An Unconventional Flavivirus and Other RNA Viruses in the Sea Cucumber (Holothuroidea; Echinodermata) Virome
Abstract
1. Introduction
2. Materials and Methods
2.1. Specimen Collection
2.2. Metavirome Preparation
2.3. Viral Metagenome Library Analyses
2.4. Viral Genome Architecture and Phylogeny
2.5. Eukaryotic 18S and 28S rRNA Analyses
3. Results and Discussion
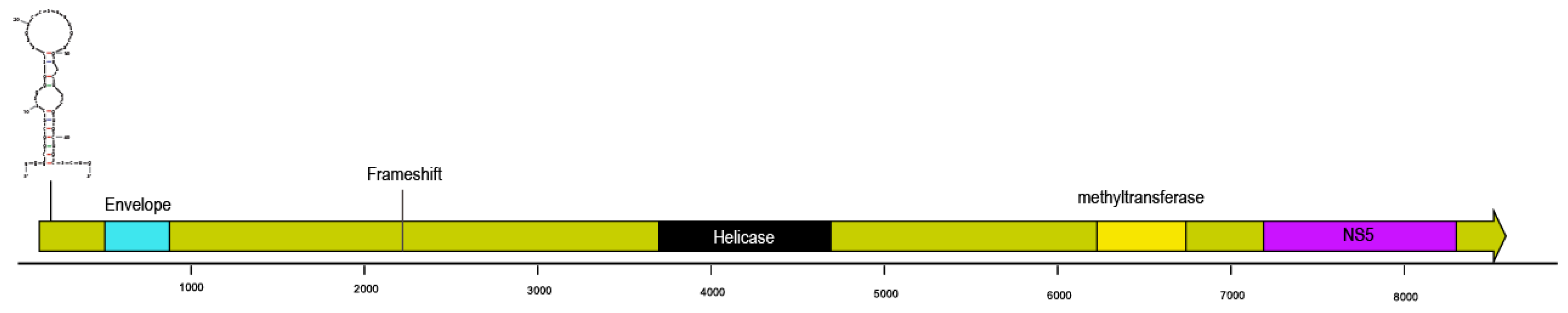
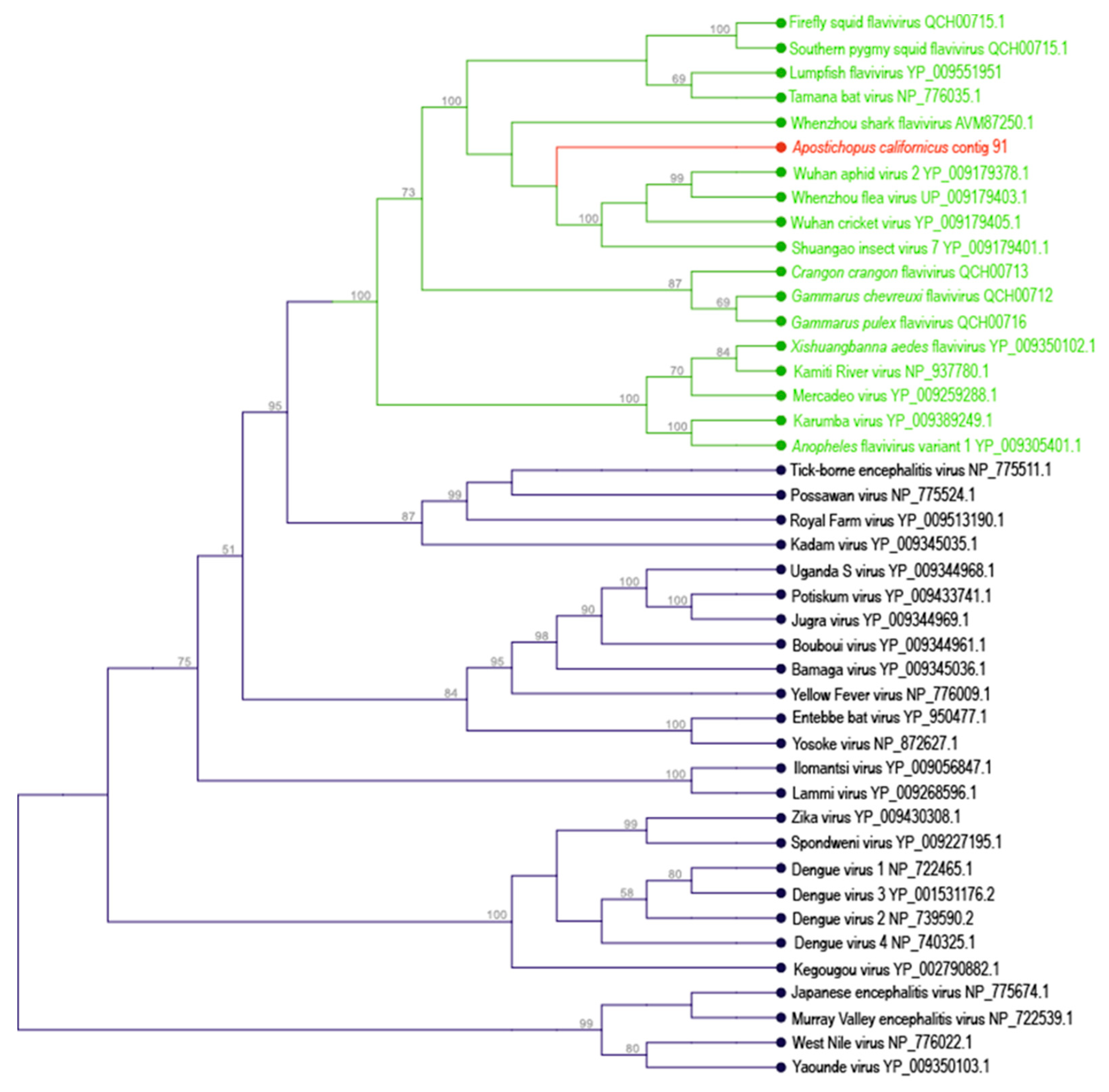
3.1. Flavivirus-Like Genome Fragment
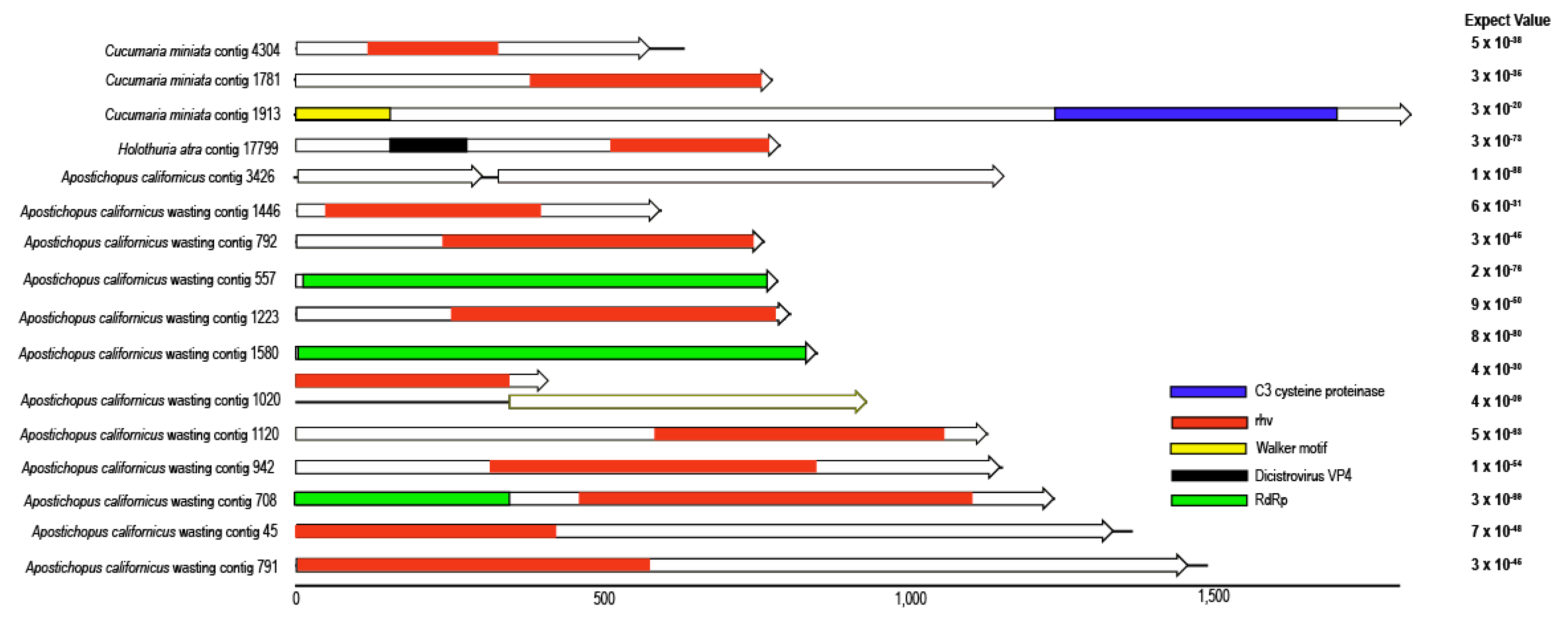
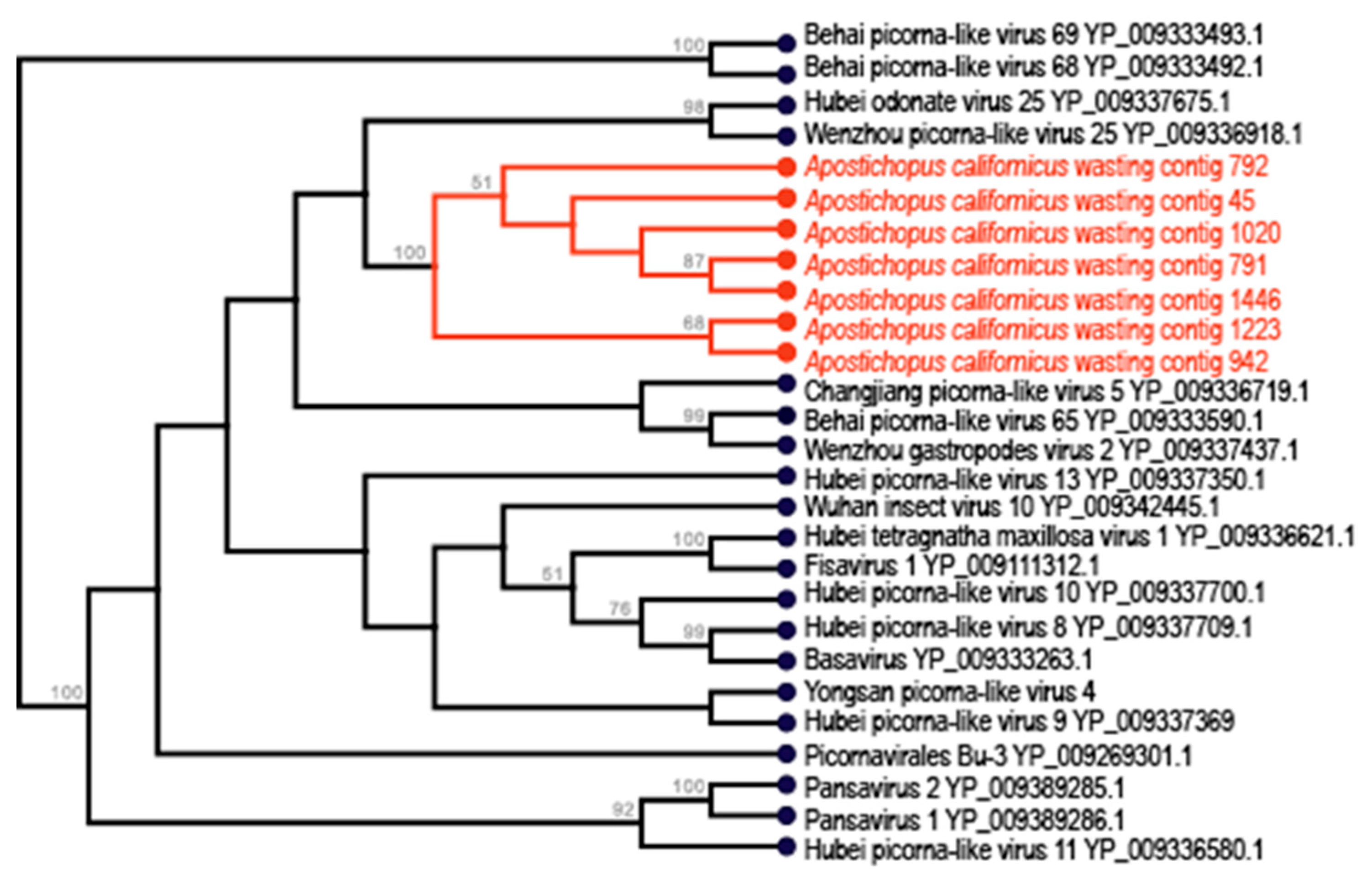
3.2. Picornavirales-Like Genome Fragments
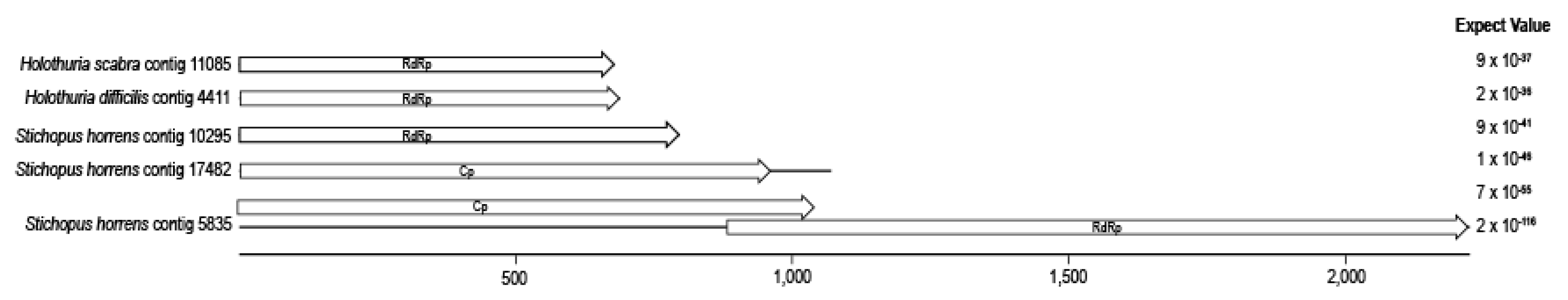
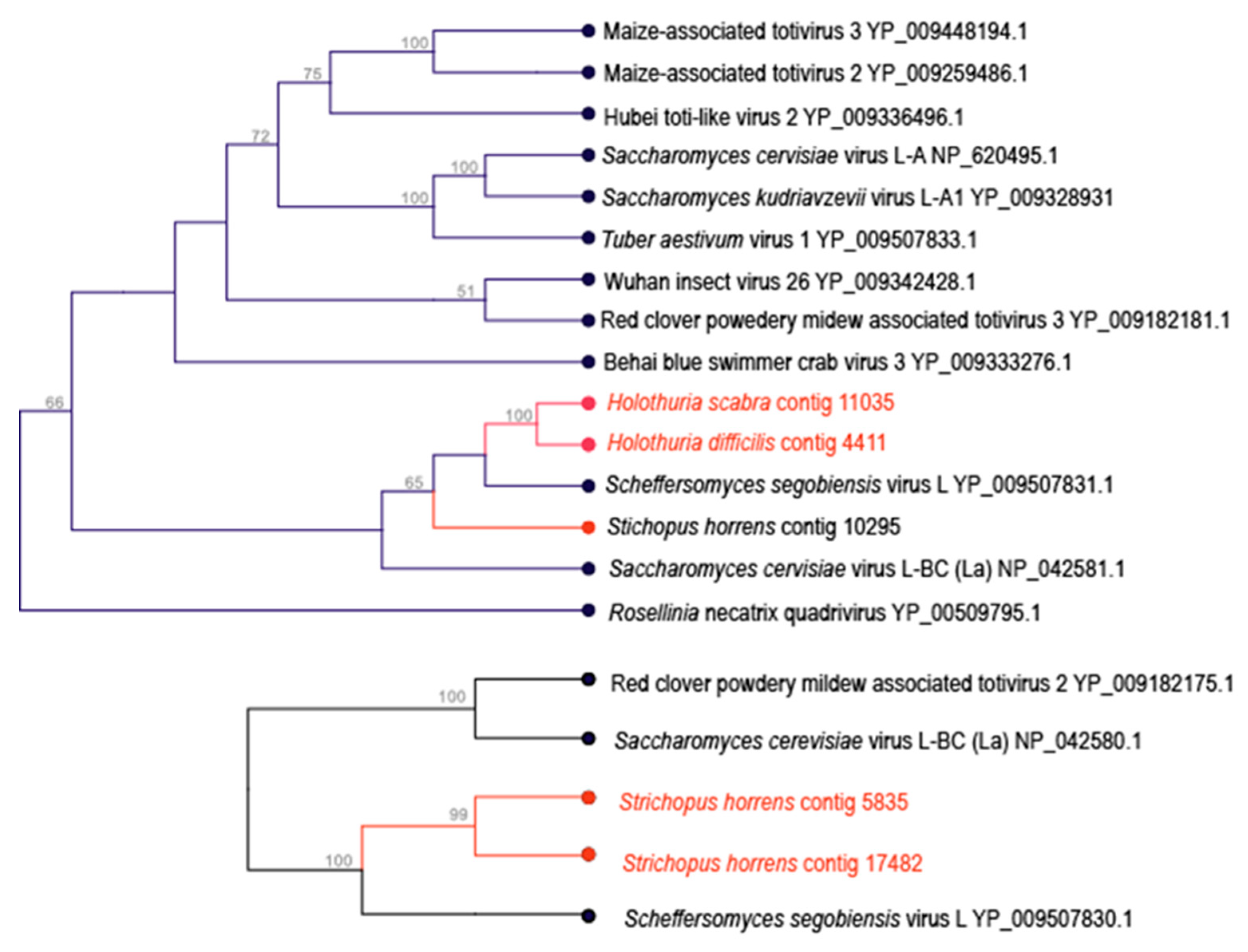
3.3. Totivirus-Like Genome Fragments
3.4. Viral Diseases in Holothurians
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Thurber, R.L.V.; Barott, K.L.; Hall, D.; Liu, H.; Rodriguez-Mueller, B.; Desnues, C.; Edwards, R.A.; Haynes, M.; Angly, F.E.; Wegley, L.; et al. Metagenomic analysis indicates that stressors induce production of herpes-like viruses in the coral Porites compressa. Proc. Natl. Acad. Sci. USA 2008, 105, 18413–18418. [Google Scholar] [CrossRef] [PubMed]
- van Oppen, M.J.H.; Leong, J.A.; Gates, R.D. Coral-virus interactions: A double-edged sword? Symbiosis 2009, 47, 1–8. [Google Scholar] [CrossRef]
- Thurber, R.L.V.; Correa, A.M.S. Viruses of reef-building scleractinian corals. J. Exp. Mar. Biol. Ecol. 2011, 408, 102–113. [Google Scholar] [CrossRef]
- Correa, A.M.S.; Welsh, R.M.; Thurber, R.L.V. Unique nucleocytoplasmic dsDNA and +ssRNA viruses are associated with the dinoflagellate endosymbionts of corals. ISME J. 2013, 7, 13–27. [Google Scholar] [CrossRef]
- Soffer, N.; Brandt, M.E.; Correa, A.M.S.; Smith, T.B.; Thurber, R.V. Potential role of viruses in white plague coral disease. ISME J. 2014, 8, 271–283. [Google Scholar] [CrossRef]
- Levin, R.A.; Voolstra, C.R.; Weynberg, K.D.; van Oppen, M.J.H. Evidence for a role of viruses in the thermal sensitivity of coral photosymbionts. ISME J. 2017, 11, 808–812. [Google Scholar] [CrossRef]
- Breitbart, M.; Benner, B.E.; Jernigan, P.E.; Rosario, K.; Birsa, L.M.; Harbeitner, R.C.; Fulford, S.; Graham, C.; Walters, A.; Goldsmith, D.B.; et al. Discovery, prevalence, and persistence of novel circular single-stranded DNA viruses in the ctenophores Mnemiopsis leidyi and Beroe ovata. Front. Microbiol. 2015, 6, 1427. [Google Scholar] [CrossRef]
- Bistolas, K.; Besemer, R.; Rudstam, L.; Hewson, I. Distribution and inferred evolutionary characteristics of a chimeric ssDNA virus associated with intertidal marine isopods. Viruses 2017, 9, 361. [Google Scholar] [CrossRef]
- Bistolas, K.S.I.; Rudstam, L.G.; Hewson, I. Gene expression of benthic amphipods (genus: Diporeia) in relation to a circular ssDNA virus across two Laurentian Great Lakes. PeerJ 2017, 5, e3810. [Google Scholar] [CrossRef]
- Dunlap, D.S.; Ng, T.F.F.; Rosario, K.; Barbosa, J.G.; Greco, A.M.; Breitbart, M.; Hewson, I. Molecular and microscopic evidence of viruses in marine copepods. Proc. Natl. Acad. Sci. USA 2013, 110, 1375–1380. [Google Scholar] [CrossRef]
- Gudenkauf, B.M.; Eaglesham, J.B.; Aragundi, W.M.; Hewson, I. Discovery of urchin-associated densoviruses (Parvoviridae) in coastal waters of the Big Island, Hawaii. J. Gen. Virol. 2014, 95, 652–658. [Google Scholar] [CrossRef]
- Hewson, I.; Button, J.B.; Gudenkauf, B.M.; Miner, B.; Newton, A.L.; Gaydos, J.K.; Wynne, J.; Groves, C.J.; Hendler, G.; Murray, M.; et al. Densovirus associated with sea-star wasting disease and mass mortality. Proc. Natl. Acad. Sci. USA 2014, 111, 17276–17283. [Google Scholar] [CrossRef]
- Gudenkauf, B.M.; Hewson, I. Comparative metagenomics of viral assemblages inhabiting four phyla of marine invertebrates. Front. Mar. Sci. 2016, 3, 23. [Google Scholar] [CrossRef]
- Jackson, E.W.; Pepe-Ranney, C.; Johnson, M.R.; Distel, D.L.; Hewson, I. A highly prevalent and pervasive densovirus discovered among sea stars from the north american Atlantic coast. Appl. Environ. Microbiol. 2020, 86, e02723-19. [Google Scholar] [CrossRef] [PubMed]
- Rosario, K.; Marinov, M.; Stainton, D.; Kraberger, S.; Wiltshire, E.J.; Collings, D.A.; Walters, M.; Martin, D.P.; Breitbart, M.; Varsani, A. Dragonfly cyclovirus, a novel single-stranded DNA virus discovered in dragonflies (Odonata: Anisoptera). J. Gen. Virol. 2011, 92, 1302–1308. [Google Scholar] [CrossRef] [PubMed]
- Hewson, I. Technical pitfalls that bias comparative microbial community analyses of aquatic disease. Dis. Aquat. Organ. 2019, 137, 109–124. [Google Scholar] [CrossRef] [PubMed]
- Paine, R.T. Food web complexity and species diversity. Am. Nat. 1966, 100, 65. [Google Scholar] [CrossRef]
- Uthicke, S. Nutrient regeneration by abundant coral reef holothurians. J. Exp. Mar. Biol. Ecol. 2001, 265, 153–170. [Google Scholar] [CrossRef]
- Billett, D.S.M.; Bett, B.J.; Rice, A.L.; Thurston, M.H.; Galeron, J.; Sibuet, M.; Wolff, G.A. Long-term change in the megabenthos of the Porcupine Abyssal Plain (NE Atlantic). Prog. Oceanogr. 2001, 50, 325–348. [Google Scholar] [CrossRef]
- Moriarty, D.J.W.; Pollard, P.C.; Hunt, W.G.; Moriarty, C.M.; Wassenberg, T.G. Productivity of bacteria and microalgae and the effect of grazing by holothurians in sediments on a coral reef flat. Mar. Biol. 1985, 85, 293–300. [Google Scholar] [CrossRef]
- Hewson, I.; Fuhrman, J.A. Spatial and vertical biogeography of coral reef sediment bacterial and diazotroph communities. Mar. Ecol. Prog. Ser. 2006, 306, 79–86. [Google Scholar] [CrossRef]
- Pernet, B. Larval Feeding: Mechanisms, Rates, and Performance in Nature; Oxford Univ Press: New York, NY, USA, 2018; pp. 87–102. [Google Scholar]
- Ameneiro, J.; Lubian, L.M.; Sangra, P.; Vazquez, E. Food-limited invertebrate larvae in the Southern Ocean: Testing a paradigm. Mar. Ecol. Prog. Ser. 2016, 554, 71–80. [Google Scholar] [CrossRef]
- Almeda, R.; Messmer, A.M.; Sampedro, N.; Gosselin, L.A. Feeding rates and abundance of marine invertebrate planktonic larvae under harmful algal bloom conditions off Vancouver Island. Harmful Algae 2011, 10, 194–206. [Google Scholar] [CrossRef]
- Toral-Granda, V.; Lovatelli, A.; Vasconcellos, M. Sea Cucumbers: A Global Review of Fisheries and Trade; Food and Agriculture Organization of the United Nations: Rome, Italy, 2008; p. 331. [Google Scholar]
- Liu, X. Chinese Sea Cucumber Farming. Available online: http://www.fao.org/in-action/globefish/fishery-information/resource-detail/en/c/1113389/ (accessed on 24 December 2018).
- Matrosovich, M.; Herrler, G.; Klenk, H.D. Sialic acid receptors of viruses. In SialoGlyco Chemistry and Biology II: Tools and Techniques to Identify and Capture Sialoglycans; Gerardy-Schahn, R., Delannoy, P., von Itzstein, M., Eds.; Springer International Publishing: Cham, Germany, 2015; pp. 1–28. [Google Scholar]
- Varki, A.; Schnaar, R.L.; Schauer, R. Sialic acids and other nonulosonic acids. In Essentials of Glycobiology; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Prestegard, J.H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2015; pp. 179–195. [Google Scholar]
- Smith, L.C.; Ghosh, J.; Buckley, K.M.; Clow, L.A.; Dheilly, N.M.; Haug, T.; Henson, J.H.; Li, C.; Lun, C.M.; Majeske, A.J.; et al. Echinoderm immunity. Adv. Exp. Med. Biol. 2010, 708, 260–301. [Google Scholar]
- Kondo, M.; Akasaka, K. Current status of echinoderm genome analysis—What do we know? Curr. Genom. 2012, 13, 134–143. [Google Scholar] [CrossRef]
- Jackson, E.W.; Bistolas, K.S.I.; Button, J.B.; Hewson, I. Novel circular single-stranded DNA viruses among an asteroid, echinoid and holothurian (Phylum: Echinodermata). PLoS ONE 2016, 11, e0166093. [Google Scholar] [CrossRef]
- Fahsbender, E.; Hewson, I.; Rosario, K.; Tuttle, A.D.; Varsani, A.; Breitbart, M. Discovery of a novel circular DNA virus in the Forbes sea star, Asterias forbesi. Arch. Virol. 2015, 160, 2349–2351. [Google Scholar] [CrossRef]
- Hewson, I.; Bistolas, K.S.I.; Quijano Carde, E.M.; Button, J.B.; Foster, P.J.; Flanzenbaum, J.M.; Kocian, J.; Lewis, C.K. Investigating the complex association between viral ecology, environment and Northeast Pacific Sea Star Wasting. Front. Mar. Sci. 2018, 5, 77. [Google Scholar] [CrossRef]
- WoRMS Apostichopus californicus (Stimpson, 1857). Available online: https://www.marinespecies.org/aphia.php?p=taxdetails&id=529363 (accessed on 14 August 2020).
- Thurber, R.V.; Haynes, M.; Breitbart, M.; Wegley, L.; Rohwer, F. Laboratory procedures to generate viral metagenomes. Nat. Protocol. 2009, 4, 470–483. [Google Scholar] [CrossRef]
- Ng, F.F.T.; Wheeler, E.; Greig, D.; Waltzek, T.B.; Gulland, F.; Breitbart, M. Metagenomic identification of a novel anellovirus in Pacific harbor seal (Phoca vitulina richardsii) lung samples and its detection in samples from multiple years. J. Gen. Virol. 2011, 92, 1318–1323. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Williams, G. GetORF; EMBOSS Wellcome Trust Genome Campus: Cambridge, UK, 2000. [Google Scholar]
- Kelley, L.A.; Sternberg, M.J.E. Protein structure prediction on the Web: A case study using the Phyre server. Nat. Protoc. 2009, 4, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef]
- McGuffin, L.J.; Bryson, K.; Jones, D.T. The PSIPRED protein structure prediction server. Bioinformatics 2000, 16, 404–405. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef]
- Parrish, C.R.; Holmes, E.C.; Morens, D.M.; Park, E.-C.; Burke, D.S.; Calisher, C.H.; Laughlin, C.A.; Saif, L.J.; Daszak, P. Cross-species virus transmission and the emergence of new epidemic diseases. Microbiol. Mol. Biol. Rev. 2008, 72, 457–470. [Google Scholar] [CrossRef]
- Junglen, S.; Korries, M.; Grasse, W.; Wieseler, J.; Kopp, A.; Hermanns, K.; León-Juárez, M.; Drosten, C.; Kümmerer, B.M. Host range restriction of insect-specific flaviviruses occurs at several levels of the viral life cycle. mSphere 2017, 2, e00375-16. [Google Scholar] [CrossRef]
- Firth, A.E.; Blitvich, B.J.; Wills, N.M.; Miller, C.L.; Atkins, J.F. Evidence for ribosomal frameshifting and a novel overlapping gene in the genomes of insect-specific flaviviruses. Virology 2010, 399, 153–166. [Google Scholar] [CrossRef]
- Clyde, K.; Harris, E. RNA secondary structure in the coding region of dengue virus type 2 directs translation start codon selection and is required for viral replication. J. Virol. 2006, 80, 2170–2182. [Google Scholar] [CrossRef]
- Clyde, K.; Barrera, J.; Harris, E. The capsid-coding region hairpin element (cHP) is a critical determinant of dengue virus and West Nile virus RNA synthesis. Virology 2008, 379, 314–323. [Google Scholar] [CrossRef]
- Jones, C.T.; Ma, L.; Burgner, J.W.; Groesch, T.D.; Post, C.B.; Kuhn, R.J. Flavivirus capsid is a dimeric alpha-helical protein. J. Virol. 2003, 77, 7143–7149. [Google Scholar] [CrossRef] [PubMed]
- Kenney, J.L.; Solberg, O.D.; Langevin, S.A.; Brault, A.C. Characterization of a novel insect-specific flavivirus from Brazil: Potential for inhibition of infection of arthropod cells with medically important flaviviruses. J. Gen. Virol. 2014, 95 Pt 12, 2796–2808. [Google Scholar] [CrossRef]
- Parry, R.; Asgari, S. Discovery of novel crustacean and cephalopod flaviviruses: Insights into the evolution and circulation of flaviviruses between marine invertebrate and vertebrate hosts. J. Virol. 2019, 93, e00432-19. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Lin, X.-D.; Vasilakis, N.; Tian, J.-H.; Li, C.-X.; Chen, L.-J.; Eastwood, G.; Diao, X.-N.; Chen, M.-H.; Chen, X.; et al. Divergent viruses discovered in arthropods and vertebrates revise the evolutionary history of the Flaviviridae and related viruses. J. Virol. 2016, 90, 659. [Google Scholar] [CrossRef]
- Mordecai, G.J.; Hewson, I. Coronaviruses in the sea. Front. Microbiol. 2020, 11, 1795. [Google Scholar] [CrossRef]
- Kuno, G. Host range specificity of flaviviruses: Correlation with in vitro replication. J. Med. Entomol. 2007, 44, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Skoge, R.H.; Brattespe, J.; Økland, A.L.; Plarre, H.; Nylund, A. New virus of the family Flaviviridae detected in lumpfish (Cyclopterus lumpus). Arch. Virol. 2018, 163, 679–685. [Google Scholar] [CrossRef]
- Soto, E.; Camus, A.; Yun, S.; Kurobe, T.; Leary, J.H.; Rosser, T.G.; Dill-Okubo, J.A.; Nyaoke, A.C.; Adkison, M.; Renger, A.; et al. First isolation of a novel aquatic flavivirus from Chinook Salmon (Oncorhynchus tshawytscha) and its in vivo replication in a piscine animal model. J. Virol. 2020, 94, e00337-20. [Google Scholar] [CrossRef]
- ICTV. Order—Picornavirales. In Virus Taxonomy; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier: San Diego, CA, USA, 2012; pp. 835–839. [Google Scholar]
- Lau, S.K.P.; Woo, P.C.Y.; Lai, K.K.Y.; Huang, Y.; Yip, C.C.Y.; Shek, C.-T.; Lee, P.; Lam, C.S.F.; Chan, K.-H.; Yuen, K.-Y. Complete genome analysis of three novel picornaviruses from diverse bat species. J. Virol. 2011, 85, 8819–8828. [Google Scholar] [CrossRef] [PubMed]
- Christian, P.D.; Scotti, P.D. Dicistroviruses. Encycl. Virol. 2008, 37–44. [Google Scholar] [CrossRef]
- Wolf, Y.I.; Silas, S.; Wang, Y.; Wu, S.; Bocek, M.; Kazlauskas, D.; Krupovic, M.; Fire, A.; Dolja, V.V.; Koonin, E.V. Doubling of the known set of RNA viruses by metagenomic analysis of an aquatic virome. Nat. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Sanborn, M.A.; Klein, T.A.; Kim, H.-C.; Fung, C.K.; Figueroa, K.L.; Yang, Y.; Asafo-adjei, E.A.; Jarman, R.G.; Hang, J. Metagenomic analysis reveals three novel and prevalent mosquito viruses from a single pool of Aedes vexans nipponii collected in the Republic of Korea. Viruses 2019, 11, 222. [Google Scholar] [CrossRef] [PubMed]
- Hewson, I.; Bistolas, K.S.I.; Button, J.B.; Jackson, E.W. Occurrence and seasonal dynamics of RNA viral genotypes in three contrasting temperate lakes. PLoS ONE 2018, 13, e0194419. [Google Scholar] [CrossRef]
- Miranda, J.A.; Culley, A.I.; Schvarcz, C.R.; Steward, G.F. RNA viruses as major contributors to Antarctic virioplankton. Environ. Microbiol. 2016, 18, 3714–3727. [Google Scholar] [CrossRef]
- Culley, A.I.; Lang, A.S.; Suttle, C.A. The complete genomes of three viruses assembled from shotgun libraries of marine RNA virus communities. Virol. J. 2007, 4, 69. [Google Scholar] [CrossRef]
- Culley, A.I.; Steward, G.F. New genera of RNA viruses in subtropical seawater, inferred from polymerase gene sequences. Appl. Environ. Microbiol. 2007, 73, 5937–5944. [Google Scholar] [CrossRef]
- Culley, A.I.; Lang, A.S.; Suttle, C.A. High diversity of unknown picorna-like viruses in the sea. Nature 2003, 424, 1054–1057. [Google Scholar] [CrossRef]
- Culley, A.I.; Lang, A.S.; Suttle, C.A. Metagenomic analysis of coastal RNA virus communities. Science 2006, 312, 1795–1798. [Google Scholar] [CrossRef]
- Le Gall, O.; Christian, P.; Fauquet, C.M.; King, A.M.Q.; Knowles, N.J.; Nakashima, N.; Stanway, G.; Gorbalenya, A.E. Picornavirales, a proposed order of positive-sense single-stranded RNA viruses with a pseudo-T = 3 virion architecture. Arch. Virol. 2008, 153, 715. [Google Scholar] [CrossRef] [PubMed]
- Ghabrial, S.A. Totiviruses. In Encyclopedia of Virology, 3rd ed.; Mahy, B.W.J., Van Regenmortel, M.H.V., Eds.; Academic Press: Oxford, UK, 2008; pp. 163–174. [Google Scholar]
- Jones, D.T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 1999, 292, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, K.; Miyazaki, N.; Larsson, D.S.D.; Kobayashi, D.; Svenda, M.; Mühlig, K.; Maia, F.R.N.C.; Gunn, L.H.; Isawa, H.; Kobayashi, M.; et al. The infectious particle of insect-borne totivirus-like Omono River virus has raised ridges and lacks fibre complexes. Sci. Rep. 2016, 6, 33170. [Google Scholar] [CrossRef]
- Yarden, O. Fungal association with sessile marine invertebrates. Front. Microbiol. 2014, 5, 228. [Google Scholar] [CrossRef]
- Alker, A.P.; Smith, G.W.; Kim, K. Characterization of Aspergillus sydowii (Thom et Church), a fungal pathogen of Caribbean sea fan corals. Hydrobiologia 2001, 460, 105–111. [Google Scholar] [CrossRef]
- Pivkin, M.V. Filamentous fungi associated with holothurians from the sea of Japan, off the primorye coast of Russia. Biol. Bull. 2000, 198, 101–109. [Google Scholar] [CrossRef]
- Zhuravleva, O.I.; Antonov, A.S.; Oleinikova, G.K.; Khudyakova, Y.V.; Popov, R.S.; Denisenko, V.A.; Pislyagin, E.A.; Chingizova, E.A.; Afiyatullov, S.S. Virescenosides from the holothurian-associated fungus Acremonium striatisporum Kmm 4401. Mar. Drugs 2019, 17, 616. [Google Scholar] [CrossRef]
- Marchese, P.; Garzoli, L.; Gnavi, G.; O’Connell, E.; Bouraoui, A.; Mehiri, M.; Murphy, J.M.; Varese, G.C. Diversity and bioactivity of fungi associated with the marine sea cucumber Holothuria poli: Disclosing the strains potential for biomedical applications. J. Appl. Microbiol. 2020, 129, 612–625. [Google Scholar] [CrossRef]
- Ismail, H.; Lemriss, S.; Ben Aoun, Z.; Mhadhebi, L.; Dellai, A.; Kacem, Y.; Boiron, P.; Bouraoui, A. Antifungal activity of aqueous and methanolic extracts from the Mediterranean sea cucumber, Holothuria polii. J. Mycol. Méd. 2008, 18, 23–26. [Google Scholar] [CrossRef]
- Adibpour, N.; Nasr, F.; Nematpour, F.; Shakouri, A.; Ameri, A. Antibacterial and antifungal activity of Holothuria leucospilota isolated from Persian Gulf and Oman Sea. Jundishapur J. Microbiol. 2014, 7, e8708. [Google Scholar] [CrossRef]
- Nerva, L.; Forgia, M.; Ciuffo, M.; Chitarra, W.; Chiapello, M.; Vallino, M.; Varese, G.C.; Turina, M. The mycovirome of a fungal collection from the sea cucumber Holothuria polii. Virus Res. 2019, 273, 197737. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, L. Wasting-like lesions occurring on California Sea Cucumbers. Dredg 2017, 57, 3. [Google Scholar]
- Fankboner, P.V.; Cameron, J.L. Seasonal atrophy of the visceral organs in a sea cucumber. Can. J. Zool. 1985, 63, 2888–2892. [Google Scholar] [CrossRef]
- García-Arrarás, J.E.; Greenberg, M.J. Visceral regeneration in holothurians. Microsc. Res. Tech. 2001, 55, 438–451. [Google Scholar] [CrossRef] [PubMed]
- Deng, H.; Zhou, Z.-C.; Wang, N.-B.; Liu, C. The syndrome of sea cucumber (Apostichopus japonicus) infected by virus and bacteria. Virol. Sin. 2008, 23, 63–67. [Google Scholar] [CrossRef]
- Wang, P.; Chang, Y.; Yu, J.; Li, C.; Xu, G. Acute peristome edema disease in juvenile and adult sea cucumbers Apostichopus japonicus (Selenka) reared in North China. J. Invertebr. Pathol. 2007, 96, 11–17. [Google Scholar] [CrossRef]
- Liu, H.; Zheng, F.; Sun, X.; Hong, X.; Dong, S.; Wang, B.; Tang, X.; Wang, Y. Identification of the pathogens associated with skin ulceration and peristome tumescence in cultured sea cucumbers Apostichopus japonicus (Selenka). J. Invertebr. Pathol. 2010, 105, 236–242. [Google Scholar] [CrossRef]
| Species | Family | Sample Location | Latitude | Longitude | Collection Date | Sequence Library Size (reads) | # Contigs | N50 Contigs | Contigs Matching Viral Genomes E < 10−20 | Reads Mapped to Viral Contigs |
|---|---|---|---|---|---|---|---|---|---|---|
| Cucumaria miniata | Cucumariidae | Salish Sea | 48.2333 N | 122.7917 W | 7 January 2016 | 1,686,849 | 5394 | 811 | 3 | 247,494 |
| Apostichopus californicus * | Stichopodidae | Ketchikan | 55.4351 N | 131.9481 W | 26 October 2016 | 897,618 | 3431 | 804 | 11 | 195,230 |
| Apostichopus californicus | Stichopodidae | Ketchikan | 55.4351 N | 131.9481 W | 26 October 2016 | 1,019,071 | 3900 | 880 | 2 | 179,175 |
| Holothuria scabra | Holothuridae | Amity Banks | 27.4115 S | 153.4344 E | 10 December 2015 | 822,244 | 21,777 | 838 | 1 | 368 |
| Synaptula recta | Synaptidae | Dunwich | 27.4952 S | 153.3985 E | 10 December 2015 | 983,151 | 19,760 | 847 | 0 | 299 |
| Holothuria difficilis | Holothuridae | Amity Banks | 27.4115 S | 153.4344 E | 10 December 2015 | 794,579 | 5803 | 846 | 1 | 301 |
| Stichopus horrens | Stichopodidae | Amity Banks | 27.4115 S | 153.4346 E | 10 December 2015 | 1,577,532 | 21,194 | 817 | 2 | 423 |
| Holothuria atra | Holothuridae | Heron Island | 23.4441 S | 151.9113 E | 19 December 2015 | 875,840 | 18,866 | 793 | 1 | 345 |
| Holothuria pardalis | Holothuridae | Dunwich | 27.4952 S | 153.3985 E | 10 December 2015 | 1,467,795 | 7424 | 896 | 0 | 306 |
| Species of Metagenome | Contig # | Viral Order | Contig Length (nt) | Holothurian Species Recruited | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sh | Ha | Hs | Hp | Hd | Sr | Cm | Ac | Ac * | ||||
| Stichopus horrens | 17,482 | Ghabrivirales | 1070 | + | ||||||||
| Stichopus horrens | 5835 | Ghabrivirales | 2224 | + | + | + | + | + | + | + | ||
| Stichopus horrens | 10,295 | Ghabrivirales | 796 | + | + | + | + | + | ||||
| Holothuria atra | 17,799 | Picornavirales | 793 | + | ||||||||
| Holothuria scabra | 11,085 | Ghabrivirales | 679 | + | ||||||||
| Holothuria difficilis | 4411 | Ghabrivirales | 688 | + | + | + | + | |||||
| Cucumaria miniata | 4304 | Picornavirales | 636 | + | ||||||||
| Cucumaria miniata | 1913 | Picornavirales | 1826 | + | ||||||||
| Cucumaria miniata | 1781 | Picornavirales | 782 | + | ||||||||
| Apostichopus californicus | 91 | Amarillovirales | 8883 | + | + | + | ||||||
| Apostichopus californicus | 3426 | Picornavirales | 1158 | + | + | |||||||
| Apostichopus californicus * | 942 | Picornavirales | 1156 | + | + | + | ||||||
| Apostichopus californicus * | 792 | Picornavirales | 767 | + | + | |||||||
| Apostichopus californicus * | 791 | Picornavirales | 1491 | + | + | |||||||
| Apostichopus californicus * | 708 | Picornavirales | 1241 | + | + | + | + | + | + | + | + | + |
| Apostichopus californicus * | 557 | Picornavirales | 789 | + | + | |||||||
| Apostichopus californicus * | 45 | Picornavirales | 1368 | + | + | |||||||
| Apostichopus californicus * | 1580 | Picornavirales | 854 | + | + | |||||||
| Apostichopus californicus * | 1446 | Picornavirales | 597 | + | + | |||||||
| Apostichopus californicus * | 1223 | Picornavirales | 810 | + | ||||||||
| Apostichopus californicus * | 1120 | Picornavirales | 1132 | + | + | |||||||
| Apostichopus californicus * | 1020 | Picornavirales | 934 | + | + | |||||||
| Group | Taxonomic Assignment | Holothurian Species | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Holothuria pardalis | Holothuria difficilis | Holothuria scabra | Holothuria atra | Synaptula recta | Stichopus horrens | Cucumaria miniata | Apostichopus californicus | Apostichopus californicus * | ||
| Unicellular Eukaryotes | Alveolata | 1 | 2 | 6 | 8 | |||||
| Apusozoa | 2 | |||||||||
| Amoebozoa | 2 | |||||||||
| Cryptomonada | 1 | 9 | ||||||||
| Heterokonta | 1 | |||||||||
| Metamonada | 1 | |||||||||
| Viridiplantae | 2 | 2 | 2 | 63 | 11 | 18 | 6 | 1 | ||
| Fungi | Ascomycota | 17 | 14 | 6 | 7 | 8 | 17 | 14 | 2 | 21 |
| Basidiomycota | 11 | 8 | 6 | 9 | 8 | 13 | 14 | 12 | 16 | |
| Fungi intercae sedis | 1 | 1 | 2 | 2 | ||||||
| Metazoa | Annelida | 1 | ||||||||
| Arthropoda | 4 | 2 | 4 | 1 | 3 | 4 | 3 | 5 | 5 | |
| Asteroidea | 2 | |||||||||
| Chordata | 3 | 1 | 2 | 2 | 4 | 14 | 3 | 3 | ||
| Cnidaria | 1 | 1 | 1 | 2 | ||||||
| Echinoidea | 1 | |||||||||
| Holothuria | 1 | 1 | 1 | 3 | 1 | 1 | ||||
| Mollusca | 1 | 2 | 1 | 1 | ||||||
| Nematoda | 3 | |||||||||
| Phoronida | 1 | |||||||||
| Platyhelminthes | 1 | |||||||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hewson, I.; Johnson, M.R.; Tibbetts, I.R. An Unconventional Flavivirus and Other RNA Viruses in the Sea Cucumber (Holothuroidea; Echinodermata) Virome. Viruses 2020, 12, 1057. https://doi.org/10.3390/v12091057
Hewson I, Johnson MR, Tibbetts IR. An Unconventional Flavivirus and Other RNA Viruses in the Sea Cucumber (Holothuroidea; Echinodermata) Virome. Viruses. 2020; 12(9):1057. https://doi.org/10.3390/v12091057
Chicago/Turabian StyleHewson, Ian, Mitchell R. Johnson, and Ian R. Tibbetts. 2020. "An Unconventional Flavivirus and Other RNA Viruses in the Sea Cucumber (Holothuroidea; Echinodermata) Virome" Viruses 12, no. 9: 1057. https://doi.org/10.3390/v12091057
APA StyleHewson, I., Johnson, M. R., & Tibbetts, I. R. (2020). An Unconventional Flavivirus and Other RNA Viruses in the Sea Cucumber (Holothuroidea; Echinodermata) Virome. Viruses, 12(9), 1057. https://doi.org/10.3390/v12091057




