Structures of Three Actinobacteriophage Capsids: Roles of Symmetry and Accessory Proteins
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacteriophages, Hosts, and Sources
2.2. Production of Bacteriophages for Cryo-Electron Microscopy and Mass Spectrometry
2.3. Preparation of Bacteriophages for Transmission Electron Microscopy
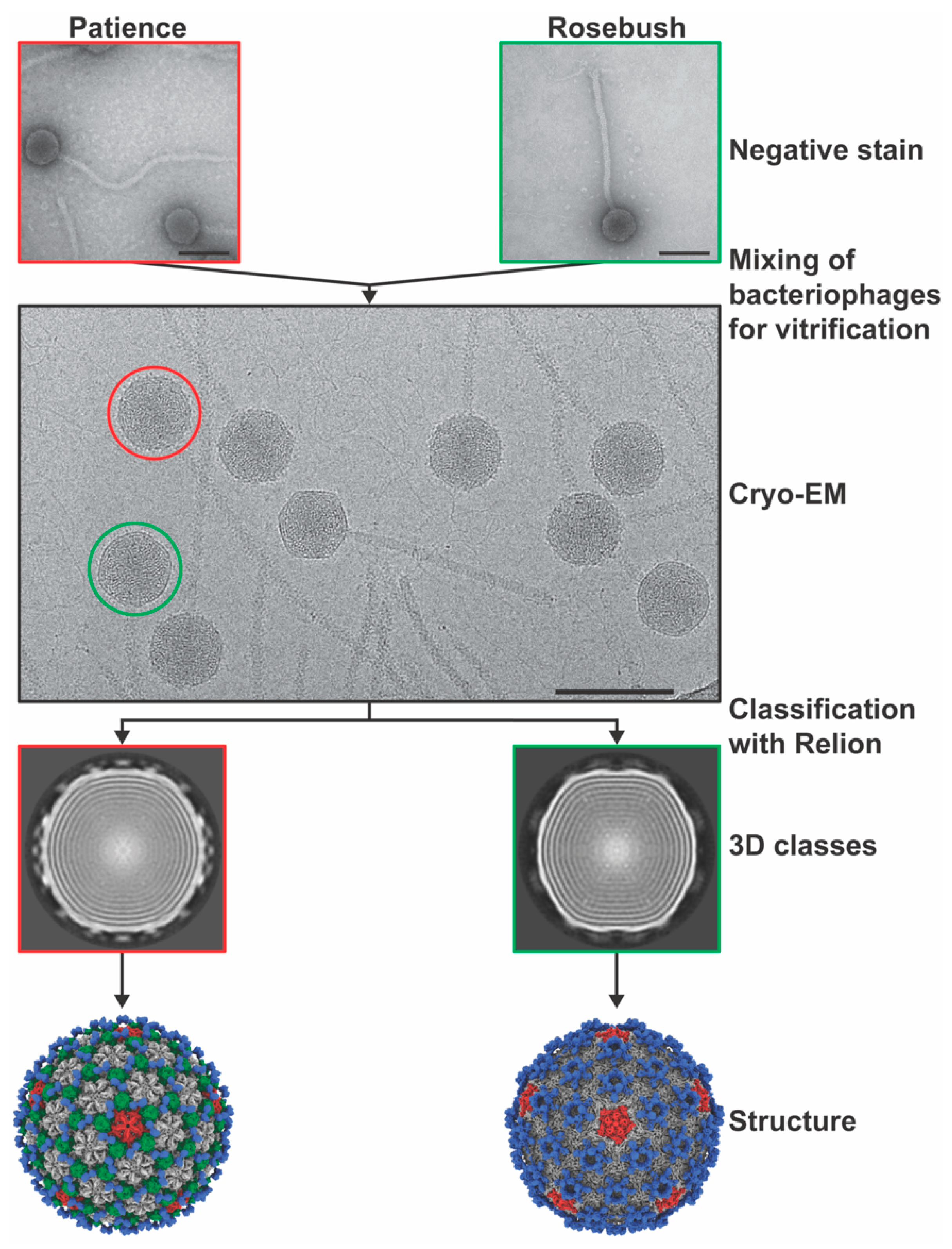
2.4. Preparation of Cryo-Electron Microscopy Grids
2.5. Cryo-Electron Microscopy
2.6. Cryo-Electron Microscopy Data Analysis
2.6.1. Data Deposition
2.7. Bioinformatics
2.8. Mass Spectrometry
2.8.1. In-Gel Digestions, Tryptic Peptide Extraction and Desalting
2.8.2. Peptide and Protein Identification Using Mass Spectrometry
3. Results
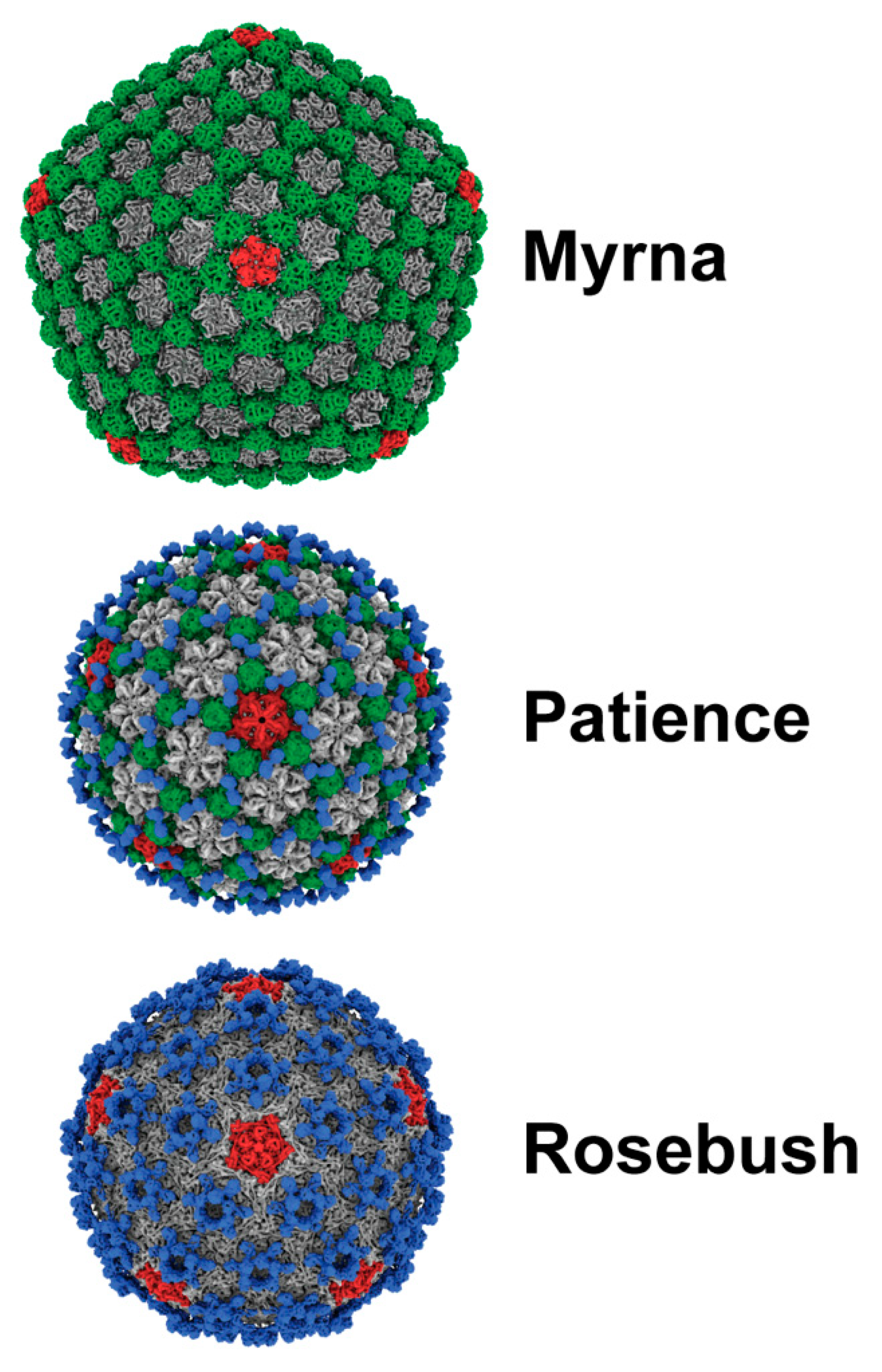
3.1. Virion Morphologies
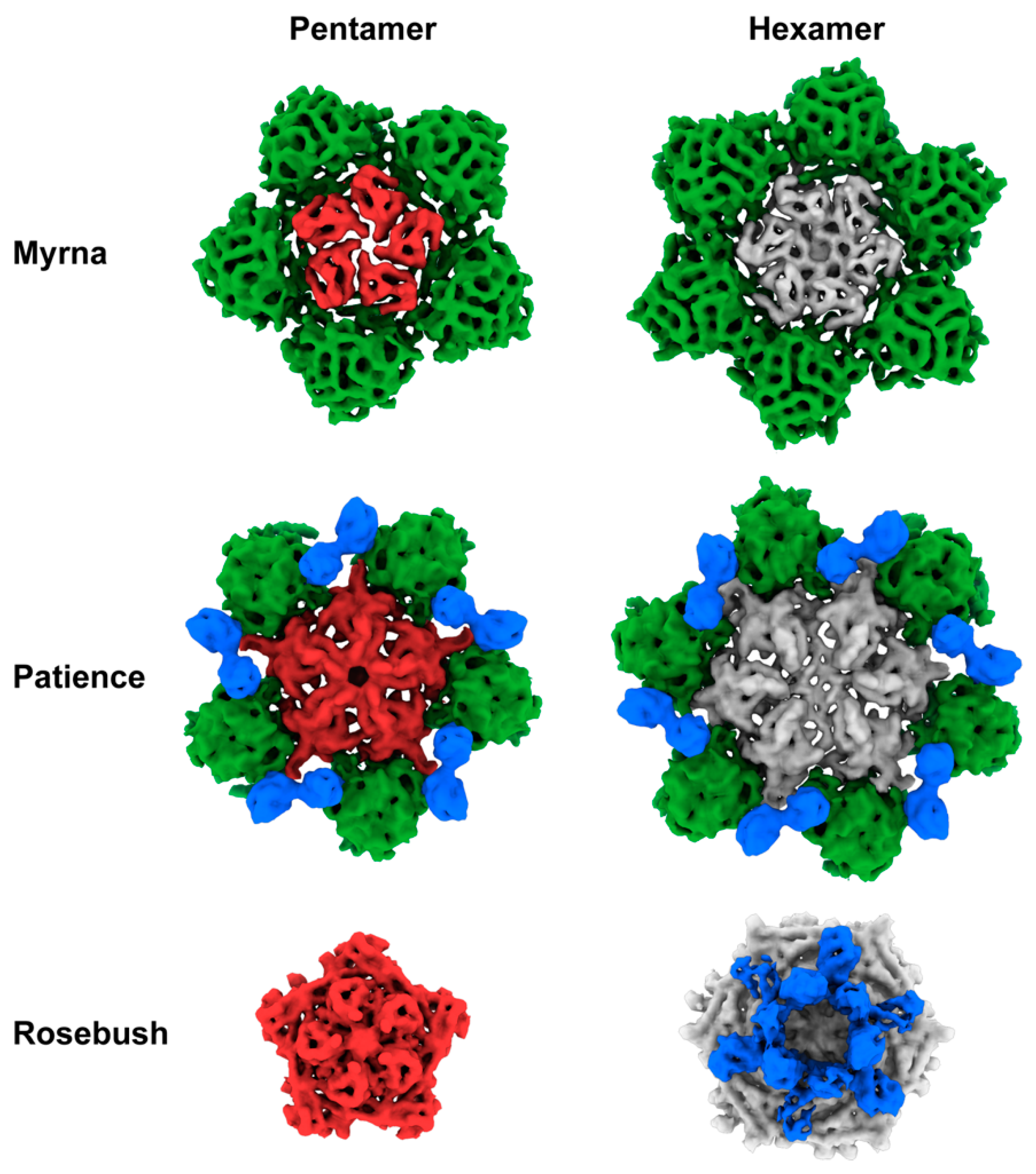
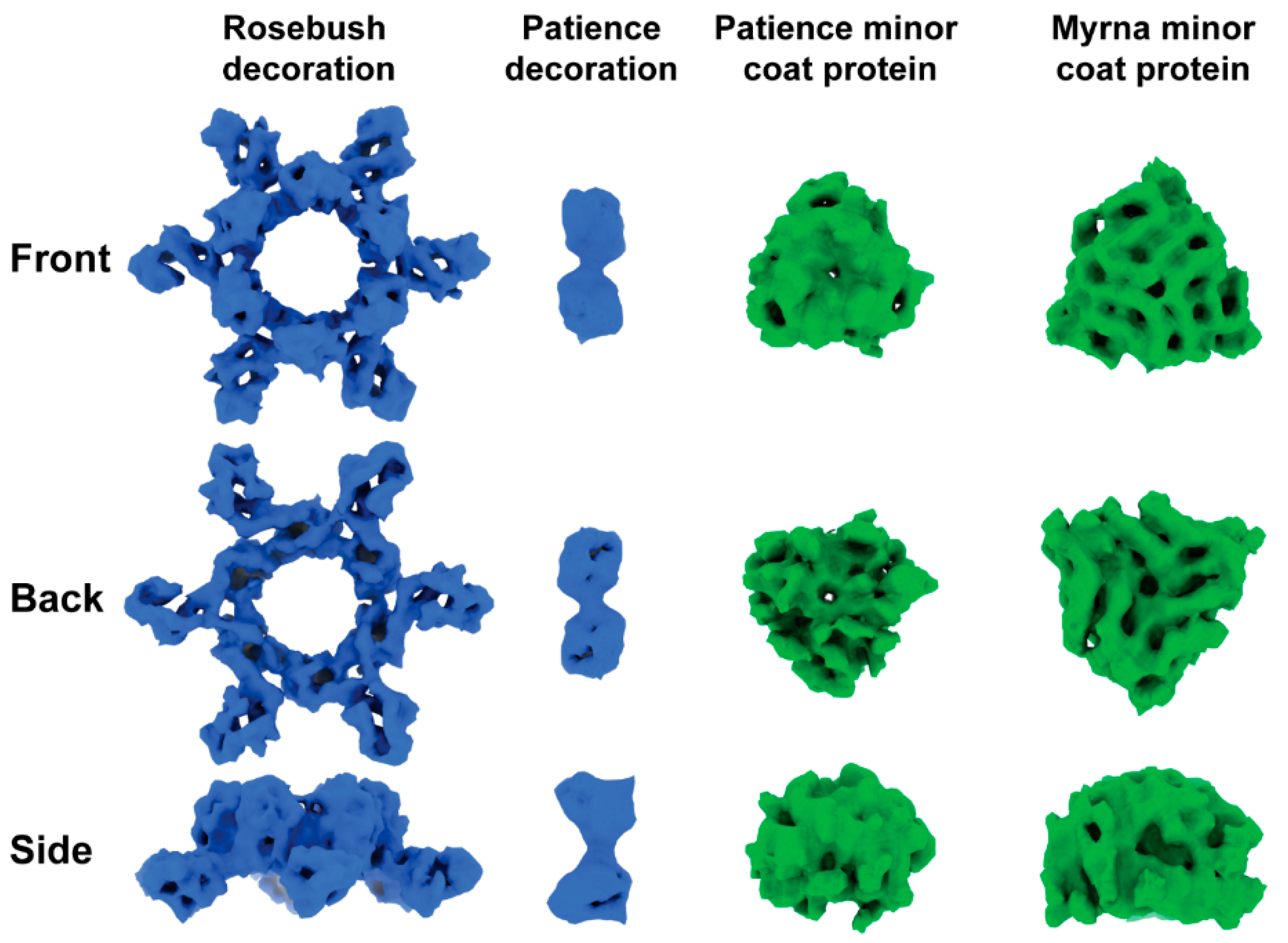
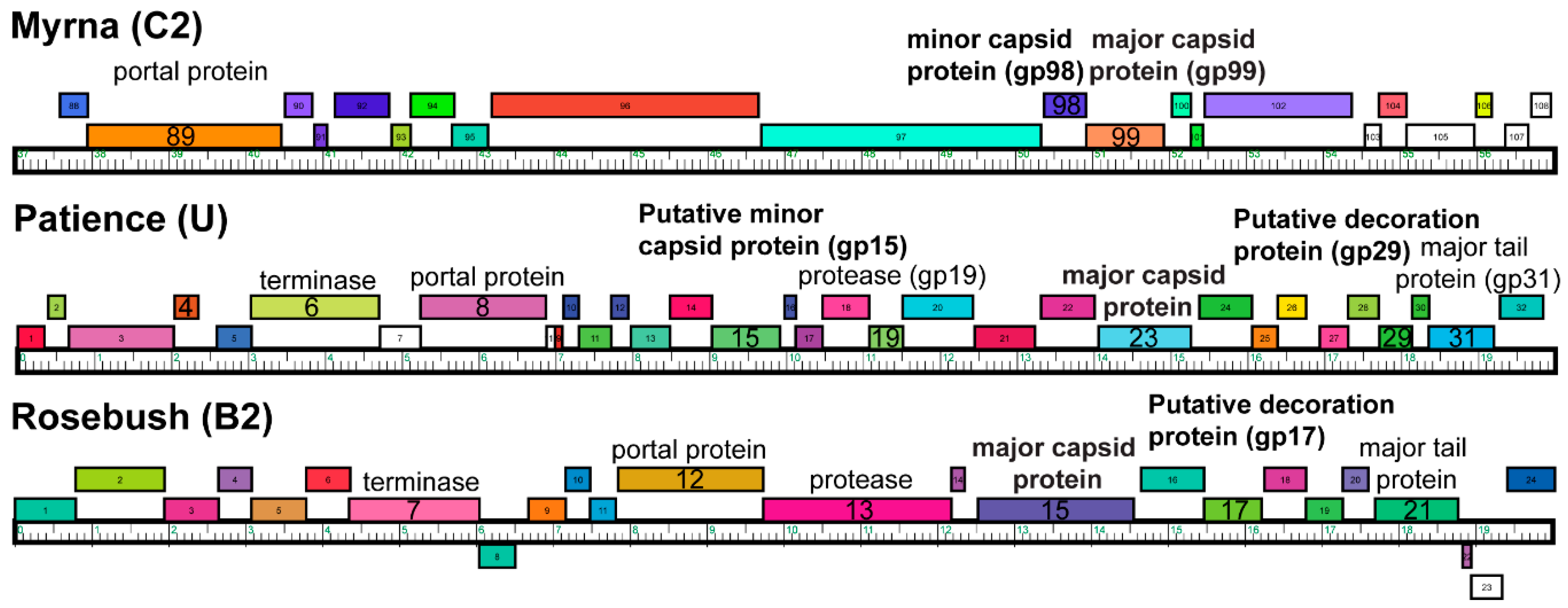
3.2. Capsid Morphologies and Accessory Proteins
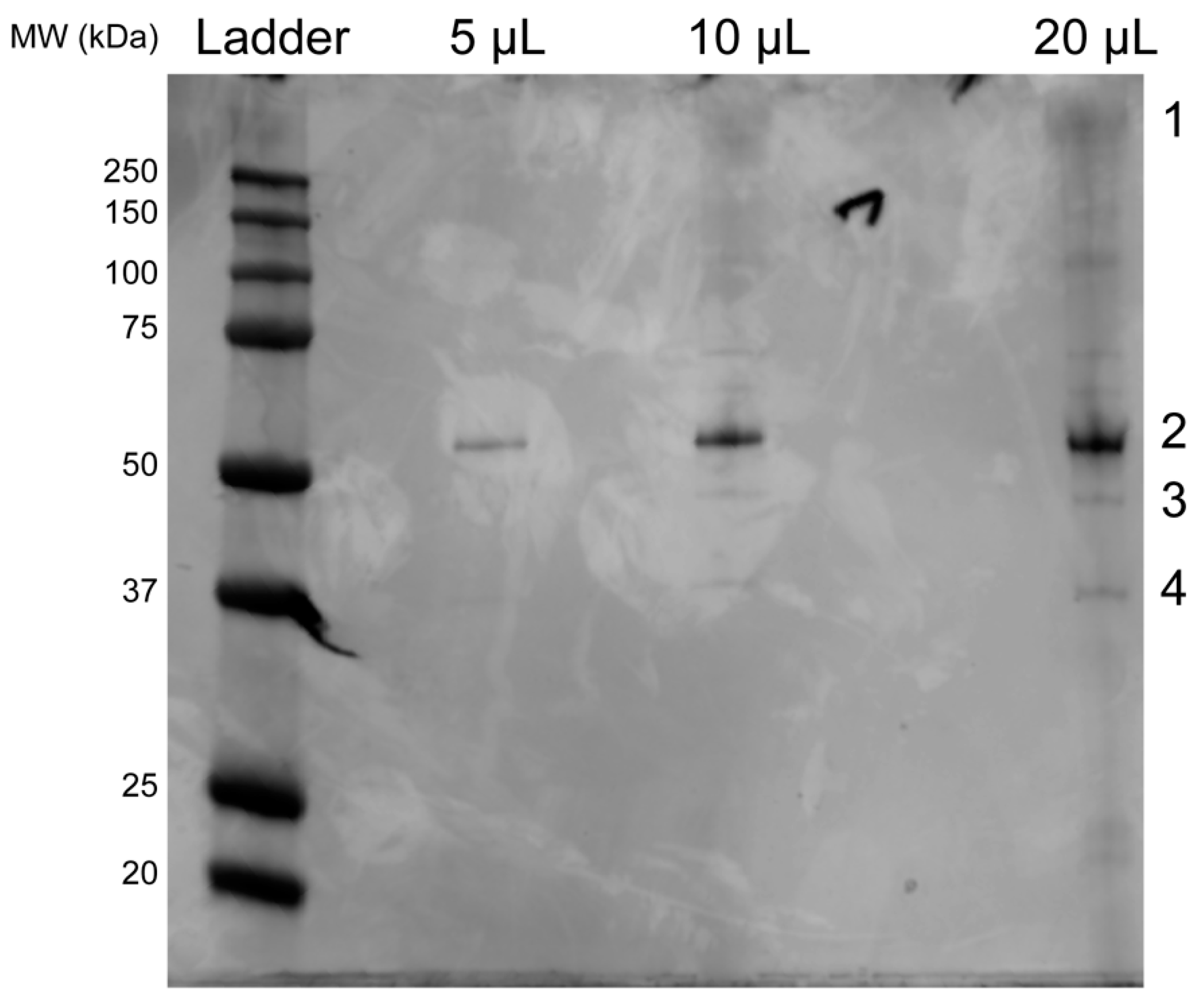
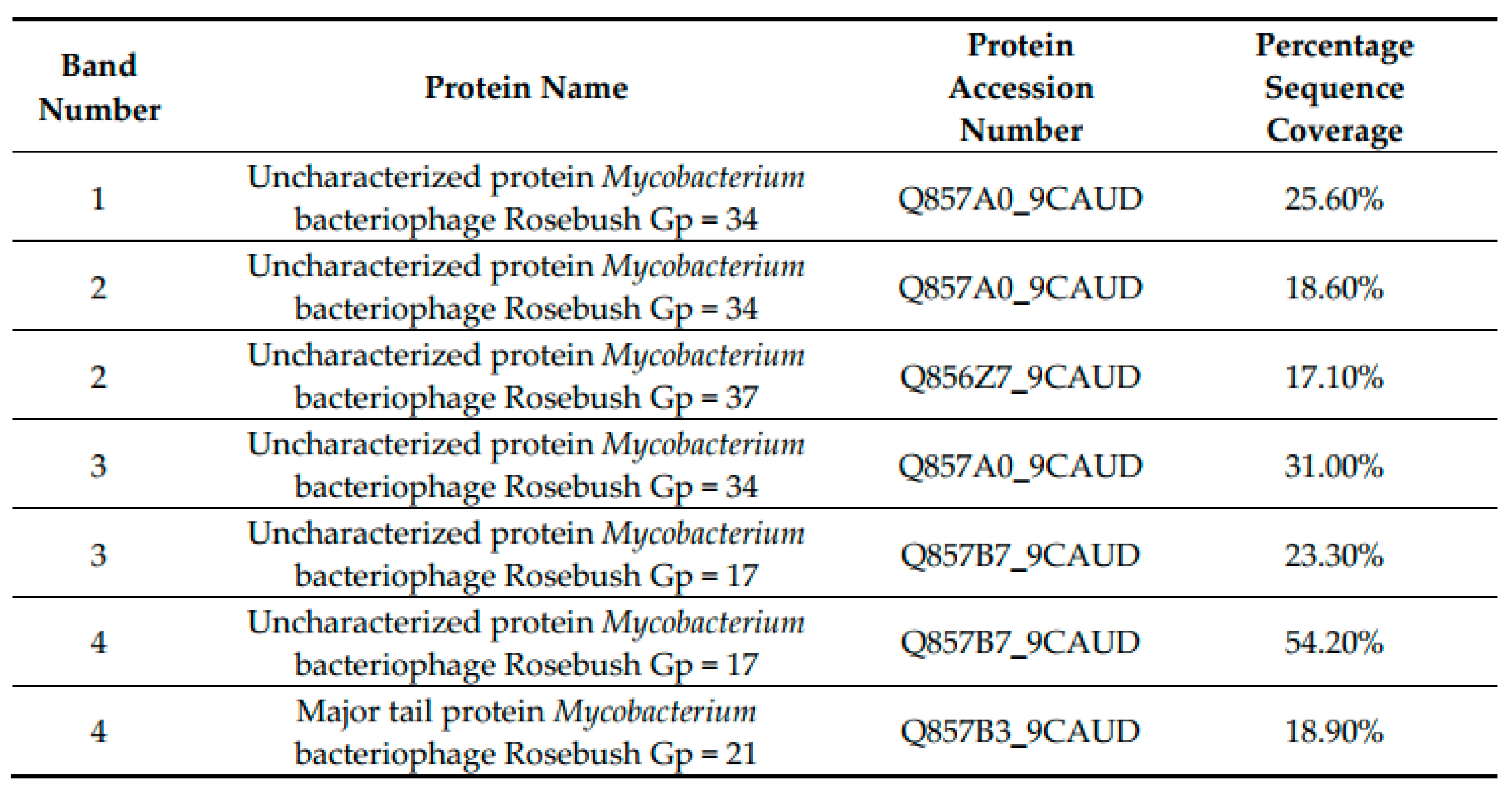
3.3. SDS-PAGE Gel and Mass Spectrometry Analysis of Rosebush
3.4. Bioinformatic Analysis of Accessory Protein gp17
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dedrick, R.M.; Guerrero-Bustamante, C.A.; Garlena, R.A.; Russell, D.A.; Ford, K.; Harris, K.; Gilmour, K.C.; Soothill, J.; Jacobs-Sera, D.; Schooley, R.T.; et al. Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat. Med. 2019, 25, 730–733. [Google Scholar] [CrossRef] [PubMed]
- Hatfull, G.F.; Pedulla, M.L.; Jacobs-Sera, D.; Cichon, P.M.; Foley, A.; Ford, M.E.; Gonda, R.M.; Houtz, J.M.; Hryckowian, A.J.; Kelchner, V.A.; et al. Exploring the mycobacteriophage metaproteome: phage genomics as an educational platform. Plos Genet. 2006, 2, e92. [Google Scholar] [CrossRef] [PubMed]
- Hanauer, D.I.; Graham, M.J.; SEA-PHAGES; Betancur, L.; Bobrownicki, A.; Cresawn, S.G.; Garlena, R.A.; Jacobs-Sera, D.; Kaufmann, N.; Pope, W.H.; et al. An inclusive Research Education Community (iREC): Impact of the SEA-PHAGES program on research outcomes and student learning. Proc. Natl. Acad. Sci. USA 2017, 114, 13531–13536. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H.-W. 5500 Phages examined in the electron microscope. Arch. Virol. 2007, 152, 227–243. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H.-W.; Prangishvili, D. Prokaryote viruses studied by electron microscopy. Arch. Virol. 2012, 157, 1843–1849. [Google Scholar] [CrossRef] [PubMed]
- Helgstrand, C.; Wikoff, W.R.; Duda, R.L.; Hendrix, R.W.; Johnson, J.E.; Liljas, L. The refined structure of a protein catenane: the HK97 bacteriophage capsid at 3.44 A resolution. J. Mol. Biol. 2003, 334, 885–899. [Google Scholar] [CrossRef] [PubMed]
- São-José, C.; de Frutos, M.; Raspaud, E.; Santos, M.A.; Tavares, P. Pressure Built by DNA Packing Inside Virions: Enough to Drive DNA Ejection in Vitro, Largely Insufficient for Delivery into the Bacterial Cytoplasm. J. Mol. Biol. 2007, 374, 346–355. [Google Scholar] [CrossRef]
- Evilevitch, A.; Lavelle, L.; Knobler, C.M.; Raspaud, E.; Gelbart, W.M. Osmotic pressure inhibition of DNA ejection from phage. Proc. Natl. Acad. Sci. USA 2003, 100, 9292–9295. [Google Scholar] [CrossRef]
- Lander, G.C.; Evilevitch, A.; Jeembaeva, M.; Potter, C.S.; Carragher, B.; Johnson, J.E. Bacteriophage lambda stabilization by auxiliary protein gpD: timing, location, and mechanism of attachment determined by cryo-EM. Structure 2008, 16, 1399–1406. [Google Scholar] [CrossRef]
- Sae-Ueng, U.; Liu, T.; Catalano, C.E.; Huffman, J.B.; Homa, F.L.; Evilevitch, A. Major capsid reinforcement by a minor protein in herpesviruses and phage. Nucleic Acids Res. 2014, 42, 9096–9107. [Google Scholar] [CrossRef]
- Sathaliyawala, T.; Islam, M.Z.; Li, Q.; Fokine, A.; Rossmann, M.G.; Rao, V.B. Functional analysis of the highly antigenic outer capsid protein, Hoc, a virus decoration protein from T4-like bacteriophages. - PubMed - NCBI. Mol. Microbiol. 2010, 77, 444–455. [Google Scholar] [CrossRef] [PubMed]
- Fokine, A.; Islam, M.Z.; Zhang, Z.; Bowman, V.D.; Rao, V.B.; Rossmann, M.G. Structure of the Three N-Terminal Immunoglobulin Domains of the Highly Immunogenic Outer Capsid Protein from a T4-Like Bacteriophage. J. Virol. 2011, 85, 8141–8148. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, R.; Nath, S.; Ranjan, A.; Khamrui, S.; Pani, B.; Sen, R.; Sen, U. The first structure of polarity suppression protein, Psu from enterobacteria phage P4, reveals a novel fold and a knotted dimer. J. Biol. Chem. 2012, 287, 44667–44675. [Google Scholar] [CrossRef] [PubMed]
- Lander, G.C.; Baudoux, A.-C.; Azam, F.; Potter, C.S.; Carragher, B.; Johnson, J.E. Capsomer dynamics and stabilization in the T = 12 marine bacteriophage SIO-2 and its procapsid studied by CryoEM. Structure 2012, 20, 498–503. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Snapper, S.B.; Melton, R.E.; Mustafa, S.; Kieser, T.; Jacobs, W.R., Jr. Isolation and characterization of efficient plasmid transformation mutants of Mycobacterium smegmatis. Mol. Microbiol. 1990, 4, 1911–1919. [Google Scholar] [CrossRef]
- Pope, W.H.; Bowman, C.A.; Russell, D.A.; Jacobs-Sera, D.; Asai, D.J.; Cresawn, S.G.; Jacobs, W.R.; Hendrix, R.W.; Lawrence, J.G.; Hatfull, G.F. Science Education Alliance Phage Hunters Advancing Genomics and Evolutionary Science; Phage Hunters Integrating Research and Education; Mycobacterial Genetics Course Whole genome comparison of a large collection of mycobacteriophages reveals a continuum of phage genetic diversity. Elife 2015, 4, e06416. [Google Scholar]
- Hatfull, G.F. Science Education Alliance Phage Hunters Advancing Genomics and Evolutionary Science Program; KwaZulu-Natal Research Institute for Tuberculosis and HIV Mycobacterial Genetics Course Students; Phage Hunters Integrating Research and Education Program Complete genome sequences of 138 mycobacteriophages. J. Virol. 2012, 86, 2382–2384. [Google Scholar]
- Pope, W.H.; Jacobs-Sera, D.; Russell, D.A.; Rubin, D.H.F.; Kajee, A.; Msibi, Z.N.P.; Larsen, M.H.; Jacobs, W.R.; Lawrence, J.G.; Hendrix, R.W.; et al. Genomics and proteomics of mycobacteriophage patience, an accidental tourist in the Mycobacterium neighborhood. MBio 2014, 5, e02145. [Google Scholar] [CrossRef]
- Jacobs-Sera, D.; Marinelli, L.J.; Bowman, C.; Broussard, G.W.; Guerrero Bustamante, C.; Boyle, M.M.; Petrova, Z.O.; Dedrick, R.M.; Pope, W.H.; Science Education Alliance Phage Hunters Advancing Genomics And Evolutionary Science Sea-Phages Program; et al. On the nature of mycobacteriophage diversity and host preference. Virology 2012, 434, 187–201. [Google Scholar] [CrossRef]
- Pedulla, M.L.; Ford, M.E.; Houtz, J.M.; Karthikeyan, T.; Wadsworth, C.; Lewis, J.A.; Jacobs-Sera, D.; Falbo, J.; Gross, J.; Pannunzio, N.R.; et al. Origins of highly mosaic mycobacteriophage genomes. Cell 2003, 113, 171–182. [Google Scholar] [CrossRef]
- Hatfull, G.F.; Jacobs-Sera, D.; Lawrence, J.G.; Pope, W.H.; Russell, D.A.; Ko, C.-C.; Weber, R.J.; Patel, M.C.; Germane, K.L.; Edgar, R.H.; et al. Comparative genomic analysis of 60 Mycobacteriophage genomes: genome clustering, gene acquisition, and gene size. J. Mol. Biol. 2010, 397, 119–143. [Google Scholar] [CrossRef] [PubMed]
- Salifu, S.P.; Valero-Rello, A.; Campbell, S.A.; Inglis, N.F.; Scortti, M.; Foley, S.; Vázquez-Boland, J.A. Genome and proteome analysis of phage E3 infecting the soil-borne actinomycete Rhodococcus equi. Env. Microbiol. Rep. 2013, 5, 170–178. [Google Scholar] [CrossRef] [PubMed]
- Lavigne, R.; Darius, P.; Summer, E.J.; Seto, D.; Mahadevan, P.; Nilsson, A.S.; Ackermann, H.-W.; Kropinski, A.M. Classification of Myoviridae bacteriophages using protein sequence similarity. BMC Microbiol. 2009, 9, 224. [Google Scholar] [CrossRef] [PubMed]
- Santiago-Rodriguez, T.M.; Fornaciari, G.; Luciani, S.; Dowd, S.E.; Toranzos, G.A.; Marota, I.; Cano, R.J. Natural mummification of the human gut preserves bacteriophage DNA. Fems Microbiol. Lett. 2016, 363, fnv219. [Google Scholar] [CrossRef]
- Catalão, M.J.; Gil, F.; Moniz-Pereira, J.; São-José, C.; Pimentel, M. Diversity in bacterial lysis systems: bacteriophages show the way. Fems Microbiol. Rev. 2013, 37, 554–571. [Google Scholar] [CrossRef]
- Zivanov, J.; Nakane, T.; Forsberg, B.O.; Kimanius, D.; Hagen, W.J.; Lindahl, E.; Scheres, S.H. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 2018, 7, e42166. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 2018, 27, 14–25. [Google Scholar] [CrossRef]
- Pintilie, G.D.; Zhang, J.; Goddard, T.D.; Chiu, W.; Gossard, D.C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 2010, 170, 427–438. [Google Scholar] [CrossRef]
- Zhang, Y. I-TASSER server for protein 3D structure prediction. Bmc Bioinform. 2008, 9, 40. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, L.; Stephens, A.; Nam, S.-Z.; Rau, D.; Kübler, J.; Lozajic, M.; Gabler, F.; Söding, J.; Lupas, A.N.; Alva, V. A Completely Reimplemented MPI Bioinformatics Toolkit with a New HHpred Server at its Core. J. Mol. Biol. 2018, 430, 2237–2243. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. - PubMed - NCBI. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef]
- Twarock, R.; Luque, A. Structural puzzles in virology solved with an overarching icosahedral design principle. Nat. Commun. 2019, 10, 4414–4419. [Google Scholar] [CrossRef]
- Dearborn, A.D.; Wall, E.A.; Kizziah, J.L.; Klenow, L.; Parker, L.K.; Manning, K.A.; Spilman, M.S.; Spear, J.M.; Christie, G.E.; Dokland, T. Competing scaffolding proteins determine capsid size during mobilization of Staphylococcus aureus pathogenicity islands. Elife 2017, 6, 429. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar]
- Dai, X.; Zhou, Z.H. Structure of the herpes simplex virus 1 capsid with associated tegument protein complexes. Science 2018, 360, eaao7298. [Google Scholar] [CrossRef]
- Wang, Z.; Hardies, S.C.; Fokine, A.; Klose, T.; Jiang, W.; Cho, B.C.; Rossmann, M.G. Structure of the Marine Siphovirus TW1: Evolution of Capsid-Stabilizing Proteins and Tail Spikes. Structure 2018, 26, 238–248.e3. [Google Scholar] [CrossRef]
- Shin, H.; Lee, J.-H.; Ahn, C.S.; Ryu, S.; Cho, B.C. Complete genome sequence of marine bacterium Pseudoalteromonas phenolica bacteriophage TW1. Arch. Virol. 2014, 159, 159–162. [Google Scholar] [CrossRef] [PubMed]
- Cresawn, S.G.; Bogel, M.; Day, N.; Jacobs-Sera, D.; Hendrix, R.W.; Hatfull, G.F. Phamerator: a bioinformatic tool for comparative bacteriophage genomics. BMC Bioinform. 2011, 12, 395. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Guo, H.; Jin, L.; Czornyj, E.; Hodes, A.; Hui, W.H.; Nieh, A.W.; Miller, J.F.; Zhou, Z.H. A new topology of the HK97-like fold revealed in Bordetella bacteriophage by cryoEM at 3.5 A resolution. Elife 2013, 2, e01299. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.L.; Hryc, C.F.; Zhang, Q.; Wu, W.; Jakana, J.; Haase-Pettingell, C.; Afonine, P.V.; Adams, P.D.; King, J.A.; Jiang, W.; et al. Validated near-atomic resolution structure of bacteriophage epsilon15 derived from cryo-EM and modeling. Proc. Natl. Acad. Sci. USA 2013, 110, 12301–12306. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Y.; Rossmann, M.G. Structure of bacteriophage phi29 head fibers has a supercoiled triple repeating helix-turn-helix motif. Proc. Natl. Acad. Sci. USA 2011, 108, 4806–4810. [Google Scholar] [CrossRef] [PubMed]
- Salas, M.; Vásquez, C.; Méndez, E.; Viñuela, E. Head fibers of bacteriophage ∅29. Virology 1972, 50, 180–188. [Google Scholar] [CrossRef]
- Grose, J.H.; Belnap, D.M.; Jensen, J.D.; Mathis, A.D.; Prince, J.T.; Merrill, B.D.; Burnett, S.H.; Breakwell, D.P. The genomes, proteomes, and structures of three novel phages that infect the Bacillus cereus group and carry putative virulence factors. J. Virol. 2014, 88, 11846–11860. [Google Scholar] [CrossRef]
- Dodonova, S.O.; Prinz, S.; Bilanchone, V.; Sandmeyer, S.; Briggs, J.A.G. Structure of the Ty3/Gypsy retrotransposon capsid and the evolution of retroviruses. Proc. Natl. Acad. Sci. USA 2019, 116, 10048–10057. [Google Scholar] [CrossRef]
- Sutter, M.; Greber, B.; Aussignargues, C.; Kerfeld, C.A. Assembly principles and structure of a 6.5-MDa bacterial microcompartment shell. Science 2017, 356, 1293–1297. [Google Scholar] [CrossRef]
- Greber, B.J.; Sutter, M.; Kerfeld, C.A. The Plasticity of Molecular Interactions Governs Bacterial Microcompartment Shell Assembly. Structure 2019, 27, 749–763.e4. [Google Scholar] [CrossRef]
- Choi, K.H.; McPartland, J.; Kaganman, I.; Bowman, V.D.; Rothman-Denes, L.B.; Rossmann, M.G. Insight into DNA and protein transport in double-stranded DNA viruses: the structure of bacteriophage N4. J. Mol. Biol. 2008, 378, 726–736. [Google Scholar] [CrossRef] [PubMed]
- Newcomer, R.L.; Schrad, J.R.; Gilcrease, E.B.; Casjens, S.R.; Feig, M.; Teschke, C.M.; Alexandrescu, A.T.; Parent, K.N. The phage L capsid decoration protein has a novel OB-fold and an unusual capsid binding strategy. Elife 2019, 8, 814. [Google Scholar] [CrossRef] [PubMed]
- Forrer, P.; Chang, C.; Ott, D.; Wlodawer, A.; Plückthun, A. Kinetic stability and crystal structure of the viral capsid protein SHP. J. Mol. Biol. 2004, 344, 179–193. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.L.; Baker, M.R.; Hryc, C.F.; Ju, T.; Chiu, W. Gorgon and pathwalking: macromolecular modeling tools for subnanometer resolution density maps. Biopolymers 2012, 97, 655–668. [Google Scholar] [CrossRef]
- Abrescia, N.G.A.; Cockburn, J.J.B.; Grimes, J.M.; Sutton, G.C.; Diprose, J.M.; Butcher, S.J.; Fuller, S.D.; Martín, C.S.; Burnett, R.M.; Stuart, D.I.; et al. Insights into assembly from structural analysis of bacteriophage PRD1. Nature 2004, 432, 68–74. [Google Scholar] [CrossRef]
- Krupovic, M.; Koonin, E.V. Multiple origins of viral capsid proteins from cellular ancestors. Proc. Natl. Acad. Sci. U.S.A. 2017, 114, E2401–E2410. [Google Scholar] [CrossRef]
- Krupovic, M.; Bamford, D.H. Double-stranded DNA viruses: 20 families and only five different architectural principles for virion assembly. Curr. Opin. Virol. 2011, 1, 118–124. [Google Scholar] [CrossRef]
- Abrescia, N.G.A.; Bamford, D.H.; Grimes, J.M.; Stuart, D.I. Structure unifies the viral universe. Annu. Rev. Biochem. 2012, 81, 795–822. [Google Scholar] [CrossRef]



| Name | Number of Genes | Cluster | Number of Phages in Cluster | Genome Length (bp) | Genome GC Content (%) | Life Cycle | Host | Morphology |
|---|---|---|---|---|---|---|---|---|
| Myrna | 229 | C2 | 2 | 164602 | 65.4 | Lytic | M. smegmatis mc2155 | Myoviridae |
| Patience | 109 | U | 2 | 70506 | 50.3 | Lytic | M. smegmatis mc2155 | Siphoviridae |
| Rosebush | 90 | B2 | 27 | 67480 | 68.9 | Lytic | M. smegmatis mc2155 | Siphoviridae |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Podgorski, J.; Calabrese, J.; Alexandrescu, L.; Jacobs-Sera, D.; Pope, W.; Hatfull, G.; White, S. Structures of Three Actinobacteriophage Capsids: Roles of Symmetry and Accessory Proteins. Viruses 2020, 12, 294. https://doi.org/10.3390/v12030294
Podgorski J, Calabrese J, Alexandrescu L, Jacobs-Sera D, Pope W, Hatfull G, White S. Structures of Three Actinobacteriophage Capsids: Roles of Symmetry and Accessory Proteins. Viruses. 2020; 12(3):294. https://doi.org/10.3390/v12030294
Chicago/Turabian StylePodgorski, Jennifer, Joshua Calabrese, Lauren Alexandrescu, Deborah Jacobs-Sera, Welkin Pope, Graham Hatfull, and Simon White. 2020. "Structures of Three Actinobacteriophage Capsids: Roles of Symmetry and Accessory Proteins" Viruses 12, no. 3: 294. https://doi.org/10.3390/v12030294
APA StylePodgorski, J., Calabrese, J., Alexandrescu, L., Jacobs-Sera, D., Pope, W., Hatfull, G., & White, S. (2020). Structures of Three Actinobacteriophage Capsids: Roles of Symmetry and Accessory Proteins. Viruses, 12(3), 294. https://doi.org/10.3390/v12030294






