High Genomic Variability in Equine Infectious Anemia Virus Obtained from Naturally Infected Horses in Pantanal, Brazil: An Endemic Region Case
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Sample Collection and RNA Purification
2.3. Library Preparation and Next-Generation Sequencing
2.4. Bioinformatics Analysis
2.5. Primer Design and Sanger Sequencing
2.6. Variability Analysis
2.7. Molecular Modelling
3. Results
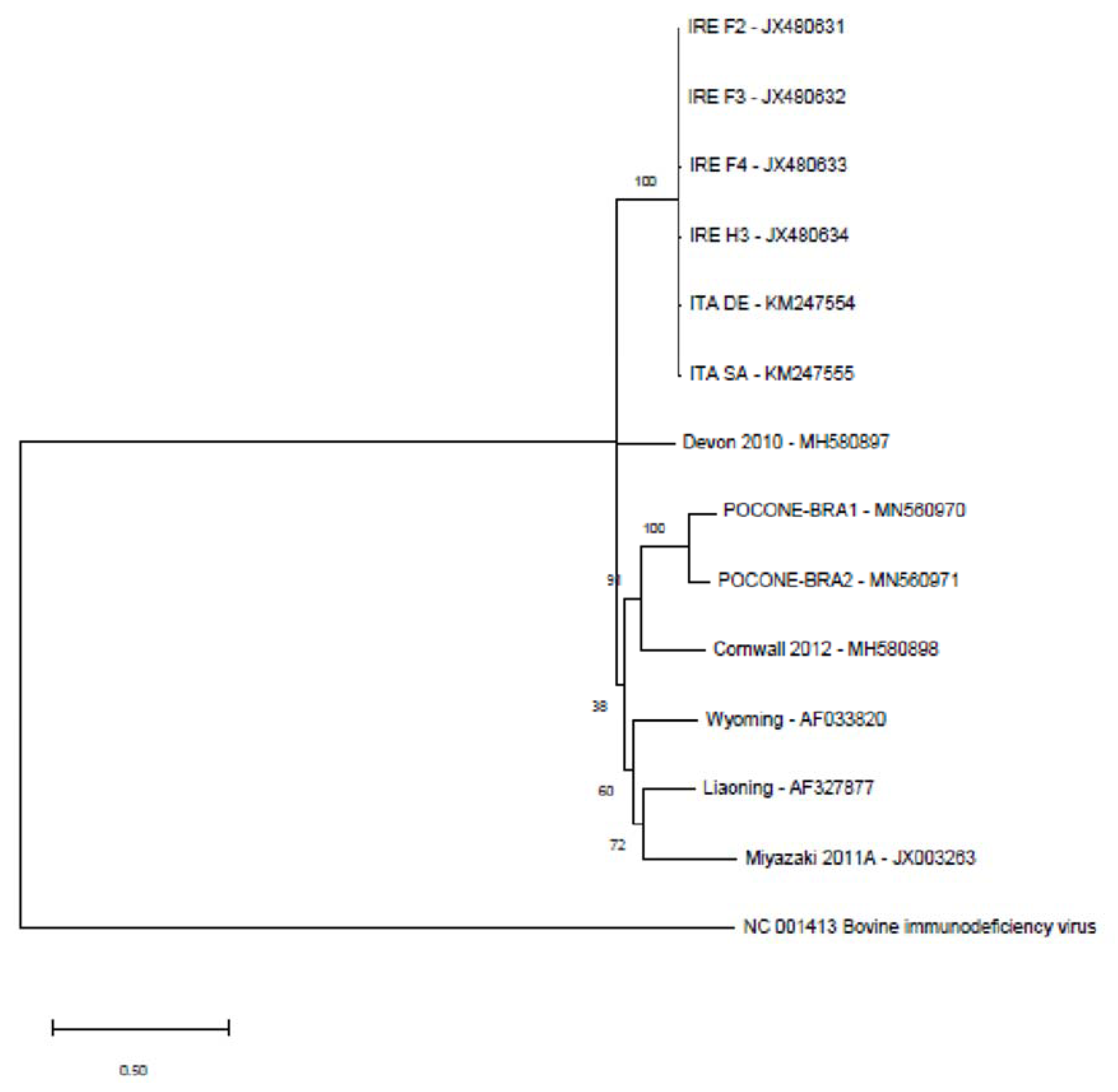
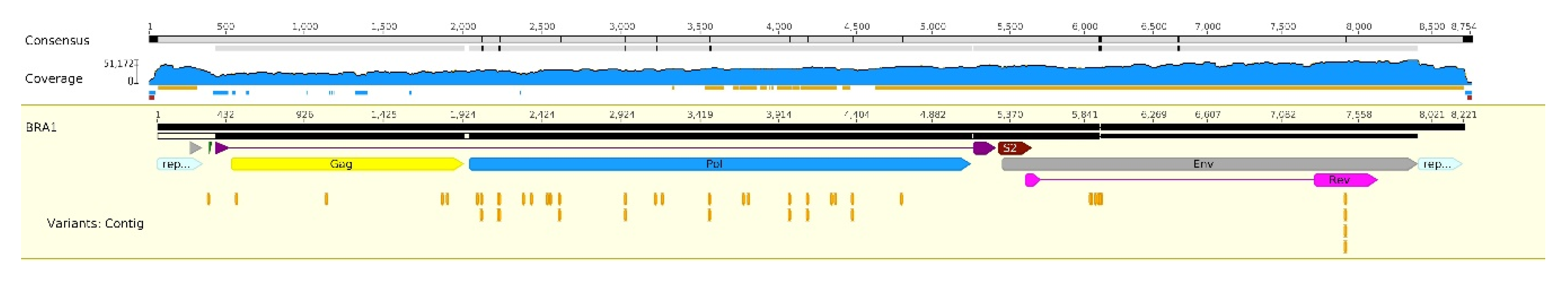
3.1. Complete Sequence of Brazilian EIAV
3.2. Polymorphisms in the EIAV Genome
3.3. LTR and Viral Genes
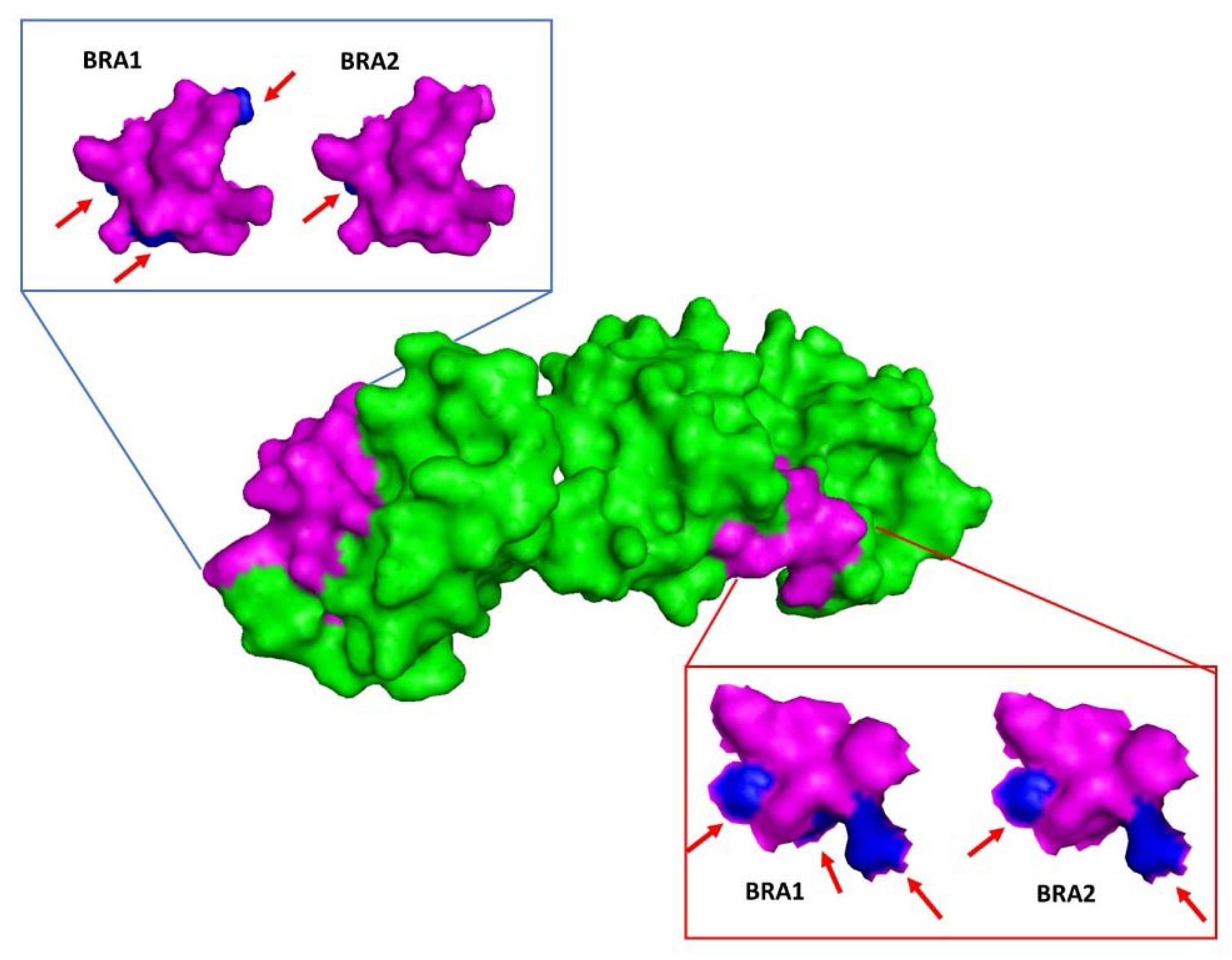
3.4. Gag Variation and p26 Molecular Modelling
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cook, R.F.; Leroux, C.; Issel, C.J. Equine infectious anemia and equine infectious anemia virus in 2013: A review. Vet. Microbiol. 2013, 167, 181–204. [Google Scholar] [CrossRef] [PubMed]
- Issel, C.J.; Cook, R.F.; Mealey, R.H.; Horohov, D.W. Equine infectious anemia in 2014: Live with it or eradicate it? Vet. Clin. North. Am.-Equine Pr. 2014, 30, 561–577. [Google Scholar] [CrossRef]
- Leroux, C.; Cadoré, J.-L.; Montelaro, R.C. Equine Infectious Anemia Virus (EIAV): What has HIV’s country cousin got to tell us? Vet. Res. 2004, 35, 485–512. [Google Scholar] [CrossRef] [PubMed]
- Leroux, C.; Craigo, J.K.; Issel, C.J.; Montelaro, R.C. Equine Infectious Anemia Virus Genomic Evolution in Progressor and Nonprogressor Ponies. J. Virol. 2001, 75, 4570–4583. [Google Scholar] [CrossRef]
- Schwartz, E.J.; Nanda, S.; Mealey, R.H. Antibody Escape Kinetics of Equine Infectious Anemia Virus Infection of Horses. J. Virol. 2015, 89, 6945–6951. [Google Scholar] [CrossRef]
- Zheng, Y.H.; Sentsui, H.; Nakaya, T.; Kono, Y.; Ikuta, K. In vivo dynamics of equine infectious anemia viruses emerging during febrile episodes: Insertions/duplications at the principal neutralizing domain. J. Virol. 1997, 71, 5031–5039. [Google Scholar] [CrossRef] [PubMed]
- Buzón, M.J.; Codoñer, F.M.; Frost, S.D.W.; Pou, C.; Puertas, M.C.; Massanella, M.; Dalmau, J.; Llibre, J.M.; Stevenson, M.; Blanco, J.; et al. Deep molecular characterization of hiv-1 dynamics under suppressive haart. PLoSPathog 2011, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.B.; Zhu, W.; Cook, F.R.; Goto, Y.; Horii, Y.; Haga, T. Identification of a novel equine infectious anemia virus field strain isolated from feral horses in southern Japan. J. Gen. Virol. 2013, 94, 360–365. [Google Scholar] [CrossRef] [PubMed]
- Quinlivan, M.; Cook, F.; Kenna, R.; Callinan, J.J.; Cullinane, A. Genetic characterization by composite sequence analysis of a new pathogenic field strain of equine infectious anemia virus from the 2006 outbreak in Ireland. J. Gen. Virol. 2013, 94, 612–622. [Google Scholar] [CrossRef]
- Dorey-Robinson, D.L.W.; Locker, N.; Steinbach, F.; Choudhury, B. Molecular characterization of equine infectious anaemia virus strains detected in England in 2010 and 2012. Transbound. Emerg. Dis. 2019, 66, 2311–2317. [Google Scholar] [CrossRef]
- Cappelli, K.; Cook, R.F.; Stefanetti, V.; Passamonti, F.; Autorino, G.L.; Scicluna, M.T.; Coletti, M.; VeriniSupplizi, A.; Capomaccio, S. Deep sequencing and variant analysis of an Italian pathogenic field strain of equine infectious anaemia virus. Transbound. Emerg. Dis. 2017, 64, 2104–2112. [Google Scholar] [CrossRef] [PubMed]
- Capomaccio, S.; Cappelli, K.; Cook, R.F.; Nardi, F.; Gifford, R.; Marenzoni, M.L.; Passamonti, F. Geographic structuring of global EIAV isolates: A single origin for New World strains? Virus Res. 2012, 163, 656–659. [Google Scholar] [CrossRef] [PubMed]
- OIE. Equine Infectious Anaemia. In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals; OIE: Paris, France, 2013; Available online: http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.05.06_EIA.pdf (accessed on 22 January 2019).
- Silva, R.A.M.S.; Dávila, A.M.R.; Iversson, L.B.; Abreu, U.G.P.D. Equine viral diseases in the Pantanal, Brazil: Studies carried out from 1990 to 1995. Rev. D’élevage Médecine Vétérinaire Pays Trop. 1999, 52, 9–12. [Google Scholar]
- Nogueira, M.F.; Oliveira, J.M.; Santos, C.J.S.; Petzold, H.V.; Aguiar, D.M.; Juliano, R.S.; Reis, J.K.P.; Abreu, U.G.P. Equine infectious anaemia in equids of Southern Pantanal, Brazil: Seroprevalence and evaluation of the adoption of a control programme. Pesqui. Vet. Bras. 2017, 37, 227–233. [Google Scholar] [CrossRef]
- Borges, A.M.C.M.; Silva, L.G.; Nogueira, M.F.; Oliveira, A.C.S.; Segri, N.J.; Ferreira, F.; Witter, R.; Aguiar, D.M. Prevalence and risk factors for Equine Infectious Anemia in Poconé municipality, northern Brazilian Pantanal. Res. Vet. Sci. 2013, 95, 76–81. [Google Scholar] [CrossRef]
- Souza, G.D.S.E.; Gomes, E.G.; Santos, S.A.; Rezende, A.S.C.D.; Andrade, D.R.D.F.; Nogueira, M.F.; Trigo, P.; Abreu, U.G.P.D. Factors affecting the performance of Pantaneiro horses. Rev. Bras. Zootec. 2018, 47. [Google Scholar] [CrossRef]
- Silva, R.A.M.S.; Abreu, U.G.P.D.; Barros, A.T.M.D. AnemiaInfecciosaEqüina: Epizootiologia, Prevenção e Controle no Pantanal. Embrapa Pantanal.-Circ. Técnica 2001, 29, 1–32. [Google Scholar]
- Gifford, R.J. Viral evolution in deep time: Lentiviruses and mammals. Trends Genet. 2012, 28, 89–100. [Google Scholar] [CrossRef]
- Capomaccio, S.; Willand, Z.A.; Cook, S.J.; Issel, C.J.; Santos, E.M.; Reis, J.K.P.; Cook, R.F. Detection, molecular characterization and phylogenetic analysis of full-length equine infectious anemia (EIAV) gag genes isolated from Shackleford Banks wild horses. Vet. Microbiol. 2012, 157, 320–332. [Google Scholar] [CrossRef]
- Coggins, L.; Norcross, N.L.; Nusbaum, S.R. Diagnosis of equine infectious anemia by immunodiffusion test. Am. J. Vet. Res. 1972, 33, 11–18. [Google Scholar]
- Reis, J.K.P.; Diniz, R.S.; Haddad, J.P.A.; Ferraz, I.B.F.; Carvalho, A.F.; Kroon, E.G.; Ferreira, P.C.P.; Leite, R.C. Recombinant envelope protein (rgp90) ELISA for equine infectious anemia virus provides comparable results to the agar gel immunodiffusion. J. Virol. Methods 2012, 180, 62–67. [Google Scholar] [CrossRef]
- Cursino, A.E.; Vilela, A.P.P.; Franco-Luiz, A.P.M.; de Oliveira, J.G.; Nogueira, M.F.; Júnior, J.P.A.; de Aguiar, D.M.; Kroon, E.G. Equine infectious anemia virus in naturally infected horses from the Brazilian Pantanal. Arch. Virol. 2018, 163, 2385–2394. [Google Scholar] [CrossRef] [PubMed]
- Ullmann, L.S.; de Camargo Tozato, C.; Malossi, C.D.; da Cruz, T.F.; Cavalcante, R.V.; Kurissio, J.K.; Cagnini, D.Q.; Rodrigues, M.V.; Biondo, A.W.; Araujo, J.P. Comparative clinical sample preparation of DNA and RNA viral nucleic acids for a commercial deep sequencing system (Illumina MiSeq(®)). J. Virol. Methods 2015, 220, 60–63. [Google Scholar] [CrossRef] [PubMed]
- Kruger, F. Trim Galore. Available online: https://github.com/FelixKrueger/TrimGalore (accessed on 15 October 2019).
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Leung, H.C.M.; Yiu, S.M.; Chin, F.Y.L. IDBA-UD: A de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 2012, 28, 1420–1428. [Google Scholar] [CrossRef]
- Bankevick, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Kerfeld, C.A.; Scott, K.M. Using BLAST to teach “E-value-tionary” concepts. Plos Biol. 2011, 9, 1–4. [Google Scholar] [CrossRef]
- Ondov, B.D.; Bergman, N.H.; Phillippy, A.M. Interactive metagenomic visualization in a Web browser. BMC Bioinform. 2011, 12, 1–9. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Rozen, S.; Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 2000, 132, 365–386. [Google Scholar] [PubMed]
- Jin, Z.; Jin, L.; Peterson, D.L.; Lawson, C.L. Model for lentivirus capsid core assembly based on crystal dimers of EIAV p26. J. Mol. Biol. 1999, 286, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Söding, J.; Biegert, A.; Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 2005, 33, 244–248. [Google Scholar] [CrossRef] [PubMed]
- Šali, A.; Potterton, L.; Yuan, F.; van Vlijmen, H.; Karplus, M. Evaluation of comparative protein modeling by Modeller. Proteins Struct. Funct. Bioinforma. 1995, 23, 318–326. [Google Scholar] [CrossRef] [PubMed]
- Chen, V.B.; Arendall, W.B.; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Cryst. Sect. D Biol. Cryst. 2010, 66, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.H.; Payne, S.L.; Issel, C.J.; Montelaro, R.C.; Rushlow, K.E. Characterization of the antigenic domains of the major core protein (p26) of equine infectious anemia virus. J. Virol. 1991, 65, 1007–1012. [Google Scholar] [CrossRef]
- Soutullo, A.; Santi, M.N.; Perin, J.C.; Beltramini, L.M.; Borel, I.M.; Frank, R.; Tonarelli, G.G. Systematic epitope analysis of the p26 EIAV core protein. J. Mol. Recognit. 2007, 20, 227–237. [Google Scholar] [CrossRef]
- Hu, Z.; Chang, H.; Chu, X.; Li, S.; Wang, M.; Wang, X. Identification and characterization of a common B-cell epitope on EIAV capsid proteins. Appl. Microbiol. Biotechnol. 2016, 100, 10531–10542. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera - A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Wang, X.-F.; Liu, Q.; Wang, Y.-H.; Wang, S.; Chen, J.; Lin, Y.-Z.; Ma, J.; Zhou, J.-H.; Wang, X. Characterization of Equine Infectious Anemia Virus Long Terminal Repeat Quasispecies In Vitro and In Vivo. J. Virol. 2018, 92, 2150–2170. [Google Scholar] [CrossRef] [PubMed]
- Tigre, D.M.; Brandão, C.F.L.; de Paula, F.L.; Chinalia, F.A.; Campos, G.S.; Sardi, S.I. Characterization of isolates of equine infectious anemia virus in Brazil. Arch. Virol. 2017, 162, 873–877. [Google Scholar] [CrossRef] [PubMed]
- Jara, M.; Frias-De-Diego, A.; Machado, G. Phylogeography of equine infectious anemia virus. bioRxiv 2019, 1, 1–19. [Google Scholar] [CrossRef]
- Dong, J.; Cook, F.R.; Zhu, W. Equine infectious anemia virus in Japan: Viral isolates V70 and V26 are of North American not Japanese origin. Vet. Microbiol. 2014, 174, 276–278. [Google Scholar] [CrossRef]
- Howe, L.; Leroux, C.; Issel, C.J.; Montelaro, R.C. Equine infectious anemia virus envelope evolution in vivo during persistent infection progressively increases resistance to in vitro serum antibody neutralization as a dominant phenotype. J. Virol. 2002, 76, 10588–10597. [Google Scholar] [CrossRef]
- Craigo, J.K.; Montelaro, R.C. EIAV envelope diversity: Shaping viral persistence and encumbering vaccine efficacy. Curr. Hiv. Res. 2010, 8, 81–86. [Google Scholar] [CrossRef]
- Autorino, G.L.; Eleni, C.; Manna, G.; Frontoso, R.; Nardini, R.; Cocumelli, C.; Rosone, F.; Caprioli, A.; Alfieri, L.; Scicluna, M.T. Evolution of equine infectious anaemia in naturally infected mules with different serological reactivity patterns prior and after immune suppression. Vet. Microbiol. 2016, 189, 15–23. [Google Scholar] [CrossRef]
- Scicluna, M.T.; Autorino, G.L.; Nogarol, C.; Ricci, I.; Frontoso, R.; Rosone, F.; Nardini, R. Validation of an indirect ELISA employing a chimeric recombinant gag and env peptide for the serological diagnosis of equine infectious anemia. J. Virol. Methods 2018, 251, 111–117. [Google Scholar] [CrossRef]
- Ricotti, S.; Garcia, M.I.; Veaute, C.; Bailat, A.; Lucca, E.; Cook, R.F.; Cook, S.J.; Soutullo, A. Serologically silent, occult equine infectious anemia virus (EIAV) infections in horses. Vet. Microbiol. 2016, 187, 41–49. [Google Scholar] [CrossRef]
- Hemelaar, J. The origin and diversity of the HIV-1 pandemic. Trends Mol. Med. 2012, 18, 182–192. [Google Scholar] [CrossRef]
- Burke, D.S. Recombination in HIV: An Important Viral Evolutionary Strategy. Emerg. Infect. Dis. 1997, 3, 253–259. [Google Scholar] [CrossRef] [PubMed]

| Primer | Sequence 5′-3′ | Genome Position | Product Size |
|---|---|---|---|
| AIE_GEN F 463-482 | CCA GAG CAC AGG ARG ACA GG | 463-482 | |
| AIE_GEN R 926-906 * | CCT CTG GGK GTT AAG GGT CGG | 926-906 | 889 bp |
| AIE_GEN R 1352-1333 | TAA GGC TCT TTK GGS CCT TG | 1352-1333 | |
| AIE_GEN F 1333-1352 | CAA GGS CCM AAA GAG CCT TA | 1333-1352 | |
| AIE_GEN R 1963-1943 * | TTA CTC CCA CAA ACT GCT CAG | 1963-1943 | 940 bp |
| AIE_GEN R 2273-2254 | GCC ATT ACC AAT TGT GCC CC | 2273-2254 | |
| AIE_GEN F 3878-3897 | GCC AGG TCA CAA GGG CAT AT | 3878-3897 | |
| AIE_GEN R 4540-4521 * | CCT GCA GGT CCA GAT CCY TG | 4540-4521 | 1096 bp |
| AIE_GEN R 4974-4955 | CCA TGG TGT TTG KCC YCC CA | 4974-4955 | |
| AIE_GEN F 5112-5132 | GGG TGA TGG TGC TGT AGT GGT | 5112-5132 | |
| AIE_GEN F 5934-5959 * | GTG TAC AGA TAG TGA TCA TTG TCA AG | 5934-5959 | 1637 bp |
| AIE_GEN R 6749-6727 | GCC CGA GAA GTA ACA GGA AAA GG | 6749-6727 | |
| AIE_GEN F 6823-6842 | GCT ATT GCT GCT AGT GCY AC | 6823-6842 | 1177 bp |
| AIE_GEN R 8000-7981 | AGA TGT AGC TGG ATT TAR CG | 8000-7981 | |
| AIE_GEN F 7751-7770 | CAA AGC GAA GGA GGA AAC AT | 7751-7770 | 526 bp |
| AIE_GEN R 8277-8258 | GGG ACT CAG ACC GCA GAA TC | 8277-8258 |
| Samples | Raw Reads | Reads at Least Q30 | Mapped Reads | Mean Coverage |
|---|---|---|---|---|
| BRA1 | 18,371,626 | 92.7% | 422,676 | 3969 |
| BRA2 | 10,374,696 | 71.2% | 6927 | 78.3 |
| Miyazaki | BRA1 | BRA2 | Cornwall | Wyoming | Devon | Liaoning | IRE H3 | IRE F4 | IRE F2 | IRE F3 | ITA SA | ITA DE | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Miyazaki - JX003263 | |||||||||||||
| BRA1 - MN560970 | 74.963 | ||||||||||||
| BRA2 - MN560971 | 74.944 | 88.507 | |||||||||||
| Cornwall 2012 - MH580898 | 72.507 | 75.171 | 75.440 | ||||||||||
| Wyoming - AF033820 | 75.156 | 76.601 | 76.614 | 74.829 | |||||||||
| Devon 2010 - MH580897 | 75.897 | 76.500 | 77.322 | 75.255 | 77.490 | ||||||||
| Liaoning - AF327877 | 76.531 | 76.751 | 77.491 | 75.115 | 77.860 | 78.906 | |||||||
| IRE H3 - JX480634 | 75.568 | 77.021 | 76.741 | 75.044 | 78.189 | 78.855 | 79.018 | ||||||
| IRE F4 - JX480633 | 75.520 | 77.058 | 76.741 | 74.994 | 78.196 | 78.951 | 79.066 | 99.190 | |||||
| IRE F2 - JX480631 | 75.348 | 77.042 | 77.015 | 75.224 | 78.393 | 79.080 | 79.297 | 98.937 | 99.118 | ||||
| IRE F3 - JX480632 | 75.612 | 77.005 | 76.741 | 75.053 | 78.149 | 78.934 | 78.990 | 99.057 | 99.299 | 99.420 | |||
| ITA SA - KM247555 | 75.455 | 76.756 | 76.823 | 75.409 | 78.338 | 79.128 | 79.252 | 98.512 | 98.792 | 99.197 | 98.983 | ||
| ITA DE - KM247554 | 75.253 | 76.630 | 76.798 | 75.282 | 78.288 | 79.014 | 79.176 | 98.461 | 98.665 | 99.133 | 98.944 | 99.324 |
| Reference | Epitope | BRA1 | BRA2 |
|---|---|---|---|
| [35] | 73NLDKIAEE80 | 73LLDKMAED80 | 73LLDKIAED80 |
| [35] | 199KNAMRHLRPEDTLEEKMYAC218 | 199RNAMRHLRPEDSLEEKLYAC218 | 199KNAMRHLRPEDTLEEKLYAC218 |
| [33] | 158KEPYPEFVDRLLSQI172 | 158KEPYPEFVDRLLSQI172 | 158KEPYPEFVDRLLSQI172 |
| [34] | 200NAMRHL205 | 200NAMRHL205 | 200NAMRHL205 |
| [34] | 215MYACRD220 | 215LYACRD220 | 215LYACRD220 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malossi, C.D.; Fioratti, E.G.; Cardoso, J.F.; Magro, A.J.; Kroon, E.G.; Aguiar, D.M.d.; Borges, A.M.C.M.; Nogueira, M.F.; Ullmann, L.S.; Araujo, J.P., Jr. High Genomic Variability in Equine Infectious Anemia Virus Obtained from Naturally Infected Horses in Pantanal, Brazil: An Endemic Region Case. Viruses 2020, 12, 207. https://doi.org/10.3390/v12020207
Malossi CD, Fioratti EG, Cardoso JF, Magro AJ, Kroon EG, Aguiar DMd, Borges AMCM, Nogueira MF, Ullmann LS, Araujo JP Jr. High Genomic Variability in Equine Infectious Anemia Virus Obtained from Naturally Infected Horses in Pantanal, Brazil: An Endemic Region Case. Viruses. 2020; 12(2):207. https://doi.org/10.3390/v12020207
Chicago/Turabian StyleMalossi, Camila Dantas, Eduardo Gorzoni Fioratti, Jedson Ferreira Cardoso, Angelo Jose Magro, Erna Geessien Kroon, Daniel Moura de Aguiar, Alice Mamede Costa Marque Borges, Marcia Furlan Nogueira, Leila Sabrina Ullmann, and João Pessoa Araujo, Jr. 2020. "High Genomic Variability in Equine Infectious Anemia Virus Obtained from Naturally Infected Horses in Pantanal, Brazil: An Endemic Region Case" Viruses 12, no. 2: 207. https://doi.org/10.3390/v12020207
APA StyleMalossi, C. D., Fioratti, E. G., Cardoso, J. F., Magro, A. J., Kroon, E. G., Aguiar, D. M. d., Borges, A. M. C. M., Nogueira, M. F., Ullmann, L. S., & Araujo, J. P., Jr. (2020). High Genomic Variability in Equine Infectious Anemia Virus Obtained from Naturally Infected Horses in Pantanal, Brazil: An Endemic Region Case. Viruses, 12(2), 207. https://doi.org/10.3390/v12020207






