Efficient Method for Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Cultures, Viral Isolation and Titration
2.2. Viral RNA Extraction and DENV Serotyping
2.3. Primer Design for Amplification of the DENV 5′ and 3′ Ends of the DENV Genome
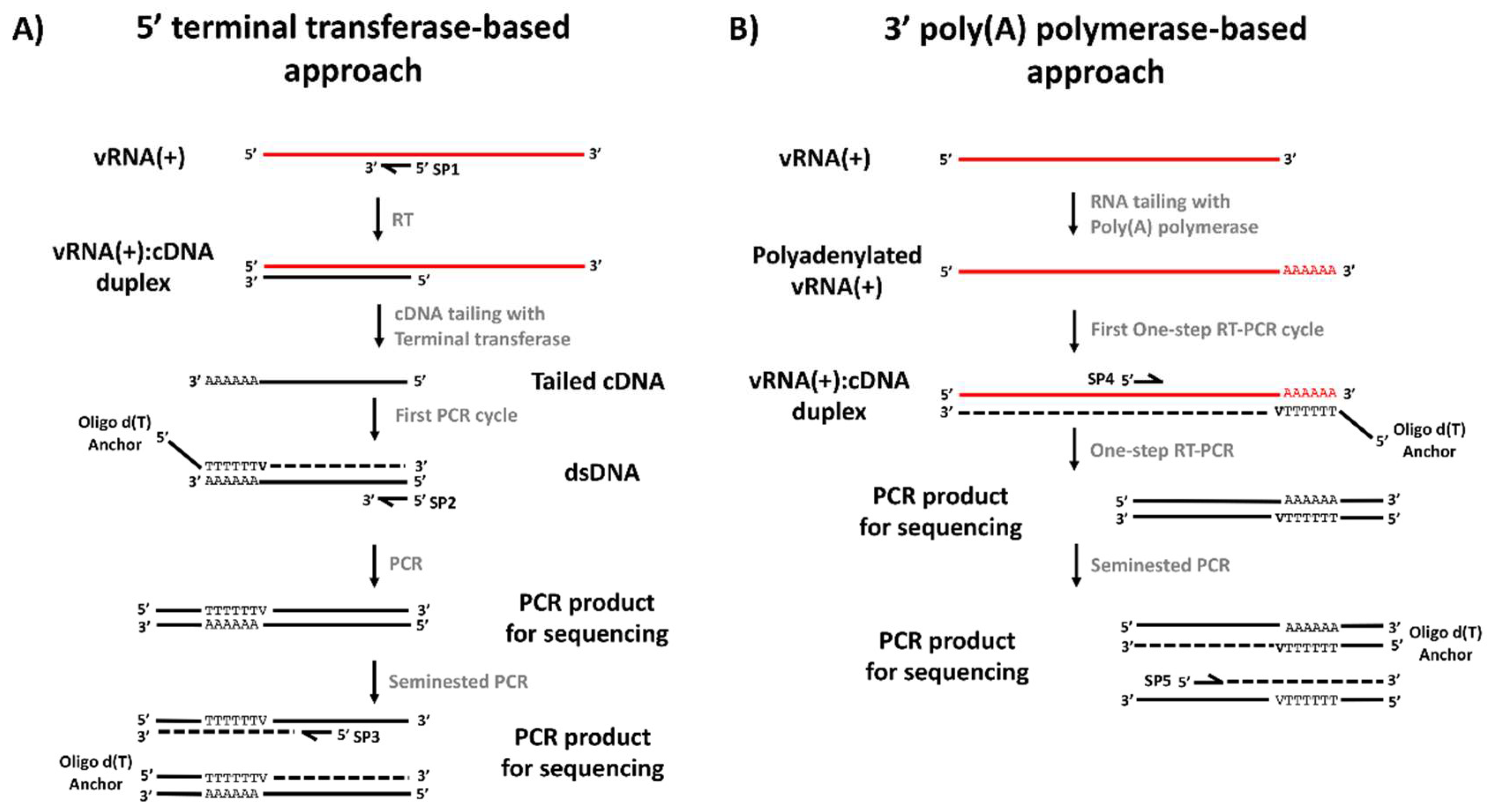
2.4. Reverse Transcription and PCR Amplification of the 5′ Region of the DENV Genome
2.5. Poly (A) Tailing, Reverse Transcription and PCR Amplification of the End of 3′ Untranslated Region
2.6. DNA Sequencing and Sequence Handling
3. Results
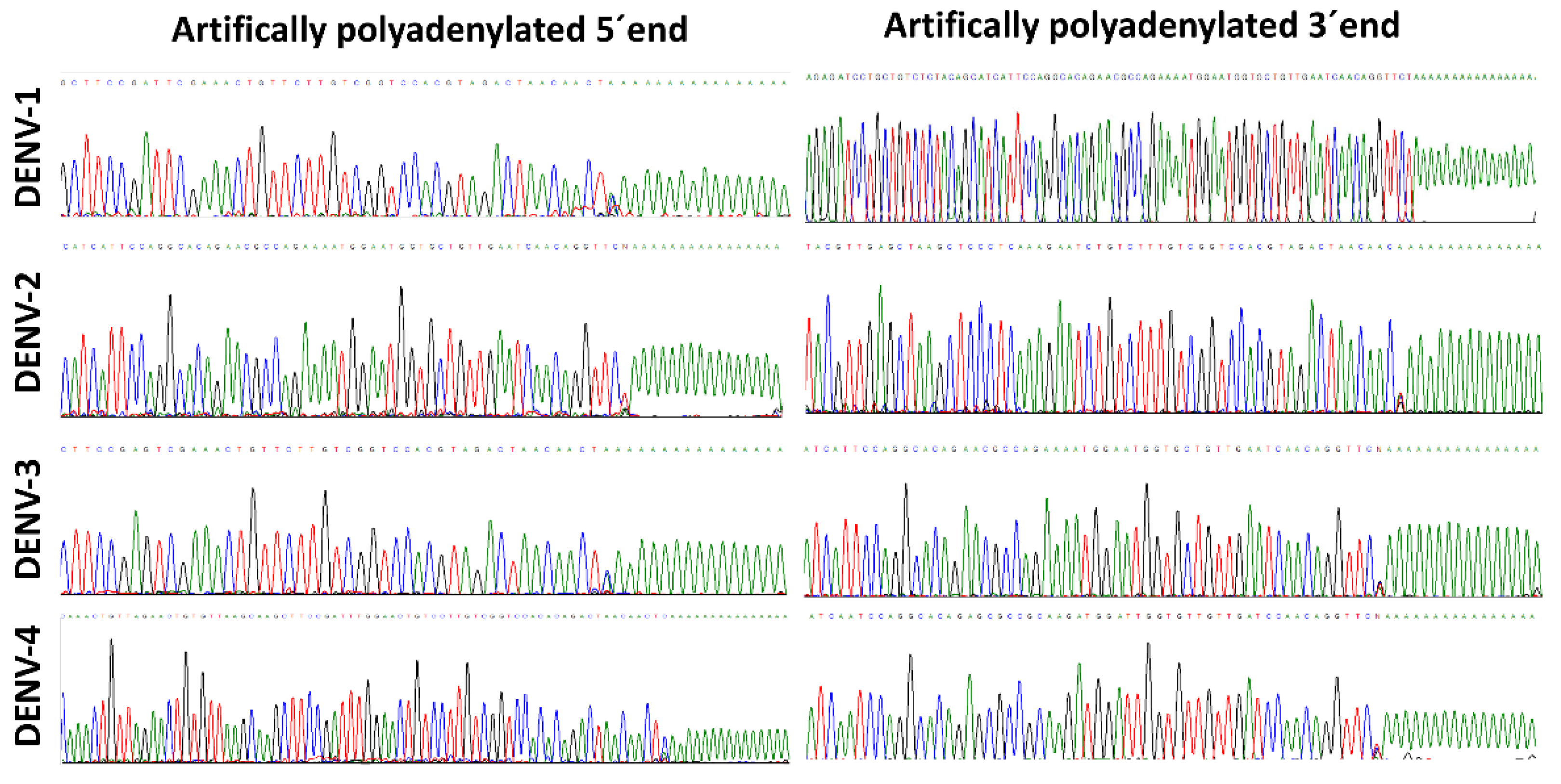
3.1. Successful Amplification of the 5′ and 3′ Regions of All DENV Serotypes
3.2. Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome
3.3. Complete Coverage at the 5′ and 3′ Ends of the DENV Genome
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brathwaite Dick, O.; San Martín, J.L.; Montoya, R.H.; del Diego, J.; Zambrano, B.; Dayan, G.H. The History of Dengue Outbreaks in the Americas. Am. J. Trop. Med. Hyg. 2012, 87, 584–593. [Google Scholar] [CrossRef]
- WHO. Dengue Guidelines for Diagnosis, Treatment, Prevention and Control; World Health Organization, Ed.; WHO: Geneva, Switzerland, 2009; pp. 1–160. [Google Scholar]
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef]
- Twiddy, S.S.; Farrar, J.J.; Vinh Chau, N.; Wills, B.; Gould, E.A.; Gritsun, T.; Lloyd, G.; Holmes, E.C. Phylogenetic relationships and differential selection pressures among genotypes of dengue-2 virus. Virology 2002, 298, 63–72. [Google Scholar] [CrossRef]
- Lindenbach, B.D.; Murray, C.L.; Thiel, H.J.; Rice, C.M. Flaviviridae. In Fields in Virology, 6th ed.; Knipe, D.M., Ed.; Williams & Wilkins Lippincott: Philadelphia, PA, USA, 2013; pp. 712–746. [Google Scholar]
- Clyde, K.; Kyle, J.L.; Harris, E. Recent advances in deciphering viral and host determinants of dengue virus replication and pathogenesis. J. Virol. 2006, 80, 11418–11431. [Google Scholar] [CrossRef]
- Filomatori, C.V.; Lodeiro, M.F.; Alvarez, D.E.; Samsa, M.M.; Pietrasanta, L.; Gamarnik, A.V. A 5′ RNA element promotes dengue virus RNA synthesis on a circular genome. Genes Dev. 2006, 20, 2238–2249. [Google Scholar] [CrossRef]
- Khromykh, A.A.; Meka, H.; Guyatt, K.J.; Westaway, E.G. Essential role of cyclization sequences in flavivirus RNA replication. J. Virol. 2001, 75, 6719–6728. [Google Scholar] [CrossRef]
- Friebe, P.; Harris, E. Interplay of RNA elements in the dengue virus 5′ and 3′ ends required for viral RNA replication. J. Virol. 2010, 84, 6103–6118. [Google Scholar] [CrossRef]
- Clyde, K.; Barrera, J.; Harris, E. The capsid-coding region hairpin element (cHP) is a critical determinant of dengue virus and West Nile virus RNA synthesis. Virology 2008, 379, 314–323. [Google Scholar] [CrossRef]
- Gebhard, L.G.; Filomatori, C.V.; Gamarnik, A.V. Functional RNA Elements in the Dengue Virus Genome. Viruses 2011, 3, 1739–1756. [Google Scholar] [CrossRef]
- Villordo, S.M.; Carballeda, J.M.; Filomatori, C.V.; Gamarnik, A.V. RNA Structure Duplications and Flavivirus Host Adaptation. Trends Microbiol. 2016, 24, 270–283. [Google Scholar] [CrossRef]
- Paranjape, S.M.; Harris, E. Y box-binding protein-1 binds to the dengue virus 3′-untranslated region and mediates antiviral effects. J. Biol. Chem. 2007, 282, 30497–30508. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Yu, M.; Zhang, H.; Wang, H.Y.; Wang, L.F. Improved rapid amplification of cDNA ends (RACE) for mapping both the 5′ and 3′ terminal sequences of paramyxovirus genomes. J. Virol. Methods 2005, 130, 154–156. [Google Scholar] [CrossRef] [PubMed]
- Miller, E. Rapid Amplification of cDNA Ends for RNA Transcript Sequencing in Staphylococcus. Methods Mol. Biol. 2016, 1373, 169–183. [Google Scholar] [CrossRef] [PubMed]
- Usme, J.; Gómez, A.; Gallego, J. Molecular detection and typing of dengue virus by RT-PCR and nested PCR using degenerated oligonucleotides. Rev. Salud Uninorte 2012, 28, 1–15. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Usme-Ciro, J.A.; Mendez, J.A.; Laiton, K.D.; Paez, A. The relevance of dengue virus genotypes surveillance at country level before vaccine approval. Hum. Vaccines Immunother. 2014, 10, 2674–2678. [Google Scholar] [CrossRef]
- IUPAC Codes. Available online: https://www.bioinformatics.org/sms2/iupac.html (accessed on 2 April 2020).
- Chu, P.W.; Westaway, E.G. Replication strategy of Kunjin virus: Evidence for recycling role of replicative form RNA as template in semiconservative and asymmetric replication. Virology 1985, 140, 68–79. [Google Scholar] [CrossRef]
- Leitmeyer, K.C.; Vaughn, D.W.; Watts, D.M.; Salas, R.; Villalobos, I.; de Ramos, C.; Rico-Hesse, R. Dengue virus structural differences that correlate with pathogenesis. J. Virol. 1999, 73, 4738–4747. [Google Scholar] [CrossRef]
- Goo, L.; VanBlargan, L.A.; Dowd, K.A.; Diamond, M.S.; Pierson, T.C. A single mutation in the envelope protein modulates flavivirus antigenicity, stability, and pathogenesis. PLoS Pathog. 2017, 13, e1006178. [Google Scholar] [CrossRef]
- Filomatori, C.V.; Carballeda, J.M.; Villordo, S.M.; Aguirre, S.; Pallarés, H.M.; Maestre, A.M.; Sánchez-Vargas, I.; Blair, C.D.; Fabri, C.; Morales, M.A.; et al. Dengue virus genomic variation associated with mosquito adaptation defines the pattern of viral non-coding RNAs and fitness in human cells. PLoS Pathog. 2017, 13, e1006265. [Google Scholar] [CrossRef]
- Sirigulpanit, W.; Kinney, R.M.; Leardkamolkarn, V. Substitution or deletion mutations between nt 54 and 70 in the 5′ non-coding region of dengue type 2 virus produce variable effects on virus viability. J. Gen. Virol. 2007, 88, 1748–1752. [Google Scholar] [CrossRef]
- Tajima, S.; Nukui, Y.; Takasaki, T.; Kurane, I. Characterization of the variable region in the 3′ non-translated region of dengue type 1 virus. J. Gen. Virol. 2007, 88, 2214–2222. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, D.E.; De Lella Ezcurra, A.L.; Fucito, S.; Gamarnik, A.V. Role of RNA structures present at the 3′UTR of dengue virus on translation, RNA synthesis, and viral replication. Virology 2005, 339, 200–212. [Google Scholar] [CrossRef] [PubMed]
- Drake, J.W.; Holland, J.J. Mutation rates among RNA viruses. Proc. Natl. Acad. Sci. USA 1999, 96, 13910–13913. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.H.; Yip, J.T.; Chen, Y.L.; Liu, W.; Harun, S.; Lystiyaningsih, E.; Heriyanto, B.; Beckett, C.G.; Mitchell, W.P.; Hibberd, M.L.; et al. Periodic re-emergence of endemic strains with strong epidemic potential-a proposed explanation for the 2004 Indonesian dengue epidemic. Infect. Genet. Evol. 2008, 8, 191–204. [Google Scholar] [CrossRef] [PubMed]
- Sasmono, R.T.; Wahid, I.; Trimarsanto, H.; Yohan, B.; Wahyuni, S.; Hertanto, M.; Yusuf, I.; Mubin, H.; Ganda, I.J.; Latief, R.; et al. Genomic analysis and growth characteristic of dengue viruses from Makassar, Indonesia. Infect. Genet. Evol. 2015, 32, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Christenbury, J.G.; Aw, P.P.; Ong, S.H.; Schreiber, M.J.; Chow, A.; Gubler, D.J.; Vasudevan, S.G.; Ooi, E.E.; Hibberd, M.L. A method for full genome sequencing of all four serotypes of the dengue virus. J. Virol. Methods 2010, 169, 202–206. [Google Scholar] [CrossRef]
- Dash, P.K.; Sharma, S.; Soni, M.; Agarwal, A.; Sahni, A.K.; Parida, M. Complete genome sequencing and evolutionary phylogeography analysis of Indian isolates of Dengue virus type 1. Virus Res. 2015, 195, 124–134. [Google Scholar] [CrossRef]
- Wongsurawat, T.; Jenjaroenpun, P.; Taylor, M.K.; Lee, J.; Tolardo, A.L.; Parvathareddy, J.; Kandel, S.; Wadley, T.D.; Kaewnapan, B.; Athipanyasilp, N.; et al. Rapid Sequencing of Multiple RNA Viruses in Their Native Form. Front. Microbiol. 2019, 10, 260. [Google Scholar] [CrossRef]
- Tan, C.C.S.; Maurer-Stroh, S.; Wan, Y.; Sessions, O.M.; de Sessions, P.F. A novel method for the capture-based purification of whole viral native RNA genomes. AMB Express 2019, 9, 45. [Google Scholar] [CrossRef]

| Serotype | Primer Name | Region | Genomic Position a | Sequence (5′-3′) b | Annealing T °C |
|---|---|---|---|---|---|
| DENV-1 | DENV1_SP1 | 5′ | 560–579 | ccgggggcatttgtaggtca | 50 for cDNA |
| DENV1_SP2 | 5′ | 431–453 | tatcatgtgtggctctccycctc | 50 | |
| DENV1_SP3 | 5′ | 287–310 | ctttgatygctccattcttcttga | 58 | |
| DENV1_SP4 | 3′ | 9949–9970 | caccaatggatgacaacagaag | 55 | |
| DENV1_SP5 | 3′ | 10,119–10,140 | cacctgggccaccaacatacaa | 55 | |
| DENV-2 | DENV2_SP1 | 5′ | 475–497 | cctttctcctgcctaccaacgat | 50 for cDNA |
| DENV2_SP2 | 5′ | 355–376 | tgttcagcatccttccaatctc | 50 | |
| DENV2_SP3 | 5′ | 287–310 | tcgttccccatcttttyagtatcc | 50 | |
| DENV2_SP4 | 3′ | 10,114–10,137 | agaacatccaaacagcaataaatc | 52 | |
| DENV2_SP5 | 3′ | 10,238–10,262 | aagggaagaggaagaggcaggtgt | 52 | |
| DENV3 | DENV3_SP1 | 5′ | 560–579 | ggtratrtgggggcatttgtaag | 50 for cDNA |
| DENV3_SP2 | 5′ | 429–448 | cggctctccatctcgtgaag | 50 | |
| DENV3_SP3 | 5′ | 276–298 | cgacttcttgaaggttccccatc | 58 | |
| DENV3_SP4 | 3′ | 10,210–10,233 | gaaggaggaggartcggaggg | 55 | |
| DENV3_SP5 | 3′ | 10,311–10,333 | gcctgtgagccccgtctaag | 50 | |
| DENV-4 | DENV4_SP1 | 5′ | 496–517 | ttgttgatcccctctgttgtyt | 50 for cDNA |
| DENV4_SP2 | 5′ | 454–477 | ccctttcatgttttgccactatca | 50 | |
| DENV4_SP3 | 5′ | 298–321 | gtgggatggaaagractcgca | 55 | |
| DENV4_SP4 | 3′ | 10,302–10,320 | gcaaaccgtgctgcctgta | 52 | |
| DENV4_SP5 | 3′ | 10,098–10,120 | ggactttcttcyagagccacctg | 52 |
| Serotype | Internal Code | Year | State | Clinical Classification | Virus titer (PFU/mL) |
|---|---|---|---|---|---|
| DENV 1 | 424735 | 2013 | Meta | D | 7.8 × 106 |
| 427493 | 2013 | Tolima | D | 6.83 × 106 | |
| 450339 | 2015 | Cundinamarca | D | 4.33 × 105 | |
| 483718 | 2016 | Cauca | D | 9.75 × 105 | |
| 484940 | 2016 | Boyaca | D | 6.83 × 104 | |
| DENV 2 | 423887 | 2013 | Putumayo | D | 1.2 × 105 |
| 427516 | 2013 | Caldas | SD | 1.7 × 107 | |
| 425334 | 2013 | Putumayo | D | 2 × 107 | |
| 425817 | 2013 | Tolima | D | 2 × 107 | |
| 425819 | 2013 | Cundinamarca | D | 1.8 × 107 | |
| 428702 | 2014 | Tolima | SD | 8.17 × 106 | |
| 424233 | 2013 | Boyacá | D | 8.5 × 106 | |
| 449418 | 2015 | Tolima | DWS | 8.83 × 106 | |
| 449308 | 2015 | Huila | SD * | 2.83 × 104 | |
| 449618 | 2015 | Huila | D | 1.5 × 106 | |
| 449510 | 2015 | Putumayo | D | 1.48 × 106 | |
| 434321 | 2014 | Meta | SD | 1.6 × 107 | |
| DENV 3 | 449683 | 2015 | Risaralda | DWS | 1.9 × 107 |
| 449499 | 2015 | Risaralda | D | 1.72 × 106 | |
| 449415 | 2015 | Putumayo | D | NA | |
| 449740 | 2015 | Sucre | D | NA | |
| 449746 | 2015 | Risaralda | D | 1 × 104 | |
| 449255 | 2015 | Quindío | D | 1.23 × 105 | |
| 449326 | 2015 | Risaralda | D | 7.33 × 106 | |
| 449334 | 2015 | Boyacá | DWS | NA | |
| 449335 | 2015 | Boyacá | DWS | 8.75 × 106 | |
| DENV 4 | 371813 | NA | NA | NA | NA |
| 37178 | NA | NA | NA | NA | |
| 426553 | 2013 | Risaralda | D | 1.03 × 104 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosales-Munar, A.; Alvarez-Diaz, D.A.; Laiton-Donato, K.; Peláez-Carvajal, D.; Usme-Ciro, J.A. Efficient Method for Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome. Viruses 2020, 12, 496. https://doi.org/10.3390/v12050496
Rosales-Munar A, Alvarez-Diaz DA, Laiton-Donato K, Peláez-Carvajal D, Usme-Ciro JA. Efficient Method for Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome. Viruses. 2020; 12(5):496. https://doi.org/10.3390/v12050496
Chicago/Turabian StyleRosales-Munar, Alicia, Diego Alejandro Alvarez-Diaz, Katherine Laiton-Donato, Dioselina Peláez-Carvajal, and Jose A. Usme-Ciro. 2020. "Efficient Method for Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome" Viruses 12, no. 5: 496. https://doi.org/10.3390/v12050496
APA StyleRosales-Munar, A., Alvarez-Diaz, D. A., Laiton-Donato, K., Peláez-Carvajal, D., & Usme-Ciro, J. A. (2020). Efficient Method for Molecular Characterization of the 5′ and 3′ Ends of the Dengue Virus Genome. Viruses, 12(5), 496. https://doi.org/10.3390/v12050496





