Evaluation of Viral RNA Recovery Methods in Vectors by Metagenomic Sequencing
Abstract
1. Introduction
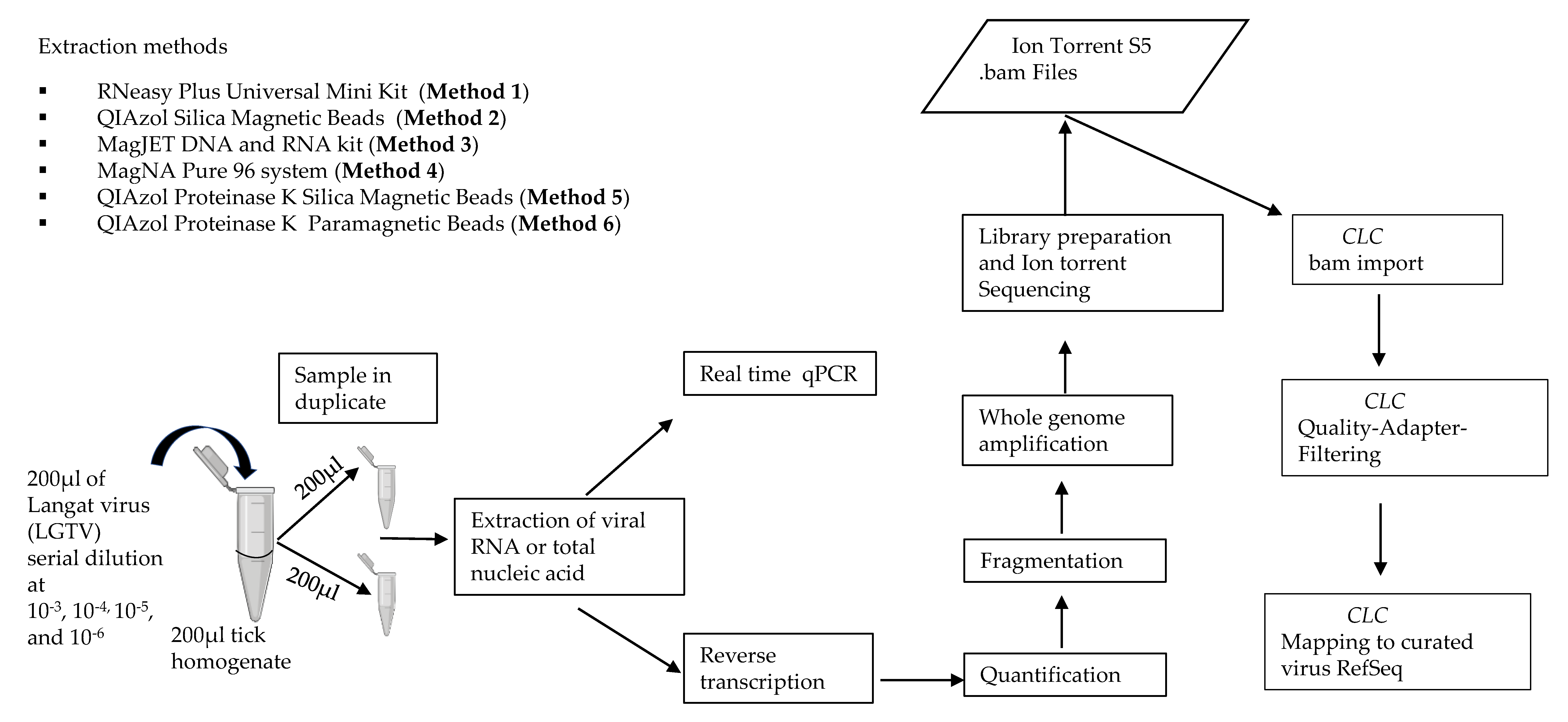
2. Materials and Methods
2.1. LGTV Viral Stock and Serial Dillutions
2.2. LGTV Spike Sample Prepartaion
2.3. Viral RNA Extraction Methods
2.4. Evaluating Performance of the Different Extraction Methods
2.5. qPCR
2.6. Real-Time PCR
2.7. Whole Genome Amplification (WGA)
2.8. Sequencing on the Ion Torrent S5 and Analyses
2.9. Statistical Analyses
3. Results
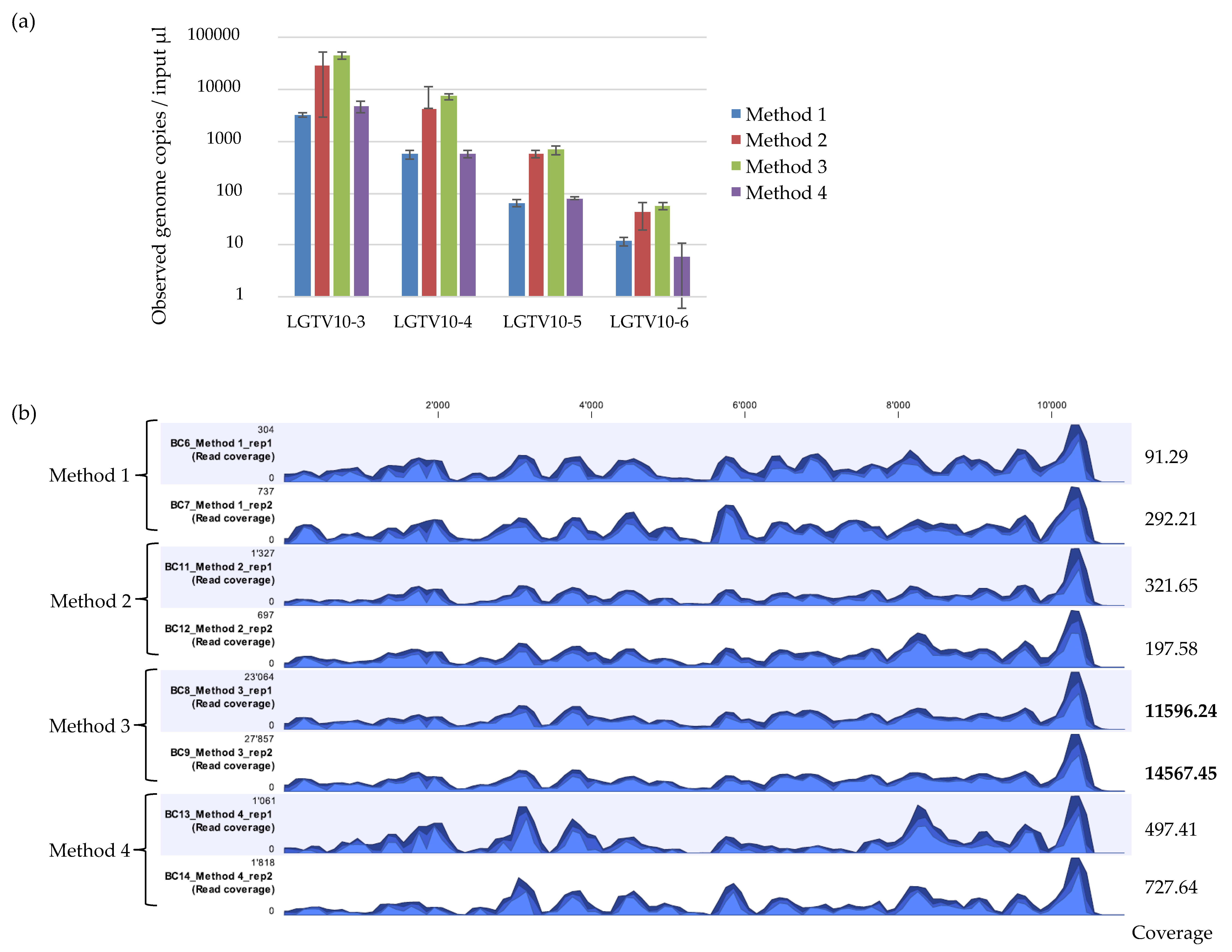
3.1. LGTV Viral Recovery Using qPCR and NGS Analysis
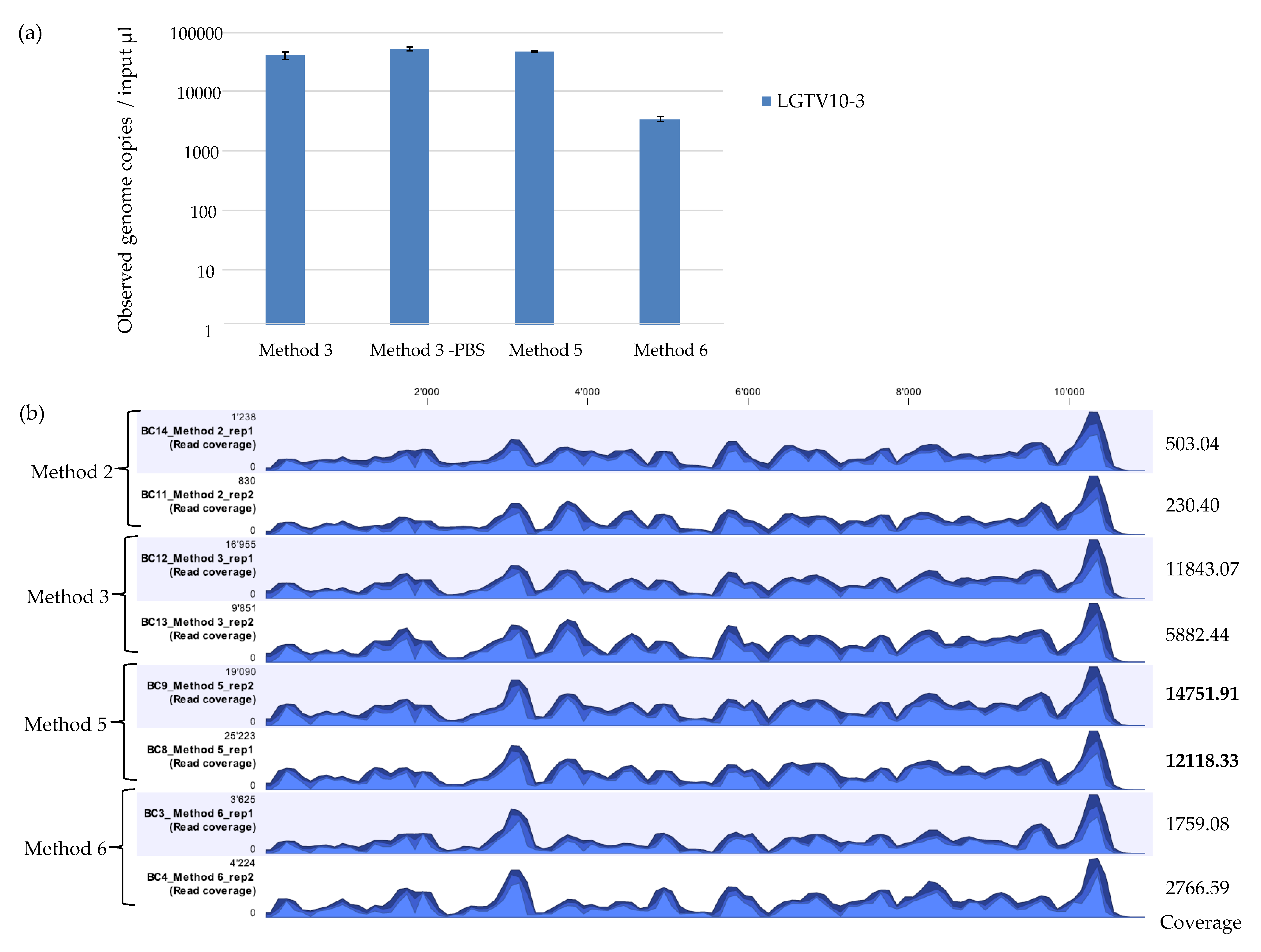
3.2. ROVIV—QIAzol Lysis Accompanied with Proteinase K and Silica Magnetic Beads Improves Viral RNA Recovery
3.3. Testing Sensitivity of the Optimized Method
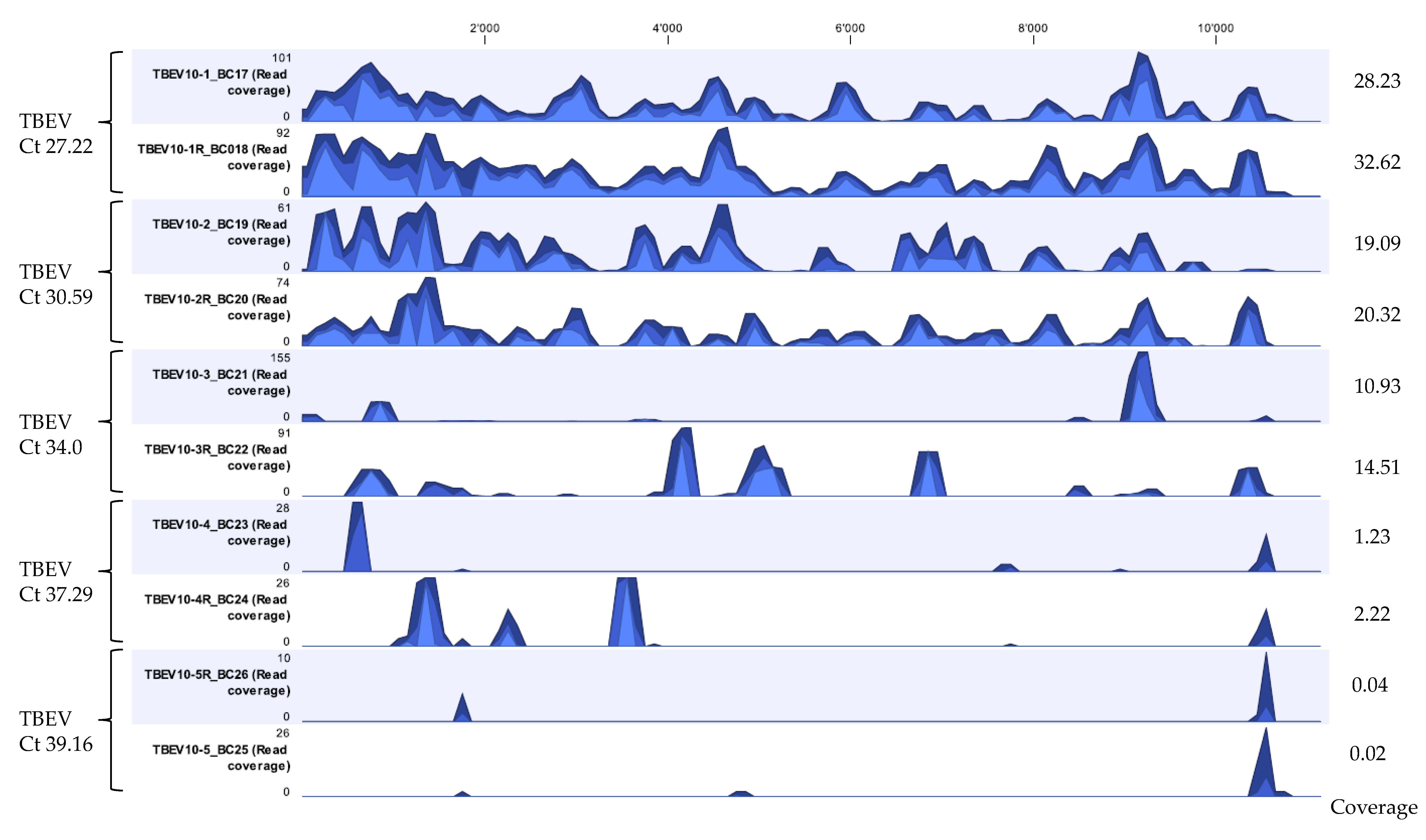
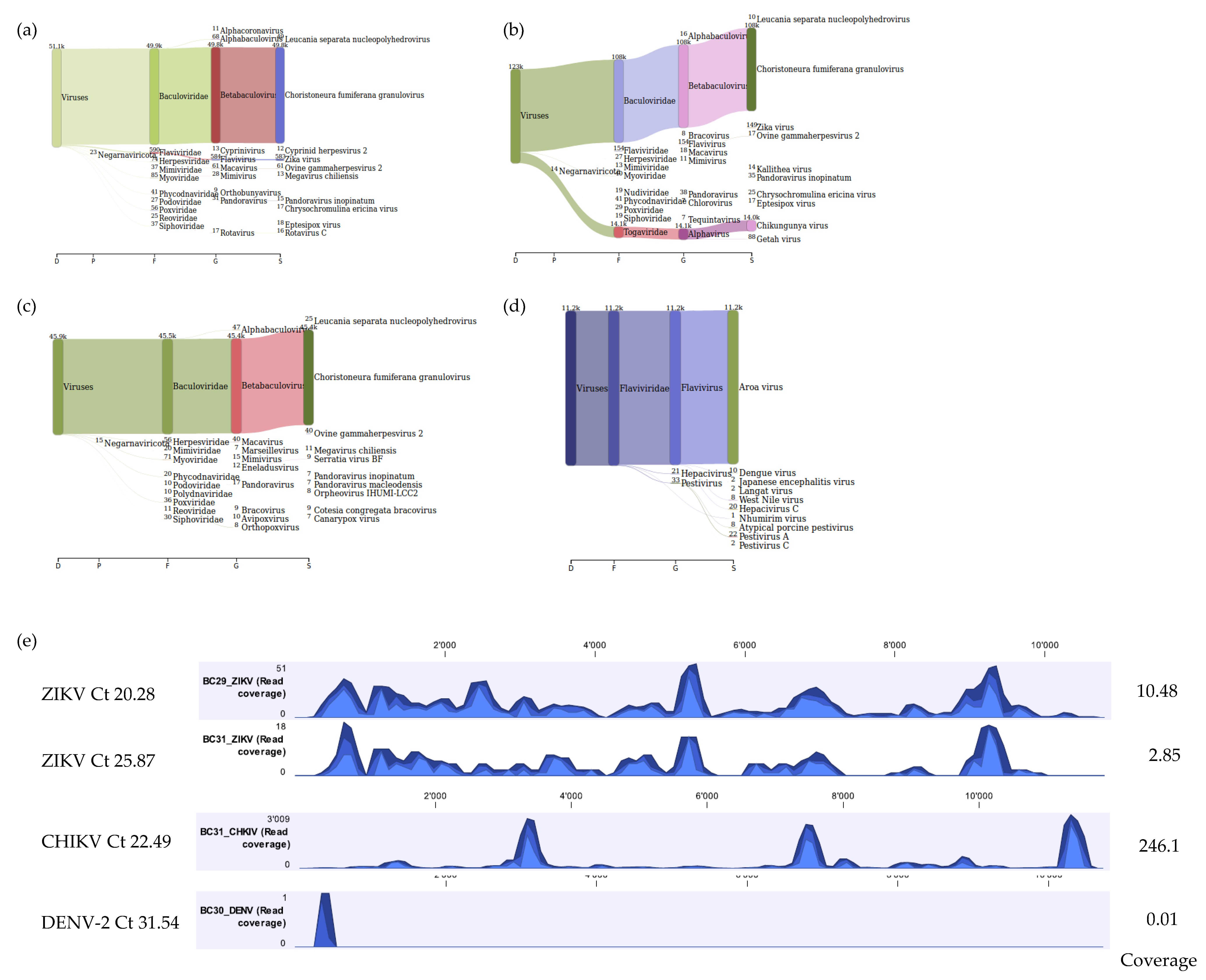
3.4. Proof of Concept of the Optimized Method
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schmidt, K.; Dressel, K.M.; Niedrig, M.; Mertens, M.; Schüle, S.A.; Groschup, M.H. Public health and vector-borne diseases-a new concept for risk governance. Zoonoses Public Health 2013, 60, 528–538. [Google Scholar] [CrossRef] [PubMed]
- Lindquist, L.; Vapalahti, O. Tick-borne encephalitis. Lancet 2008, 371, 1861–1871. [Google Scholar] [CrossRef]
- Madison-Antenucci, S.; Kramer, L.D.; Gebhardt, L.L.; Kauffman, E. Emerging Tick-Borne Diseases. Clin. Microbiol. Rev. 2020, 33, e00083-18. [Google Scholar] [PubMed]
- Kraemer, M.U.; Sinka, M.E.; Duda, K.A.; Mylne, A.Q.; Shearer, F.M.; Barker, C.M.; Moore, C.G.; Carvalho, R.G.; Coelho, G.E.; Van Bortel, W.; et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. Elife 2015, 4, e08347. [Google Scholar] [CrossRef] [PubMed]
- Conway, M.J.; Colpitts, T.M.; Fikrig, E. Role of the Vector in Arbovirus Transmission. Annu. Rev. Virol. 2014, 1, 71–88. [Google Scholar] [CrossRef] [PubMed]
- Mayer, S.V.; Tesh, R.B.; Vasilakis, N. The emergence of arthropod-borne viral diseases: A global prospective on dengue, chikungunya and zika fevers. Acta Trop. 2017, 166, 155–163. [Google Scholar]
- Asghar, N.; Pettersson, J.H.; Dinnétz, P.; Åndreassen, A.; Johansson, M. Deep sequencing analysis of tick-borne encephalitis virus from questing ticks at natural foci reveals similarities between quasispecies pools of the virus. J. Gen. Virol. 2017, 98, 413–421. [Google Scholar] [CrossRef]
- Ayres, C.F.J.; Guedes, D.R.D.; Paiva, M.H.S.; Morais-Sobral, M.C.; Krokovsky, L.; Machado, L.C.; Cardoso, O.A. Zika virus detection, isolation and genome sequencing through Culicidae sampling during the epidemic in Vitoria, Espirito Santo, Brazil. Parasit Vectors 2019, 12, 220. [Google Scholar] [CrossRef]
- Machado, L.C.; De Morais-Sobral, M.C.; Campos, T.L.; Pereira, M.R.; de Albuquerque, M.F.P.M.; Gilbert, C.; Franca, R.F.O.; Wallau, G.L. Genome sequencing reveals coinfection by multiple chikungunya virus genotypes in a recent outbreak in Brazil. PLOS Neglected Trop. Dis. 2019, 13, e0007332. [Google Scholar] [CrossRef]
- Miura, T.; Masago, Y.; Sano, D.; Omura, T. Development of an effective method for recovery of viral genomic RNA from environmental silty sediments for quantitative molecular detection. Appl. Environ. Microbiol. 2011, 77, 3975–3981. [Google Scholar] [CrossRef]
- Hill, V.R.; Narayanan, J.; Gallen, R.R.; Ferdinand, K.L.; Cromeans, T.; Vinjé, J. Development of a nucleic Acid extraction procedure for simultaneous recovery of DNA and RNA from diverse microbes in water. Pathogens 2015, 4, 335–354. [Google Scholar] [CrossRef] [PubMed]
- Ali, N.; Rampazzo, R.C.P.; Costa, A.D.T.; Krieger, M.A. Current Nucleic Acid Extraction Methods and Their Implications to Point-of-Care Diagnostics. BioMed Res. Int. 2017, 2017, 9306564. [Google Scholar] [CrossRef] [PubMed]
- Sarnecka, A.K.; Nawrat, D.; Piwowar, M.; Ligęza, J.; Swadźba, J.; Wójcik, P. DNA extraction from FFPE tissue samples—A comparison of three procedures. Contemp. Oncol. (Pozn) 2019, 23, 52–58. [Google Scholar] [CrossRef]
- Bergallo, M.; Costa, C.; Gribaudo, G.; Tarallo, S.; Baro, S.; Negro Ponzi, A.; Cavallo, R. Evaluation of six methods for extraction and purification of viral DNA from urine and serum samples. New Microbiol. 2006, 29, 111–119. [Google Scholar] [PubMed]
- Hjelmso, M.H.; Hellmér, M.; Fernandez-Cassi, X.; Timoneda, N.; Lukjancenko, O.; Seidel, M.; Elsässer, D.; Aarestrup, F.M.; Löfström, C.; Bofill-Mas, S.; et al. Evaluation of Methods for the Concentration and Extraction of Viruses from Sewage in the Context of Metagenomic Sequencing. PLOS ONE 2017, 12, e0170199. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Lou, X.; Yan, H.; Pan, J.; Mao, H.; Tang, H.; Shu, Y.; Zhao, Y.; Liu, L.; Li, J.; et al. Metagenomic analysis of viral nucleic acid extraction methods in respiratory clinical samples. BMC Genom. 2018, 19, 773. [Google Scholar] [CrossRef]
- Sagar, K.; Singh, S.P.; Goutam, K.K.; Konwar, B.K. Assessment of five soil DNA extraction methods and a rapid laboratory-developed method for quality soil DNA extraction for 16S rDNA-based amplification and library construction. J. Microbiol. Methods 2014, 97, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Gaumann, R.; Muhlemann, K.; Strasser, M.; Beuret, C. High-throughput procedure for tick surveys of tick-borne encephalitis virus and its application in a national surveillance study in Switzerland. Appl. Environ. Microbiol. 2010, 76, 4241–4249. [Google Scholar] [CrossRef]
- Pagani, S.C.; Malossa, S.F.; Klaus, C.; Hoffmann, D.; Beretta, O.; Bomio-Pacciorini, N.; Lazzaro, M.; Merlani, G.; Ackermann, R.; Beuret, C. First detection of TBE virus in ticks and sero-reactivity in goats in a non-endemic region in the southern part of Switzerland (Canton of Ticino). Ticks Tick-Borne Dis. 2019, 10, 868–874. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Boil. 2019, 20, 257. [Google Scholar] [CrossRef]
- Breitwieser, F.P.; Salzberg, S.L. Pavian: Interactive analysis of metagenomics data for microbiomics and pathogen identification. BioRxiv 2016, 084715. [Google Scholar] [CrossRef]
- Grubaugh, N.D.; Sharma, S.; Krajacich, B.J.; Fakoli, L.S., III; Bolay, F.K.; Diclaro, J.W., II; Johnson, W.E.; Ebel, G.D.; Foy, B.D.; Brackney, D.E. Xenosurveillance: A novel mosquito-based approach for examining the human-pathogen landscape. PLoS Neglected Trop. Dis. 2015, 9, e0003628. [Google Scholar] [CrossRef]
- Ng, T.F.; Willner, D.L.; Lim, Y.W.; Schmieder, R.; Chau, B.; Nilsson, C.; Anthony, S.; Ruan, Y.; Rohwer, F.; Breitbart, M. Broad surveys of DNA viral diversity obtained through viral metagenomics of mosquitoes. PLoS ONE 2011, 6, e20579. [Google Scholar] [CrossRef] [PubMed]
- Bouquet, J.; Melgar, M.; Swei, A.; Delwart, E.; Lane, R.S.; Chiu, C.Y. Metagenomic-based Surveillance of Pacific Coast tick Dermacentor occidentalis Identifies Two Novel Bunyaviruses and an Emerging Human Ricksettsial Pathogen. Sci. Rep. 2017, 7, 12234. [Google Scholar] [CrossRef] [PubMed]
- Holding, M.; Dowall, S.D.; Medlock, J.M.; Carter, D.P.; McGinley, L.; Curran-French, M.; Pullan, S.T.; Chamberlain, J.; Hansford, K.M.; Baylis, M.; et al. Detection of new endemic focus of tick-borne encephalitis virus (TBEV), Hampshire/Dorset border, England, September 2019. Eurosurveillance 2019, 24, 1900658. [Google Scholar] [CrossRef]
- Harvey, E.; Rose, K.; Eden, J.S.; Lo, N.; Abeyasuriya, T.; Shi, M.; Doggett, S.L.; Holmes, E.C. Extensive Diversity of RNA Viruses in Australian Ticks. J. Virol. 2018, 93, e01358-18. [Google Scholar] [CrossRef] [PubMed]
- Blomstrom, A.L.; Luz, H.R.; Öhlund, P.; Lukenge, M.; Brandão, P.E.; Labruna, M.B.; Berg, M. Novel Viruses Found in Antricola Ticks Collected in Bat Caves in the Western Amazonia of Brazil. Viruses 2019, 12, 48. [Google Scholar] [CrossRef]
- Birnberg, L.; Temmam, S.; Aranda, C.; Correa-Fiz, F.; Talavera, S.; Bigot, T.; Eloit, M.; Busquets, N. Viromics on Honey-Baited FTA Cards as a New Tool for the Detection of Circulating Viruses in Mosquitoes. Viruses 2020, 12, 274. [Google Scholar] [CrossRef]
- Xia, H.; Hu, C.; Zhang, D.; Tang, S.; Zhang, Z.; Kou, Z.; Fan, Z.; Bente, D.; Zeng, C.; Li, T. Metagenomic profile of the viral communities in Rhipicephalus spp. ticks from Yunnan, China. PLoS ONE 2015, 10, e0121609. [Google Scholar] [CrossRef]
- Hall-Mendelin, S.; Allcock, R.; Kresoje, N.; van den Hurk, A.F.; Warrilow, D. Detection of arboviruses and other micro-organisms in experimentally infected mosquitoes using massively parallel sequencing. PLoS ONE 2013, 8, e58026. [Google Scholar] [CrossRef]
- Conceicao-Neto; Zeller, M.; Lefrère, H.; De Bruyn, P.; Beller, L.; Deboutte, W.; Yinda, C.K.; Lavigne, R.; Maes, P.; Van Ranst, M.; et al. Modular approach to customise sample preparation procedures for viral metagenomics: A reproducible protocol for virome analysis. Sci. Rep. 2015, 5, 16532. [Google Scholar] [CrossRef]
- Wylezich, C.; Papa, A.; Beer, M.; Höper, D. A Versatile Sample Processing Workflow for Metagenomic Pathogen Detection. Sci. Rep. 2018, 8, 13108. [Google Scholar] [CrossRef]
- Gupta, S.; Preet, S. Protocol optimization for genomic DNA extraction and RAPD-PCR in mosquito larvae (Diptera: Culicidae). Ann. Biol. Res. 2012, 3, 1553–1561. [Google Scholar]
- de la Cruz-Ramos, J.M.; Hernández-Triana, L.M.; García-De La Peña, C.; González-Álvarez, V.H.; Weger-Lucarelli, J.; Siller-Rodríguez, Q.K.; Rámos, F.J.S.; Rodríguez, A.D.; Ortega-Morales, A.I. Comparison of two DNA extraction methods from larvae, pupae, and adults of Aedes aegypti. Heliyon 2019, 5, e02660. [Google Scholar] [CrossRef]
- Sathiamoorthy, S.; Malott, R.J.; Gisonni-Lex, L.; Ng, S.H.S. Selection and evaluation of an efficient method for the recovery of viral nucleic acids from complex biologicals [corrected]. NPJ Vaccines 2018, 3, 31. [Google Scholar] [CrossRef]
- Woolhouse, M.E.; Brierley, L.; McCaffery, C.; Lycett, S. Assessing the Epidemic Potential of RNA and DNA Viruses. Emerg. Infect. Dis. 2016, 22, 2037–2044. [Google Scholar] [CrossRef]
- He, H.; Li, R.; Chen, Y.; Pan, P.; Tong, W.; Dong, X.; Chen, Y.; Yu, D. Integrated DNA and RNA extraction using magnetic beads from viral pathogens causing acute respiratory infections. Sci. Rep. 2017, 7, 45199. [Google Scholar] [CrossRef]
- Cholleti, H.; Hayer, J.; Abilio, A.P.; Mulandane, F.C.; Verner-Carlsson, J.; Falk, K.I.; Fafetine, J.M.; Berg, M.; Blomström, A.-L. Discovery of Novel Viruses in Mosquitoes from the Zambezi Valley of Mozambique. PLoS ONE 2016, 11, e0162751. [Google Scholar] [CrossRef]

| Extraction Method | Chemistry | Vendor/Supplier | Reason for Inclusion in the Study |
|---|---|---|---|
| RNeasy Universal Plus Mini Kit (Method 1) | Silica column | Qiagen | RNA extraction |
| QIAzol Silica Magnetic Beads (Method 2) | Silica magnetic beads | our lab-optimized protocol | RNA extraction, no clogging by cell debris |
| MagJET DNA and RNA Kit (Method 3) | Paramagnetic beads | ThermoFisher | Total nucleic acid extraction, no clogging by cell debris |
| MagNA Pure 96 System (Method 4) | Magnetic glass particles | Roche | Total nucleic acid extraction, high throughput |
| QIAzol Proteinase K Silica Magnetic Beads (Method 5) | Silica magnetic beads | our lab-optimized protocol | RNA extraction, no clogging by cell debris |
| QIAzol Proteinase K Paramagnetic beads (Method 6) | Paramagnetic beads | our lab-optimized protocol | RNA extraction, no clogging by cell debris |
| Spiked Serial Dilution | Method 2 | Method 3 | Method 4 |
|---|---|---|---|
| LGTV10−3 | 7.73 | 12.85 | 0.49 |
| LGTV10−4 | 6.59 | 12.14 | 0.03 |
| LGTV10−5 | 7.81 | 9.42 | 0.21 |
| LGTV10−6 | 2.65 | 3.73 | −0.49 |
| Extraction | Sample | Ct Value | Total Reads | Mapped Reads N (%) |
|---|---|---|---|---|
| Method 1 | LGTV10−3 replicate 1 | 26.38 | 4,829,403 | 5367 (0.11%) |
| LGTV10−3 replicate 2 | 4,528,711 | 17,536 (0.39%) | ||
| Method 2 | LGTV10−3 replicate 1 | 22.98 | 6,876,209 | 20,016 (0.29%) |
| LGTV10−3 replicate 2 | 4,398,740 | 11,821 (0.27%) | ||
| Method 3 | LGTV10−3 replicate 1 | 22.04 | 3,588,781 | 703,931 (19.61%) |
| LGTV10−3 replicate 2 | 4,426,097 | 882,244 (19.93%) | ||
| Method 4 | LGTV10−3 replicate 1 | 25.63 | 3,748,017 | 31,925 (0.85%) |
| LGTV10−3 replicate 2 | 2,526,069 | 46,636 (1.85%) |
| Extraction | Sample | Ct Value | Total Reads | Mapped Reads N (%) |
|---|---|---|---|---|
| Method 2 | LGTV10−3 replicate 1 | 23.53 | 4,236,072 | 27,391 (0.65%) |
| LGTV10−3 replicate 2 | 2,087,367 | 12,816 (0.61%) | ||
| Method 3 | LGTV10−3 replicate 1 | 22.77 | 5,766,461 | 688,769 (11.94%) |
| LGTV10−3 replicate 2 | 3,053,077 | 352,335 (11.54%) | ||
| Method 5 | LGTV10−3 replicate 1 | 22.72 | 2,796,041 | 844,300 (30.20%) |
| LGTV10−3 replicate 2 | 2,355,912 | 693,898 (29.45%) | ||
| Method 6 | LGTV10−3 replicate 1 | 26.58 | 3,320,147 | 100,494 (3.03%) |
| LGTV10−3 replicate 2 | 5,036,970 | 153,703 (3.05%) |
| Sample | Ct Value | Total Number Reads | % Classified Reads | % Viral Reads | % Bacterial Reads | % Chordate Reads | % Protozoan Reads | % Fungal Reads |
|---|---|---|---|---|---|---|---|---|
| TBEV10−1 | 27.22 | 3,607,592 | 47.4 | 0.21 | 13.4 | 32.2 | 0.0 | 0.0 |
| TBEV10−1 R | 3,465,868 | 38.3 | 0.228 | 11.1 | 25.3 | 0.0 | 0.0 | |
| TBEV10−2 | 30.59 | 3,198,211 | 34.8 | 0.148 | 10.9 | 22.4 | 0.0 | 0.0 |
| TBEV10−2 R | 4,262,864 | 35.1 | 0.129 | 9.0 | 24.6 | 0.0 | 0.0 | |
| TBEV10−3 | 34 | 3,155,979 | 58.9 | 0.193 | 31.5 | 26.3 | 0.0 | 0.0 |
| TBEV10−3 R | 2,852,515 | 48.8 | 0.151 | 25.4 | 21.9 | 0.0 | 0.0 | |
| TBEV10−4 | 37.29 | 3,222,131 | 70.3 | 0.038 | 43.5 | 26.4 | 0.0 | 0.0 |
| TBEV10−4 R | 3,082,738 | 70.9 | 0.0507 | 42.8 | 27.8 | 0.0 | 0.0 | |
| TBEV10−5 | 39.16 | 3,196,999 | 62.7 | 0.0701 | 26.5 | 35.9 | 0.0 | 0.0 |
| TBEV10−5 R | 2,572,154 | 49.4 | 0.117 | 39.6 | 9.32 | 0.0 | 0.0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akello, J.O.; Leib, S.L.; Engler, O.; Beuret, C. Evaluation of Viral RNA Recovery Methods in Vectors by Metagenomic Sequencing. Viruses 2020, 12, 562. https://doi.org/10.3390/v12050562
Akello JO, Leib SL, Engler O, Beuret C. Evaluation of Viral RNA Recovery Methods in Vectors by Metagenomic Sequencing. Viruses. 2020; 12(5):562. https://doi.org/10.3390/v12050562
Chicago/Turabian StyleAkello, Joyce Odeke, Stephen L. Leib, Olivier Engler, and Christian Beuret. 2020. "Evaluation of Viral RNA Recovery Methods in Vectors by Metagenomic Sequencing" Viruses 12, no. 5: 562. https://doi.org/10.3390/v12050562
APA StyleAkello, J. O., Leib, S. L., Engler, O., & Beuret, C. (2020). Evaluation of Viral RNA Recovery Methods in Vectors by Metagenomic Sequencing. Viruses, 12(5), 562. https://doi.org/10.3390/v12050562





