Novel Circoviruses Detected in Feces of Sonoran Felids
Abstract
1. Introduction
2. Material and Methods
2.1. Sample Collection and Source Identification
2.2. Fecal Viral Metagenomics
2.3. Recovery of Circovirus Genomes
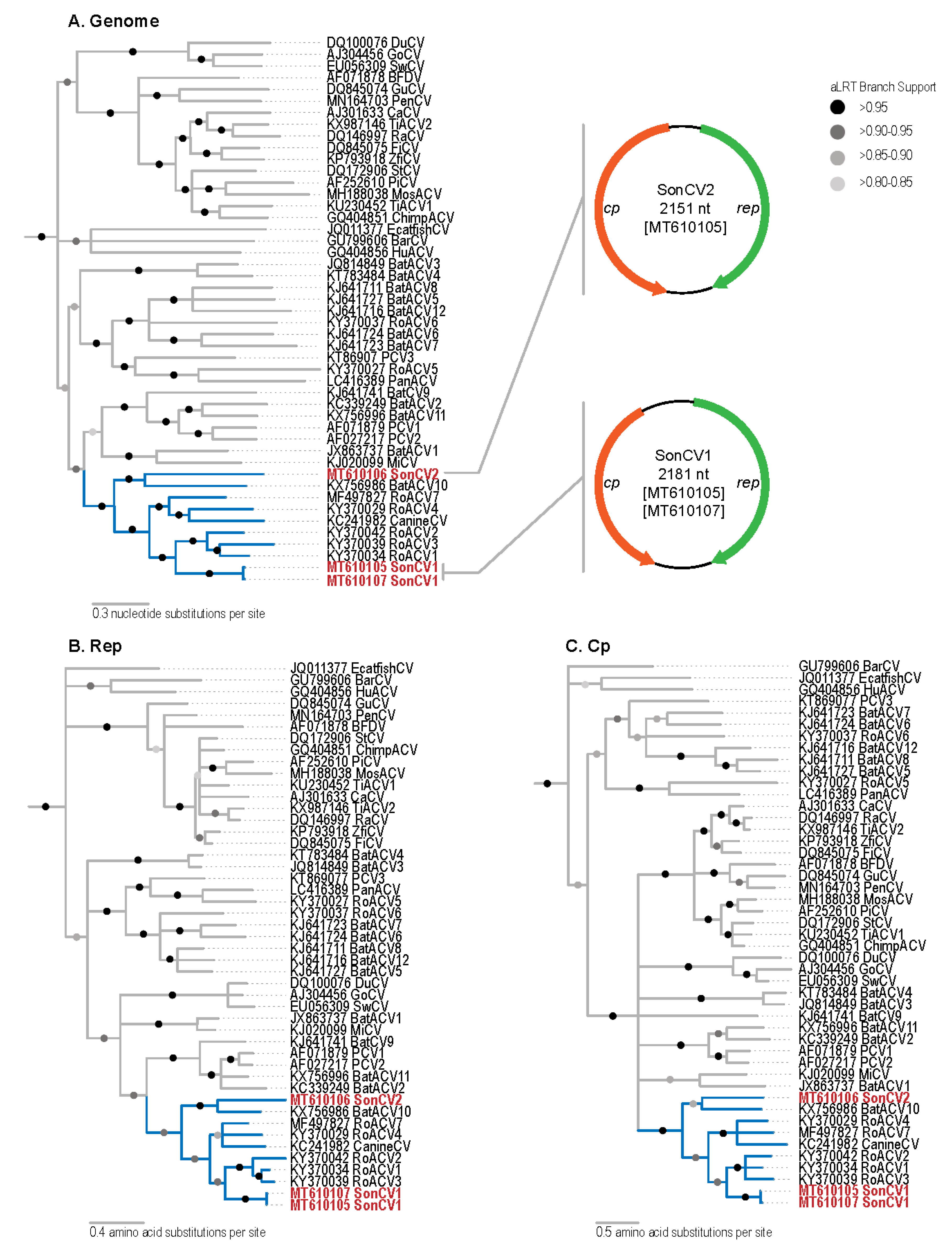
2.4. Sequence Analyses
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Avila-Villegas, S.; Lamberton-Moreno, J. Wildlife Survey and Monitoring in the Sky Island Region with an Emphasis on Neotropical Felids. USDA For. Serv. Proc. 2013, 67, 441–447. [Google Scholar]
- Rosas-Rosas, O.C.; Valdez, R.; Bender, L.C.; Daniel, D. Food Habits of Pumas in Northwestern Sonora, Mexico. Wildl. Soc. Bull. 2003, 31, 528–535. [Google Scholar]
- Luna-Soria, H.; López González, C.A. Abundance and food habits of cougars and bobcats in the Sierra San Luis, Sonora, Mexico. USDA For. Serv. Proc. 2005, 36, 416–420. [Google Scholar]
- Cassaigne, I.; Medellín, R.A.; Thompson, R.W.; Culver, M.; Ochoa, A.; Vargas, K.; Childs, J.L.; Sanderson, J.; List, R.; Torres-Gómez, A. Diet of Pumas (Puma concolor) in Sonora, Mexico, as Determined by GPS Kill Sites and Molecular Identified Scat, with Comments on Jaguar (Panthera onca) Diet. Southwest. Nat. 2016, 61, 125–132. [Google Scholar] [CrossRef]
- De Villa Meza, A.; Martinez Meyer, E.; López González, C.A. Ocelot (Leopardus pardalis) Food Habits in a Tropical Deciduous Forest of Jalisco, Mexico. Am. Midl. Nat. 2002, 148, 146–154. [Google Scholar] [CrossRef]
- McKinney, T.; Smith, T.W. Diets of Sympatric Bobcats and Coyotes During Years of Varying Rainfall in Central Arizona. West North Am. Nat. 2007, 67, 8–15. [Google Scholar] [CrossRef]
- Booth-Binczik, S.D.; Bradley, R.D.; Thompson, C.W.; Bender, L.C.; Huntley, J.W.; Harvey, J.A.; Laack, L.L.; Mays, J.L. Food Habits of Ocelots and Potential for Competition with Bobcats in Southern Texas. Southwest. Nat. 2013, 58, 403–410. [Google Scholar] [CrossRef]
- Fish and Wildlife Service, Interior. Endangered and Threatened Wildlife and Plants; Final Rule To Extend Endangered Status for the Jaguar in the United States. Federal Register 1997, 62, 39147–39157. [Google Scholar]
- SEMARNAT. NORMA Oficial Mexicana NOM-059-SEMARNAT-2010, Protección Ambiental-Especies Nativas de México de Flora y Fauna Silvestres-Categorías de Riesgo y Especificaciones Para su Inclusión, Exclusión o Cambio-Lista de Especies en Riesgo. Available online: https://www.dof.gob.mx/normasOficiales/4254/semarnat/semarnat.htm (accessed on 11 September 2020).
- US Fish and Wildlife Service. Recovery Plan for the Ocelot (Leopardus pardalis) First Revision; US Fish and Wildlife Service Southwest Region Albuquerque: Albuquerque, NM, USA, 2016; p. 217. [Google Scholar]
- Hallack-Alegria, M.; Watkins, D.W. Annual and warm season drought intensity-duration-frequency analysis for Sonora, Mexico. J. Clim. 2007, 20, 1897–1909. [Google Scholar] [CrossRef]
- Thompson, R. (Primero Conservation, Pinetop, AZ, USA). Personal communication, 2020.
- Nicholson, K.L.; Noon, T.H.; Krausman, P.R. Serosurvey of mountain lions in soutern Arizona. Wildl. Soc. Bull 2012, 36, 615–620. [Google Scholar] [CrossRef]
- Rogalski, M.A.; Gowler, C.D.; Shaw, C.L.; Hufbauer, R.A.; Duffy, M.A. Human drivers of ecological and evolutionary dynamics in emerging and disappearing infectious disease systems. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372, 20160043. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, A.A.; Tabor, G.M. Global Factors Driving Emerging Infectious Diseases. Ann. N. Y. Acad. Sci. 2008, 1149, 1–3. [Google Scholar] [CrossRef]
- Chiu, E.S.; Kraberger, S.; Cunningham, M.; Cusack, L.; Roelke, M.; VandeWoude, S. Multiple introductions of domestic cat feline leukemia virus in endangered Florida Panthers. Emerg. Infect. Dis. 2019, 25, 92–101. [Google Scholar] [CrossRef]
- Roelke-Parker, M.E.; Munson, L.; Packer, C.; Kock, R.; Cleaveland, S.; Carpenter, M.; O’Brien, S.J.; Pospischil, A.; Hofmann-Lehmann, R.; Lutz, H.; et al. A canine distemper virus epidemic in Serengeti lions (Panthera leo). Nature 1996, 379, 441–445. [Google Scholar] [CrossRef] [PubMed]
- Czupryna, A.M.; Brown, J.S.; Bigambo, M.A.; Whelan, C.J.; Mehta, S.D.; Santymire, R.M.; Lankester, F.J.; Faust, L.J. Cross-species transmission and evolutionary dynamics of canine distemper virus during a spillover in African lions of Serengeti National Park. Mol. Ecol. 2020, 11, e0167092. [Google Scholar]
- O’Brien, S.J.; Evermann, J.F. Interactive influence of infectious disease and genetic diversity in natural populations. Trends Ecol. Evol. 1988, 3, 254–259. [Google Scholar] [CrossRef]
- Lacy, R.C. Importance of Genetic Variation to the Viability of Mammalian Populations. J. Mammal. 1997, 78, 320–335. [Google Scholar] [CrossRef]
- Heard, M.J.; Smith, K.F.; Ripp, K.J.; Berger, M.; Chen, J.; Dittmeier, J.; Goter, M.; Mcgarvey, S.T.; Ryan, E. The Threat of Disease Increases as Species Move Toward Extinction. Conserv Biol. 2013. [Google Scholar] [CrossRef]
- Krupovic, M.; Varsani, A.; Kazlauskas, D.; Breitbart, M.; Delwart, E.; Rosario, K.; Yutin, N.; Wolf, Y.I.; Harrach, B.; Zerbini, F.M.; et al. Cressdnaviricota: A virus phylum unifying 7 families of Rep-encoding viruses with single-stranded, circular DNA genomes. J. Virol. 2020. [Google Scholar] [CrossRef]
- Zhang, W.; Li, L.; Deng, X.; Kapusinszky, B.; Pesavento, P.A.; Delwart, E. Faecal virome of cats in an animal shelter. J. Gen. Virol. 2014, 95, 2553–2564. [Google Scholar] [CrossRef]
- Takano, T.; Yanai, Y.; Hiramatsu, K.; Doki, T.; Hohdatsu, T. Novel single-stranded, circular DNA virus identified in cats in Japan. Arch. Virol. 2018, 163, 3389–3393. [Google Scholar] [CrossRef] [PubMed]
- Kraberger, S.; Serieys, L.; Fountain-Jones, N.; Packer, C.; Riley, S.; Varsani, A. Novel smacoviruses identified in the faeces of two wild felids: North American bobcat and African lion. Arch. Virol. 2019, 164, 2395–2399. [Google Scholar] [CrossRef] [PubMed]
- Fontenele, R.S.; Lacorte, C.; Lamas, N.S.; Schmidlin, K.; Varsani, A.; Ribeiro, S.G. Single stranded dna viruses associated with capybara faeces sampled in brazil. Viruses 2019, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Chong, R.; Shi, M.; Grueber, C.E.; Holmes, E.C.; Hogg, C.J.; Belov, K.; Barrs, V.R. Fecal Viral Diversity of Captive and Wild Tasmanian Devils Characterized Using Virion-Enriched Metagenomics and Metatranscriptomics. J. Virol. 2019, 93, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Rosario, K.; Breitbart, M.; Harrach, B.; Segalés, J.; Delwart, E.; Biagini, P.; Varsani, A. Revisiting the taxonomy of the family Circoviridae: Establishment of the genus Cyclovirus and removal of the genus Gyrovirus. Arch. Virol. 2017, 162, 1447–1463. [Google Scholar] [CrossRef]
- Li, L.; McGraw, S.; Zhu, K.; Leutenegger, C.M.; Marks, S.L.; Kubiski, S.; Gaffney, P.; Cruz, F.N., Jr.; Wang, C.; Delwart, E.; et al. Circovirus in tissues of dogs with vasculitis and hemorrhage. Emerg. Infect. Dis. 2013, 19, 534–541. [Google Scholar] [CrossRef]
- Decaro, N.; Martella, V.; Desario, C.; Lanave, G.; Circella, E.; Cavalli, A.; Elia, G.; Camero, M.; Buonavoglia, C. Genomic characterization of a circovirus associated with fatal hemorrhagic enteritis in dog, Italy. PLoS ONE 2014, 9, e105909. [Google Scholar] [CrossRef]
- Kotsias, F.; Bucafusco, D.; Nuñez, D.A.; Lago Borisovsky, L.A.; Rodriguez, M.; Bratanich, A.C. Genomic characterization of canine circovirus associated with fatal disease in dogs in South America. PLoS ONE 2019, 14, e0218735. [Google Scholar] [CrossRef]
- Ritchie, B.W.; Niagro, F.D.; Lukert, P.D.; Steffens, W.L.; Latimer, K.S. Characterization of a new virus from cockatoos with psittacine beak and feather disease. Virology 1989, 171, 83–88. [Google Scholar] [CrossRef]
- Chae, C. A review of porcine circovirus 2-associated syndromes and diseases. Vet. J. 2005, 169, 326–336. [Google Scholar] [CrossRef]
- Segalés, J.; Allan, G.M.; Domingo, M. Porcine circovirus diseases. Anim. Heal. Res. Rev. 2005, 6, 119–142. [Google Scholar] [CrossRef] [PubMed]
- Allan, G.M.; Kennedy, S.; McNeilly, F.; Foster, J.C.; Ellis, J.A.; Krakowka, S.J.; Meehan, B.M.; Adair, B.M. Experimental reproduction of severe wasting disease by co-infection of pigs with porcine circovirus and porcine parvovirus. J. Comp. Pathol. 1999, 121, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Opriessnig, T.; Halbur, P.G. Concurrent infections are important for expression of porcine circovirus associated disease. Virus Res. 2012, 164, 20–32. [Google Scholar] [CrossRef] [PubMed]
- Zaccaria, G.; Malatesta, D.; Scipioni, G.; Di Felice, E.; Campolo, M.; Casaccia, C.; Savini, G.; Di Sabatino, D.; Lorusso, A. Circovirus in domestic and wild carnivores: An important opportunistic agent? Virology 2016, 490, 69–74. [Google Scholar] [CrossRef]
- Anderson, A.; Hartmann, K.; Leutenegger, C.M.; Proksch, A.L.; Mueller, R.S.; Unterer, S. Role of canine circovirus in dogs with acute haemorrhagic diarrhoea. Vet. Rec. 2017, 180, 542. [Google Scholar] [CrossRef]
- Dowgier, G.; Lorusso, E.; Decaro, N.; Desario, C.; Mari, V.; Lucente, M.S.; Lanave, G.; Buonavoglia, C.; Elia, G. A molecular survey for selected viral enteropathogens revealed a limited role of Canine circovirus in the development of canine acute gastroenteritis. Vet. Microbiol. 2017, 204, 54–58. [Google Scholar] [CrossRef]
- Katzourakis, A.; Gifford, R.J. Endogenous viral elements in animal genomes. PLoS Genet 2010, 6. [Google Scholar] [CrossRef]
- Dennis, T.P.; de Souza, W.M.; Marsile-Medun, S.; Singer, J.B.; Wilson, S.J.; Gifford, R.J. The evolution, distribution and diversity of endogenous circoviral elements in vertebrate genomes. Virus. Res. 2019, 262, 15–23. [Google Scholar] [CrossRef]
- Verma, S.K.; Singh, L. Novel universal primers establish identity of an enormous number of animal species for forensic application. Mol. Ecol. Notes 2002, 3, 28–31. [Google Scholar] [CrossRef]
- Naidu, A.; Smythe, L.A.; Thompson, R.W.; Culver, M. Genetic analysis of scats reveals minimum number and sex of recently documented mountain lions. J. Fish Wildl. Manag. 2011, 2, 106–111. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinforma. Appl. Note 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic. Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A Virus Classification Tool Based on Pairwise Sequence Alignment and Identity Calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A Simple, Fast, and Accurate Algorithm to Estimate Large Phylogenies by Maximum Likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. JModelTest 2, More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3, fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef]
- Anisimova, M.; Gascuel, O. Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. Syst. Biol. 2006, 55, 539–552. [Google Scholar] [CrossRef] [PubMed]
- Heibl, C. PHYLOCH: R Language Tree Plotting Tools and Interfaces to Diverse Phylogenetic Software Packages. 2008. Available online: http://www.christophheibl.de/Rpackages.html (accessed on 11 September 2020).
- Paradis, E.; Schliep, K. ape 5.0, an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2018, 35, 526–528. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing. 2019. Available online: https://www.r-project.org/ (accessed on 11 September 2020).
- Rosario, K.; Duffy, S.; Breitbart, M. A field guide to eukaryotic circular single-stranded DNA viruses: Insights gained from metagenomics. Arch. Virol. 2012, 157, 1851–1871. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Payne, N.; Kraberger, S.; Fontenele, R.S.; Schmidlin, K.; Bergeman, M.H.; Cassaigne, I.; Culver, M.; Varsani, A.; Van Doorslaer, K. Novel Circoviruses Detected in Feces of Sonoran Felids. Viruses 2020, 12, 1027. https://doi.org/10.3390/v12091027
Payne N, Kraberger S, Fontenele RS, Schmidlin K, Bergeman MH, Cassaigne I, Culver M, Varsani A, Van Doorslaer K. Novel Circoviruses Detected in Feces of Sonoran Felids. Viruses. 2020; 12(9):1027. https://doi.org/10.3390/v12091027
Chicago/Turabian StylePayne, Natalie, Simona Kraberger, Rafaela S Fontenele, Kara Schmidlin, Melissa H Bergeman, Ivonne Cassaigne, Melanie Culver, Arvind Varsani, and Koenraad Van Doorslaer. 2020. "Novel Circoviruses Detected in Feces of Sonoran Felids" Viruses 12, no. 9: 1027. https://doi.org/10.3390/v12091027
APA StylePayne, N., Kraberger, S., Fontenele, R. S., Schmidlin, K., Bergeman, M. H., Cassaigne, I., Culver, M., Varsani, A., & Van Doorslaer, K. (2020). Novel Circoviruses Detected in Feces of Sonoran Felids. Viruses, 12(9), 1027. https://doi.org/10.3390/v12091027





