In Vitro Inhibition of Zika Virus Replication with Poly(Sodium 4-Styrenesulfonate)
Abstract
1. Introduction
2. Materials and Methods
2.1. Polymeric Inhibitors
2.2. Cells and Virus
2.3. Cell Viability
2.4. RNA Isolation and RT-qPCR
2.5. Mechanism of Action Assays
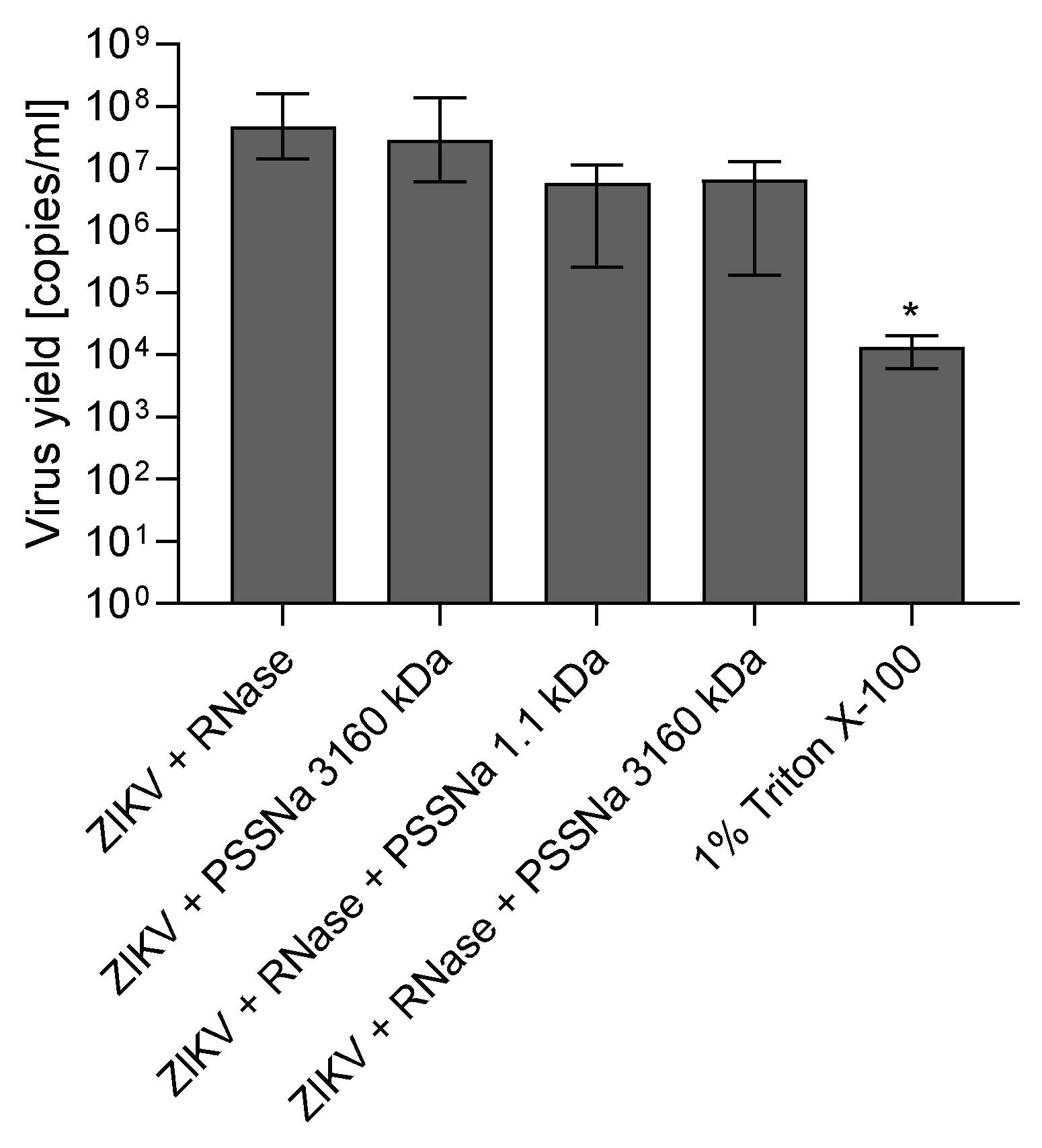
2.6. RNA Digestion Assay
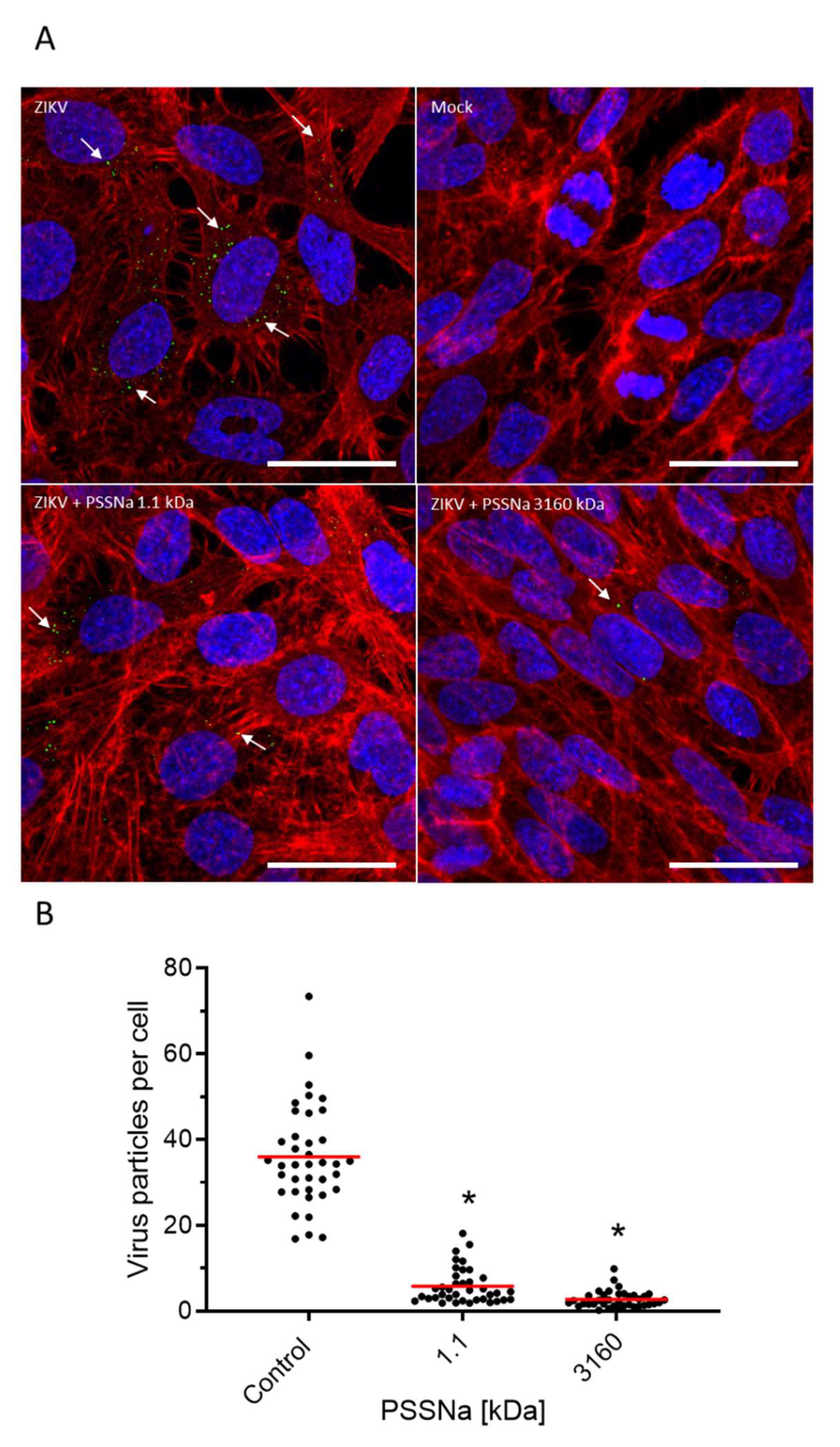
2.7. Confocal Microscopy
2.8. Surface Binding Assay
2.9. Statistics
3. Results
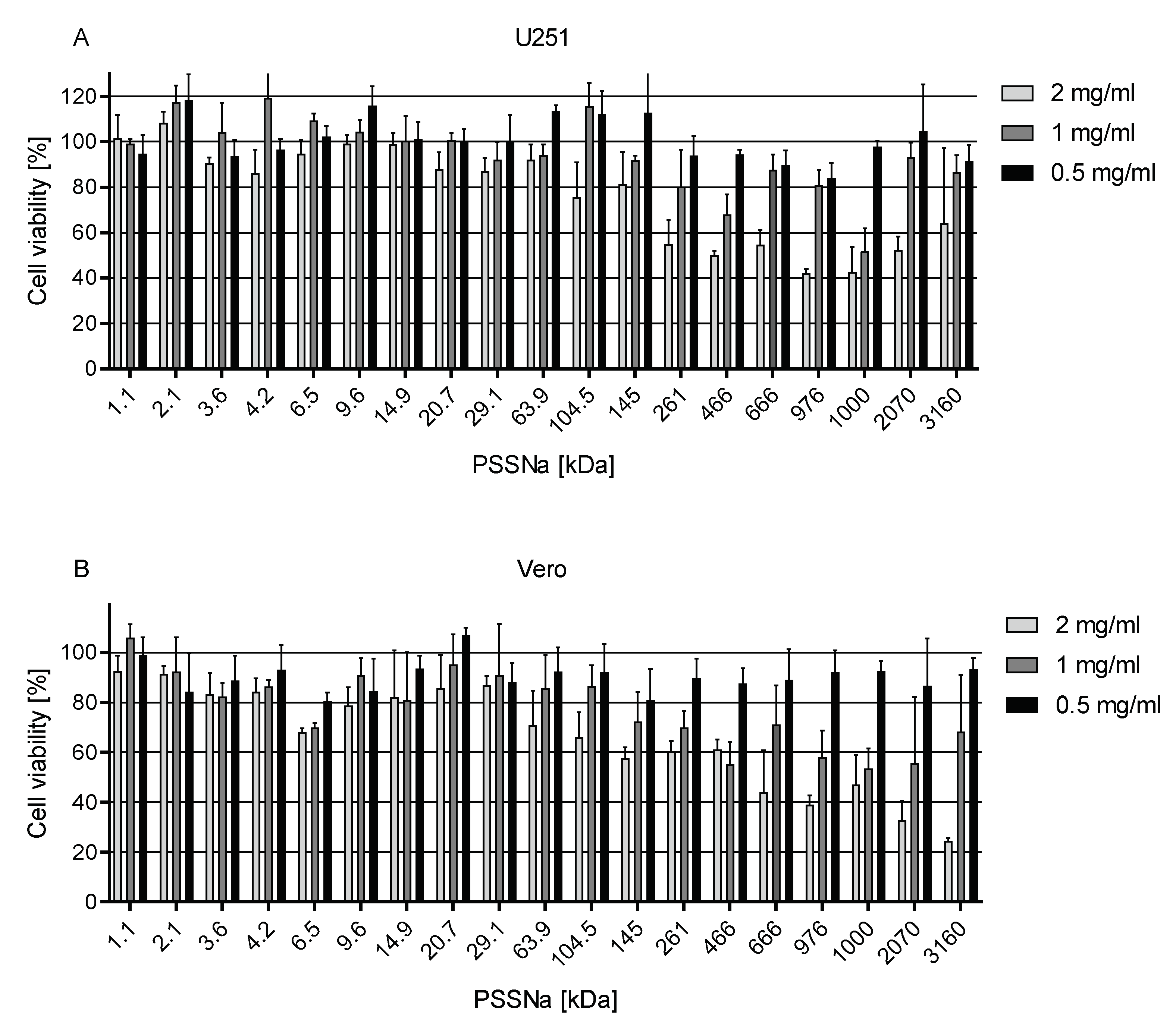
3.1. PSSNa Shows No Toxicity Towards Somatic, Glioblastoma and Primary Cells
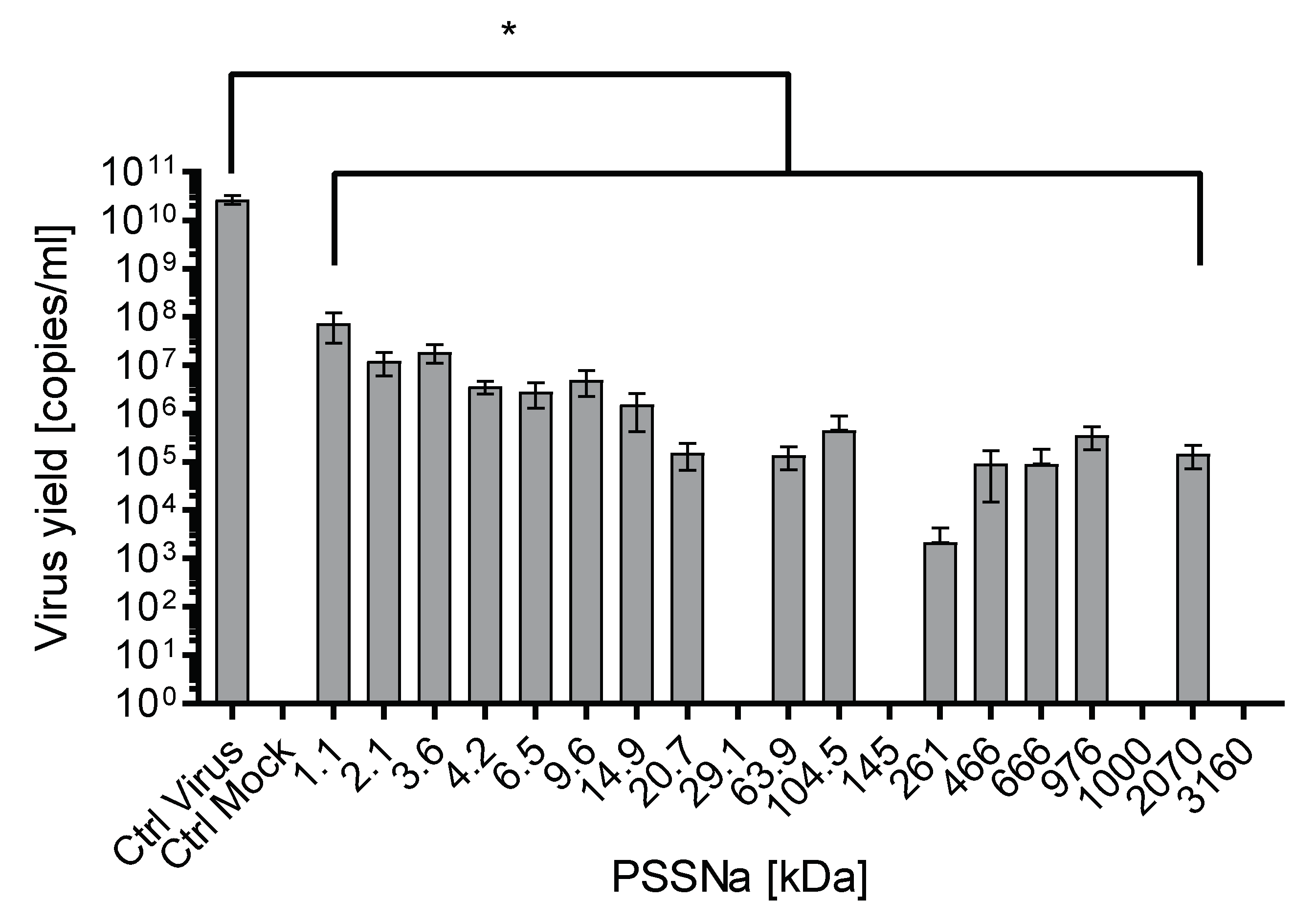
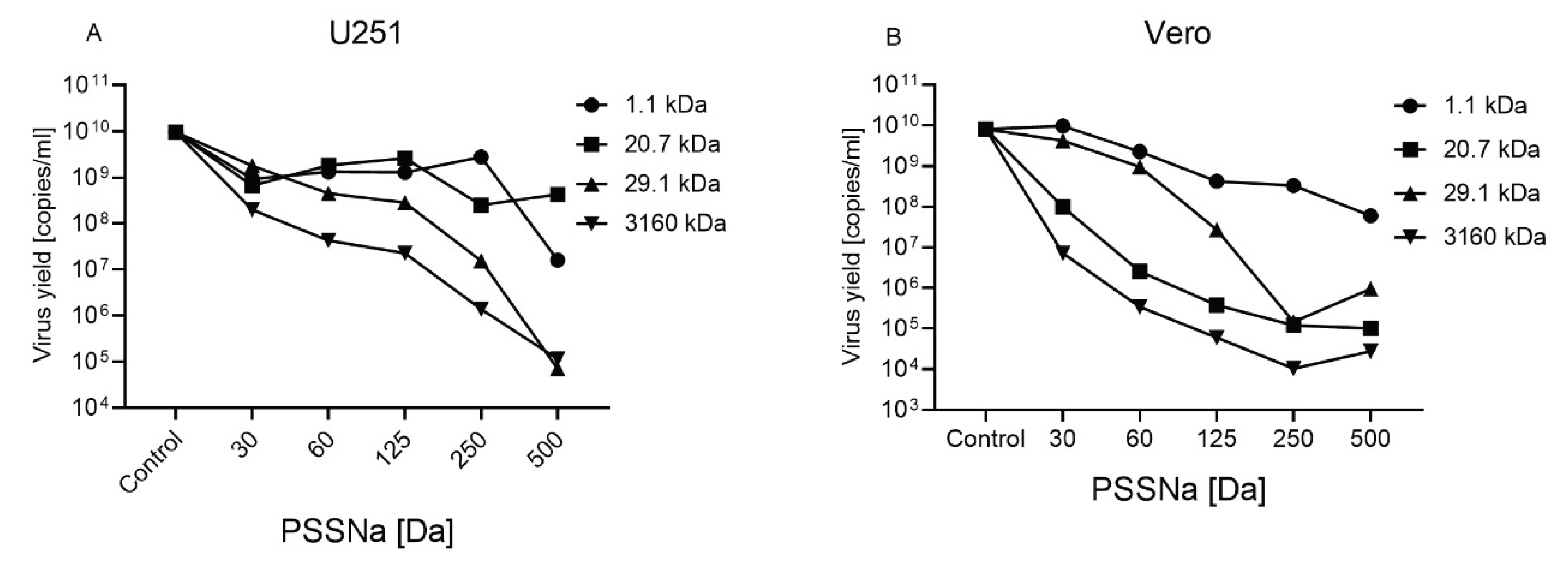
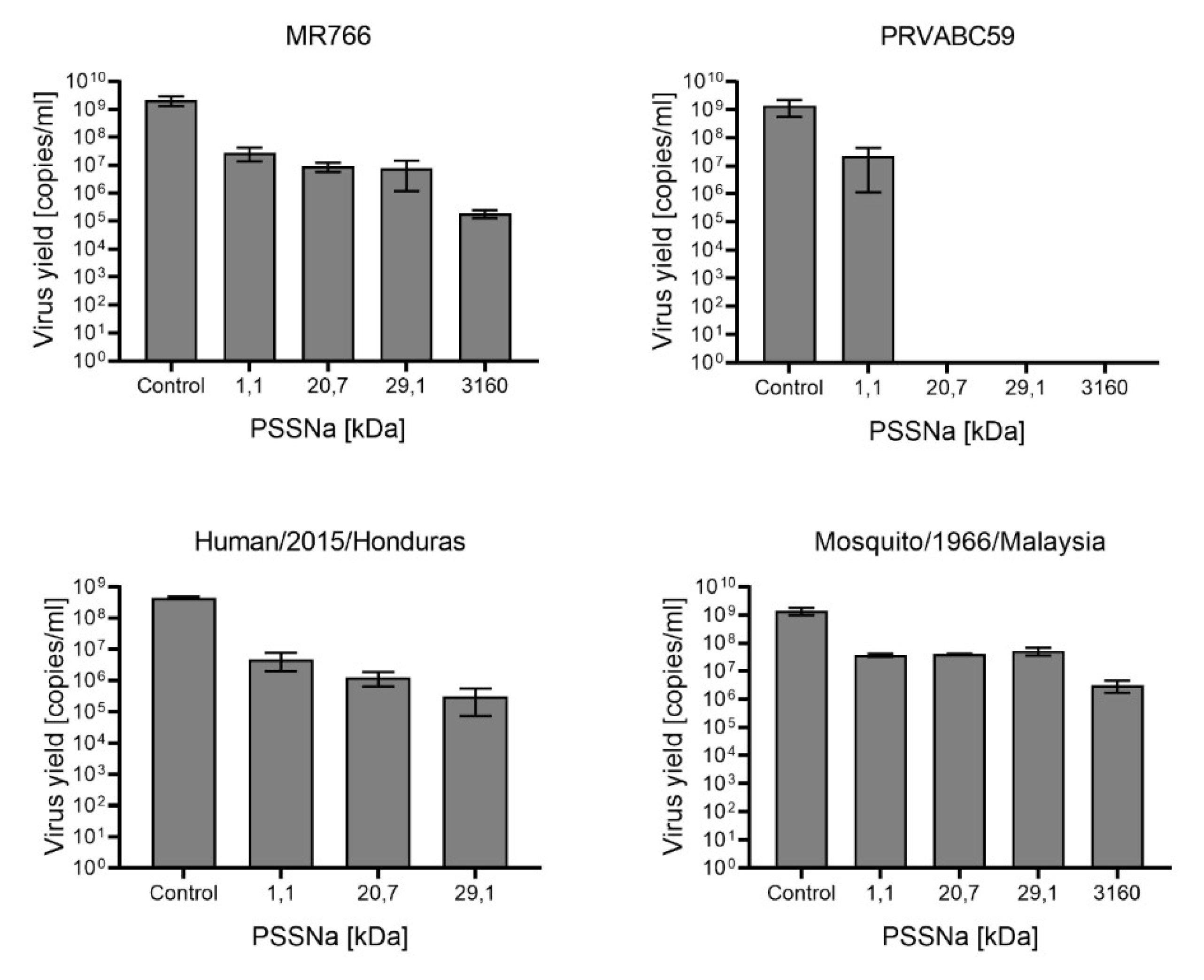
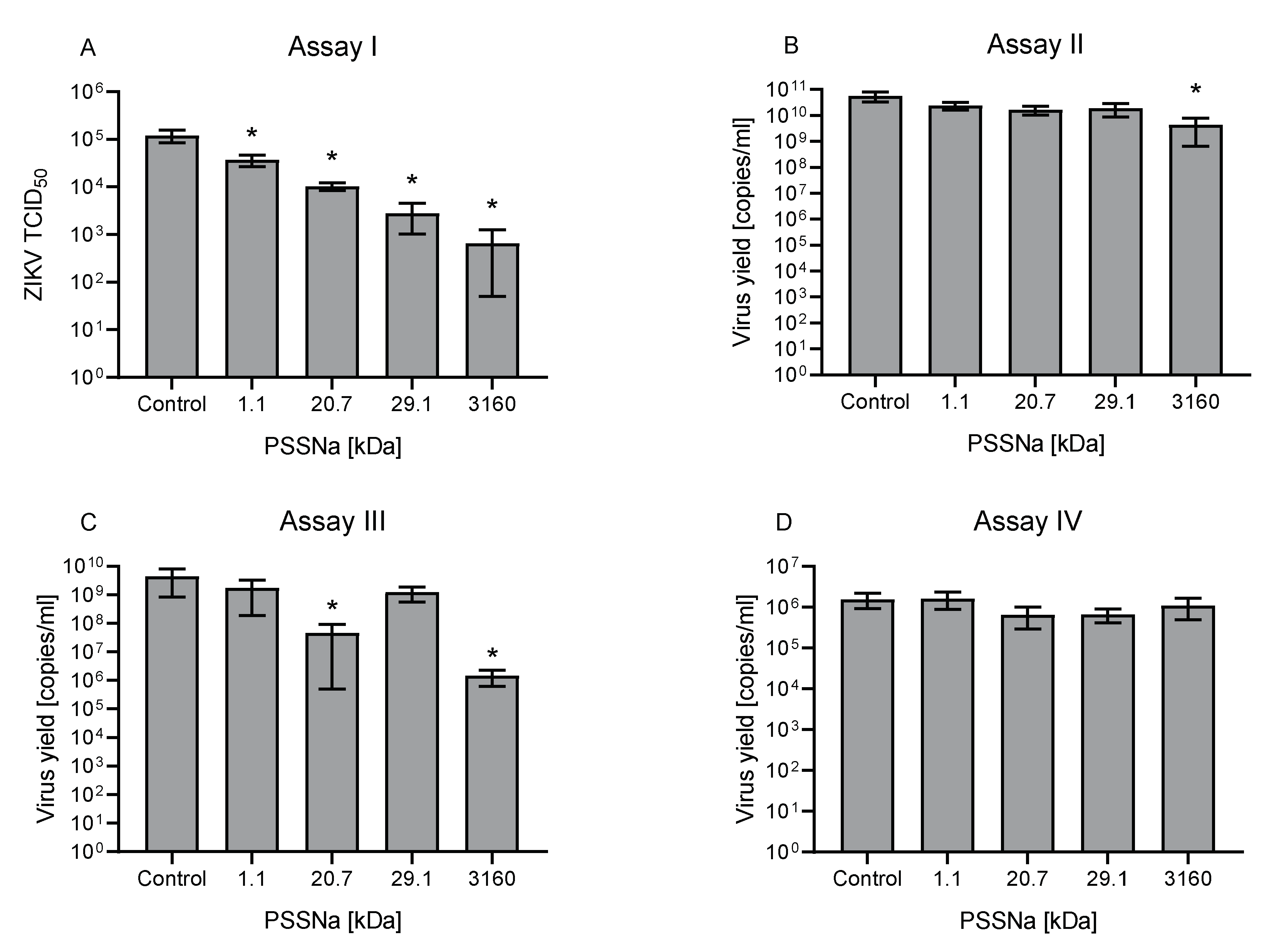
3.2. PSSNa Exhibit Strong Anti-ZIKV Properties
3.3. PSSNa Interacts Directly with ZIKV Virions, Preventing their Attachment to the Host Cell
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lanciotti, R.S.; Kosoy, O.L.; Laven, J.J.; Velez, J.O.; Lambert, A.J.; Johnson, A.J.; Stanfield, S.M.; Duffy, M.R. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 2008, 14, 1232–1239. [Google Scholar] [CrossRef]
- Bueno, M.G.; Martinez, N.; Abdalla, L.; Duarte dos Santos, C.N.; Chame, M. Animals in the Zika Virus Life Cycle: What to Expect from Megadiverse Latin American Countries. PLoS Negl. Trop. Dis. 2016, 10, e0005073. [Google Scholar] [CrossRef]
- World Health Organisation. Countries and Territories with Current or Previous Zika Virus Transmission by WHO Regional Office. Available online: https://www.who.int/emergencies/diseases/zika/countries-with-zika-and-vectors-table.pdf?ua=1 (accessed on 23 August 2020).
- Brady, O.J.; Hay, S.I. The first local cases of Zika virus in Europe. Lancet 2019, 394, 1991–1992. [Google Scholar] [CrossRef]
- Duffy, M.R.; Chen, T.H.; Hancock, W.T.; Powers, A.M.; Kool, J.L.; Lanciotti, R.S.; Pretrick, M.; Marfel, M.; Holzbauer, S.; Dubray, C.; et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. N. Engl. J. Med. 2009, 360, 2536–2543. [Google Scholar] [CrossRef] [PubMed]
- Karkhah, A.; Nouri, H.R.; Javanian, M.; Koppolu, V.; Masrour-Roudsari, J.; Kazemi, S.; Ebrahimpour, S. Zika virus: Epidemiology, clinical aspects, diagnosis, and control of infection. Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 2035–2043. [Google Scholar] [CrossRef] [PubMed]
- Honein, M.A.; Dawson, A.L.; Petersen, E.E.; Jones, A.M.; Lee, E.H.; Yazdy, M.M.; Ahmad, N.; Macdonald, J.; Evert, N.; Bingham, A.; et al. Birth defects among fetuses and infants of US women with evidence of possible zika virus infection during pregnancy. JAMA J. Am. Med. Assoc. 2017, 317, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Robinson, C.L.; Chong, A.C.N.; Ashbrook, A.W.; Jeng, G.; Jin, J.; Chen, H.; Tang, E.I.; Martin, L.A.; Kim, R.S.; Kenyon, R.M.; et al. Male germ cells support long-term propagation of Zika virus. Nat. Commun. 2018, 9, 2090. [Google Scholar] [CrossRef] [PubMed]
- Adachi, K.; Nielsen-Saines, K. Zika clinical updates: Implications for pediatrics. Curr. Opin. Pediatr. 2018, 30, 105–116. [Google Scholar] [CrossRef]
- Brasil, P.; Pereira, J.P.; Moreira, M.E.; Nogueira, R.M.R.; Damasceno, L.; Wakimoto, M.; Rabello, R.S.; Valderramos, S.G.; Halai, U.A.; Salles, T.S.; et al. Zika virus infection in pregnant women in rio de janeiro. N. Engl. J. Med. 2016, 375, 2321–2334. [Google Scholar] [CrossRef]
- Barbi, L.; Coelho, A.V.C.; de Alencar, L.C.A.; Crovella, S. Prevalence of Guillain-Barré syndrome among Zika virus infected cases: A systematic review and meta-analysis. Braz. J. Infect. Dis. 2018, 22, 137–141. [Google Scholar] [CrossRef]
- Parra, B.; Lizarazo, J.; Jiménez-Arango, J.A.; Zea-Vera, A.F.; González-Manrique, G.; Vargas, J.; Angarita, J.A.; Zuñiga, G.; Lopez-Gonzalez, R.; Beltran, C.L.; et al. Guillain-Barré syndrome associated with Zika virus infection in Colombia. N. Engl. J. Med. 2016, 375, 1513–1523. [Google Scholar] [CrossRef] [PubMed]
- Randazzo, W.; Fabra, M.J.; Falcó, I.; López-Rubio, A.; Sánchez, G. Polymers and Biopolymers with Antiviral Activity: Potential Applications for Improving Food Safety. Compr. Rev. Food Sci. Food Saf. 2018, 17, 754–768. [Google Scholar] [CrossRef]
- Ciejka, J.; Milewska, A.; Wytrwal, M.; Wojarski, J.; Golda, A.; Ochman, M.; Nowakowska, M.; Szczubialka, K.; Pyrc, K. Novel polyanions inhibiting replication of influenza viruses. Antimicrob. Agents Chemother. 2016, 60, 1955–1966. [Google Scholar] [CrossRef] [PubMed]
- Riber, C.F.; Hinton, T.M.; Gajda, P.; Zuwala, K.; Tolstrup, M.; Stewart, C.; Zelikin, A.N. Macromolecular Prodrugs of Ribavirin: Structure-Function Correlation as Inhibitors of Influenza Infectivity. Mol. Pharm. 2017, 14, 234–241. [Google Scholar] [CrossRef]
- Carlucci, M.J.; Scolaro, L.A.; Damonte, E.B. Herpes simplex virus type 1 variants arising after selection with an antiviral carrageenan: Lack of correlation between drug susceptibility and syn phenotype. J. Med. Virol. 2002, 68, 92–98. [Google Scholar] [CrossRef]
- Diogo, J.V.; Novo, S.G.; González, M.J.; Ciancia, M.; Bratanich, A.C. Antiviral activity of lambda-carrageenan prepared from red seaweed (Gigartina skottsbergii) against BoHV-1 and SuHV-1. Res. Vet. Sci. 2015, 98, 142–144. [Google Scholar] [CrossRef]
- Nahmias, A.J.; Kibrick, S. Inhibitory effect of heparin on herpes simplex virus. J. Bacteriol. 1964, 87, 1060–1066. [Google Scholar] [CrossRef]
- Pachota, M.; Klysik, K.; Synowiec, A.; Ciejka, J.; Szczubiałka, K.; Pyrć, K.; Nowakowska, M. Inhibition of Herpes Simplex Viruses by Cationic Dextran Derivatives. J. Med. Chem. 2017, 60, 8620–8630. [Google Scholar] [CrossRef]
- Gong, Y.; Matthews, B.; Cheung, D.; Tam, T.; Gadawski, I.; Leung, D.; Holan, G.; Raff, J.; Sacks, S. Evidence of dual sites of action of dendrimers: SPL-2999 inhibits both virus entry and late stages of herpes simplex virus replication. Antivir. Res. 2002, 55, 319–329. [Google Scholar] [CrossRef]
- Herold, B.C.; Bourne, N.; Marcellino, D.; Kirkpatrick, R.; Strauss, D.M.; Zaneveld, L.J.D.; Waller, D.P.; Anderson, R.A.; Chany, C.J.; Barham, B.J.; et al. Poly(sodium 4-styrene sulfonate): An effective candidate topical antimicrobial for the prevention of sexually transmitted diseases. J. Infect. Dis. 2000, 181, 770–773. [Google Scholar] [CrossRef]
- Synowiec, A.; Gryniuk, I.; Pachota, M.; Strzelec, Ł.; Roman, O.; Kłysik-Trzciańska, K.; Zając, M.; Drebot, I.; Gula, K.; Andruchowicz, A.; et al. Cat flu: Broad spectrum polymeric antivirals. Antivir. Res. 2019, 170, 104563. [Google Scholar] [CrossRef] [PubMed]
- Qiu, M.; Chen, Y.; Song, S.; Song, H.; Chu, Y.; Yuan, Z.; Cheng, L.; Zheng, D.; Chen, Z.; Wu, Z. Poly (4-styrenesulfonic acid-co-maleic acid) is an entry inhibitor against both HIV-1 and HSV infections—Potential as a dual functional microbicide. Antivir. Res. 2012, 96, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Carlucci, M.J.; Scolaro, L.A.; Noseda, M.D.; Cerezo, A.S.; Damonte, E.B. Protective effect of a natural carrageenan on genital herpes simplex virus infection in mice. Antivir. Res 2004, 64, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Pachota, M.; Kłysik-Trzciańska, K.; Synowiec, A.; Yukioka, S.; Yusa, S.I.; Zajac, M.; Zawilinska, B.; Dzieciatkowski, T.; Szczubialka, K.; Pyrc, K.; et al. Highly Effective and Safe Polymeric Inhibitors of Herpes Simplex Virus in Vitro and in Vivo. ACS Appl. Mater. Interfaces 2019, 11, 26745–26752. [Google Scholar] [CrossRef] [PubMed]
- Leydet, A.; Barragan, V.; Boyer, B.; Montéro, J.L.; Roque, J.P.; Witvrouw, M.; Este, J.; Snoeck, R.; Andrei, G.; De Clercq, E. Polyanion inhibitors of human immunodeficiency virus and other viruses. 5. Telomerized anionic surfactants derived from amino acids. J. Med. Chem. 1997, 40, 342–349. [Google Scholar] [CrossRef]
- Raghuraman, A.; Tiwari, V.; Zhao, Q.; Shukla, D.; Debnath, A.K.; Desai, U.R. Viral inhibition studies on sulfated lignin, a chemically modified biopolymer and a potential mimic heparan sulfate. Biomacromolecules 2007, 8, 1759–1763. [Google Scholar] [CrossRef]
- Ciejka, J.; Botwina, P.; Nowakowska, M.; Szczubiałka, K.; Pyrc, K. Synthetic sulfonated derivatives of poly (allylamine hydrochloride) as inhibitors of human metapneumovirus. PLoS ONE 2019, 14, e0214646. [Google Scholar] [CrossRef]
- Talarico, L.B.; Noseda, M.D.; Ducatti, D.R.B.; Duarte, M.E.R.; Damonte, E.B. Differential inhibition of dengue virus infection in mammalian and mosquito cells by iota-carrageenan. J. Gen. Virol. 2011, 92, 1332–1342. [Google Scholar] [CrossRef]
- Wang, W.; Wang, S.X.; Guan, H.S. The antiviral activities and mechanisms of marine polysaccharides: An overview. Mar. Drugs 2012, 10, 2795–2816. [Google Scholar] [CrossRef]
- Buck, C.B.; Thompson, C.D.; Roberts, J.N.; Müller, M.; Lowy, D.R.; Schiller, J.T. Carrageenan is a potent inhibitor of papillomavirus infection. PLoS Pathog. 2006, 2, 0671–0680. [Google Scholar] [CrossRef]
- Zoppe, J.O.; Ruottinen, V.; Ruotsalainen, J.; Rönkkö, S.; Johansson, L.S.; Hinkkanen, A.; Järvinen, K.; Seppälä, J. Synthesis of cellulose nanocrystals carrying tyrosine sulfate mimetic ligands and inhibition of alphavirus infection. Biomacromolecules 2014, 15, 1534–1542. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Ciejka, J.; Kaminski, K.; Karewicz, A.; Bielska, D.; Zeglen, S.; Karolak, W.; Nowakowska, M.; Potempa, J.; Bosch, B.J.; et al. Novel polymeric inhibitors of HCoV-NL63. Antivir. Res. 2013, 97, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Hosoya, M.; Balzarini, J.; Shigeta, S.; De Clercq, E. Differential inhibitory effects of sulfated polysaccharides and polymers on the replication of various myxoviruses and retroviruses, depending on the composition of the target amino acid sequences of the viral envelope glycoproteins. Antimicrob. Agents Chemother. 1991, 35, 2515–2520. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.W.; Sam, I.C.; Chong, W.L.; Lee, V.S.; Chan, Y.F. Polysulfonate suramin inhibits Zika virus infection. Antivir. Res. 2017, 143, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Botwina, P.; Owczarek, K.; Rajfur, Z.; Ochman, M.; Urlik, M.; Nowakowska, M.; Szczubiałka, K.; Pyrc, K. Berberine hampers influenza a replication through inhibition of MAPK/ERK pathway. Viruses 2020, 12, 344. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Bolte, S.; Cordelières, F.P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 2006, 224, 213–232. [Google Scholar] [CrossRef]
- Burrell, C.J.; Howard, C.R.; Murphy, F.A.; Burrell, C.J.; Howard, C.R.; Murphy, F.A. Chapter 36—Flaviviruses. Fenner White’s Med. Virol. 2017, 493–518. [Google Scholar]
- Dai, L.; Song, J.; Lu, X.; Deng, Y.Q.; Musyoki, A.M.; Cheng, H.; Zhang, Y.; Yuan, Y.; Song, H.; Haywood, J.; et al. Structures of the Zika Virus Envelope Protein and Its Complex with a Flavivirus Broadly Protective Antibody. Cell Host Microbe 2016, 19, 696–704. [Google Scholar] [CrossRef]
- Milewska, A.; Kaminski, K.; Ciejka, J.; Kosowicz, K.; Zeglen, S.; Wojarski, J.; Nowakowska, M.; Szczubiałka, K.; Pyrc, K. HTCC: Broad range inhibitor of coronavirus entry. PLoS ONE 2016, 11, e0156552. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S.; Gerding, D.N.; Louie, T.J.; Ruiz, N.M.; Gorbachc, S.L. Sustained clinical response as an endpoint in treatment trials of Clostridium difficile-associated diarrhea. Antimicrob. Agents Chemother. 2012, 56, 4043–4045. [Google Scholar] [CrossRef] [PubMed][Green Version]
| U251 Cells | ||||
| 1.1 kDa | 20.7 kDa | 29.1 kDa | 3160 kDa | |
| IC50 (µg/mL) | 9.0 | 8.1 | 14.4 | 8.8 |
| TC50 (µg/mL) | - | 2335.0 | 8495.6 | 2938.3 |
| SI | - | 288.3 | 590.0 | 333.9 |
| Vero Cells | ||||
| 1.1 kDa | 20.7 kDa | 29.1 kDa | 3160 kDa | |
| IC50 (µg/mL) | 53.5 | 13.1 | 30.9 | 8.4 |
| TC50 (µg/mL) | 2530.8 | 4345.5 | - | 1828.0 |
| SI | 47.3 | 331.0 | - | 217.6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Botwina, P.; Obłoza, M.; Szczepański, A.; Szczubiałka, K.; Nowakowska, M.; Pyrć, K. In Vitro Inhibition of Zika Virus Replication with Poly(Sodium 4-Styrenesulfonate). Viruses 2020, 12, 926. https://doi.org/10.3390/v12090926
Botwina P, Obłoza M, Szczepański A, Szczubiałka K, Nowakowska M, Pyrć K. In Vitro Inhibition of Zika Virus Replication with Poly(Sodium 4-Styrenesulfonate). Viruses. 2020; 12(9):926. https://doi.org/10.3390/v12090926
Chicago/Turabian StyleBotwina, Paweł, Magdalena Obłoza, Artur Szczepański, Krzysztof Szczubiałka, Maria Nowakowska, and Krzysztof Pyrć. 2020. "In Vitro Inhibition of Zika Virus Replication with Poly(Sodium 4-Styrenesulfonate)" Viruses 12, no. 9: 926. https://doi.org/10.3390/v12090926
APA StyleBotwina, P., Obłoza, M., Szczepański, A., Szczubiałka, K., Nowakowska, M., & Pyrć, K. (2020). In Vitro Inhibition of Zika Virus Replication with Poly(Sodium 4-Styrenesulfonate). Viruses, 12(9), 926. https://doi.org/10.3390/v12090926






