Distinct New York City Aedes albopictus Mosquito Populations Display Differences in Salivary Gland Protein D7 Diversity and Chikungunya Virus Replication
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture and Generation of Viral Stocks
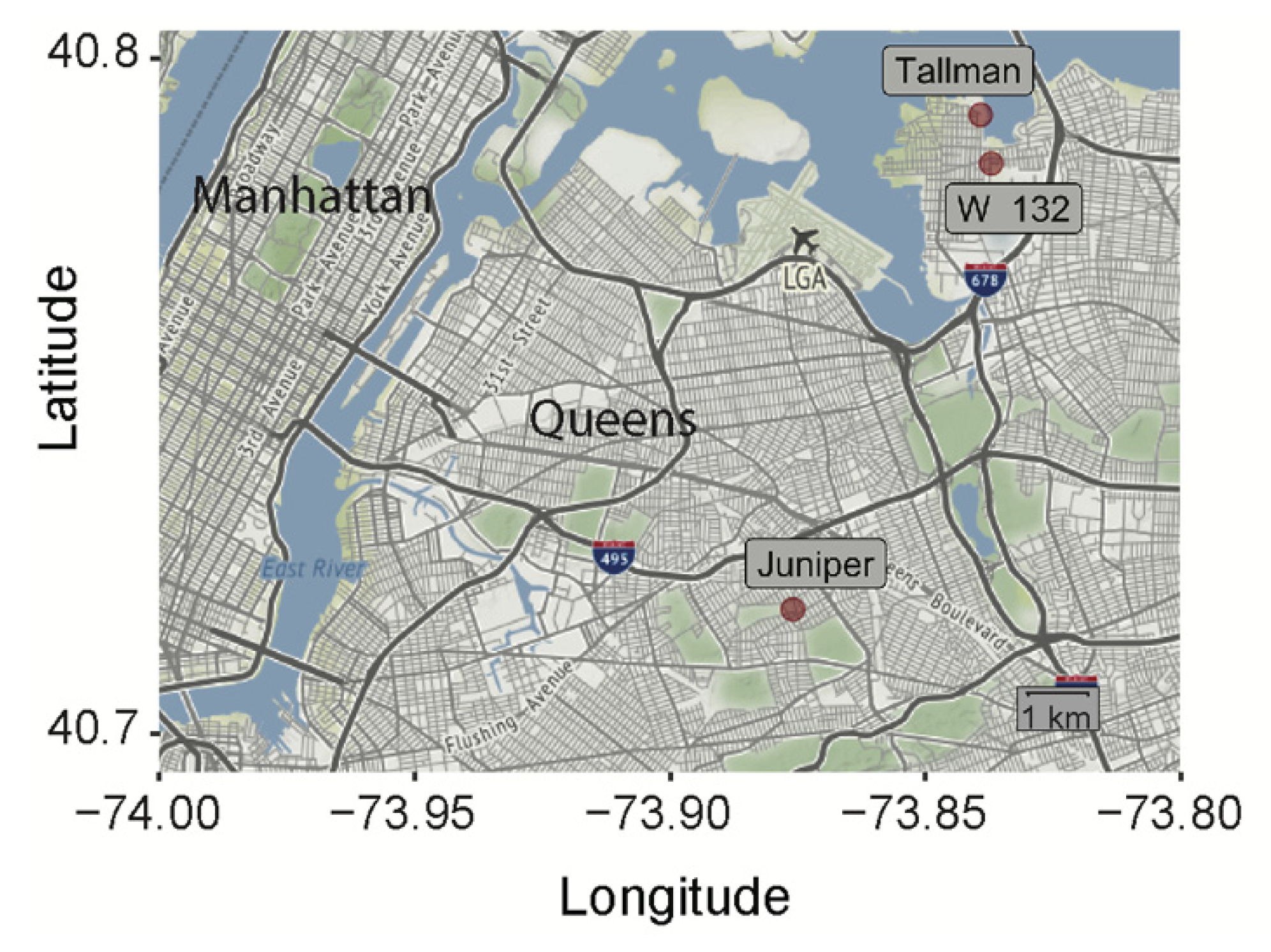
2.2. Mosquitoes
2.3. Mosquito Infections
2.4. Mouse Transmission Studies
2.5. Plaque Assay
2.6. RNA Extraction and RT-qPCR
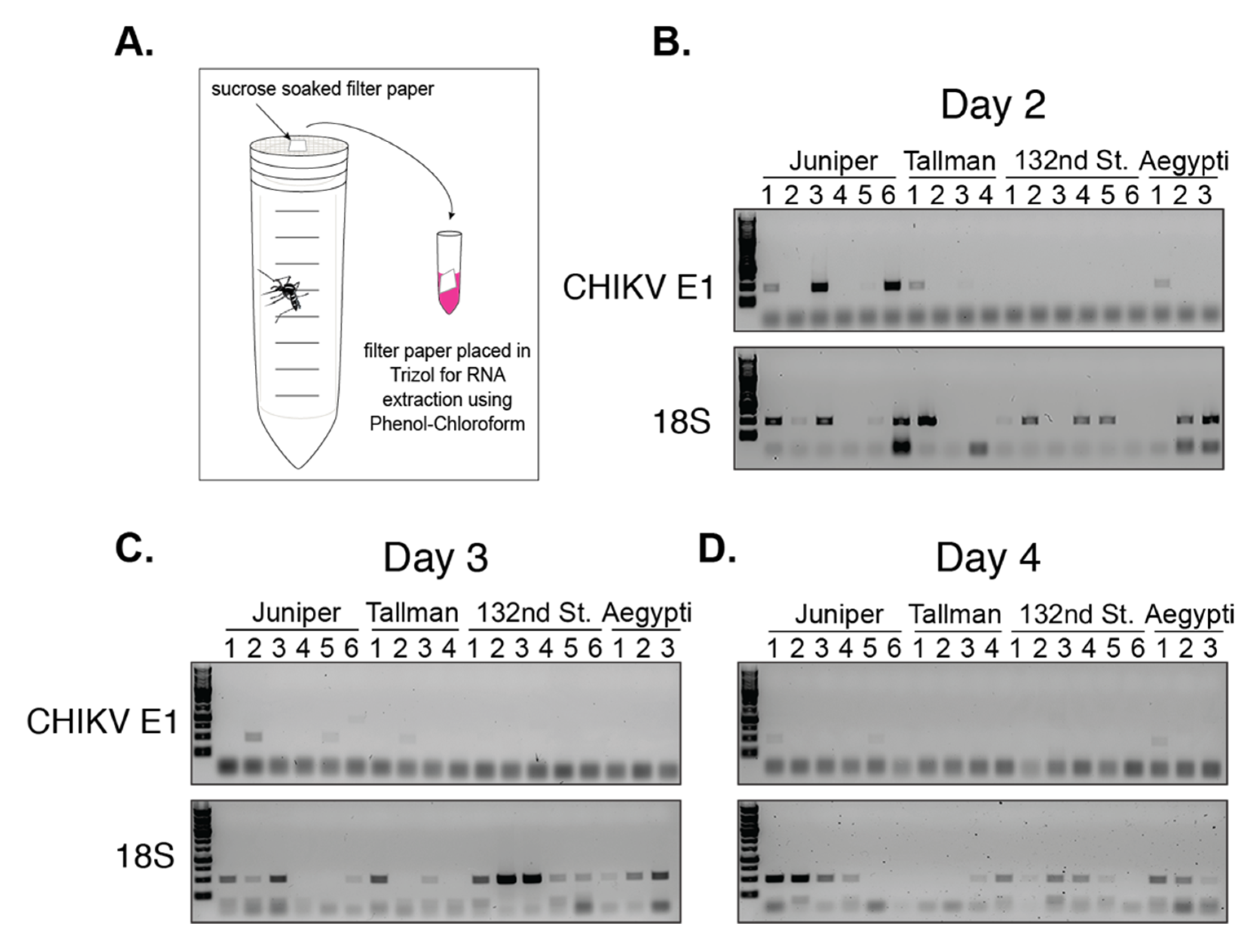
2.7. Filter Paper Assay
2.8. Mosquito Saliva Sequencing
2.9. D7 Sequencing from Mosquito Salivary Glands
2.10. Virus and D7 Sequence Analysis
2.11. Protein Structure Analysis
2.12. Linear Modeling
2.13. Data Availability
2.14. Data analysis and Statistics
3. Results
3.1. The D7 Long form Locus Reveals That NYC Ae. albopictus Populations Harbor Genetically Distinct Alleles
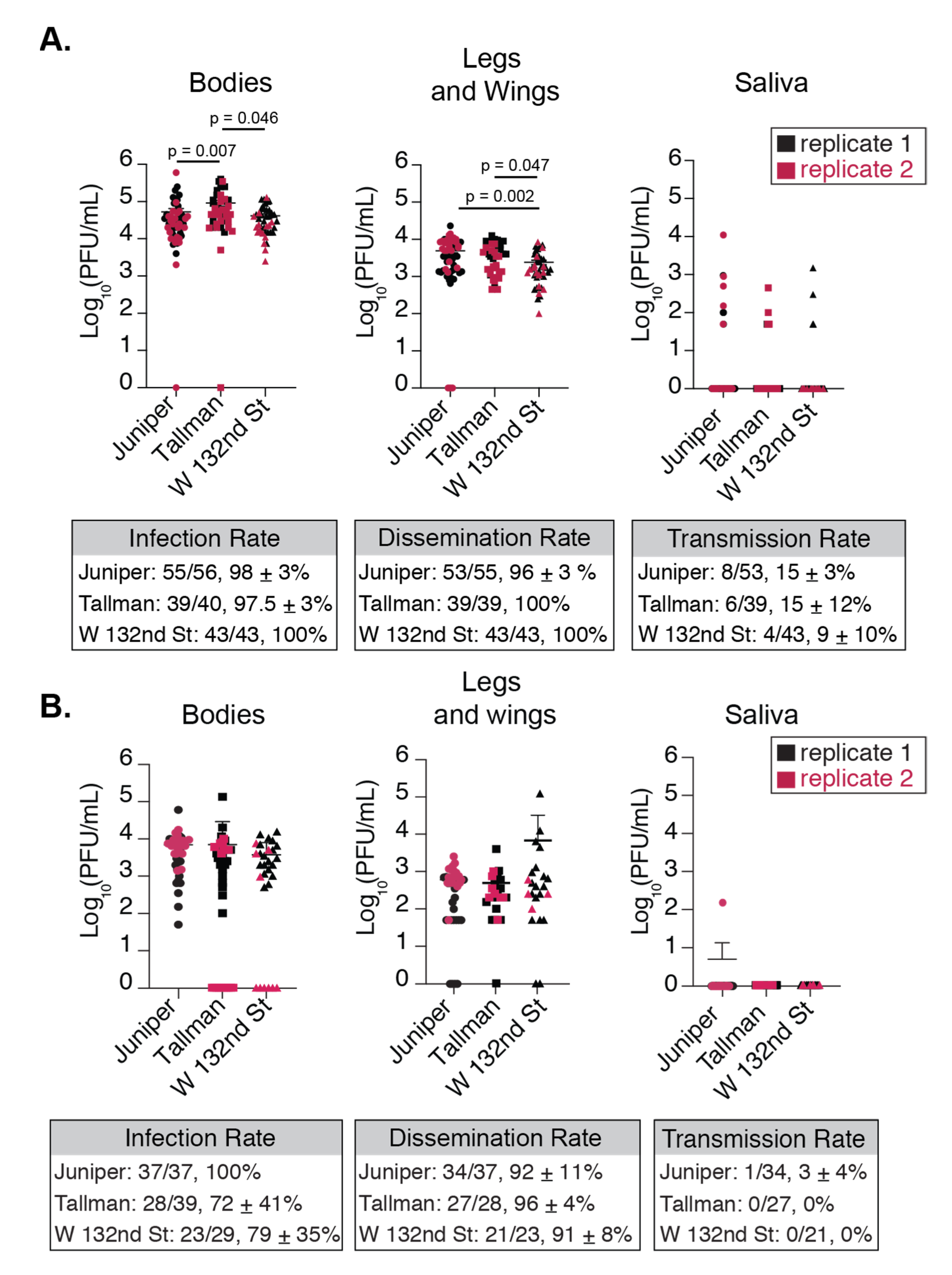
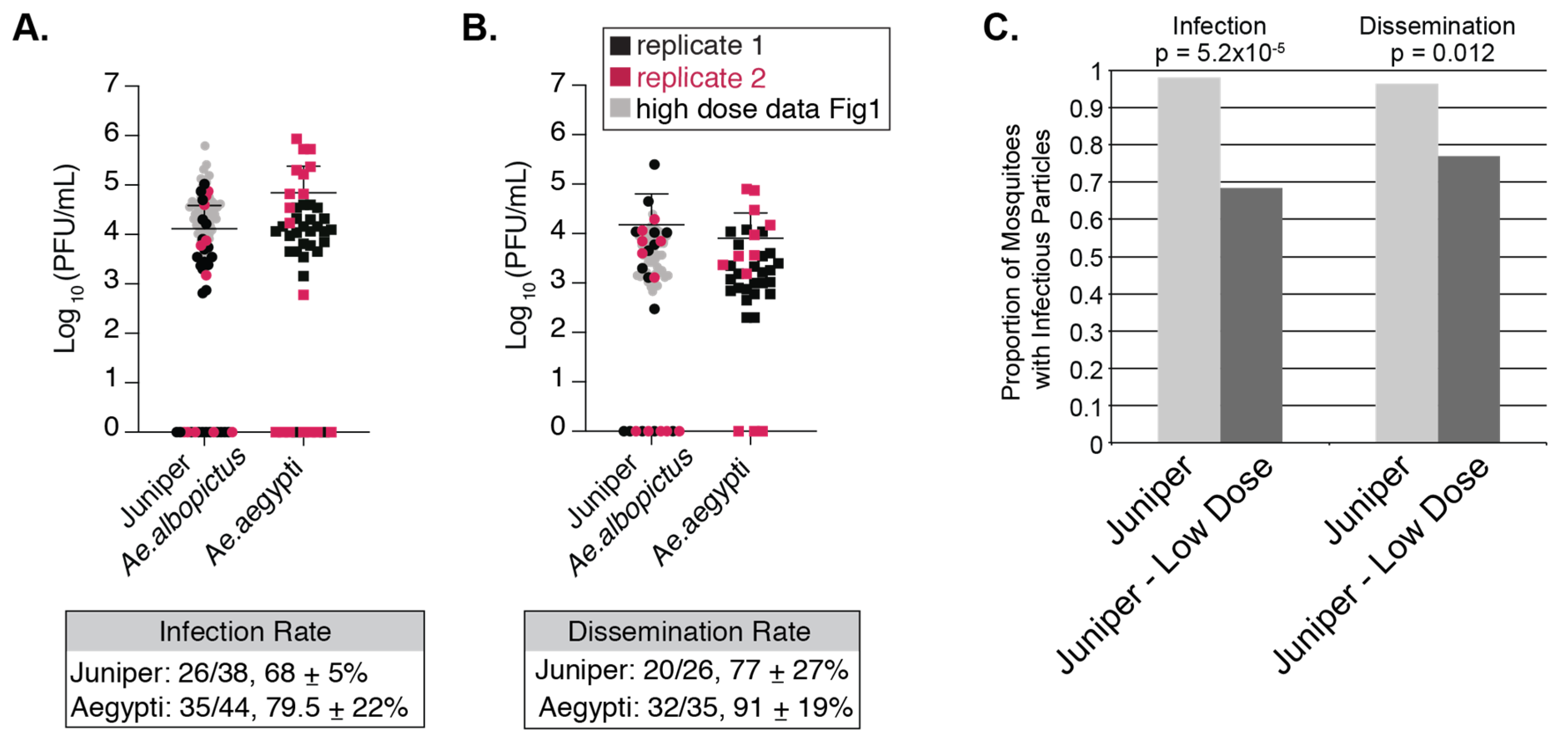
3.2. NYC Ae. albopictus Mosquitoes from Different Locations are Competent for CHIKV Infection
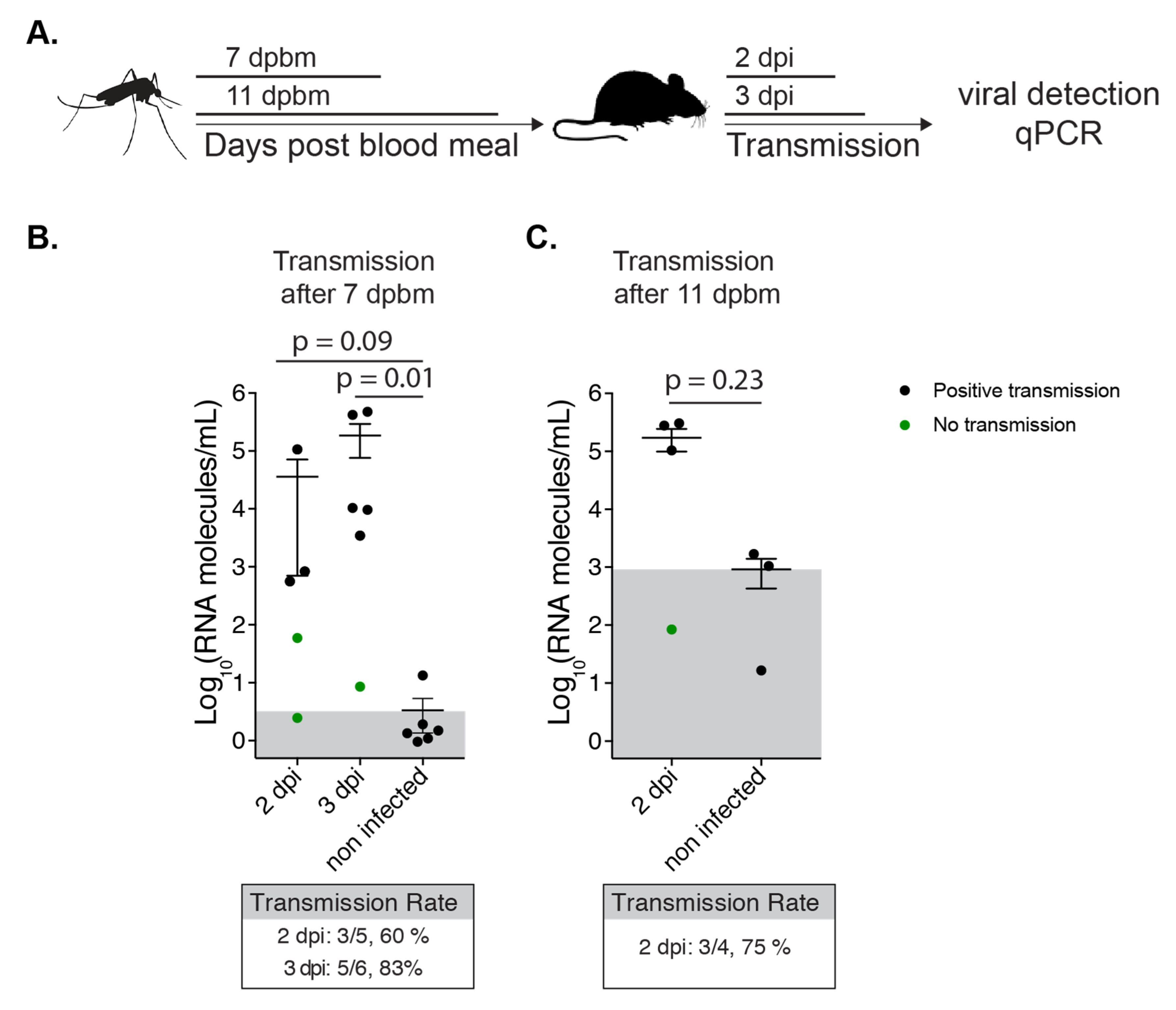
3.3. NYC Ae. albopictus Mosquitoes Can Transmit CHIKV
3.4. Within-Host Evolution of CHIKV in NYC Ae. albopictus Reveals Previously Identified Transmission Variants
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Coffey, L.L.; Failloux, A.B.; Weaver, S.C. Chikungunya virus-vector interactions. Viruses 2014, 6, 4628–4663. [Google Scholar] [CrossRef]
- Morrison, C.R.; Plante, K.S.; Heise, M.T. Chikungunya Virus: Current Perspectives on a Reemerging Virus. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Chikungunya Fever in the Americas. Number of reported cases. PAHO Public Health Emergency Department, 2018. Available online: www.paho.org/en/topics/chikungunya (accessed on 10 April 2020).
- Weaver, S.C. Arrival of chikungunya virus in the new world: Prospects for spread and impact on public health. PLoS Negl. Trop. Dis. 2014, 8, e2921. [Google Scholar] [CrossRef] [PubMed]
- Coffey, L.L.; Vignuzzi, M. Host alternation of chikungunya virus increases fitness while restricting population diversity and adaptability to novel selective pressures. J. Virol. 2011, 85, 1025–1035. [Google Scholar] [CrossRef] [PubMed]
- Riemersma, K.K.; Coffey, L.L. Chikungunya virus populations experience diversity- dependent attenuation and purifying intra-vector selection in Californian Aedes aegypti mosquitoes. PLoS Negl. Trop. Dis. 2019, 13, e0007853. [Google Scholar] [CrossRef] [PubMed]
- Schuffenecker, I.; Iteman, I.; Michault, A.; Murri, S.; Frangeul, L.; Vaney, M.C.; Lavenir, R.; Pardigon, N.; Reynes, J.M.; Pettinelli, F.; et al. Genome microevolution of chikungunya viruses causing the Indian Ocean outbreak. PLoS Med. 2006, 3, e263. [Google Scholar] [CrossRef]
- Zouache, K.; Failloux, A.B. Insect-pathogen interactions: Contribution of viral adaptation to the emergence of vector-borne diseases, the example of chikungunya. Curr. Opin. Insect. Sci. 2015, 10, 14–21. [Google Scholar] [CrossRef]
- Tsetsarkin, K.A.; Vanlandingham, D.L.; McGee, C.E.; Higgs, S. A single mutation in chikungunya virus affects vector specificity and epidemic potential. PLoS Pathog. 2007, 3, e201. [Google Scholar] [CrossRef]
- Mariconti, M.; Obadia, T.; Mousson, L.; Malacrida, A.; Gasperi, G.; Failloux, A.B.; Yen, P.S. Estimating the risk of arbovirus transmission in Southern Europe using vector competence data. Sci. Rep. 2019, 9, 17852. [Google Scholar] [CrossRef]
- Vega-Rua, A.; Zouache, K.; Girod, R.; Failloux, A.B.; Lourenco-de-Oliveira, R. High level of vector competence of Aedes aegypti and Aedes albopictus from ten American countries as a crucial factor in the spread of Chikungunya virus. J. Virol. 2014, 88, 6294–6306. [Google Scholar] [CrossRef] [PubMed]
- Vega-Rua, A.; Zouache, K.; Caro, V.; Diancourt, L.; Delaunay, P.; Grandadam, M.; Failloux, A.B. High efficiency of temperate Aedes albopictus to transmit chikungunya and dengue viruses in the Southeast of France. PLoS ONE 2013, 8, e59716. [Google Scholar] [CrossRef] [PubMed]
- Agha, S.B.; Chepkorir, E.; Mulwa, F.; Tigoi, C.; Arum, S.; Guarido, M.M.; Ambala, P.; Chelangat, B.; Lutomiah, J.; Tchouassi, D.P.; et al. Vector competence of populations of Aedes aegypti from three distinct cities in Kenya for chikungunya virus. PLoS Negl. Trop. Dis. 2017, 11, e0005860. [Google Scholar] [CrossRef]
- Sanchez-Vargas, I.; Harrington, L.C.; Black, W.C.T.; Olson, K.E. Analysis of Salivary Glands and Saliva from Aedes albopictus and Aedes aegypti Infected with Chikungunya Viruses. Insects 2019, 10, 39. [Google Scholar] [CrossRef]
- Pesko, K.; Westbrook, C.J.; Mores, C.N.; Lounibos, L.P.; Reiskind, M.H. Effects of infectious virus dose and bloodmeal delivery method on susceptibility of Aedes aegypti and Aedes albopictus to chikungunya virus. J. Med. Entomol. 2009, 46, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Honorio, N.A.; Wiggins, K.; Eastmond, B.; Camara, D.C.P.; Alto, B.W. Experimental Vertical Transmission of Chikungunya Virus by Brazilian and Florida Aedes Albopictus Populations. Viruses 2019, 11, 353. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.L.; Anderson, S.L.; Smartt, C.T. Vector competence of Florida mosquitoes for chikungunya virus. J. Vector Ecol. 2010, 35, 439–443. [Google Scholar] [CrossRef] [PubMed]
- Forrester, N.L.; Coffey, L.L.; Weaver, S.C. Arboviral bottlenecks and challenges to maintaining diversity and fitness during mosquito transmission. Viruses 2014, 6, 3991–4004. [Google Scholar] [CrossRef]
- Vega-Rua, A.; Schmitt, C.; Bonne, I.; Krijnse Locker, J.; Failloux, A.B. Chikungunya Virus Replication in Salivary Glands of the Mosquito Aedes albopictus. Viruses 2015, 7, 5902–5907. [Google Scholar] [CrossRef] [PubMed]
- White, G.S.; Symmes, K.; Sun, P.; Fang, Y.; Garcia, S.; Steiner, C.; Smith, K.; Reisen, W.K.; Coffey, L.L. Reemergence of St. Louis Encephalitis Virus, California, 2015. Emerg. Infect. Dis. 2016, 22, 2185–2188. [Google Scholar] [CrossRef] [PubMed]
- Main, B.J.; Nicholson, J.; Winokur, O.C.; Steiner, C.; Riemersma, K.K.; Stuart, J.; Takeshita, R.; Krasnec, M.; Barker, C.M.; Coffey, L.L. Vector competence of Aedes aegypti, Culex tarsalis, and Culex quinquefasciatus from California for Zika virus. PLoS Negl. Trop. Dis. 2018, 12, e0006524. [Google Scholar] [CrossRef]
- Brustolin, M.; Pujhari, S.; Henderson, C.A.; Rasgon, J.L. Anopheles mosquitoes may drive invasion and transmission of Mayaro virus across geographically diverse regions. PLoS Negl. Trop. Dis. 2018, 12, e0006895. [Google Scholar] [CrossRef] [PubMed]
- Dodson, B.L.; Rasgon, J.L. Vector competence of Anopheles and Culex mosquitoes for Zika virus. PeerJ 2017, 5, e3096. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pereira, T.N.; Carvalho, F.D.; De Mendonca, S.F.; Rocha, M.N.; Moreira, L.A. Vector competence of Aedes aegypti, Aedes albopictus, and Culex quinquefasciatus mosquitoes for Mayaro virus. PLoS Negl. Trop. Dis. 2020, 14, e0007518. [Google Scholar] [CrossRef] [PubMed]
- Ciota, A.T.; Chin, P.A.; Ehrbar, D.J.; Micieli, M.V.; Fonseca, D.M.; Kramer, L.D. Differential Effects of Temperature and Mosquito Genetics Determine Transmissibility of Arboviruses by Aedes aegypti in Argentina. Am. J. Trop. Med. Hyg. 2018, 99, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Lopez, R.; Bialosuknia, S.M.; Ciota, A.T.; Montalvo, T.; Martinez-de la Puente, J.; Gangoso, L.; Figuerola, J.; Kramer, L.D. Vector Competence of Aedes caspius and Ae. albopictus Mosquitoes for Zika Virus, Spain. Emerg. Infect. Dis. 2019, 25, 346–348. [Google Scholar] [CrossRef]
- Calvo, E.; Mans, B.J.; Ribeiro, J.M.; Andersen, J.F. Multifunctionality and mechanism of ligand binding in a mosquito antiinflammatory protein. Proc. Natl. Acad. Sci. USA 2009, 106, 3728–3733. [Google Scholar] [CrossRef]
- Conway, M.J.; Londono-Renteria, B.; Troupin, A.; Watson, A.M.; Klimstra, W.B.; Fikrig, E.; Colpitts, T.M. Aedes aegypti D7 Saliva Protein Inhibits Dengue Virus Infection. PLoS Negl. Trop. Dis. 2016, 10, e0004941. [Google Scholar] [CrossRef]
- Ruckert, C.; Weger-Lucarelli, J.; Garcia-Luna, S.M.; Young, M.C.; Byas, A.D.; Murrieta, R.A.; Fauver, J.R.; Ebel, G.D. Impact of simultaneous exposure to arboviruses on infection and transmission by Aedes aegypti mosquitoes. Nat. Commun. 2017, 8, 15412. [Google Scholar] [CrossRef]
- Noval, M.G.; Rodriguez-Rodriguez, B.A.; Rangel, M.V.; Stapleford, K.A. Evolution-Driven Attenuation of Alphaviruses Highlights Key Glycoprotein Determinants Regulating Viral Infectivity and Dissemination. Cell Rep. 2019, 28, 460–471.e465. [Google Scholar] [CrossRef]
- Stapleford, K.A.; Coffey, L.L.; Lay, S.; Borderia, A.V.; Duong, V.; Isakov, O.; Rozen-Gagnon, K.; Arias-Goeta, C.; Blanc, H.; Beaucourt, S.; et al. Emergence and transmission of arbovirus evolutionary intermediates with epidemic potential. Cell Host Microbe 2014, 15, 706–716. [Google Scholar] [CrossRef]
- Carrau, L.; Rezelj, V.V.; Noval, M.G.; Levi, L.I.; Megrian, D.; Blanc, H.; Weger-Lucarelli, J.; Moratorio, G.; Stapleford, K.A.; Vignuzzi, M. Chikungunya Virus Vaccine Candidates with Decreased Mutational Robustness Are Attenuated In Vivo and Have Compromised Transmissibility. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Grubaugh, N.D.; Fauver, J.R.; Ruckert, C.; Weger-Lucarelli, J.; Garcia-Luna, S.; Murrieta, R.A.; Gendernalik, A.; Smith, D.R.; Brackney, D.E.; Ebel, G.D. Mosquitoes Transmit Unique West Nile Virus Populations during Each Feeding Episode. Cell Rep. 2017, 19, 709–718. [Google Scholar] [CrossRef] [PubMed]
- Giraldo-Calderon, G.I.; Emrich, S.J.; MacCallum, R.M.; Maslen, G.; Dialynas, E.; Topalis, P.; Ho, N.; Gesing, S.; VectorBase, C.; Madey, G.; et al. VectorBase: An updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res. 2015, 43, D707–D713. [Google Scholar] [CrossRef] [PubMed]
- Verdonschot, P.F.M.; Besse-Lototskaya, A.A. Flight distance of mosquitoes (Culicidae): A metadata analysis to support the management of barrier zones around rewetted and newly constructed wetlands. Limnologica 2014, 45, 69–79. [Google Scholar] [CrossRef]
- Marini, F.; Caputo, B.; Pombi, M.; Tarsitani, G.; della Torre, A. Study of Aedes albopictus dispersal in Rome, Italy, using sticky traps in mark-release-recapture experiments. Med. Vet. Entomol. 2010, 24, 361–368. [Google Scholar] [CrossRef]
- Liew, C.; Curtis, C.F. Horizontal and vertical dispersal of dengue vector mosquitoes, Aedes aegypti and Aedes albopictus, in Singapore. Med. Vet. Entomol. 2004, 18, 351–360. [Google Scholar] [CrossRef]
- Jablonka, W.; Kim, I.H.; Alvarenga, P.H.; Valenzuela, J.G.; Ribeiro, J.M.C.; Andersen, J.F. Functional and structural similarities of D7 proteins in the independently-evolved salivary secretions of sand flies and mosquitoes. Sci. Rep. 2019, 9, 5340. [Google Scholar] [CrossRef]
- Valenzuela, J.G.; Charlab, R.; Gonzalez, E.C.; de Miranda-Santos, I.K.; Marinotti, O.; Francischetti, I.M.; Ribeiro, J.M. The D7 family of salivary proteins in blood sucking diptera. Insect Mol. Biol. 2002, 11, 149–155. [Google Scholar] [CrossRef]
- Waggoner, J.J.; Gresh, L.; Vargas, M.J.; Ballesteros, G.; Tellez, Y.; Soda, K.J.; Sahoo, M.K.; Nunez, A.; Balmaseda, A.; Harris, E.; et al. Viremia and Clinical Presentation in Nicaraguan Patients Infected with Zika Virus, Chikungunya Virus, and Dengue Virus. Clin. Infect. Dis. 2016, 63, 1584–1590. [Google Scholar] [CrossRef]
- Dubrulle, M.; Mousson, L.; Moutailler, S.; Vazeille, M.; Failloux, A.B. Chikungunya virus and Aedes mosquitoes: Saliva is infectious as soon as two days after oral infection. PLoS ONE 2009, 4, e5895. [Google Scholar] [CrossRef]
- Vazeille, M.; Moutailler, S.; Coudrier, D.; Rousseaux, C.; Khun, H.; Huerre, M.; Thiria, J.; Dehecq, J.S.; Fontenille, D.; Schuffenecker, I.; et al. Two Chikungunya isolates from the outbreak of La Reunion (Indian Ocean) exhibit different patterns of infection in the mosquito, Aedes albopictus. PLoS ONE 2007, 2, e1168. [Google Scholar] [CrossRef] [PubMed]
- Alto, B.W.; Wiggins, K.; Eastmond, B.; Velez, D.; Lounibos, L.P.; Lord, C.C. Transmission risk of two chikungunya lineages by invasive mosquito vectors from Florida and the Dominican Republic. PLoS Negl. Trop. Dis. 2017, 11, e0005724. [Google Scholar] [CrossRef]
- Mbaika, S.; Lutomiah, J.; Chepkorir, E.; Mulwa, F.; Khayeka-Wandabwa, C.; Tigoi, C.; Oyoo-Okoth, E.; Mutisya, J.; Ng’ang’a, Z.; Sang, R. Vector competence of Aedes aegypti in transmitting Chikungunya virus: Effects and implications of extrinsic incubation temperature on dissemination and infection rates. Virol. J. 2016, 13, 114. [Google Scholar] [CrossRef] [PubMed]
- Rudolph, K.E.; Lessler, J.; Moloney, R.M.; Kmush, B.; Cummings, D.A. Incubation periods of mosquito-borne viral infections: A systematic review. Am. J. Trop. Med. Hyg. 2014, 90, 882–891. [Google Scholar] [CrossRef] [PubMed]
- Staples, J.E.; Fischer, M. Chikungunya virus in the Americas--what a vectorborne pathogen can do. N. Engl. J. Med. 2014, 371, 887–889. [Google Scholar] [CrossRef] [PubMed]
- Souza-Neto, J.A.; Powell, J.R.; Bonizzoni, M. Aedes aegypti vector competence studies: A review. Infect. Genet. Evol. 2019, 67, 191–209. [Google Scholar] [CrossRef] [PubMed]
- Azar, S.R.; Weaver, S.C. Vector Competence: What Has Zika Virus Taught Us? Viruses 2019, 11, 867. [Google Scholar] [CrossRef] [PubMed]
- Dodson, B.L.; Kramer, L.D.; Rasgon, J.L. Effects of larval rearing temperature on immature development and West Nile virus vector competence of Culex tarsalis. Parasites Vectors 2012, 5, 199. [Google Scholar] [CrossRef]
- Hegde, S.; Rasgon, J.L.; Hughes, G.L. The microbiome modulates arbovirus transmission in mosquitoes. Curr. Opin. Virol. 2015, 15, 97–102. [Google Scholar] [CrossRef]
- Juhn, J.; Naeem-Ullah, U.; Maciel Guedes, B.A.; Majid, A.; Coleman, J.; Paolucci Pimenta, P.F.; Akram, W.; James, A.A.; Marinotti, O. Spatial mapping of gene expression in the salivary glands of the dengue vector mosquito, Aedes aegypti. Parasites Vectors 2011, 4, 1. [Google Scholar] [CrossRef]
- Ciano, K.A.; Saredy, J.J.; Bowers, D.F. Heparan sulfate proteoglycan: An arbovirus attachment factor integral to mosquito salivary gland ducts. Viruses 2014, 6, 5182–5197. [Google Scholar] [CrossRef] [PubMed]
- Tchankouo-Nguetcheu, S.; Bourguet, E.; Lenormand, P.; Rousselle, J.C.; Namane, A.; Choumet, V. Infection by chikungunya virus modulates the expression of several proteins in Aedes aegypti salivary glands. Parasites Vectors 2012, 5, 264. [Google Scholar] [CrossRef] [PubMed]

| Mosquito #—Fragment Amplified | Location | Gene | Amino Acid | Codon |
|---|---|---|---|---|
| 1—F1, F2 | Juniper | E1 | S25 | Syn |
| E1 | V80I | GTC to ATC | ||
| E1 | A129F | GCA to TTT | ||
| E1 | A129F | GCA to TTT | ||
| E1 | A129F | GCA to TTT | ||
| 3’UTR | n/a | n/a | ||
| 2—F1 | Tallman | E3 | T34 | Syn |
| E2 | P128L | CCA to CTA | ||
| E2 | G137D | GGT to GAT | ||
| E2 | R393Q | CGA to CAA | ||
| 3—F2 | Juniper | E1 | V80I | GTC to ATC |
| E1 | I162V | ATT to GTT | ||
| 4—F2 | W 132 | E1 | V80I | GTC to ATC |
| E1 | S111 | Syn | ||
| 5—F2 | Tallman | E1 | A129W | GCA to TGG |
| E1 | A129W | GCA to TGG | ||
| E1 | A129W | GCA to TGG |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaczmarek, M.E.; Herzog, N.L.; Noval, M.G.; Zuzworsky, J.; Shah, Z.; Bajwa, W.I.; Stapleford, K.A. Distinct New York City Aedes albopictus Mosquito Populations Display Differences in Salivary Gland Protein D7 Diversity and Chikungunya Virus Replication. Viruses 2020, 12, 698. https://doi.org/10.3390/v12070698
Kaczmarek ME, Herzog NL, Noval MG, Zuzworsky J, Shah Z, Bajwa WI, Stapleford KA. Distinct New York City Aedes albopictus Mosquito Populations Display Differences in Salivary Gland Protein D7 Diversity and Chikungunya Virus Replication. Viruses. 2020; 12(7):698. https://doi.org/10.3390/v12070698
Chicago/Turabian StyleKaczmarek, Maria E., Nora L. Herzog, Maria G. Noval, John Zuzworsky, Zahir Shah, Waheed I. Bajwa, and Kenneth A. Stapleford. 2020. "Distinct New York City Aedes albopictus Mosquito Populations Display Differences in Salivary Gland Protein D7 Diversity and Chikungunya Virus Replication" Viruses 12, no. 7: 698. https://doi.org/10.3390/v12070698
APA StyleKaczmarek, M. E., Herzog, N. L., Noval, M. G., Zuzworsky, J., Shah, Z., Bajwa, W. I., & Stapleford, K. A. (2020). Distinct New York City Aedes albopictus Mosquito Populations Display Differences in Salivary Gland Protein D7 Diversity and Chikungunya Virus Replication. Viruses, 12(7), 698. https://doi.org/10.3390/v12070698






