Virome of a Feline Outbreak of Diarrhea and Vomiting Includes Bocaviruses and a Novel Chapparvovirus
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Pathogen Screening
2.2. Viral Metagenomics Analysis
2.3. Genome Assembly of Novel Chaphamaparvovirus
2.4. Diagnostic PCR and Prevalence
2.5. Phylogenetic Analysis
2.6. Ethics Statement
2.7. Data Availability Statement
3. Results
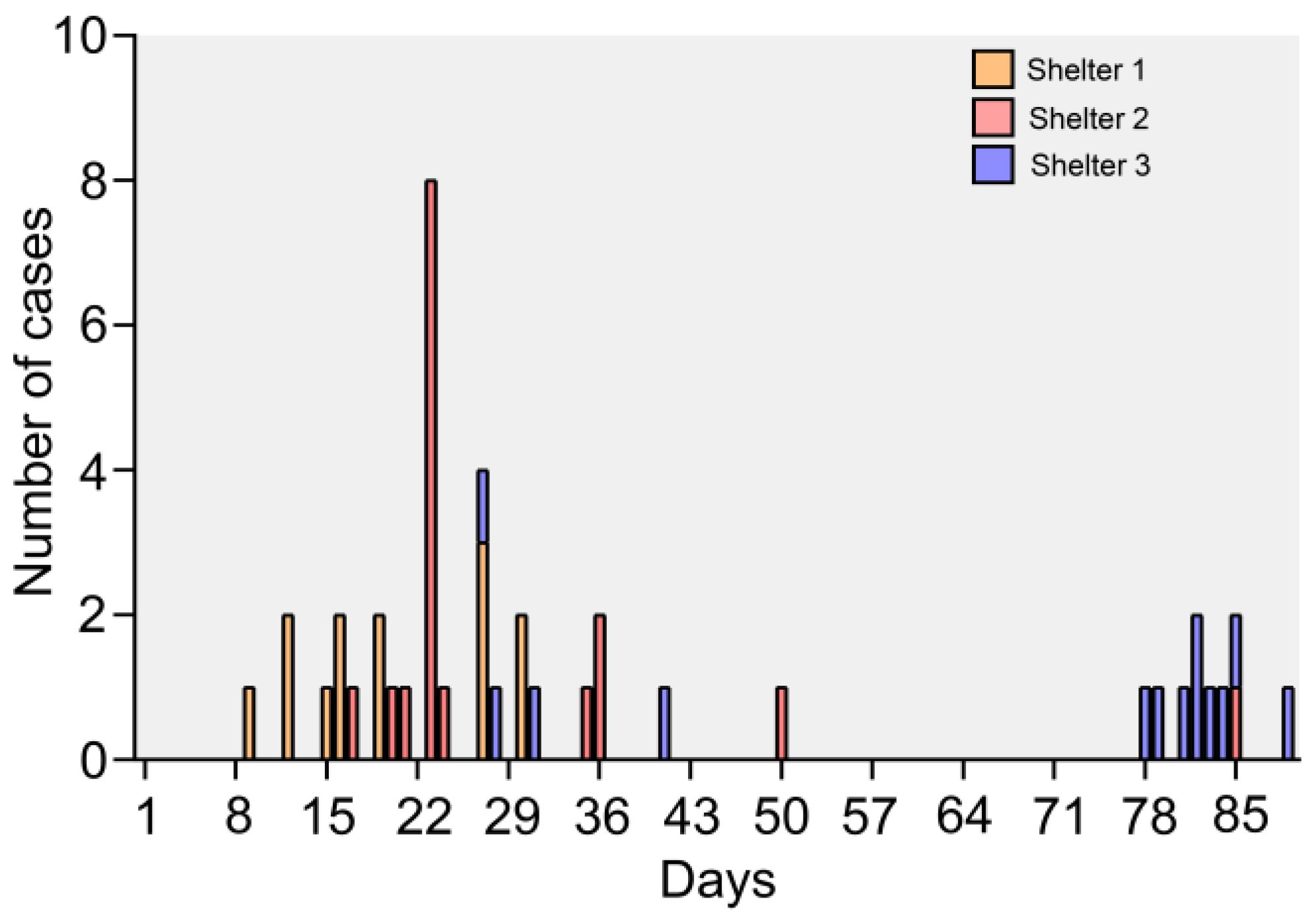
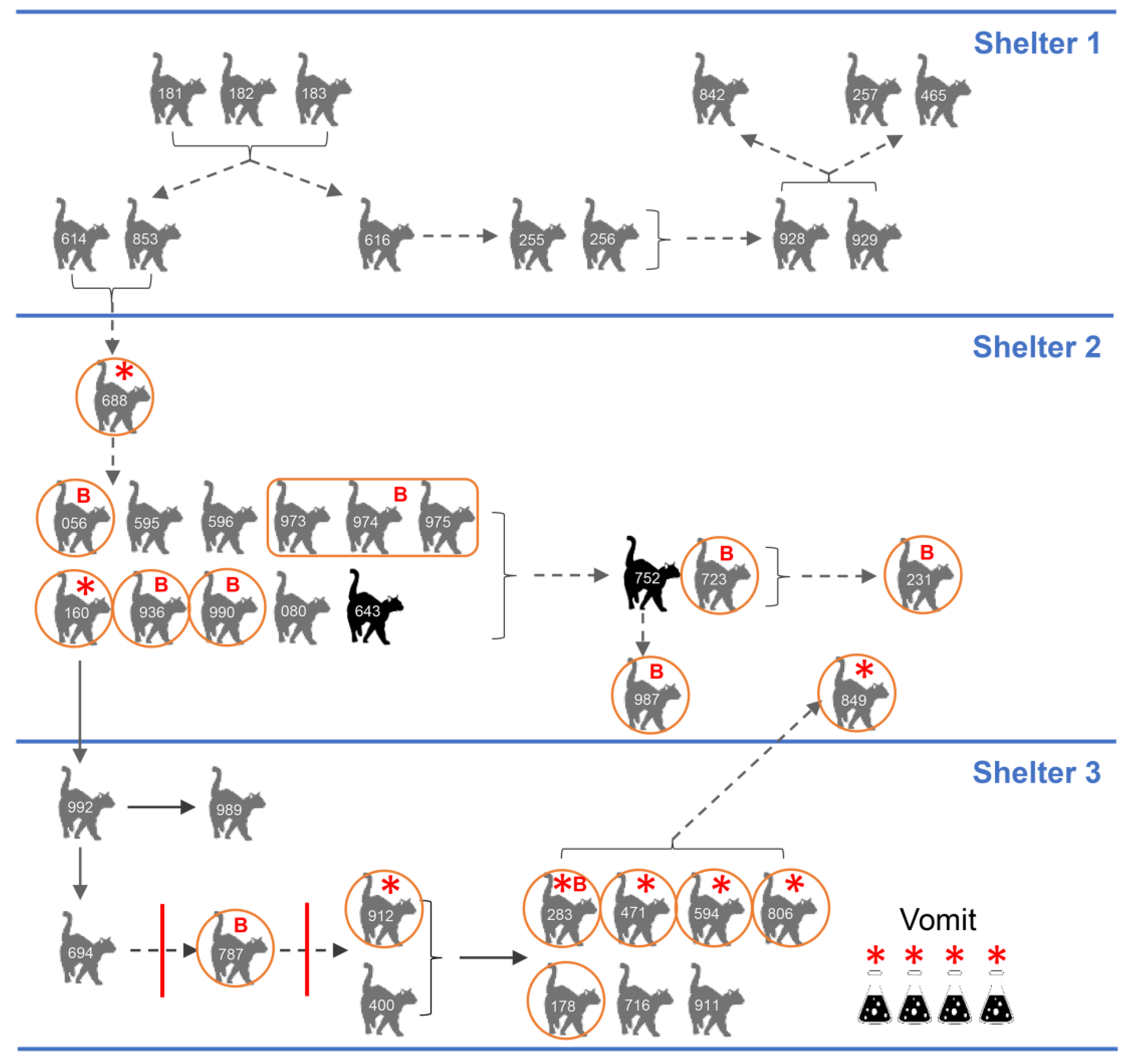
3.1. Epidemic and Clinical Data
3.2. PCR Pathogen Screening
3.3. Viral Metagenomics Analysis
3.4. Anelloviruses, Papillomaviruses and Polyomavirus
3.5. Dietary Contamination
3.6. Vaccine-Derived Sequences
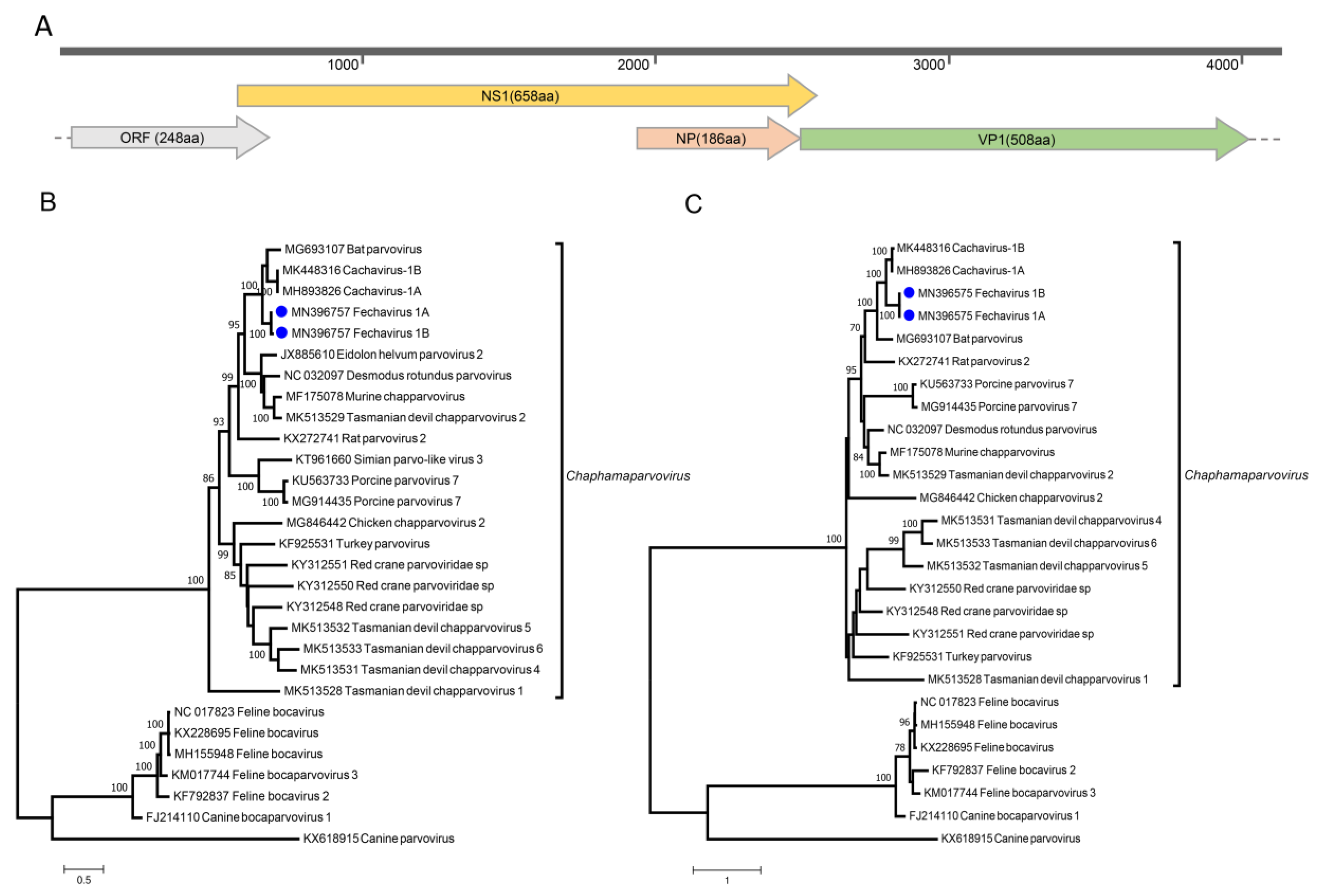
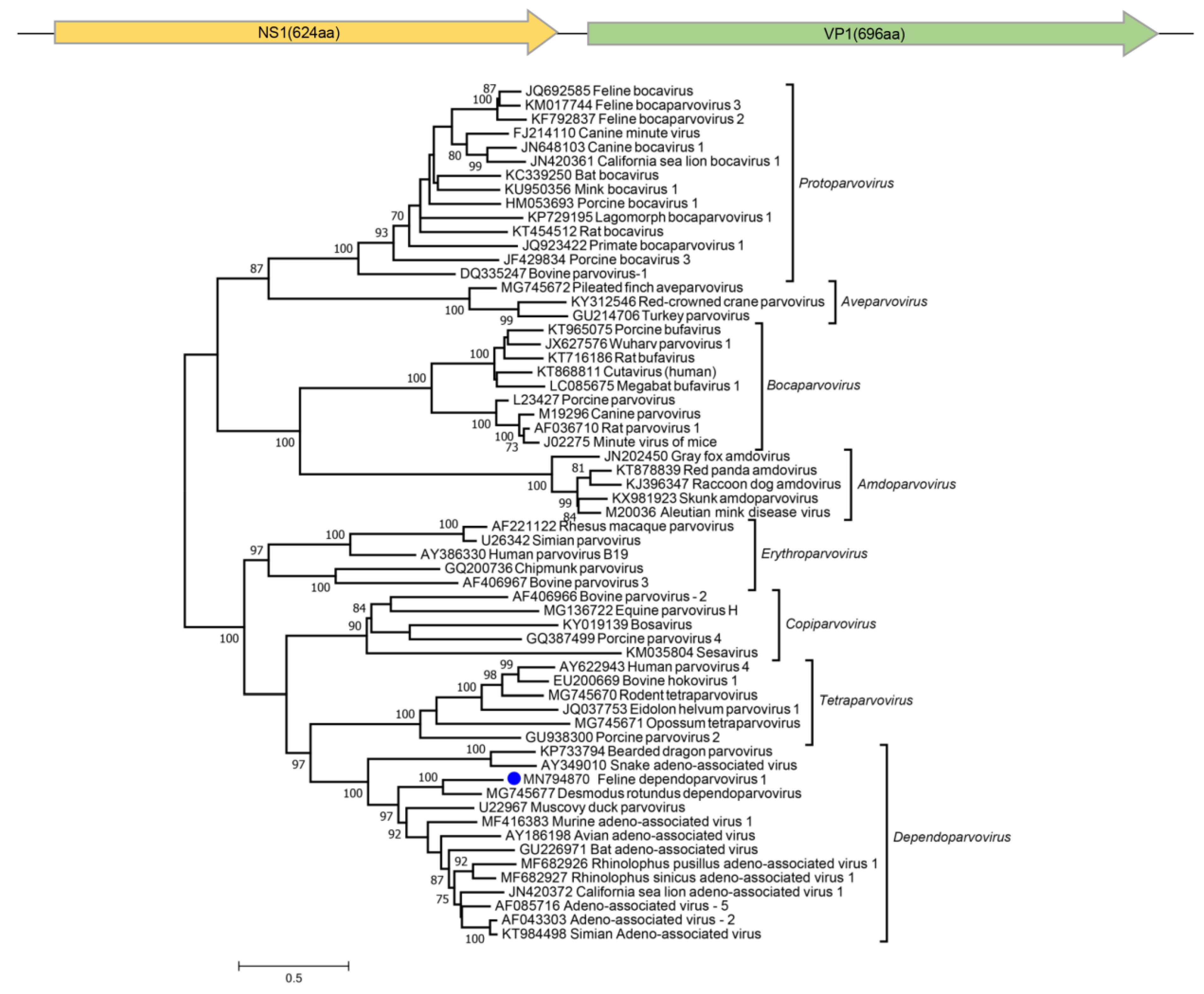
3.7. Parvoviruses
3.8. Epidemiology
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Diakoudi, G.; Lanave, G.; Capozza, P.; Di Profio, F.; Melegari, I.; Di Martino, B.; Pennisi, M.G.; Elia, G.; Cavalli, A.; Tempesta, M.; et al. Identification of a novel parvovirus in domestic cats. Vet. Microbiol. 2019, 228, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Pesavento, P.A.; Murphy, B.G. Common and emerging infectious diseases in the animal shelter. Vet. Pathol. 2014, 51, 478–491. [Google Scholar] [CrossRef] [PubMed]
- Ng, T.F.; Mesquita, J.R.; Nascimento, M.S.; Kondov, N.O.; Wong, W.; Reuter, G.; Knowles, N.J.; Vega, E.; Esona, M.D.; Deng, X.; et al. Feline fecal virome reveals novel and prevalent enteric viruses. Vet. Microbiol. 2014, 171, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, B.; Di Profio, F.; Melegari, I.; Marsilio, F. Feline Virome-A Review of Novel Enteric Viruses Detected in Cats. Viruses 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, L.; Deng, X.; Kapusinszky, B.; Pesavento, P.A.; Delwart, E. Faecal virome of cats in an animal shelter. J. Gen. Virol. 2014, 95, 2553–2564. [Google Scholar] [CrossRef] [PubMed]
- Otto, P.H.; Rosenhain, S.; Elschner, M.C.; Hotzel, H.; Machnowska, P.; Trojnar, E.; Hoffmann, K.; Johne, R. Detection of rotavirus species A, B and C in domestic mammalian animals with diarrhoea and genotyping of bovine species A rotavirus strains. Vet. Microbiol. 2015, 179, 168–176. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Tattersall, P. Parvoviruses: Small Does Not Mean Simple. Annu. Rev. Virol. 2014, 1, 517–537. [Google Scholar] [CrossRef]
- Mietzsch, M.; Penzes, J.J.; Agbandje-McKenna, M. Twenty-Five Years of Structural Parvovirology. Viruses 2019, 11. [Google Scholar] [CrossRef]
- Cotmore, S.F.; Agbandje-McKenna, M.; Canuti, M.; Chiorini, J.A.; Eis-Hubinger, A.M.; Hughes, J.; Mietzsch, M.; Modha, S.; Ogliastro, M.; Penzes, J.J.; et al. ICTV Virus Taxonomy Profile: Parvoviridae. J. Gen. Virol. 2019, 100, 367–368. [Google Scholar] [CrossRef]
- Penzes, J.J.; de Souza, W.M.; Agbandje-McKenna, M.; Gifford, R.J. An Ancient Lineage of Highly Divergent Parvoviruses Infects both Vertebrate and Invertebrate Hosts. Viruses 2019, 11. [Google Scholar] [CrossRef]
- Souza, W.M.; Romeiro, M.F.; Fumagalli, M.J.; Modha, S.; de Araujo, J.; Queiroz, L.H.; Durigon, E.L.; Figueiredo, L.T.; Murcia, P.R.; Gifford, R.J. Chapparvoviruses occur in at least three vertebrate classes and have a broad biogeographic distribution. J. Gen. Virol. 2017, 98, 225–229. [Google Scholar] [CrossRef]
- Yang, S.; Liu, Z.; Wang, Y.; Li, W.; Fu, X.; Lin, Y.; Shen, Q.; Wang, X.; Wang, H.; Zhang, W. A novel rodent Chapparvovirus in feces of wild rats. Virol. J. 2016, 13, 133. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.H.; Che, X.; Garcia, J.A.; Klena, J.D.; Lee, B.; Muller, D.; Ulrich, W.; Corrigan, R.M.; Nichol, S.; Jain, K.; et al. Viral Diversity of House Mice in New York City. MBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Yinda, C.K.; Ghogomu, S.M.; Conceicao-Neto, N.; Beller, L.; Deboutte, W.; Vanhulle, E.; Maes, P.; Van Ranst, M.; Matthijnssens, J. Cameroonian fruit bats harbor divergent viruses, including rotavirus H, bastroviruses, and picobirnaviruses using an alternative genetic code. Virus Evol. 2018, 4, vey008. [Google Scholar] [CrossRef] [PubMed]
- Baker, K.S.; Leggett, R.M.; Bexfield, N.H.; Alston, M.; Daly, G.; Todd, S.; Tachedjian, M.; Holmes, C.E.; Crameri, S.; Wang, L.F.; et al. Metagenomic study of the viruses of African straw-coloured fruit bats: Detection of a chiropteran poxvirus and isolation of a novel adenovirus. Virology 2013, 441, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Kapusinszky, B.; Ardeshir, A.; Mulvaney, U.; Deng, X.; Delwart, E. Case-Control Comparison of Enteric Viromes in Captive Rhesus Macaques with Acute or Idiopathic Chronic Diarrhea. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Sawaswong, V.; Fahsbender, E.; Altan, E.; Kemthong, T.; Deng, X.; Malaivijitnond, S.; Payungporn, S.; Delwart, E. High Diversity and Novel Enteric Viruses in Fecal Viromes of Healthy Wild and Captive Thai Cynomolgus Macaques (Macaca fascicularis). Viruses 2019, 11. [Google Scholar] [CrossRef]
- Fahsbender, E.; Altan, E.; Seguin, M.A.; Young, P.; Estrada, M.; Leutenegger, C.; Delwart, E. Chapparvovirus DNA Found in 4% of Dogs with Diarrhea. Viruses 2019, 11. [Google Scholar] [CrossRef]
- Palinski, R.M.; Mitra, N.; Hause, B.M. Discovery of a novel Parvovirinae virus, porcine parvovirus 7, by metagenomic sequencing of porcine rectal swabs. Virus Genes 2016, 52, 564–567. [Google Scholar] [CrossRef]
- Chong, R.; Shi, M.; Grueber, C.E.; Holmes, E.C.; Hogg, C.J.; Belov, K.; Barrs, V.R. Fecal Viral Diversity of Captive and Wild Tasmanian Devils Characterized Using Virion-Enriched Metagenomics and Metatranscriptomics. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, S.; Liu, D.; Zhou, C.; Li, W.; Lin, Y.; Wang, X.; Shen, Q.; Wang, H.; Li, C.; et al. The fecal virome of red-crowned cranes. Arch. Virol. 2019, 164, 3–16. [Google Scholar] [CrossRef] [PubMed]
- Lima, D.A.; Cibulski, S.P.; Tochetto, C.; Varela, A.P.M.; Finkler, F.; Teixeira, T.F.; Loiko, M.R.; Cerva, C.; Junqueira, D.M.; Mayer, F.Q.; et al. The intestinal virome of malabsorption syndrome-affected and unaffected broilers through shotgun metagenomics. Virus Res. 2019, 261, 9–20. [Google Scholar] [CrossRef]
- Reuter, G.; Boros, A.; Delwart, E.; Pankovics, P. Novel circular single-stranded DNA virus from turkey faeces. Arch. Virol. 2014, 159, 2161–2164. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Wang, W.; Chan, J.F.; Wang, G.; Huang, Y.; Yi, Y.; Zhu, Z.; Peng, R.; Hu, X.; Wu, Y.; et al. Identification of a Novel Ichthyic Parvovirus in Marine Species in Hainan Island, China. Front. Microbiol. 2019, 10, 2815. [Google Scholar] [CrossRef] [PubMed]
- Roediger, B.; Lee, Q.; Tikoo, S.; Cobbin, J.C.A.; Henderson, J.M.; Jormakka, M.; O’Rourke, M.B.; Padula, M.P.; Pinello, N.; Henry, M.; et al. An Atypical Parvovirus Drives Chronic Tubulointerstitial Nephropathy and Kidney Fibrosis. Cell 2018, 175, 530–543.e524. [Google Scholar] [CrossRef]
- Lee, Q.; Padula, M.P.; Pinello, N.; Williams, S.H.; O’Rourke, M.B.; Fumagalli, M.J.; Orkin, J.D.; Song, R.; Shaban, B.; Brenner, O.; et al. Murine and related chapparvoviruses are nephro-tropic and produce novel accessory proteins in infected kidneys. PLoS Pathog. 2020, 16, e1008262. [Google Scholar] [CrossRef]
- Andersen, L.A.; Levy, J.K.; McManus, C.M.; McGorray, S.P.; Leutenegger, C.M.; Piccione, J.; Blackwelder, L.K.; Tucker, S.J. Prevalence of enteropathogens in cats with and without diarrhea in four different management models for unowned cats in the southeast United States. Vet. J. 2018, 236, 49–55. [Google Scholar] [CrossRef]
- Li, L.; Deng, X.; Mee, E.T.; Collot-Teixeira, S.; Anderson, R.; Schepelmann, S.; Minor, P.D.; Delwart, E. Comparing viral metagenomics methods using a highly multiplexed human viral pathogens reagent. J. Virol. Methods 2015, 213, 139–146. [Google Scholar] [CrossRef]
- Ye, J.; McGinnis, S.; Madden, T.L. BLAST: Improvements for better sequence analysis. Nucleic Acids Res. 2006, 34, W6–W9. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Deng, X.; Naccache, S.N.; Ng, T.; Federman, S.; Li, L.; Chiu, C.Y.; Delwart, E.L. An ensemble strategy that significantly improves de novo assembly of microbial genomes from metagenomic next-generation sequencing data. Nucleic Acids Res. 2015, 43, e46. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Niu, J.; Yi, S.; Dong, G.; Yu, D.; Guo, Y.; Huang, H.; Hu, G. Development and application of a multiplex PCR method for the simultaneous detection and differentiation of feline panleukopenia virus, feline bocavirus, and feline astrovirus. Arch. Virol. 2019, 164, 2761–2768. [Google Scholar] [CrossRef] [PubMed]
- Omidbakhsh, N.; Sattar, S.A. Broad-spectrum microbicidal activity, toxicologic assessment, and materials compatibility of a new generation of accelerated hydrogen peroxide-based environmental surface disinfectant. Am. J. Infect. Control 2006, 34, 251–257. [Google Scholar] [CrossRef]
- Marks, S.L.; Rankin, S.C.; Byrne, B.A.; Weese, J.S. Enteropathogenic bacteria in dogs and cats: Diagnosis, epidemiology, treatment, and control. J. Vet. Intern. Med. 2011, 25, 1195–1208. [Google Scholar] [CrossRef]
- Gizzi, A.B.; Oliveira, S.T.; Leutenegger, C.M.; Estrada, M.; Kozemjakin, D.A.; Stedile, R.; Marcondes, M.; Biondo, A.W. Presence of infectious agents and co-infections in diarrheic dogs determined with a real-time polymerase chain reaction-based panel. BMC Vet. Res. 2014, 10, 23. [Google Scholar] [CrossRef]
- Adam, F. Infectious Diseases of the Dog and Cat, 4th edition. J. Small Anim. Pract. 2014, 55, E4. [Google Scholar] [CrossRef][Green Version]
- Janeczko, S.; Griffin, B. Giardia infection in cats. Compend. Contin. Educ. Vet. 2010, 32, E4. [Google Scholar]
- Moustafa, A.; Xie, C.; Kirkness, E.; Biggs, W.; Wong, E.; Turpaz, Y.; Bloom, K.; Delwart, E.; Nelson, K.E.; Venter, J.C.; et al. The blood DNA virome in 8,000 humans. PLoS Pathog. 2017, 13, e1006292. [Google Scholar] [CrossRef]
- Li, Y.; Fu, X.; Ma, J.; Zhang, J.; Hu, Y.; Dong, W.; Wan, Z.; Li, Q.; Kuang, Y.Q.; Lan, K.; et al. Altered respiratory virome and serum cytokine profile associated with recurrent respiratory tract infections in children. Nat. Commun. 2019, 10, 2288. [Google Scholar] [CrossRef]
- Fahsbender, E.; Altan, E.; Estrada, M.; Seguin, M.A.; Young, P.; Leutenegger, C.M.; Delwart, E. Lyon-IARC Polyomavirus DNA in Feces of Diarrheic Cats. Microbiol. Resour. Announc. 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Schat, K.A. Chicken anemia virus. Curr. Top. Microbiol. Immunol. 2009, 331, 151–183. [Google Scholar] [CrossRef] [PubMed]
- Gia Phan, T.; Phung Vo, N.; Sdiri-Loulizi, K.; Aouni, M.; Pothier, P.; Ambert-Balay, K.; Deng, X.; Delwart, E. Divergent gyroviruses in the feces of Tunisian children. Virology 2013, 446, 346–348. [Google Scholar] [CrossRef] [PubMed]
- Niu, J.T.; Yi, S.S.; Dong, G.Y.; Guo, Y.B.; Zhao, Y.L.; Huang, H.L.; Wang, K.; Hu, G.X.; Dong, H. Genomic Characterization of Diverse Gyroviruses Identified in the Feces of Domestic Cats. Sci. Rep. 2019, 9, 13303. [Google Scholar] [CrossRef]
- Xiao, C.T.; Gimenez-Lirola, L.G.; Jiang, Y.H.; Halbur, P.G.; Opriessnig, T. Characterization of a novel porcine parvovirus tentatively designated PPV5. PLoS ONE 2013, 8, e65312. [Google Scholar] [CrossRef]
- Radford, A.D.; Coyne, K.P.; Dawson, S.; Porter, C.J.; Gaskell, R.M. Feline calicivirus. Vet. Res. 2007, 38, 319–335. [Google Scholar] [CrossRef]
- Balboni, A.; Bassi, F.; De Arcangeli, S.; Zobba, R.; Dedola, C.; Alberti, A.; Battilani, M. Molecular analysis of carnivore Protoparvovirus detected in white blood cells of naturally infected cats. BMC Vet. Res. 2018, 14, 41. [Google Scholar] [CrossRef]
- Battilani, M.; Balboni, A.; Ustulin, M.; Giunti, M.; Scagliarini, A.; Prosperi, S. Genetic complexity and multiple infections with more Parvovirus species in naturally infected cats. Vet. Res. 2011, 42, 43. [Google Scholar] [CrossRef]
- Abd-Eldaim, M.; Potgieter, L.; Kennedy, M. Genetic analysis of feline caliciviruses associated with a hemorrhagic-like disease. J. Vet. Diagn. Investig. 2005, 17, 420–429. [Google Scholar] [CrossRef]
- Hou, J.; Sanchez-Vizcaino, F.; McGahie, D.; Lesbros, C.; Almeras, T.; Howarth, D.; O’Hara, V.; Dawson, S.; Radford, A.D. European molecular epidemiology and strain diversity of feline calicivirus. Vet. Rec. 2016, 178, 114–115. [Google Scholar] [CrossRef]
- Takano, T.; Takadate, Y.; Doki, T.; Hohdatsu, T. Genetic characterization of feline bocavirus detected in cats in Japan. Arch. Virol. 2016, 161, 2825–2828. [Google Scholar] [CrossRef] [PubMed]
- Yi, S.; Niu, J.; Wang, H.; Dong, G.; Zhao, Y.; Dong, H.; Guo, Y.; Wang, K.; Hu, G. Detection and genetic characterization of feline bocavirus in Northeast China. Virol. J. 2018, 15, 125. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Woo, P.C.; Yeung, H.C.; Teng, J.L.; Wu, Y.; Bai, R.; Fan, R.Y.; Chan, K.H.; Yuen, K.Y. Identification and characterization of bocaviruses in cats and dogs reveals a novel feline bocavirus and a novel genetic group of canine bocavirus. J. Gen. Virol. 2012, 93, 1573–1582. [Google Scholar] [CrossRef] [PubMed]
- Jartti, T.; Hedman, K.; Jartti, L.; Ruuskanen, O.; Allander, T.; Söderlund-Venermo, M. Human bocavirus-the first 5 years. Rev. Med. Virol. 2012, 22, 46–64. [Google Scholar] [CrossRef]
- Ong, D.S.; Schuurman, R.; Heikens, E. Human bocavirus in stool: A true pathogen or an innocent bystander? J. Clin. Virol. 2016, 74, 45–49. [Google Scholar] [CrossRef]
- Kapoor, A.; Simmonds, P.; Slikas, E.; Li, L.; Bodhidatta, L.; Sethabutr, O.; Triki, H.; Bahri, O.; Oderinde, B.S.; Baba, M.M.; et al. Human bocaviruses are highly diverse, dispersed, recombination prone, and prevalent in enteric infections. J. Infect. Dis. 2010, 201, 1633–1643. [Google Scholar] [CrossRef]
- Piewbang, C.; Kasantikul, T.; Pringproa, K.; Techangamsuwan, S. Feline bocavirus-1 associated with outbreaks of hemorrhagic enteritis in household cats: Potential first evidence of a pathological role, viral tropism and natural genetic recombination. Sci. Rep. 2019, 9, 16367. [Google Scholar] [CrossRef]
- Barrs, V.R. Feline Panleukopenia: A Re-emergent Disease. Vet. Clin. N. Am. Small Anim. Pract. 2019. [Google Scholar] [CrossRef]
- Truyen, U.; Parrish, C.R. Feline panleukopenia virus: Its interesting evolution and current problems in immunoprophylaxis against a serious pathogen. Vet. Microbiol. 2013, 165, 29–32. [Google Scholar] [CrossRef]
- Stuetzer, B.; Hartmann, K. Feline parvovirus infection and associated diseases. Vet. J. 2014, 201, 150–155. [Google Scholar] [CrossRef]
- Martella, V.; Lanave, G.; Mihalov-Kovacs, E.; Marton, S.; Varga-Kugler, R.; Kaszab, E.; Di Martino, B.; Camero, M.; Decaro, N.; Buonavoglia, C.; et al. Novel Parvovirus Related to Primate Bufaviruses in Dogs. Emerg. Infect. Dis. 2018, 24, 1061–1068. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Cui, L.; Deng, X.; Yu, X.; Zhang, Z.; Yang, Z.; Delwart, E.; Zhang, W.; Hua, X. Canine bufavirus in faeces and plasma of dogs with diarrhoea, China. Emerg. Microbes Infect. 2019, 8, 245–247. [Google Scholar] [CrossRef] [PubMed]
- Berns, K.I. Parvovirus replication. Microbiol. Rev. 1990, 54, 316–329. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hoelzer, K.; Shackelton, L.A.; Parrish, C.R.; Holmes, E.C. Phylogenetic analysis reveals the emergence, evolution and dispersal of carnivore parvoviruses. J. Gen. Virol. 2008, 89, 2280–2289. [Google Scholar] [CrossRef]
- Allison, A.B.; Kohler, D.J.; Fox, K.A.; Brown, J.D.; Gerhold, R.W.; Shearn-Bochsler, V.I.; Dubovi, E.J.; Parrish, C.R.; Holmes, E.C. Frequent cross-species transmission of parvoviruses among diverse carnivore hosts. J. Virol. 2013, 87, 2342–2347. [Google Scholar] [CrossRef]
- Bergmann, M.; Schwertler, S.; Speck, S.; Truyen, U.; Reese, S.; Hartmann, K. Faecal shedding of parvovirus deoxyribonucleic acid following modified live feline panleucopenia virus vaccination in healthy cats. Vet. Rec. 2019, 185, 83. [Google Scholar] [CrossRef]
| Exposed (Presumptive) | ||||
|---|---|---|---|---|
| Location | Sick | Not Sick | Total | AR% |
| Shelter 2 * | 8 | 4 | 12 | 66.7% |
| Shelter 3 ** | 10 | 2 | 12 | 83.3% |
| Clinical Signs | #of Cats (n = 43) | Percentage |
|---|---|---|
| Diarrhea | 35 | 81.4% |
| Vomiting | 29 | 67.4% |
| Inappetence | 11 | 25.6% |
| Lethargy | 5 | 11.6% |
| Required veterinary visit | 29 | 67.4% |
| Viral Read Numbers | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Shelter 2 | Shelter 3 | ||||||||||||||||||
| Cat ID | 688 | 056 | 160 | 973–975 | 990 | 936 | 723 | 987 | 231 | 787 | 912 | 283 | 178 | 806 | 471 | 594 | 849 | ||
| Total reads (106) | 0.76 | 0.77 | 1.24 | 1.11 | 1.35 | 1.02 | 3.16 | 1.88 | 0.68 | 1.29 | 1.23 | 1.15 | 1.41 | 1.20 | 1.76 | 1.21 | 1.36 | ||
| Family | Genus/Species | ||||||||||||||||||
| Anelloviridae | Gyrovirus | ||||||||||||||||||
| Chicken anemia virus | 2 | ||||||||||||||||||
| Gyrovirus 4 | 14 | ||||||||||||||||||
| Gyrovirus 6 | 2 | ||||||||||||||||||
| Feline anellovirus | 4 | 6 | 70 | ||||||||||||||||
| Other anellovirus | 2 | 44 | 402 | 46 | |||||||||||||||
| Parvoviridae | Bocaparvovirus | ||||||||||||||||||
| FeBoV1 | 34 | 2 | 1547 | 1547 | |||||||||||||||
| FeBoV2 | 63,826 | 8191 | 2 | ||||||||||||||||
| FeBoV3 | 1350 | 12 | |||||||||||||||||
| Chaphamaparvovirus | |||||||||||||||||||
| Fechavirus | 2 | 8 | 789 | 17 | 1597 | 6511 | |||||||||||||
| Copiparvovirus | |||||||||||||||||||
| Porcine parvovirus 5 | 2 | ||||||||||||||||||
| Dependoparvovirus | |||||||||||||||||||
| Feline dependoparvovirus | 19 | 8315 | |||||||||||||||||
| Protoparvovirus | |||||||||||||||||||
| Felocell 3 vaccine FPV | 112 | 2086 | |||||||||||||||||
| Papillomaviridae | Dyothetapapillomavirus | 7 | 93 | ||||||||||||||||
| Polyomaviridae | LIPyV | 838 | |||||||||||||||||
| Caliciviridae | Felocell 3 vaccine calicivirus | 1 | 2 | 4 | 3 | ||||||||||||||
| PCR test | |||||||||||||||||||
| Fechavirus | + | - | + | − | − | − | − | − | − | − | + | + | − | + | + | + | + | ||
| FeBoV | − | + | − | + | + | + | − | − | + | + | − | + | − | − | − | − | − | ||
| IDEXX diarrhea panel | − | − | − | FPV | + a | − | ND | ND | ND | + b | + a | + c | ND | + b | − | + a | ND | ||
| Rotavirus | − | − | − | − | − | − | ND | − | ND | ND | − | − | − | − | − | − | − | ||
| ID | D1 | D2 | D3 | D4 | D5 | D6 | D7 | D8 | D9 | D10 | D11 | D12 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 688 | + | − | ||||||||||
| 912 | + | − | − | − | ||||||||
| 594 | + | + | + | + | + | + | + | + | ||||
| 283 | − | + | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Gordon, E.; Idle, A.; Altan, E.; Seguin, M.A.; Estrada, M.; Deng, X.; Delwart, E. Virome of a Feline Outbreak of Diarrhea and Vomiting Includes Bocaviruses and a Novel Chapparvovirus. Viruses 2020, 12, 506. https://doi.org/10.3390/v12050506
Li Y, Gordon E, Idle A, Altan E, Seguin MA, Estrada M, Deng X, Delwart E. Virome of a Feline Outbreak of Diarrhea and Vomiting Includes Bocaviruses and a Novel Chapparvovirus. Viruses. 2020; 12(5):506. https://doi.org/10.3390/v12050506
Chicago/Turabian StyleLi, Yanpeng, Emilia Gordon, Amanda Idle, Eda Altan, M. Alexis Seguin, Marko Estrada, Xutao Deng, and Eric Delwart. 2020. "Virome of a Feline Outbreak of Diarrhea and Vomiting Includes Bocaviruses and a Novel Chapparvovirus" Viruses 12, no. 5: 506. https://doi.org/10.3390/v12050506
APA StyleLi, Y., Gordon, E., Idle, A., Altan, E., Seguin, M. A., Estrada, M., Deng, X., & Delwart, E. (2020). Virome of a Feline Outbreak of Diarrhea and Vomiting Includes Bocaviruses and a Novel Chapparvovirus. Viruses, 12(5), 506. https://doi.org/10.3390/v12050506






