Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adélie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula
Abstract
1. Introduction
2. Materials and Methods
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Barbosa, A.; Palacios, M.J. Health of Antarctic birds: A review of their parasites, pathogens and diseases. Polar Biol. 2009, 32, 1095. [Google Scholar] [CrossRef]
- Grimaldi, W.W.; Seddon, P.J.; Lyver, P.O.; Nakagawa, S.; Tompkins, D.M. Infectious diseases of Antarctic penguins: Current status and future threats. Polar Biol. 2015, 38, 591–606. [Google Scholar] [CrossRef]
- Morandini, V.; Dugger, K.M.; Ballard, G.; Elrod, M.; Schmidt, A.; Ruoppolo, V.; Lescroel, A.; Jongsomjit, D.; Massaro, M.; Pennycook, J.; et al. Identification of a novel adelie penguin circovirus at cape crozier (Ross Island, Antarctica). Viruses 2019, 11, 1088. [Google Scholar] [CrossRef]
- Neira, V.; Tapia, R.; Verdugo, C.; Barriga, G.; Mor, S.; Ng, T.F.F.; Garcia, V.; Del Rio, J.; Rodrigues, P.; Briceno, C.; et al. Novel avulaviruses in penguins, Antarctica. Emerg. Infect. Dis. 2017, 23, 1212–1214. [Google Scholar] [CrossRef]
- Olivares, F.; Tapia, R.; Galvez, C.; Meza, F.; Barriga, G.P.; Borras-Chavez, R.; Mena-Vasquez, J.; Medina, R.A.; Neira, V. Novel penguin Avian avulaviruses 17, 18 and 19 are widely distributed in the Antarctic Peninsula. Transbound. Emerg. Dis. 2019, 66, 2227–2232. [Google Scholar] [CrossRef] [PubMed]
- Smeele, Z.E.; Ainley, D.G.; Varsani, A. Viruses associated with Antarctic wildlife: From serology based detection to identification of genomes using high throughput sequencing. Virus Res. 2018, 243, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Wille, M.; Aban, M.; Wang, J.; Moore, N.; Shan, S.; Marshall, J.; Gonzalez-Acuna, D.; Vijaykrishna, D.; Butler, J.; Wang, J.; et al. Antarctic penguins as reservoirs of diversity for avian avulaviruses. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Yinda, C.K.; Esefeld, J.; Peter, H.U.; Matthijnssens, J.; Zell, R. Penguin megrivirus, a novel picornavirus from an Adelie penguin (Pygoscelis adeliae). Arch. Virol. 2019, 164, 2887–2890. [Google Scholar] [CrossRef]
- Alexander, D.J.; Manvell, R.J.; Collins, M.S.; Brockman, S.J.; Westbury, H.A.; Morgan, I.; Austin, F.J. Characterization of paramyxoviruses isolated from penguins in Antarctica and sub-Antarctica during 1976-1979. Arch. Virol. 1989, 109, 135–143. [Google Scholar] [CrossRef]
- Austin, F.J.; Webster, R.G. Evidence of ortho- and paramyxoviruses in fauna from Antarctica. J. Wildl. Dis. 1993, 29, 568–571. [Google Scholar] [CrossRef]
- Baumeister, E.; Leotta, G.; Pontoriero, A.; Campos, A.; Montalti, D.; Vigo, G.; Pecoraro, M.; Savy, V. Serological evidences of influenza A virus infection in Antarctica migratory birds. Int. Congr. Ser. 2004, 1263, 737–740. [Google Scholar] [CrossRef]
- Gardner, H.; Kerry, K.; Riddle, M.; Brouwer, S.; Gleeson, L. Poultry virus infection in Antarctic penguins. Nature 1997, 387, 245. [Google Scholar] [CrossRef] [PubMed]
- Mac Donald, J.W.; Conroy, J.W.H. Virus disease resembling puffinosis in the gentoo penguin Pygoscelis papua on Signy Island South Orkney Islands. Br. Antarct. Surv. Bull. 1971, 80–82. [Google Scholar]
- Morgan, I.R.; Westbury, H.A. Virological studies of Adelie Penguins (Pygoscelis adeliae) in Antarctica. Avian Dis. 1981, 25, 1019–1026. [Google Scholar] [CrossRef] [PubMed]
- Morgan, I.R.; Westbury, H.A. Studies of viruses in penguins in the Vestfold hills. Hydrobiologia 1988, 165, 263–269. [Google Scholar] [CrossRef]
- Morgan, I.R.; Westbury, H.A.; Caple, I.W.; Campbell, J. A survey of virus infection in sub-Antarctic penguins on Macquarie Island, Southern Ocean. Aust. Vet. J. 1981, 57, 333–335. [Google Scholar] [CrossRef] [PubMed]
- Hurt, A.C.; Vijaykrishna, D.; Butler, J.; Baas, C.; Maurer-Stroh, S.; Silva-de-la-Fuente, M.C.; Medina-Vogel, G.; Olsen, B.; Kelso, A.; Barr, I.G.; et al. Detection of evolutionarily distinct avian influenza a viruses in Antarctica. mBio 2014, 5, e01098-14. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Kim, J.H.; Park, Y.M.; Shin, O.S.; Kim, H.; Choi, H.G.; Song, J.W. A novel adenovirus in Chinstrap penguins (Pygoscelis Antarctica) in Antarctica. Viruses 2014, 6, 2052–2061. [Google Scholar] [CrossRef]
- Lee, S.Y.; Kim, J.H.; Seo, T.K.; No, J.S.; Kim, H.; Kim, W.K.; Choi, H.G.; Kang, S.H.; Song, J.W. Genetic and molecular epidemiological characterization of a novel adenovirus in Antarctic penguins collected between 2008 and 2013. PLoS ONE 2016, 11, e0157032. [Google Scholar] [CrossRef]
- Thomazelli, L.M.; Araujo, J.; Oliveira, D.B.; Sanfilippo, L.; Ferreira, C.S.; Brentano, L.; Pelizari, V.H.; Nakayama, C.; Duarte, R.; Hurtado, R.; et al. Newcastle disease virus in penguins from King George Island on the Antarctic region. Vet. Microbiol. 2010, 146, 155–160. [Google Scholar] [CrossRef]
- Varsani, A.; Kraberger, S.; Jennings, S.; Porzig, E.L.; Julian, L.; Massaro, M.; Pollard, A.; Ballard, G.; Ainley, D.G. A novel papillomavirus in Adelie penguin (Pygoscelis adeliae) faeces sampled at the Cape Crozier colony, Antarctica. J. Gen. Virol. 2014, 95, 1352–1365. [Google Scholar] [CrossRef]
- Varsani, A.; Porzig, E.L.; Jennings, S.; Kraberger, S.; Farkas, K.; Julian, L.; Massaro, M.; Ballard, G.; Ainley, D.G. Identification of an avian polyomavirus associated with Adelie penguins (Pygoscelis adeliae). J. Gen. Virol. 2015, 96, 851–857. [Google Scholar] [CrossRef] [PubMed]
- Breitbart, M.; Delwart, E.; Rosario, K.; Segales, J.; Varsani, A.; Ictv Report, C. ICTV virus taxonomy profile: Circoviridae. J. Gen. Virol. 2017, 98, 1997–1998. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, A.; Colominas-Ciuro, R.; Coria, N.; Centurion, M.; Sandler, R.; Negri, A.; Santos, M. First record of feather-loss disorder in Antarctic penguins. Antarct. Sci. 2015, 27, 69–70. [Google Scholar] [CrossRef]
- Gavryushkina, A.; Heath, T.A.; Ksepka, D.T.; Stadler, T.; Welch, D.; Drummond, A.J. Bayesian total-evidence dating reveals the recent crown radiation of Penguins. Syst. Biol. 2017, 66, 57–73. [Google Scholar] [CrossRef]
- Lynch, H.J.; Naveen, R.; Fagan, W.F. Censuses of penguin, blue-eyed shag Phalacrocorax atriceps and southern giant petrel Macronectes giganteus populations on the Antarctic Peninsula, 2001–2007. Mar. Ornithol. 2008, 36, 83–97. [Google Scholar]
- Humphries, G.R.W.; Naveen, R.; Schwaller, M.; Che-Castaldo, C.; McDowall, P.; Schrimpf, M.; Lynch, H.J. Mapping Application for Penguin Populations and Projected Dynamics (MAPPPD): Data and tools for dynamic management and decision support. Polar Rec. 2017, 53, 160–166. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- O’Leary, N.A.; Wright, M.W.; Brister, J.R.; Ciufo, S.; Haddad, D.; McVeigh, R.; Rajput, B.; Robbertse, B.; Smith-White, B.; Ako-Adjei, D.; et al. Reference sequence (RefSeq) database at NCBI: Current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016, 44, D733–D745. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Muscle: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Posada, D. jModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef]
- Stover, B.C.; Muller, K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010, 11, 7. [Google Scholar] [CrossRef]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef]
- Yang, Z.; Wong, W.S.; Nielsen, R. Bayes empirical bayes inference of amino acid sites under positive selection. Mol. Biol. Evol. 2005, 22, 1107–1118. [Google Scholar] [CrossRef] [PubMed]
- Rosario, K.; Breitbart, M.; Harrach, B.; Segales, J.; Delwart, E.; Biagini, P.; Varsani, A. Revisiting the taxonomy of the family Circoviridae: Establishment of the genus Cyclovirus and removal of the genus Gyrovirus. Arch. Virol. 2017, 162, 1447–1463. [Google Scholar] [CrossRef] [PubMed]
- Naveen, R.; Forrest, S.C.; Dagit, R.G.; Blight, L.K.; Trivelpiece, W.Z.; Trivelpiece, S.G. Censuses of penguin, blue-eyed shag, and southern giant petrel populations in the Antarctic Peninsula region, 1994–2000. Polar Rec. 2000, 36, 323–334. [Google Scholar] [CrossRef]
- Julian, L.; Piasecki, T.; Chrzastek, K.; Walters, M.; Muhire, B.; Harkins, G.W.; Martin, D.P.; Varsani, A. Extensive recombination detected among beak and feather disease virus isolates from breeding facilities in Poland. J. Gen. Virol. 2013, 94, 1086–1095. [Google Scholar] [CrossRef] [PubMed]
- Smyth, J.A.; Todd, D.; Scott, A.; Beckett, A.; Twentyman, C.M.; Brojer, C.; Uhlhorn, H.; Gavier-Widen, D. Identification of circovirus infection in three species of gull. Vet. Rec. 2006, 159, 212–214. [Google Scholar] [CrossRef] [PubMed]
- Cimino, M.A.; Lynch, H.J.; Saba, V.S.; Oliver, M.J. Projected asymmetric response of Adelie penguins to Antarctic climate change. Sci. Rep. 2016, 6, 28785. [Google Scholar] [CrossRef]
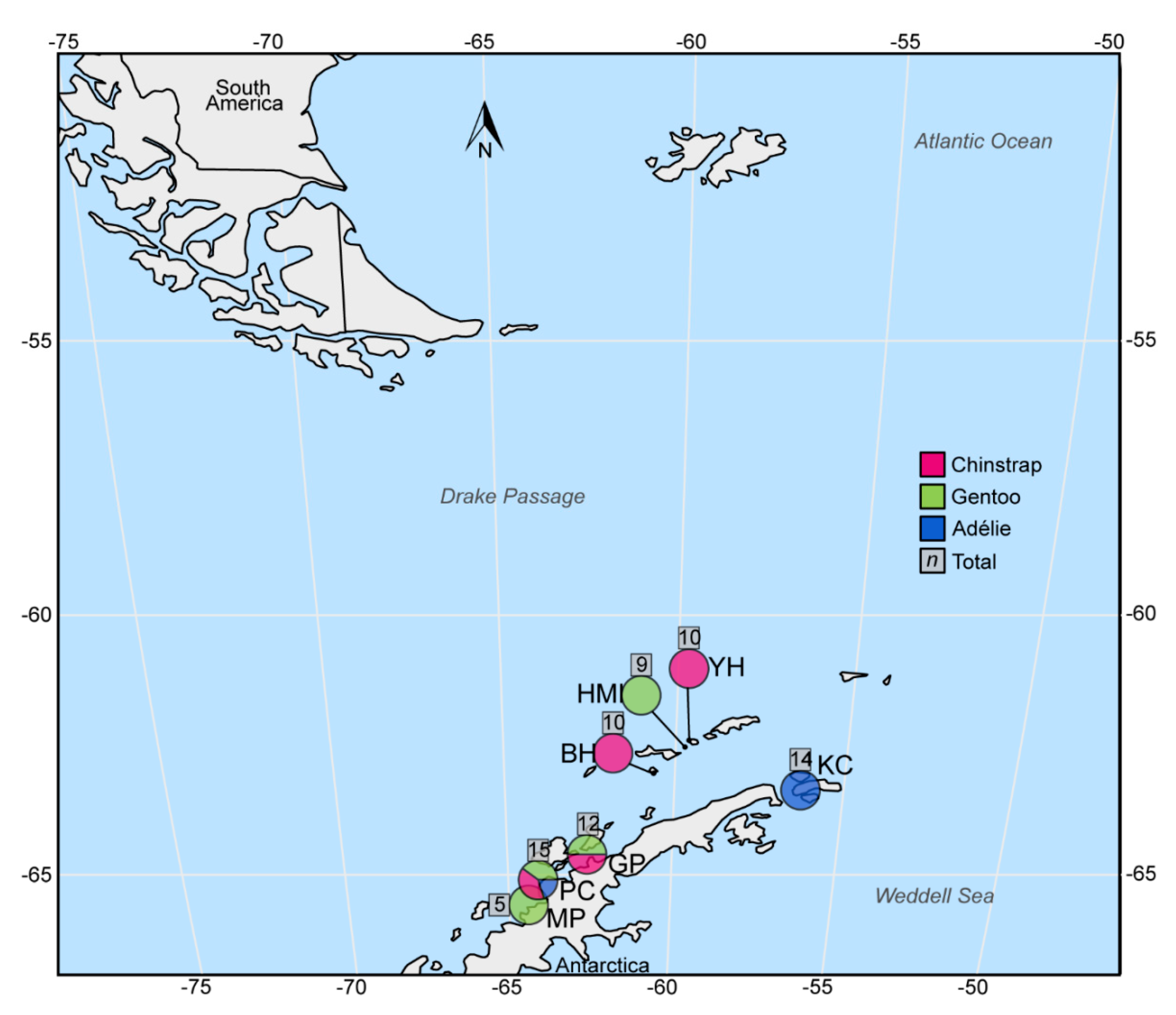
| Colony and Antarctic Site Inventory Subarea | Coordinates | Penguin Species (Individuals Sampled) | PenCV Dentification in Adults Sampled | PenCV Identification in Chicks Sampled |
|---|---|---|---|---|
| Half Moon Island (HMI), South Shetland Islands | 60°36′ S, 59°55′ W | Chinstrap (10) | 0/10 | - |
| Yankee Harbor (YH), Greenwich Island, South Shetland Islands | 62°32′ S, 59°47′ W | Gentoo (9) | 0/6 | 0/3 |
| Baily Head (BH), Deception Island, South Shetland Islands | 62°58′ S, 60°30′ W | Chinstrap (10) | 0/5 | 0/5 |
| Kinnes Cove (KC, Madder Cliff), Joinville Island, Northeast Antarctic Peninsula | 63°18′ S, 56°29′ W | Adélie (14) | 0/7 | 0/7 |
| Georges Point (GP), Ronge Island, Northwest Antarctic Peninsula | 64°40′ S, 62°40′ W | Chinstrap (6) | 0/3 | 0/3 |
| Gentoo (6) | 0/3 | 0/3 | ||
| Port Charcot (PC), Booth Island, Southwest Antarctic Peninsula | 65°05′ S, 64°00′ W | Adélie (3) | 1/2 | 0/1 |
| Chinstrap (6) | 1/3 | 0/3 | ||
| Gentoo (6) | 0/3 | 0/3 | ||
| Moot Point (MP), Southwest Antarctic Peninsula | 65°12′ S, 64°06′ W | Gentoo (5) | 0/4 | 0/1 |
| Total by Species | Adélie (17) | 1/9 | 0/8 | |
| Chinstrap (32) | 1/21 | 0/11 | ||
| Gentoo (26) | 0/16 | 0/10 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Levy, H.; Fiddaman, S.R.; Djurhuus, A.; Black, C.E.; Kraberger, S.; Smith, A.L.; Hart, T.; Varsani, A. Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adélie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula. Viruses 2020, 12, 858. https://doi.org/10.3390/v12080858
Levy H, Fiddaman SR, Djurhuus A, Black CE, Kraberger S, Smith AL, Hart T, Varsani A. Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adélie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula. Viruses. 2020; 12(8):858. https://doi.org/10.3390/v12080858
Chicago/Turabian StyleLevy, Hila, Steven R. Fiddaman, Anni Djurhuus, Caitlin E. Black, Simona Kraberger, Adrian L. Smith, Tom Hart, and Arvind Varsani. 2020. "Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adélie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula" Viruses 12, no. 8: 858. https://doi.org/10.3390/v12080858
APA StyleLevy, H., Fiddaman, S. R., Djurhuus, A., Black, C. E., Kraberger, S., Smith, A. L., Hart, T., & Varsani, A. (2020). Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adélie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula. Viruses, 12(8), 858. https://doi.org/10.3390/v12080858







