Intrinsic Immune Mechanisms Restricting Human Cytomegalovirus Replication
Abstract
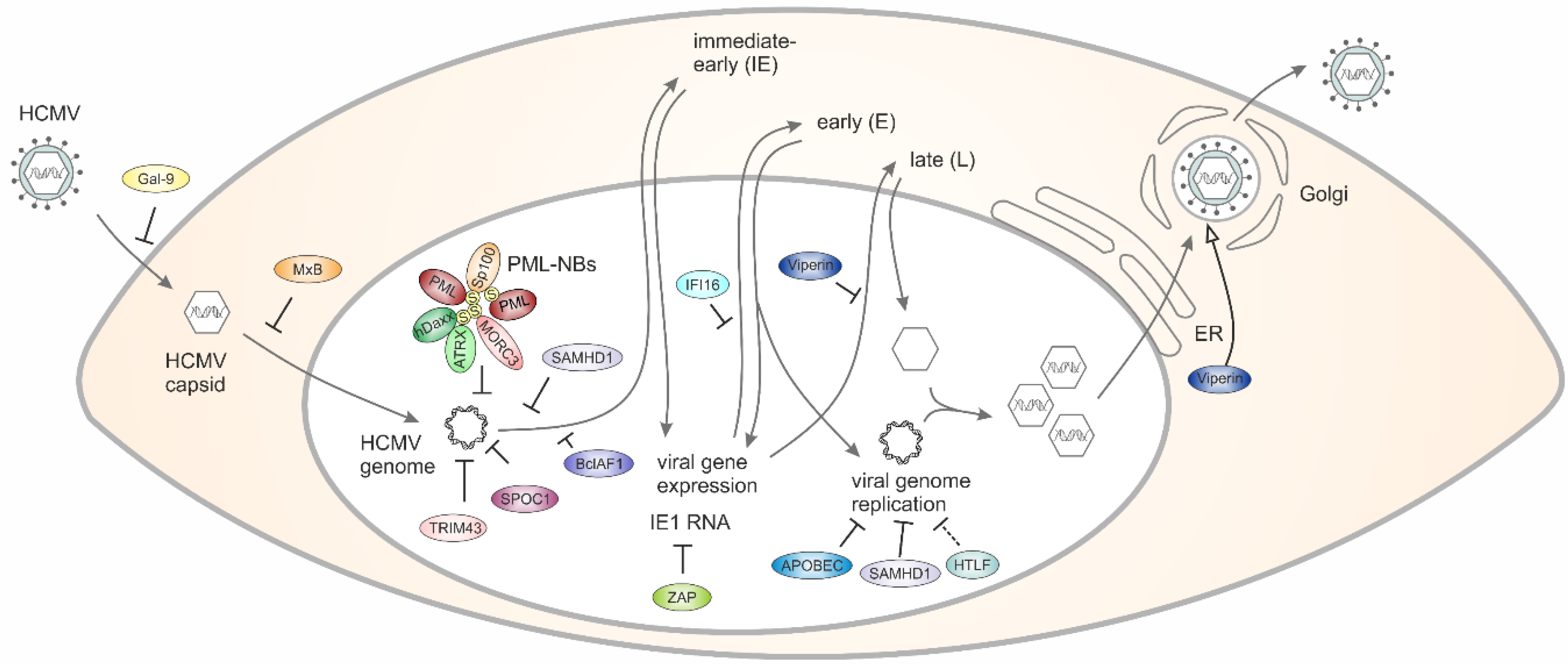
1. Introduction
2. Galectin-9 Restricts the Entry of HCMV into Host Cells
3. MxB Interferes with the Nuclear Delivery of HCMV Genomes
4. PML Nuclear Bodies Associate with Parental HCMV Genomes to Induce Epigenetic Silencing
5. MORC3 Affects the Recruitment of PML-NB Components to Viral DNA
6. TRIM43 Represses Active Viral Chromatin States via Ubiquitination and Degradation of Pericentrin
7. SPOC1 Associates with the HCMV Major Immediate Early Promoter to Induce Chromatin Compaction
8. IFI16 Targets Incoming HCMV DNA to Regulate Viral Promoter Activities
9. BclAF1 Restricts Herpesviral Replication via Regulation of the Type I Interferon Response
10. HCMV Evades ZAP Detection by Suppressing CpG Dinucleotides in the Major Immediate Early 1 RNA
11. HTLF is Degraded during HCMV Infection to Avoid Post-Replicative DNA Repair
12. APOBEC3 Proteins Induce Cytidine-Deaminase Mediated Hypermutation of the HCMV Genome in a Cell-Type Specific Manner
13. SAMHD1 Restricts HCMV Replication by Limiting NF-kB Activation and Intracellular Deoxynucleoside Triphosphate Pools
14. HCMV Redirects the RF Viperin to Enhance Viral Infectivity
15. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bieniasz, P.D. Intrinsic immunity: A front-line defense against viral attack. Nat. Immunol. 2004, 5, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Paludan, S.R.; Pradeu, T.; Masters, S.L.; Mogensen, T.H. Constitutive immune mechanisms: Mediators of host defence and immune regulation. Nat. Rev. Immunol. 2020, 2020, 1–14. [Google Scholar] [CrossRef]
- Blanco-Melo, D.; Venkatesh, S.; Bieniasz, P.D. Intrinsic cellular defenses against human immunodeficiency viruses. Immunity 2012, 37, 399–411. [Google Scholar] [CrossRef]
- Hrecka, K.; Hao, C.; Gierszewska, M.; Swanson, S.K.; Kesik-Brodacka, M.; Srivastava, S.; Florens, L.; Washburn, M.P.; Skowronski, J. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 2011, 474, 658–661. [Google Scholar] [CrossRef] [PubMed]
- Neil, S.J.; Zang, T.; Bieniasz, P.D. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature 2008, 451, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Sheehy, A.M.; Gaddis, N.C.; Choi, J.D.; Malim, M.H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 2002, 418, 646–650. [Google Scholar] [CrossRef]
- Simon, V.; Bloch, N.; Landau, N.R. Intrinsic host restrictions to HIV-1 and mechanisms of viral escape. Nat. Immunol. 2015, 16, 546–553. [Google Scholar] [CrossRef]
- Stremlau, M.; Owens, C.M.; Perron, M.J.; Kiessling, M.; Autissier, P.; Sodroski, J. The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature 2004, 427, 848–853. [Google Scholar] [CrossRef]
- Yan, N.; Chen, Z.J. Intrinsic antiviral immunity. Nat. Immunol. 2012, 13, 214–222. [Google Scholar] [CrossRef]
- Nakayama, E.E.; Shioda, T. TRIM5α and Species Tropism of HIV/SIV. Front. Microbiol. 2012, 3, 13. [Google Scholar] [CrossRef]
- Brass, A.L.; Huang, I.C.; Benita, Y.; John, S.P.; Krishnan, M.N.; Feeley, E.M.; Ryan, B.J.; Weyer, J.L.; van der Weyden, L.; Fikrig, E.; et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 2009, 139, 1243–1254. [Google Scholar] [CrossRef] [PubMed]
- Daffis, S.; Szretter, K.J.; Schriewer, J.; Li, J.; Youn, S.; Errett, J.; Lin, T.Y.; Schneller, S.; Zust, R.; Dong, H.; et al. 2′-O methylation of the viral mRNA cap evades host restriction by IFIT family members. Nature 2010, 468, 452–456. [Google Scholar] [CrossRef] [PubMed]
- Pavlovic, J.; Haller, O.; Staeheli, P. Human and mouse Mx proteins inhibit different steps of the influenza virus multiplication cycle. J. Virol. 1992, 66, 2564–2569. [Google Scholar] [CrossRef] [PubMed]
- Shapira, S.D.; Gat-Viks, I.; Shum, B.O.; Dricot, A.; de Grace, M.M.; Wu, L.; Gupta, P.B.; Hao, T.; Silver, S.J.; Root, D.E.; et al. A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell 2009, 139, 1255–1267. [Google Scholar] [CrossRef] [PubMed]
- Aiello, A.; Accardi, G.; Candore, G.; Caruso, C.; Colomba, C.; Di Bona, D.; Duro, G.; Gambino, C.M.; Ligotti, M.E.; Pandey, J.P. Role of Immunogenetics in the Outcome of HCMV Infection: Implications for Ageing. Int. J. Mol. Sci. 2019, 20, 685. [Google Scholar] [CrossRef]
- Jackson, S.E.; Mason, G.M.; Wills, M.R. Human cytomegalovirus immunity and immune evasion. Virus Res. 2011, 157, 151–160. [Google Scholar] [CrossRef]
- Berry, R.; Watson, G.M.; Jonjic, S.; Degli-Esposti, M.A.; Rossjohn, J. Modulation of innate and adaptive immunity by cytomegaloviruses. Nat. Rev. Immunol. 2020, 20, 113–127. [Google Scholar] [CrossRef]
- Mocarski, E.S., Jr. Betaherpes viral genes and their functions. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University PressCopyright © Cambridge University Press: Cambridge, UK, 2007. [Google Scholar]
- Tavalai, N.; Papior, P.; Rechter, S.; Leis, M.; Stamminger, T. Evidence for a role of the cellular ND10 protein PML in mediating intrinsic immunity against human cytomegalovirus infections. J. Virol. 2006, 80, 8006–8018. [Google Scholar] [CrossRef]
- Tavalai, N.; Stamminger, T. Interplay between Herpesvirus Infection and Host Defense by PML Nuclear Bodies. Viruses 2009, 1, 1240–1264. [Google Scholar] [CrossRef]
- John, S.; Mishra, R. Galectin-9: From cell biology to complex disease dynamics. J. Biosci. 2016, 41, 507–534. [Google Scholar] [CrossRef]
- McSharry, B.P.; Forbes, S.K.; Cao, J.Z.; Avdic, S.; Machala, E.A.; Gottlieb, D.J.; Abendroth, A.; Slobedman, B. Human cytomegalovirus upregulates expression of the lectin galectin 9 via induction of beta interferon. J. Virol. 2014, 88, 10990–10994. [Google Scholar] [CrossRef] [PubMed]
- Machala, E.A.; Avdic, S.; Stern, L.; Zajonc, D.M.; Benedict, C.A.; Blyth, E.; Gottlieb, D.J.; Abendroth, A.; McSharry, B.P.; Slobedman, B. Restriction of Human Cytomegalovirus Infection by Galectin-9. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Haller, O.; Staeheli, P.; Schwemmle, M.; Kochs, G. Mx GTPases: Dynamin-like antiviral machines of innate immunity. Trends Microbiol. 2015, 23, 154–163. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.Y.; Sanchez, D.J.; Aliyari, R.; Lu, S.; Cheng, G. Systematic identification of type I and type II interferon-induced antiviral factors. Proc. Natl. Acad. Sci. USA 2012, 109, 4239–4244. [Google Scholar] [CrossRef]
- Crameri, M.; Bauer, M.; Caduff, N.; Walker, R.; Steiner, F.; Franzoso, F.D.; Gujer, C.; Boucke, K.; Kucera, T.; Zbinden, A.; et al. MxB is an interferon-induced restriction factor of human herpesviruses. Nat. Commun. 2018, 9, 1980. [Google Scholar] [CrossRef] [PubMed]
- Schilling, M.; Bulli, L.; Weigang, S.; Graf, L.; Naumann, S.; Patzina, C.; Wagner, V.; Bauersfeld, L.; Goujon, C.; Hengel, H.; et al. Human MxB Protein Is a Pan-herpesvirus Restriction Factor. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Jaguva Vasudevan, A.A.; Bähr, A.; Grothmann, R.; Singer, A.; Häussinger, D.; Zimmermann, A.; Münk, C. MXB inhibits murine cytomegalovirus. Virology 2018, 522, 158–167. [Google Scholar] [CrossRef]
- Staeheli, P.; Haller, O. Human MX2/MxB: A Potent Interferon-Induced Postentry Inhibitor of Herpesviruses and HIV-1. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Bernardi, R.; Pandolfi, P.P. Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies. Nat. Rev. Mol. Cell Biol. 2007, 8, 1006–1016. [Google Scholar] [CrossRef]
- Ishov, A.M.; Maul, G.G. The periphery of nuclear domain 10 (ND10) as site of DNA virus deposition. J. Cell Biol. 1996, 134, 815–826. [Google Scholar] [CrossRef]
- Ishov, A.M.; Stenberg, R.M.; Maul, G.G. Human cytomegalovirus immediate early interaction with host nuclear structures: Definition of an immediate transcript environment. J. Cell Biol. 1997, 138, 5–16. [Google Scholar] [CrossRef] [PubMed]
- Full, F.; Ensser, A. Early Nuclear Events after Herpesviral Infection. J. Clin. Med. 2019, 8, 1408. [Google Scholar] [CrossRef] [PubMed]
- Lukashchuk, V.; McFarlane, S.; Everett, R.D.; Preston, C.M. Human cytomegalovirus protein pp71 displaces the chromatin-associated factor ATRX from nuclear domain 10 at early stages of infection. J. Virol. 2008, 82, 12543–12554. [Google Scholar] [CrossRef] [PubMed]
- Saffert, R.T.; Kalejta, R.F. Inactivating a cellular intrinsic immune defense mediated by Daxx is the mechanism through which the human cytomegalovirus pp71 protein stimulates viral immediate-early gene expression. J. Virol. 2006, 80, 3863–3871. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Kalejta, R.F. Human cytomegalovirus protein pp71 induces Daxx SUMOylation. J. Virol. 2009, 83, 6591–6598. [Google Scholar] [CrossRef] [PubMed]
- Korioth, F.; Maul, G.G.; Plachter, B.; Stamminger, T.; Frey, J. The nuclear domain 10 (ND10) is disrupted by the human cytomegalovirus gene product IE1. Exp. Cell Res. 1996, 229, 155–158. [Google Scholar] [CrossRef] [PubMed]
- Scherer, M.; Klingl, S.; Sevvana, M.; Otto, V.; Schilling, E.M.; Stump, J.D.; Müller, R.; Reuter, N.; Sticht, H.; Muller, Y.A.; et al. Crystal structure of cytomegalovirus IE1 protein reveals targeting of TRIM family member PML via coiled-coil interactions. PLoS Pathog. 2014, 10, e1004512. [Google Scholar] [CrossRef]
- Schilling, E.M.; Scherer, M.; Reuter, N.; Schweininger, J.; Muller, Y.A.; Stamminger, T. The Human Cytomegalovirus IE1 Protein Antagonizes PML Nuclear Body-Mediated Intrinsic Immunity via the Inhibition of PML De Novo SUMOylation. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Poole, E.L.; Kew, V.G.; Lau, J.C.H.; Murray, M.J.; Stamminger, T.; Sinclair, J.H.; Reeves, M.B. A Virally Encoded DeSUMOylase Activity Is Required for Cytomegalovirus Reactivation from Latency. Cell Rep. 2018, 24, 594–606. [Google Scholar] [CrossRef]
- Ulbricht, T.; Alzrigat, M.; Horch, A.; Reuter, N.; von Mikecz, A.; Steimle, V.; Schmitt, E.; Krämer, O.H.; Stamminger, T.; Hemmerich, P. PML promotes MHC class II gene expression by stabilizing the class II transactivator. J. Cell Biol. 2012, 199, 49–63. [Google Scholar] [CrossRef]
- El Asmi, F.; Maroui, M.A.; Dutrieux, J.; Blondel, D.; Nisole, S.; Chelbi-Alix, M.K. Implication of PMLIV in both intrinsic and innate immunity. PLoS Pathog. 2014, 10, e1003975. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.E.; Ahn, J.H. Positive role of promyelocytic leukemia protein in type I interferon response and its regulation by human cytomegalovirus. PLoS Pathog. 2015, 11, e1004785. [Google Scholar] [CrossRef] [PubMed]
- Scherer, M.; Otto, V.; Stump, J.D.; Klingl, S.; Müller, R.; Reuter, N.; Muller, Y.A.; Sticht, H.; Stamminger, T. Characterization of Recombinant Human Cytomegaloviruses Encoding IE1 Mutants L174P and 1-382 Reveals that Viral Targeting of PML Bodies Perturbs both Intrinsic and Innate Immune Responses. J. Virol. 2016, 90, 1190–1205. [Google Scholar] [CrossRef] [PubMed]
- Paulus, C.; Harwardt, T.; Walter, B.; Marxreiter, A.; Zenger, M.; Reuschel, E.; Nevels, M.M. Revisiting promyelocytic leukemia protein targeting by human cytomegalovirus immediate-early protein 1. PLoS Pathog. 2020, 16, e1008537. [Google Scholar] [CrossRef]
- Sloan, E.; Orr, A.; Everett, R.D. MORC3, a Component of PML Nuclear Bodies, Has a Role in Restricting Herpes Simplex Virus 1 and Human Cytomegalovirus. J. Virol. 2016, 90, 8621–8633. [Google Scholar] [CrossRef]
- Mimura, Y.; Takahashi, K.; Kawata, K.; Akazawa, T.; Inoue, N. Two-step colocalization of MORC3 with PML nuclear bodies. J. Cell Sci. 2010, 123 Pt 12, 2014–2024. [Google Scholar] [CrossRef]
- Sloan, E.; Tatham, M.H.; Groslambert, M.; Glass, M.; Orr, A.; Hay, R.T.; Everett, R.D. Analysis of the SUMO2 Proteome during HSV-1 Infection. PLoS Pathog. 2015, 11, e1005059. [Google Scholar] [CrossRef]
- Nightingale, K.; Lin, K.M.; Ravenhill, B.J.; Davies, C.; Nobre, L.; Fielding, C.A.; Ruckova, E.; Fletcher-Etherington, A.; Soday, L.; Nichols, H.; et al. High-Definition Analysis of Host Protein Stability during Human Cytomegalovirus Infection Reveals Antiviral Factors and Viral Evasion Mechanisms. Cell Host Microbe 2018, 24, 447–460.e11. [Google Scholar] [CrossRef]
- Zhang, Y.; Bertulat, B.; Tencer, A.H.; Ren, X.; Wright, G.M.; Black, J.; Cardoso, M.C.; Kutateladze, T.G. MORC3 Forms Nuclear Condensates through Phase Separation. iScience 2019, 17, 182–189. [Google Scholar] [CrossRef]
- Full, F.; van Gent, M.; Sparrer, K.M.J.; Chiang, C.; Zurenski, M.A.; Scherer, M.; Brockmeyer, N.H.; Heinzerling, L.; Stürzl, M.; Korn, K.; et al. Centrosomal protein TRIM43 restricts herpesvirus infection by regulating nuclear lamina integrity. Nat. Microbiol. 2019, 4, 164–176. [Google Scholar] [CrossRef]
- Mohrmann, G.; Hengstler, J.G.; Hofmann, T.G.; Endele, S.U.; Lee, B.; Stelzer, C.; Zabel, B.; Brieger, J.; Hasenclever, D.; Tanner, B.; et al. SPOC1, a novel PHD-finger protein: Association with residual disease and survival in ovarian cancer. Int. J. Cancer 2005, 116, 547–554. [Google Scholar] [CrossRef] [PubMed]
- Mund, A.; Schubert, T.; Staege, H.; Kinkley, S.; Reumann, K.; Kriegs, M.; Fritsch, L.; Battisti, V.; Ait-Si-Ali, S.; Hoffbeck, A.S.; et al. SPOC1 modulates DNA repair by regulating key determinants of chromatin compaction and DNA damage response. Nucleic Acids Res. 2012, 40, 11363–11379. [Google Scholar] [CrossRef] [PubMed]
- Kinkley, S.; Staege, H.; Mohrmann, G.; Rohaly, G.; Schaub, T.; Kremmer, E.; Winterpacht, A.; Will, H. SPOC1: A novel PHD-containing protein modulating chromatin structure and mitotic chromosome condensation. J. Cell Sci. 2009, 122 Pt 16, 2946–2956. [Google Scholar] [CrossRef]
- Schreiner, S.; Kinkley, S.; Bürck, C.; Mund, A.; Wimmer, P.; Schubert, T.; Groitl, P.; Will, H.; Dobner, T. SPOC1-mediated antiviral host cell response is antagonized early in human adenovirus type 5 infection. PLoS Pathog. 2013, 9, e1003775. [Google Scholar] [CrossRef] [PubMed]
- Reichel, A.; Stilp, A.C.; Scherer, M.; Reuter, N.; Lukassen, S.; Kasmapour, B.; Schreiner, S.; Cicin-Sain, L.; Winterpacht, A.; Stamminger, T. Chromatin-Remodeling Factor SPOC1 Acts as a Cellular Restriction Factor against Human Cytomegalovirus by Repressing the Major Immediate Early Promoter. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Park, Y.H.; Jeong, M.S.; Jang, S.B. Structural insights of homotypic interaction domains in the ligand-receptor signal transduction of tumor necrosis factor (TNF). BMB Rep. 2016, 49, 159–166. [Google Scholar] [CrossRef]
- Yan, H.; Dalal, K.; Hon, B.K.; Youkharibache, P.; Lau, D.; Pio, F. RPA nucleic acid-binding properties of IFI16-HIN200. Biochim. Biophys. Acta 2008, 1784, 1087–1097. [Google Scholar] [CrossRef]
- Unterholzner, L.; Keating, S.E.; Baran, M.; Horan, K.A.; Jensen, S.B.; Sharma, S.; Sirois, C.M.; Jin, T.; Latz, E.; Xiao, T.S.; et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 2010, 11, 997–1004. [Google Scholar] [CrossRef]
- Dell’Oste, V.; Gatti, D.; Gugliesi, F.; De Andrea, M.; Bawadekar, M.; Lo Cigno, I.; Biolatti, M.; Vallino, M.; Marschall, M.; Gariglio, M.; et al. Innate nuclear sensor IFI16 translocates into the cytoplasm during the early stage of in vitro human cytomegalovirus infection and is entrapped in the egressing virions during the late stage. J. Virol. 2014, 88, 6970–6982. [Google Scholar] [CrossRef]
- Diner, B.A.; Lum, K.K.; Toettcher, J.E.; Cristea, I.M. Viral DNA Sensors IFI16 and Cyclic GMP-AMP Synthase Possess Distinct Functions in Regulating Viral Gene Expression, Immune Defenses, and Apoptotic Responses during Herpesvirus Infection. mBio 2016, 7. [Google Scholar] [CrossRef]
- Biolatti, M.; Dell’Oste, V.; Pautasso, S.; von Einem, J.; Marschall, M.; Plachter, B.; Gariglio, M.; De Andrea, M.; Landolfo, S. Regulatory Interaction between the Cellular Restriction Factor IFI16 and Viral pp65 (pUL83) Modulates Viral Gene Expression and IFI16 Protein Stability. J. Virol. 2016, 90, 8238–8250. [Google Scholar] [CrossRef] [PubMed]
- Cristea, I.M.; Moorman, N.J.; Terhune, S.S.; Cuevas, C.D.; O’Keefe, E.S.; Rout, M.P.; Chait, B.T.; Shenk, T. Human cytomegalovirus pUL83 stimulates activity of the viral immediate-early promoter through its interaction with the cellular IFI16 protein. J. Virol. 2010, 84, 7803–7814. [Google Scholar] [CrossRef] [PubMed]
- Gariano, G.R.; Dell’Oste, V.; Bronzini, M.; Gatti, D.; Luganini, A.; De Andrea, M.; Gribaudo, G.; Gariglio, M.; Landolfo, S. The intracellular DNA sensor IFI16 gene acts as restriction factor for human cytomegalovirus replication. PLoS Pathog. 2012, 8, e1002498. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Chen, J.; Cristea, I.M. Human cytomegalovirus tegument protein pUL83 inhibits IFI16-mediated DNA sensing for immune evasion. Cell Host Microbe 2013, 14, 591–599. [Google Scholar] [CrossRef]
- Kasof, G.M.; Goyal, L.; White, E. Btf, a novel death-promoting transcriptional repressor that interacts with Bcl-2-related proteins. Mol. Cell. Biol. 1999, 19, 4390–4404. [Google Scholar] [CrossRef]
- Sarras, H.; Alizadeh Azami, S.; McPherson, J.P. In search of a function for BCLAF1. Sci. World J. 2010, 10, 1450–1461. [Google Scholar] [CrossRef]
- Lee, S.H.; Kalejta, R.F.; Kerry, J.; Semmes, O.J.; O’Connor, C.M.; Khan, Z.; Garcia, B.A.; Shenk, T.; Murphy, E. BclAF1 restriction factor is neutralized by proteasomal degradation and microRNA repression during human cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 2012, 109, 9575–9580. [Google Scholar] [CrossRef]
- Qin, C.; Zhang, R.; Lang, Y.; Shao, A.; Xu, A.; Feng, W.; Han, J.; Wang, M.; He, W.; Yu, C.; et al. Bclaf1 critically regulates the type I interferon response and is degraded by alphaherpesvirus US3. PLoS Pathog. 2019, 15, e1007559. [Google Scholar] [CrossRef]
- Karlin, S.; Doerfler, W.; Cardon, L.R. Why is CpG suppressed in the genomes of virtually all small eukaryotic viruses but not in those of large eukaryotic viruses? J. Virol. 1994, 68, 2889–2897. [Google Scholar] [CrossRef]
- Rima, B.K.; McFerran, N.V. Dinucleotide and stop codon frequencies in single-stranded RNA viruses. J. Gen. Virol. 1997, 78 Pt 11, 2859–2870. [Google Scholar] [CrossRef]
- Simmonds, P.; Xia, W.; Baillie, J.K.; McKinnon, K. Modelling mutational and selection pressures on dinucleotides in eukaryotic phyla--selection against CpG and UpA in cytoplasmically expressed RNA and in RNA viruses. BMC Genom. 2013, 14, 610. [Google Scholar] [CrossRef]
- Gaunt, E.; Wise, H.M.; Zhang, H.; Lee, L.N.; Atkinson, N.J.; Nicol, M.Q.; Highton, A.J.; Klenerman, P.; Beard, P.M.; Dutia, B.M.; et al. Elevation of CpG frequencies in influenza A genome attenuates pathogenicity but enhances host response to infection. eLife 2016, 5, e12735. [Google Scholar] [CrossRef] [PubMed]
- Goff, S.P. Evolution: Zapping viral RNAs. Nature 2017, 550, 46–47. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.T.; Chiweshe, S.; McCormick, D.; Raper, A.; Wickenhagen, A.; DeFillipis, V.; Gaunt, E.; Simmonds, P.; Wilson, S.J.; Grey, F. Human cytomegalovirus evades ZAP detection by suppressing CpG dinucleotides in the major immediate early 1 gene. PLoS Pathog. 2020, 16, e1008844. [Google Scholar] [CrossRef] [PubMed]
- Honess, R.W.; Gompels, U.A.; Barrell, B.G.; Craxton, M.; Cameron, K.R.; Staden, R.; Chang, Y.N.; Hayward, G.S. Deviations from expected frequencies of CpG dinucleotides in herpesvirus DNAs may be diagnostic of differences in the states of their latent genomes. J. Gen. Virol. 1989, 70 Pt 4, 837–855. [Google Scholar] [CrossRef]
- Dhont, L.; Mascaux, C.; Belayew, A. The helicase-like transcription factor (HLTF) in cancer: Loss of function or oncomorphic conversion of a tumor suppressor? Cell. Mol. Life Sci. CMLS 2016, 73, 129–147. [Google Scholar] [CrossRef]
- Unk, I.; Hajdú, I.; Fátyol, K.; Hurwitz, J.; Yoon, J.H.; Prakash, L.; Prakash, S.; Haracska, L. Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. Proc. Natl. Acad. Sci. USA 2008, 105, 3768–3773. [Google Scholar] [CrossRef]
- Zhou, X.; DeLucia, M.; Hao, C.; Hrecka, K.; Monnie, C.; Skowronski, J.; Ahn, J. HIV-1 Vpr protein directly loads helicase-like transcription factor (HLTF) onto the CRL4-DCAF1 E3 ubiquitin ligase. J. Biol. Chem. 2017, 292, 21117–21127. [Google Scholar] [CrossRef]
- Yan, J.; Shun, M.C.; Zhang, Y.; Hao, C.; Skowronski, J. HIV-1 Vpr counteracts HLTF-mediated restriction of HIV-1 infection in T cells. Proc. Natl. Acad. Sci. USA 2019, 116, 9568–9577. [Google Scholar] [CrossRef]
- Yu, Q.; König, R.; Pillai, S.; Chiles, K.; Kearney, M.; Palmer, S.; Richman, D.; Coffin, J.M.; Landau, N.R. Single-strand specificity of APOBEC3G accounts for minus-strand deamination of the HIV genome. Nat. Struct. Mol. Biol. 2004, 11, 435–442. [Google Scholar] [CrossRef]
- Harris, R.S.; Dudley, J.P. APOBECs and virus restriction. Virology 2015, 479–480, 131–145. [Google Scholar] [CrossRef] [PubMed]
- Weisblum, Y.; Oiknine-Djian, E.; Zakay-Rones, Z.; Vorontsov, O.; Haimov-Kochman, R.; Nevo, Y.; Stockheim, D.; Yagel, S.; Panet, A.; Wolf, D.G. APOBEC3A Is Upregulated by Human Cytomegalovirus (HCMV) in the Maternal-Fetal Interface, Acting as an Innate Anti-HCMV Effector. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Goila-Gaur, R.; Strebel, K. HIV-1 Vif, APOBEC, and intrinsic immunity. Retrovirology 2008, 5, 51. [Google Scholar] [CrossRef]
- Pautasso, S.; Galitska, G.; Dell’Oste, V.; Biolatti, M.; Cagliani, R.; Forni, D.; De Andrea, M.; Gariglio, M.; Sironi, M.; Landolfo, S. Strategy of Human Cytomegalovirus To Escape Interferon Beta-Induced APOBEC3G Editing Activity. J. Virol. 2018, 92. [Google Scholar] [CrossRef] [PubMed]
- Baldauf, H.M.; Pan, X.; Erikson, E.; Schmidt, S.; Daddacha, W.; Burggraf, M.; Schenkova, K.; Ambiel, I.; Wabnitz, G.; Gramberg, T.; et al. SAMHD1 restricts HIV-1 infection in resting CD4(+) T cells. Nat. Med. 2012, 18, 1682–1687. [Google Scholar] [CrossRef]
- Descours, B.; Cribier, A.; Chable-Bessia, C.; Ayinde, D.; Rice, G.; Crow, Y.; Yatim, A.; Schwartz, O.; Laguette, N.; Benkirane, M. SAMHD1 restricts HIV-1 reverse transcription in quiescent CD4(+) T-cells. Retrovirology 2012, 9, 87. [Google Scholar] [CrossRef]
- Laguette, N.; Sobhian, B.; Casartelli, N.; Ringeard, M.; Chable-Bessia, C.; Ségéral, E.; Yatim, A.; Emiliani, S.; Schwartz, O.; Benkirane, M. SAMHD1 is the dendritic- and myeloid-cell-specific HIV-1 restriction factor counteracted by Vpx. Nature 2011, 474, 654–657. [Google Scholar] [CrossRef]
- Hollenbaugh, J.A.; Gee, P.; Baker, J.; Daly, M.B.; Amie, S.M.; Tate, J.; Kasai, N.; Kanemura, Y.; Kim, D.H.; Ward, B.M.; et al. Host factor SAMHD1 restricts DNA viruses in non-dividing myeloid cells. PLoS Pathog. 2013, 9, e1003481. [Google Scholar] [CrossRef]
- Kim, E.T.; White, T.E.; Brandariz-Núñez, A.; Diaz-Griffero, F.; Weitzman, M.D. SAMHD1 restricts herpes simplex virus 1 in macrophages by limiting DNA replication. J. Virol. 2013, 87, 12949–12956. [Google Scholar] [CrossRef]
- Businger, R.; Deutschmann, J.; Gruska, I.; Milbradt, J.; Wiebusch, L.; Gramberg, T.; Schindler, M. Human cytomegalovirus overcomes SAMHD1 restriction in macrophages via pUL97. Nat. Microbiol. 2019, 4, 2260–2272. [Google Scholar] [CrossRef]
- Zhang, K.; Lv, D.W.; Li, R. Conserved Herpesvirus Protein Kinases Target SAMHD1 to Facilitate Virus Replication. Cell Rep. 2019, 28, 449–459.e5. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.T.; Roche, K.L.; Kulej, K.; Spruce, L.A.; Seeholzer, S.H.; Coen, D.M.; Diaz-Griffero, F.; Murphy, E.A.; Weitzman, M.D. SAMHD1 Modulates Early Steps during Human Cytomegalovirus Infection by Limiting NF-κB Activation. Cell Rep. 2019, 28, 434–448.e6. [Google Scholar] [CrossRef] [PubMed]
- Deutschmann, J.; Schneider, A.; Gruska, I.; Vetter, B.; Thomas, D.; Kießling, M.; Wittmann, S.; Herrmann, A.; Schindler, M.; Milbradt, J.; et al. A viral kinase counteracts in vivo restriction of murine cytomegalovirus by SAMHD1. Nat. Microbiol. 2019, 4, 2273–2284. [Google Scholar] [CrossRef] [PubMed]
- Hyeon, S.; Lee, M.K.; Kim, Y.E.; Lee, G.M.; Ahn, J.H. Degradation of SAMHD1 Restriction Factor Through Cullin-Ring E3 Ligase Complexes During Human Cytomegalovirus Infection. Front. Cell. Infect. Microbiol. 2020, 10, 391. [Google Scholar] [CrossRef]
- Honarmand Ebrahimi, K. A unifying view of the broad-spectrum antiviral activity of RSAD2 (viperin) based on its radical-SAM chemistry. Met. Integr. Biometal Sci. 2018, 10, 539–552. [Google Scholar] [CrossRef]
- Honarmand Ebrahimi, K.; Vowles, J.; Browne, C.; McCullagh, J.; James, W.S. ddhCTP produced by the radical-SAM activity of RSAD2 (viperin) inhibits the NAD(+) -dependent activity of enzymes to modulate metabolism. FEBS Lett. 2020, 594, 1631–1644. [Google Scholar] [CrossRef]
- Chin, K.C.; Cresswell, P. Viperin (cig5), an IFN-inducible antiviral protein directly induced by human cytomegalovirus. Proc. Natl. Acad. Sci. USA 2001, 98, 15125–15130. [Google Scholar] [CrossRef]
- Seo, J.Y.; Yaneva, R.; Hinson, E.R.; Cresswell, P. Human cytomegalovirus directly induces the antiviral protein viperin to enhance infectivity. Science 2011, 332, 1093–1097. [Google Scholar] [CrossRef]
- Seo, J.Y.; Cresswell, P. Viperin regulates cellular lipid metabolism during human cytomegalovirus infection. PLoS Pathog. 2013, 9, e1003497. [Google Scholar] [CrossRef]
| Restriction Factor | Important Cellular Functions | Major Mechanism of Interference with HCMV Replication | Viral Antagonization—HCMV Effector Protein | |
|---|---|---|---|---|
| Acronym | Detailed Denomination | |||
| Gal-9 | Galectin-9 | modulation of immune cell responses | entry inhibition | ? |
| MxB | Myxovirus resistance B | integrity of mitochondria | nuclear import of viral DNA | ? |
| PML | Promyelocytic leukemia | tumor suppressor, regulation of stress and DNA damage response | epigenetic silencing | PML deSUMOylation, PML NB-disperal—IE1 (UL123) |
| Sp100 | Speckled protein of 100 kDa | chromatin regulatory protein | epigenetic silencing | PML NB-dispersal, degradation—IE1 (UL123) |
| Daxx | Death domain associated protein 6 | histone H3.3 chaperone (together with ATRX) | epigenetic silencing | proteasomal degradation—pp71 (UL83) |
| ATRX | Alpha Thalassemia Retardation X-linked | histone H3.3 chaperone (together with Daxx) | epigenetic silencing | displacement from PML NBs —pp71 (UL83) |
| MORC3 | Microorchidia 3 | DNA-dependent ATPase, chromatin modulation | recruitment of PML-NB proteins | proteasomal degradation—? |
| TRIM43 | Tripartite Motif Protein 43 | ubiquitin E3 ligase, regulation of centrosomal integrity | nuclear lamina alteration to repress active viral chromatin | ? |
| SPOC1 | Survival-time associated PHD protein in ovarian cancer-1 | H3K4me3 binding, DNA repair, chromatin compaction | silencing of IE transcription | ? |
| IFI16 | Interferon gamma inducible protein 16 | intracellular DNA sensor, regulation of cellular proliferation and differentiation | regulation of HCMV promoter activities (e.g., downregulation of UL54 expression) | cytoplasmic translocation—pp65 (UL82), UL97 |
| BclAF1 | Bcl-2-associated transcription factor 1 | regulation of apoptosis, repressor of transcription | regulation of IFN-mediated antiviral response 1 | proteasomal degradation—pp71 (UL83); microRNA repression—miR-UL112-1 |
| ZAP | Zinc finger antiviral protein | RNA binding, resolution of IFN response, suppression of tumors | degradation of HCMV RNA | CpG suppression in IE1 RNA |
| HTLF | Helicase-like transcription factor | ubiquitin E3 ligase for PCNA, post- replication repair at DNA forks, tumor suppression | processing of viral replicative intermediates 1 | proteasomal degradation—UL145 |
| APOBEC3 | apolipoprotein B editing enzyme catalytic subunit 3 | cytidine deaminase, antiviral defense, cancer mutagenesis | cytidine deaminase mediated hypermutation of HCMV DNA | limited presence of APOBEC3G hotspots in viral genes |
| SAMHD1 | sterile alpha motif and histidine-aspartate domain-containing protein 1 | dNTP triphosphohydrolase activity, suppression of autoimmunity, tumor suppressor | depletion of dNTPs, reduced NFkB activation | phosphorylation-mediated inactivation—UL97; proteasomal degradation |
| Viperin | virus inhibitory protein, endoplasmic reticulum-associated, interferon-inducible | radical SAM enzyme, regulation of cellular metabolism | downregulation of viral late proteins, facilitation of virion morphogenesis | vMIA (UL37x1) co-opts viperin to facilitate viral egress |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schilling, E.-M.; Scherer, M.; Stamminger, T. Intrinsic Immune Mechanisms Restricting Human Cytomegalovirus Replication. Viruses 2021, 13, 179. https://doi.org/10.3390/v13020179
Schilling E-M, Scherer M, Stamminger T. Intrinsic Immune Mechanisms Restricting Human Cytomegalovirus Replication. Viruses. 2021; 13(2):179. https://doi.org/10.3390/v13020179
Chicago/Turabian StyleSchilling, Eva-Maria, Myriam Scherer, and Thomas Stamminger. 2021. "Intrinsic Immune Mechanisms Restricting Human Cytomegalovirus Replication" Viruses 13, no. 2: 179. https://doi.org/10.3390/v13020179
APA StyleSchilling, E.-M., Scherer, M., & Stamminger, T. (2021). Intrinsic Immune Mechanisms Restricting Human Cytomegalovirus Replication. Viruses, 13(2), 179. https://doi.org/10.3390/v13020179






