 outbreak investigation in a German meat processing plant. EMBO Mol. Med. 2020, 12, e13296. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Garrison, E.; Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv 2012, arXiv:1207.3907v2. [Google Scholar]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids. Res. 2010, 38, e164. [Google Scholar] [CrossRef] [PubMed]
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef]
- Elbe, S.; Buckland-Merrett, G. Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob. Chall. 2017, 1, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Pflüger, L.S.; Bannasch, J.H.; Brehm, T.T.; Pfefferle, S.; Hoffmann, A.; Nörz, D.; van der Meirschen, M.; Kluge, S.; Haddad, M.; Pischke, S.; et al. Clinical evaluation of five different automated SARS-CoV-2 serology assays in a cohort of hospitalized COVID-19 patients. J. Clin. Virol. 2020, 130, 104549. [Google Scholar] [CrossRef]
- Nörz, D.; Fischer, N.; Schultze, A.; Kluge, S.; Mayer-Runge, U.; Aepfelbacher, M.; Pfefferle, S.; Lütgehetmann, M. Clinical evaluation of a SARS-CoV-2 RT-PCR assay on a fully automated system for rapid on-demand testing in the hospital setting. J. Clin. Virol. 2020, 128, 104390. [Google Scholar] [CrossRef]
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020, 5, 1403–1407. [Google Scholar] [CrossRef]
- Callow, K.A.; Parry, H.F.; Sergeant, M.; Tyrrell, D.A. The time course of the immune response to experimental coronavirus infection of man. Epidemiol. Infect. 1990, 105, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Galanti, M.; Shaman, J. Direct Observation of Repeated Infections With Endemic Coronaviruses. J. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lumley, S.F.; O’Donnell, D.; Stoesser, N.E.; Matthews, P.C.; Howarth, A.; Hatch, S.B.; Marsden, B.D.; Cox, S.; James, T.; Warren, F.; et al. Antibody Status and Incidence of SARS-CoV-2 Infection in Health Care Workers. N. Engl. J. Med. 2021, 384, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Dan, J.M.; Mateus, J.; Kato, Y.; Hastie, K.M.; Yu, E.D.; Faliti, C.E.; Grifoni, A.; Ramirez, S.I.; Haupt, S.; Frazier, A.; et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science 2021, 371. [Google Scholar] [CrossRef]
- Yahav, D.; Yelin, D.; Eckerle, I.; Eberhardt, C.S.; Wang, J.; Cao, B.; Kaiser, L. Definitions for coronavirus disease 2019 reinfection, relapse and PCR re-positivity. Clin. Microbiol. Infect. 2020. [Google Scholar] [CrossRef]
- Li, W.; Su, Y.Y.; Zhi, S.S.; Huang, J.; Zhuang, C.L.; Bai, W.Z.; Wan, Y.; Meng, X.R.; Zhang, L.; Zhou, Y.B.; et al. Virus shedding dynamics in asymptomatic and mildly symptomatic patients infected with SARS-CoV-2. Clin. Microbiol. Infect. 2020, 26, 1556.e1–1556.e6. [Google Scholar] [CrossRef] [PubMed]
- Molina, L.P.; Chow, S.K.; Nickel, A.; Love, J.E. Prolonged Detection of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) RNA in an Obstetric Patient With Antibody Seroconversion. Obstet. Gynecol. 2020, 136, 838–841. [Google Scholar] [CrossRef]
- Abu Jabal, K.; Ben-Amram, H.; Beiruti, K.; Batheesh, Y.; Sussan, C.; Zarka, S.; Edelstein, M. Impact of age, ethnicity, sex and prior infection status on immunogenicity following a single dose of the BNT162b2 mRNA COVID-19 vaccine: Real-world evidence from healthcare workers, Israel, December 2020 to January 2021. Eurosurveillance 2021, 26. [Google Scholar] [CrossRef]
- Brehm, T.T.; Schwinge, D.; Lampalzer, S.; Schlicker, V.; Küchen, J.; Thompson, M.; Ullrich, F.; Huber, S.; Schmiedel, S.; Addo, M.M.; et al. Seroprevalence of SARS-CoV-2 antibodies among hospital workers in a German tertiary care center: A sequential follow-up study. Int. J. Hyg. Environ. Health 2021, 232, 113671. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brehm, T.T.; Pfefferle, S.; von Possel, R.; Kobbe, R.; Nörz, D.; Schmiedel, S.; Grundhoff, A.; Olearo, F.; Emmerich, P.; Robitaille, A.; et al. SARS-CoV-2 Reinfection in a Healthcare Worker Despite the Presence of Detectable Neutralizing Antibodies. Viruses 2021, 13, 661. https://doi.org/10.3390/v13040661
Brehm TT, Pfefferle S, von Possel R, Kobbe R, Nörz D, Schmiedel S, Grundhoff A, Olearo F, Emmerich P, Robitaille A, et al. SARS-CoV-2 Reinfection in a Healthcare Worker Despite the Presence of Detectable Neutralizing Antibodies. Viruses. 2021; 13(4):661. https://doi.org/10.3390/v13040661
Chicago/Turabian StyleBrehm, Thomas Theo, Susanne Pfefferle, Ronald von Possel, Robin Kobbe, Dominik Nörz, Stefan Schmiedel, Adam Grundhoff, Flaminia Olearo, Petra Emmerich, Alexis Robitaille, and et al. 2021. "SARS-CoV-2 Reinfection in a Healthcare Worker Despite the Presence of Detectable Neutralizing Antibodies" Viruses 13, no. 4: 661. https://doi.org/10.3390/v13040661
APA StyleBrehm, T. T., Pfefferle, S., von Possel, R., Kobbe, R., Nörz, D., Schmiedel, S., Grundhoff, A., Olearo, F., Emmerich, P., Robitaille, A., Günther, T., Braun, P., Andersen, G., Knobloch, J. K., Addo, M. M., Lohse, A. W., Aepfelbacher, M., Fischer, N., Schulze zur Wiesch, J., & Lütgehetmann, M. (2021). SARS-CoV-2 Reinfection in a Healthcare Worker Despite the Presence of Detectable Neutralizing Antibodies. Viruses, 13(4), 661. https://doi.org/10.3390/v13040661





