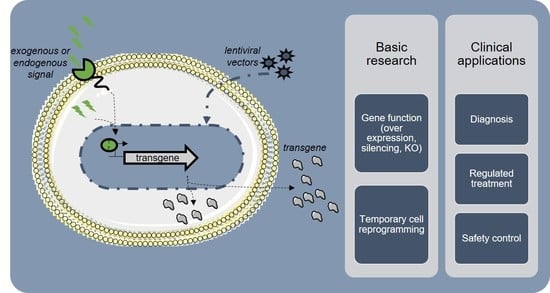
Toward Tightly Tuned Gene Expression Following Lentiviral Vector Transduction
Abstract
:1. Introduction
2. Induction of Transgene Expression by External Stimuli
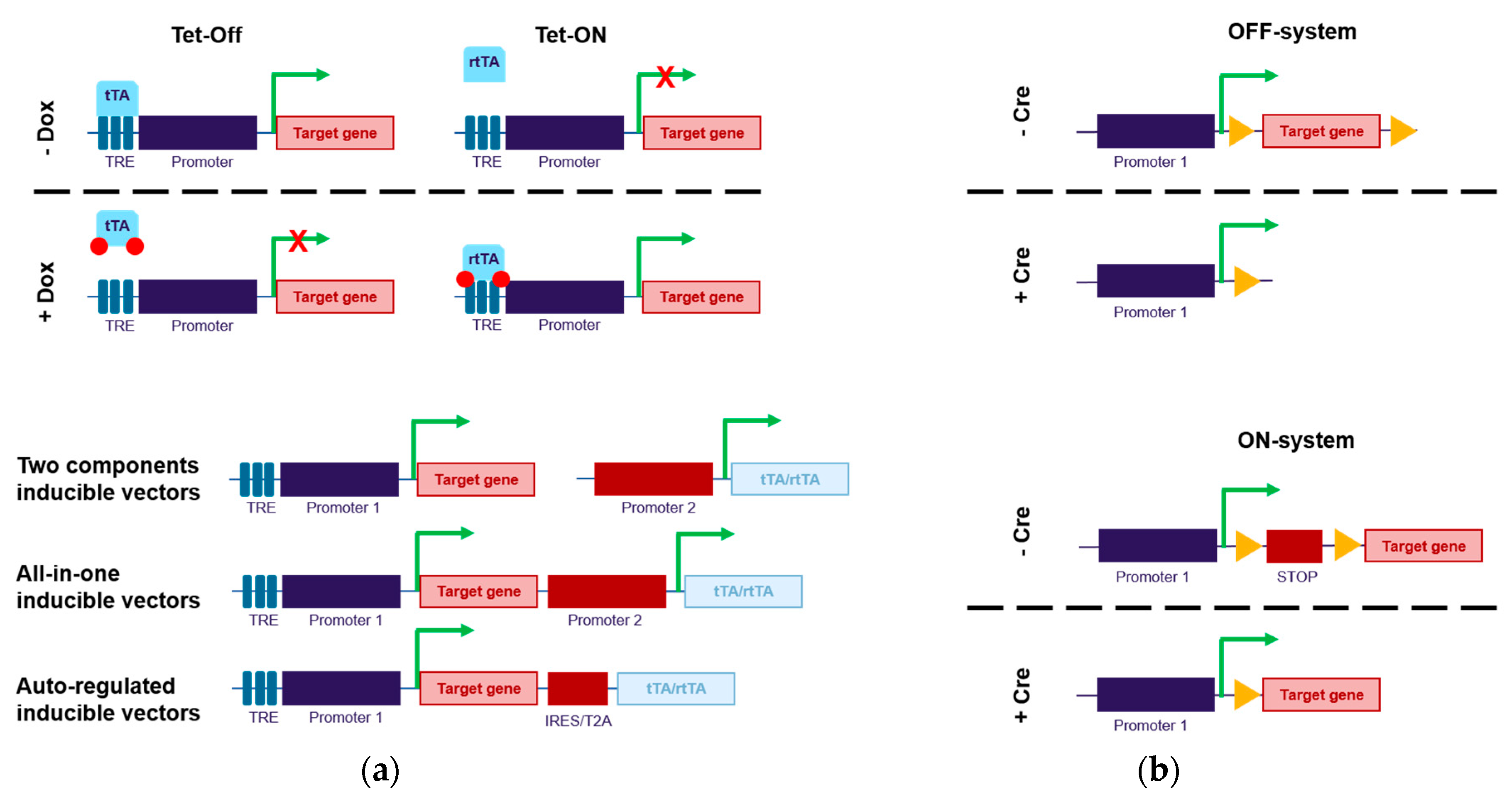
2.1. Direct Control of Transgene Expression (Through Promoter Regulation)
2.2. Indirect Control of Transgene Expression
2.3. Conclusions
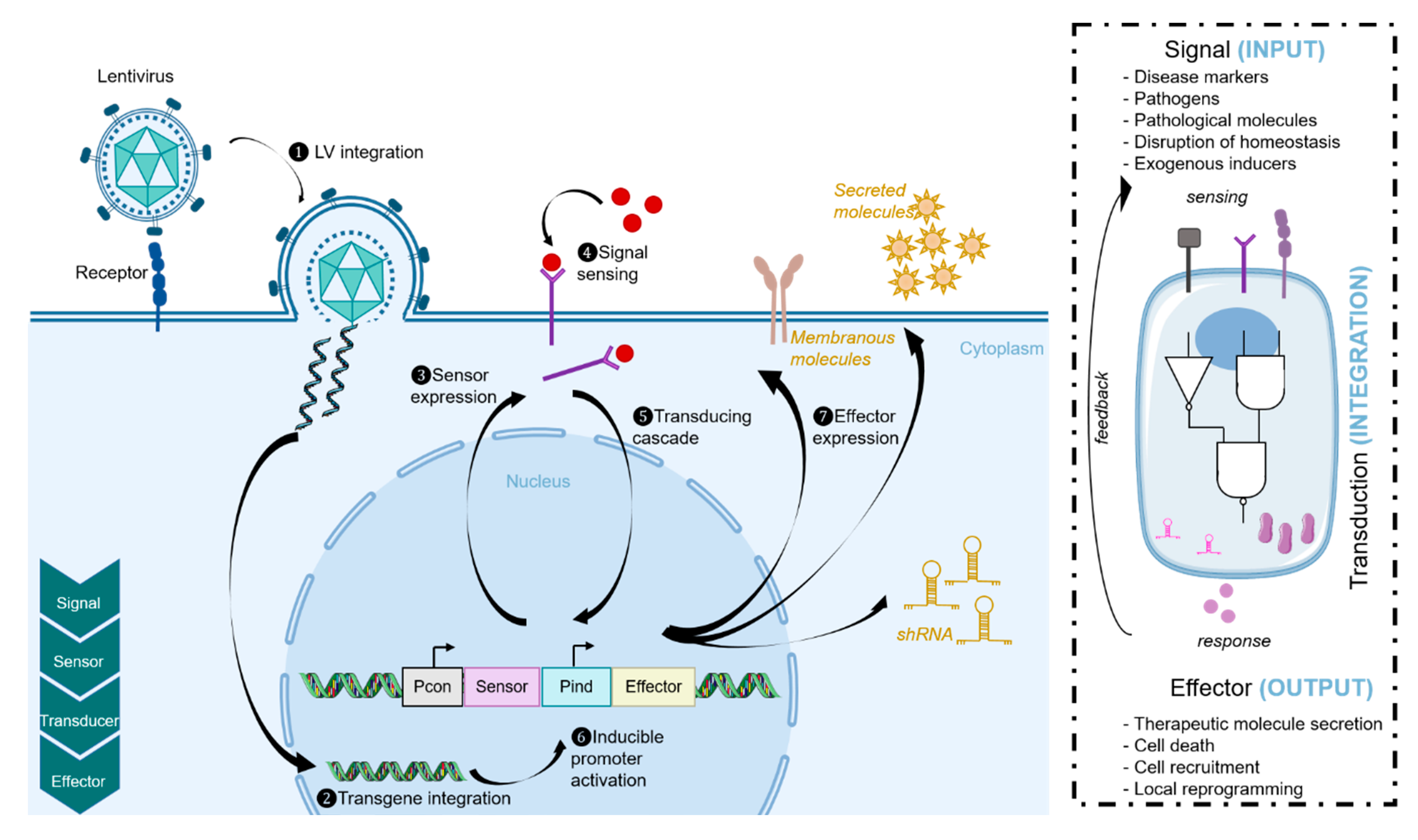
3. Internal Induction by Endogenous Stimuli
3.1. Small Molecule Sensitive Promoters
3.2. Synthetic Biology Approaches
4. Applications of Inducible LVs
4.1. Gene Function
4.2. Cell Reprogramming
4.3. Transient Therapeutic Transgene Expression
4.4. Safety Switches
5. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Frecha, C.; Costa, C.; Lévy, C.; Nègre, D.; Russell, S.J.; Maisner, A.; Salles, G.; Peng, K.-W.; Cosset, F.-L.; Verhoeyen, E. Efficient and Stable Transduction of Resting B Lymphocytes and Primary Chronic Lymphocyte Leukemia Cells Using Measles Virus gp Displaying Lentiviral Vectors. Blood 2009, 114, 3173–3180. [Google Scholar] [CrossRef] [PubMed]
- Girard-Gagnepain, A.; Amirache, F.; Costa, C.; Lévy, C.; Frecha, C.; Fusil, F.; Nègre, D.; Lavillette, D.; Cosset, F.-L.; Verhoeyen, E. Baboon Envelope Pseudotyped LVs Outperform VSV-G-LVs for Gene Transfer into Early-Cytokine-Stimulated and Resting HSCs. Blood 2014, 124, 1221–1231. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sandrin, V.; Boson, B.; Salmon, P.; Gay, W.; Nègre, D.; Le Grand, R.; Trono, D.; Cosset, F.-L. Lentiviral Vectors Pseudotyped with a Modified RD114 Envelope Glycoprotein Show Increased Stability in Sera and Augmented Transduction of Primary Lymphocytes and CD34+ Cells Derived from Human and Nonhuman Primates. Blood 2002, 100, 823–832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duvergé, A.; Negroni, M. Pseudotyping Lentiviral Vectors: When the Clothes Make the Virus. Viruses 2020, 12, 1311. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.M.; Braun, A.H.; Scheib, L.; Agarwal, S.; Schneider, I.C.; Fusil, F.; Perian, S.; Sahin, U.; Thalheimer, F.B.; Verhoeyen, E.; et al. Combining T-cell-specific Activation and In Vivo Gene Delivery Through CD3-Targeted Lentiviral Vectors. Blood Adv. 2020, 4, 5702–5715. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Guerrero, A.; Cosset, F.-L.; Verhoeyen, E. Lentiviral Vector Pseudotypes: Precious Tools to Improve Gene Modification of Hematopoietic Cells for Research and Gene Therapy. Viruses 2020, 12, 1016. [Google Scholar] [CrossRef]
- Hatziioannou, T.; Delahaye, E.; Martin, F.; Russell, S.J.; Cosset, F.-L. Retroviral Display of Functional Binding Domains Fused to the Amino Terminus of Influenza Hemagglutinin. Hum. Gene Ther. 1999, 10, 1533–1544. [Google Scholar] [CrossRef]
- Durand, S.; Cimarelli, A. The Inside Out of Lentiviral Vectors. Viruses 2011, 3, 132–159. [Google Scholar] [CrossRef]
- Naldini, L.; Blomer, U.; Gallay, P.; Ory, D.; Mulligan, R.; Gage, F.H.; Verma, I.M.; Trono, D. In Vivo Gene Delivery and Stable Transduction of Nondividing Cells by a Lentiviral Vector. Science 1996, 272, 263–267. [Google Scholar] [CrossRef] [Green Version]
- Dull, T.; Zufferey, R.; Kelly, M.; Mandel, R.J.; Nguyen, M.; Trono, D.; Naldini, L. A Third-Generation Lentivirus Vector with a Conditional Packaging System. J. Virol. 1998, 72, 8463–8471. [Google Scholar] [CrossRef] [Green Version]
- Zufferey, R.; Donello, J.E.; Trono, D.; Hope, T.J. Woodchuck Hepatitis Virus Posttranscriptional Regulatory Element Enhances Expression of Transgenes Delivered by Retroviral Vectors. J. Virol. 1999, 73, 2886–2892. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miyoshi, H.; Blömer, U.; Takahashi, M.; Gage, F.H.; Verma, I.M. Development of a Self-Inactivating Lentivirus Vector. J. Virol. 1998, 72, 8150–8157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benabdellah, K.; Muñoz, P.; Cobo, M.; Gutierrez-Guerrero, A.; Sánchez-Hernández, S.; Garcia-Perez, A.; Anderson, P.; Carrillo-Gálvez, A.B.; Toscano, M.G.; Martin, F. Lent-On-Plus Lentiviral Vectors for Conditional Expression in Human Stem Cells. Sci. Rep. 2016, 6, 37289. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhan, X.; D’Costa, J.; Tanavde, V.M.; Ye, Z.; Peng, T.; Malehorn, M.T.; Yang, X.; Civin, C.I.; Cheng, L. Lentiviral Vectors with Two Independent Internal Promoters Transfer High-Level Expression of Multiple Transgenes to Human Hematopoietic Stem-Progenitor Cells. Mol. Ther. 2003, 7, 827–838. [Google Scholar] [CrossRef]
- Park, S.K.; Hwang, B.J.; Kee, Y. Promoter Cross-Talk Affects the Inducible Expression of Intronic shRNAs from the Tetracycline Response Element. Genes Genom. 2019, 41, 483–490. [Google Scholar] [CrossRef] [PubMed]
- Kelm, J.M.; Kramer, B.P.; Gonzalez-Nicolini, V.; Ley, B.; Fussenegger, M. Synergies of Microtissue Design, Viral Transduction and Adjustable Transgene Expression for Regenerative Medicine. Biotechnol. Appl. Biochem. 2004, 39, 3. [Google Scholar] [CrossRef]
- Gossen, M.; Bujard, H. Tight Control of Gene Expression in Mammalian Cells by Tetracycline-Responsive Promoters. Proc. Natl. Acad. Sci. USA 1992, 89, 5547–5551. [Google Scholar] [CrossRef] [Green Version]
- Markusic, D.; Seppen, J. Doxycycline Regulated Lentiviral Vectors. In Lentivirus Gene Engineering Protocols; Federico, M., Ed.; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2010; Volume 614, pp. 69–76. ISBN 978-1-60761-532-3. [Google Scholar]
- Chait, R.; Palmer, A.C.; Yelin, I.; Kishony, R. Pervasive Selection for and against Antibiotic Resistance in Inhomogeneous Multistress Environments. Nat. Commun. 2016, 7, 10333. [Google Scholar] [CrossRef] [Green Version]
- Koponen, J.K.; Kankkonen, H.; Kannasto, J.; Wirth, T.; Hillen, W.; Bujard, H.; Ylä-Herttuala, S. Doxycycline-Regulated Lentiviral Vector System with a Novel Reverse Transactivator rtTA2S-M2 Shows a Tight Control of Gene Expression In Vitro and In Vivo. Gene Ther. 2003, 10, 459–466. [Google Scholar] [CrossRef] [Green Version]
- Urlinger, S.; Baron, U.; Thellmann, M.; Hasan, M.T.; Bujard, H.; Hillen, W. Exploring the Sequence Space for Tetracycline-Dependent Transcriptional Activators: Novel Mutations Yield Expanded Range and Sensitivity. Proc. Natl. Acad. Sci. USA 2000, 97, 7963–7968. [Google Scholar] [CrossRef] [Green Version]
- Loew, R.; Heinz, N.; Hampf, M.; Bujard, H.; Gossen, M. Improved Tet-Responsive Promoters with Minimized Background Expression. BMC Biotechnol. 2010, 10, 81. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baron, U.; Freundlieb, S.; Gossen, M.; Bujard, H. Co-Regulation of Two Gene Activities by Tetracycline via a Bidirectional Promoter. Nucleic Acids Res. 1995, 23, 3605–3606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loew, R.; Vigna, E.; Lindemann, D. Retroviral Vectors Containing Tet-Controlled Bidirectional Transcription Units for Simultaneous Regulation of Two Gene Activities. J. Mol. Genet. Med. 2006, 2, 107–118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vieyra, D.S.; Goodell, M.A. Pluripotentiality and Conditional Transgene Regulation in Human Embryonic Stem Cells Expressing Insulated Tetracycline-ON Transactivator. Stem Cells 2007, 25, 2559–2566. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Burrows, C.; Park, J.H. Development of a Doxycycline-Inducible Lentiviral Plasmid with an Instant Regulatory Feature. Plasmid 2014, 72, 29–35. [Google Scholar] [CrossRef]
- Barde, I.; Zanta-Boussif, M.A.; Paisant, S.; Leboeuf, M.; Rameau, P.; Delenda, C.; Danos, O. Efficient Control of Gene Expression in the Hematopoietic System Using a Single Tet-on Inducible Lentiviral Vector. Mol. Ther. 2006, 13, 382–390. [Google Scholar] [CrossRef]
- Giry-Laterrière, M.; Cherpin, O.; Kim, Y.-S.; Jensen, J.; Salmon, P. Polyswitch Lentivectors: “All-in-One” Lentiviral Vectors for Drug-Inducible Gene Expression, Live Selection, and Recombination Cloning. Hum. Gene Ther. 2011, 22, 1255–1267. [Google Scholar] [CrossRef] [Green Version]
- De Groote, P.; Grootjans, S.; Lippens, S.; Eichperger, C.; Leurs, K.; Kahr, I.; Tanghe, G.; Bruggeman, I.; De Schamphelaire, W.; Urwyler, C.; et al. Generation of a New Gateway-Compatible Inducible Lentiviral Vector Platform Allowing Easy Derivation of Co-transduced Cells. BioTechniques 2016, 60, 252–259. [Google Scholar] [CrossRef] [Green Version]
- Kafri, T.; van Praag, H.; Gage, F.H.; Verma, I.M. Lentiviral Vectors: Regulated Gene Expression. Mol. Ther. 2000, 1, 516–521. [Google Scholar] [CrossRef]
- Huang, Y.; Zhen, R.; Jiang, M.; Yang, J.; Yang, Y.; Huang, Z.; Lin, Y. Development of All-in-One Multicistronic Tet-On Lentiviral Vectors for Inducible Co-expression of Two Transgenes: All-in-One Multicistronic Tet-on Lentiviral Vectors. Biotechnol. Appl. Biochem. 2015, 62, 48–54. [Google Scholar] [CrossRef]
- Ogueta, S.B.; Yao, F.; Marasco, W.A. Design and In Vitro Characterization of a Single Regulatory Module for Efficient Control of Gene Expression in Both Plasmid DNA and a Self-Inactivating Lentiviral Vector. Mol. Med. 2001, 7, 569–579. [Google Scholar] [CrossRef] [PubMed]
- Centlivre, M.; Zhou, X.; Pouw, S.M.; Weijer, K.; Kleibeuker, W.; Das, A.T.; Blom, B.; Seppen, J.; Berkhout, B.; Legrand, N. Autoregulatory Lentiviral Vectors Allow Multiple Cycles of Doxycycline-Inducible Gene Expression in Human Hematopoietic Cells In Vivo. Gene Ther. 2010, 17, 14–25. [Google Scholar] [CrossRef] [PubMed]
- Benabdellah, K.; Cobo, M.; Muñoz, P.; Toscano, M.G.; Martin, F. Development of an All-in-One Lentiviral Vector System Based on the Original TetR for the Easy Generation of Tet-ON Cell Lines. PLoS ONE 2011, 6, e23734. [Google Scholar] [CrossRef] [PubMed]
- Alexeyev, M.F.; Fayzulin, R.; Shokolenko, I.N.; Pastukh, V. A Retro-Lentiviral System for Doxycycline-Inducible Gene Expression and Gene Knockdown in Cells with Limited Proliferative Capacity. Mol. Biol. Rep. 2010, 37, 1987–1991. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoyng, S.A.; Gnavi, S.; de Winter, F.; Eggers, R.; Ozawa, T.; Zaldumbide, A.; Hoeben, R.C.; Malessy, M.J.A.; Verhaagen, J. Developing a Potentially Immunologically Inert Tetracycline-Regulatable Viral Vector for Gene Therapy in the Peripheral Nerve. Gene Ther. 2014, 21, 549–557. [Google Scholar] [CrossRef] [PubMed]
- Reiser, J.; Lai, Z.; Zhang, X.-Y.; Brady, R.O. Development of Multigene and Regulated Lentivirus Vectors. J. Virol. 2000, 74, 10589–10599. [Google Scholar] [CrossRef] [Green Version]
- Mullick, A.; Xu, Y.; Warren, R.; Koutroumanis, M.; Guilbault, C.; Broussau, S.; Malenfant, F.; Bourget, L.; Lamoureux, L.; Lo, R.; et al. The Cumate Gene-Switch: A System for Regulated Expression in Mammalian Cells. BMC Biotechnol. 2006, 6, 43. [Google Scholar] [CrossRef] [Green Version]
- Malphettes, L. A Novel Mammalian Expression System Derived from Components Coordinating Nicotine Degradation in Arthrobacter Nicotinovorans pAO1. Nucleic Acids Res. 2005, 33, e107. [Google Scholar] [CrossRef]
- Fux, C.; Fussenegger, M. Bidirectional Expression Units Enable Streptogramin-Adjustable Gene Expression in Mammalian Cells. Biotechnol. Bioeng. 2003, 83, 618–625. [Google Scholar] [CrossRef]
- Fux, C.; Weber, W.; Daoud-El Baba, M.; Heinzen, C.; Aubel, D.; Fussenegger, M. Novel Macrolide-Adjustable Bidirectional Expression Modules for Coordinated Expression of Two Different Transgenes in Mice. J. Gene Med. 2003, 5, 1067–1079. [Google Scholar] [CrossRef]
- Mitta, B.; Weber, C.C.; Fussenegger, M. In Vivo Transduction of HIV-1-Derived Lentiviral Particles Engineered for Macrolide-Adjustable Transgene Expression. J. Gene Med. 2005, 7, 1400–1408. [Google Scholar] [CrossRef] [PubMed]
- Mitta, B. Design and In Vivo Characterization of Self-Inactivating Human and Non-Human Lentiviral Expression Vectors Engineered for Streptogramin-Adjustable Transgene Expression. Nucleic Acids Res. 2004, 32, e106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fux, C.; Fussenegger, M. Toward Higher Order Control Modalities in Mammalian Cells-Independent Adjustment of Two Different Gene Activities. Biotechnol. Prog. 2003, 19, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Kramer, B.P.; Fischer, C.; Fussenegger, M. BioLogic Gates Enable Logical Transcription Control in Mammalian Cells. Biotechnol. Bioeng. 2004, 87, 478–484. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bojar, D.; Scheller, L.; Hamri, G.C.-E.; Xie, M.; Fussenegger, M. Caffeine-Inducible Gene Switches Controlling Experimental Diabetes. Nat. Commun. 2018, 9, 2318. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, P.; Liu, Y.; Xue, S.; Hamri, G.C.-E.; Saxena, P.; Ye, H.; Xie, M.; Fussenegger, M. A Fully Human Transgene Switch to Regulate Therapeutic Protein Production by Cooling Sensation. Nat. Med. 2019, 25, 1266–1273. [Google Scholar] [CrossRef]
- Gitzinger, M.; Kemmer, C.; El-Baba, M.D.; Weber, W.; Fussenegger, M. Controlling Transgene Expression in Subcutaneous Implants Using a Skin Lotion Containing the Apple Metabolite Phloretin. Proc. Natl. Acad. Sci. USA 2009, 106, 10638–10643. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Xie, M.; Charpin-El Hamri, G.; Ye, H.; Fussenegger, M. Treatment of Chronic Pain by Designer Cells Controlled by Spearmint Aromatherapy. Nat. Biomed. Eng. 2018, 2, 114–123. [Google Scholar] [CrossRef]
- Xie, M.; Ye, H.; Hamri, G.C.-E.; Fussenegger, M. Antagonistic Control of a Dual-Input Mammalian Gene Switch by Food Additives. Nucleic Acids Res. 2014, 42, e116. [Google Scholar] [CrossRef]
- Gitzinger, M.; Kemmer, C.; Fluri, D.A.; Daoud El-Baba, M.; Weber, W.; Fussenegger, M. The Food Additive Vanillic Acid Controls Transgene Expression in Mammalian Cells and Mice. Nucleic Acids Res. 2012, 40, e37. [Google Scholar] [CrossRef]
- Li, S.; Ma, L.; Ou, M.; Feng, J.; Liao, Y.; Wang, G.; Tang, L. A Novel Inducible Lentiviral System for Multi-Gene Expression with Human HSP70 Promoter and Tetracycline-Induced Promoter. Appl. Microbiol. Biotechnol. 2017, 101, 3689–3702. [Google Scholar] [CrossRef] [PubMed]
- Marignol, L.; Coffey, M.; Hollywood, D.; Lawler, M. Radiation to Control Transgene Expression in Tumors. Cancer Biol. Ther. 2007, 6, 1005–1012. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Datta, R.; Rubin, E.; Sukhatme, V.; Qureshi, S.; Hallahan, D.; Weichselbaum, R.R.; Kufe, D.W. Ionizing Radiation Activates Transcription of the EGR1 Gene via CArG Elements. Proc. Natl. Acad. Sci. USA 1992, 89, 10149–10153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eggers, R.; de Winter, F.; Hoyng, S.A.; Roet, K.C.D.; Ehlert, E.M.; Malessy, M.J.A.; Verhaagen, J.; Tannemaat, M.R. Lentiviral Vector-Mediated Gradients of GDNF in the Injured Peripheral Nerve: Effects on Nerve Coil Formation, Schwann Cell Maturation and Myelination. PLoS ONE 2013, 8, e71076. [Google Scholar] [CrossRef] [PubMed]
- Ventura, A.; Meissner, A.; Dillon, C.P.; McManus, M.; Sharp, P.A.; Van Parijs, L.; Jaenisch, R.; Jacks, T. Cre-Lox-Regulated Conditional RNA Interference from Transgenes. Proc. Natl. Acad. Sci. USA 2004, 101, 10380–10385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Papapetrou, E.P.; Sadelain, M. Generation of Transgene-Free Human Induced Pluripotent Stem Cells with an Excisable Single Polycistronic Vector. Nat. Protoc. 2011, 6, 1251–1273. [Google Scholar] [CrossRef]
- Tiscornia, G.; Tergaonkar, V.; Galimi, F.; Verma, I.M. From the Cover: CRE Recombinase-Inducible RNA Interference Mediated by Lentiviral Vectors. Proc. Natl. Acad. Sci. USA 2004, 101, 7347–7351. [Google Scholar] [CrossRef] [Green Version]
- Bougioukli, S.; Vakhshori, V.; Ortega, B.; Sugiyama, O.; Lieberman, J. Regulated Ex Vivo Regional Gene Therapy for Bone Repair Using an Inducible Caspase-9 Suicide Gene System. Gene Ther. 2019, 26, 230–239. [Google Scholar] [CrossRef]
- Zhou, X.; Di Stasi, A.; Tey, S.-K.; Krance, R.A.; Martinez, C.; Leung, K.S.; Durett, A.G.; Wu, M.-F.; Liu, H.; Leen, A.M.; et al. Long-Term Outcome after Haploidentical Stem Cell Transplant and Infusion of T Cells Expressing the Inducible Caspase 9 Safety Transgene. Blood 2014, 123, 3895–3905. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Wang, S.; Brenner, M.; Paton, J.F.R.; Kasparov, S. Enhancement of Cell-Specific Transgene Expression from a Tet-Off Regulatory System Using a Transcriptional Amplification Strategy in the Rat Brain. J. Gene Med. 2008, 10, 583–592. [Google Scholar] [CrossRef] [Green Version]
- Gascón, S.; Paez-Gomez, J.A.; Díaz-Guerra, M.; Scheiffele, P.; Scholl, F.G. Dual-Promoter Lentiviral Vectors for Constitutive and Regulated Gene Expression in Neurons. J. Neurosci. Methods 2008, 168, 104–112. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Hill, P.J.; Karamitri, A.; Ryan, K.J.P.; Chen, H.Y.; Lomax, M.A. Construction of a Doxycycline Inducible Adipogenic Lentiviral Expression System. Plasmid 2013, 69, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Kramer, M.G.; Fernandez-Ruiz, V.; Kawa, M.P.; Huang, X.; Liu, Z.; Prieto, J.; Qian, C. Development of Endothelial-Specific Single Inducible Lentiviral Vectors for Genetic Engineering of Endothelial Progenitor Cells. Sci. Rep. 2015, 5, 17166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Broeren, M.G.A.; de Vries, M.; Bennink, M.B.; Arntz, O.J.; Blom, A.B.; Koenders, M.I.; van Lent, P.L.E.M.; van der Kraan, P.M.; van den Berg, W.B.; van de Loo, F.A.J. Disease-Regulated Gene Therapy with Anti-Inflammatory Interleukin-10 under the Control of the CXCL10 Promoter for the Treatment of Rheumatoid Arthritis. Hum. Gene Ther. 2016, 27, 244–254. [Google Scholar] [CrossRef] [PubMed]
- Broeren, M.G.A.; de Vries, M.; Bennink, M.B.; Arntz, O.J.; van Lent, P.L.E.M.; van der Kraan, P.M.; van den Berg, W.B.; van den Hoogen, F.H.J.; Koenders, M.I.; van de Loo, F.A.J. Suppression of the Inflammatory Response by Disease-Inducible Interleukin-10 Gene Therapy in a Three-Dimensional Micromass Model of the Human Synovial Membrane. Arthritis Res. Ther. 2016, 18, 186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garaulet, G.; Alfranca, A.; Torrente, M.; Escolano, A.; López-Fontal, R.; Hortelano, S.; Redondo, J.M.; Rodríguez, A. IL10 Released by a New Inflammation-Regulated Lentiviral System Efficiently Attenuates Zymosan-Induced Arthritis. Mol. Ther. 2013, 21, 119–130. [Google Scholar] [CrossRef] [Green Version]
- Henningsson, L.; Eneljung, T.; Jirholt, P.; Tengvall, S.; Lidberg, U.; van den Berg, W.B.; van de Loo, F.A.; Gjertsson, I. Disease-Dependent Local IL-10 Production Ameliorates Collagen Induced Arthritis in Mice. PLoS ONE 2012, 7, e49731. [Google Scholar] [CrossRef]
- Gabner, S.; Hlavaty, J.; Velde, K.; Renner, M.; Jenner, F.; Egerbacher, M. Inflammation-Induced Transgene Expression in Genetically Engineered Equine Mesenchymal Stem Cells: Inducible Expression in Equine MSCs. J. Gene Med. 2016, 18, 154–164. [Google Scholar] [CrossRef]
- Romero, Z.; Torres, S.; Cobo, M.; Muñoz, P.; Unciti, J.D.; Martín, F.; Molina, I.J. A Tissue-Specific, Activation-Inducible, Lentiviral Vector Regulated by Human CD40L Proximal Promoter Sequences. Gene Ther. 2011, 18, 364–371. [Google Scholar] [CrossRef]
- Fernández-Rubio, P.; Torres-Rusillo, S.; Molina, I.J. Regulated Expression of Murine CD40L by a Lentiviral Vector Transcriptionally Targeted Through Its Endogenous Promoter. J. Gene Med. 2015, 17, 219–228. [Google Scholar] [CrossRef] [Green Version]
- Vermeij, E.A.; Broeren, M.G.A.; Bennink, M.B.; Arntz, O.J.; Gjertsson, I.; van Lent, P.L.E.M.; van den Berg, W.B.; Koenders, M.I.; van de Loo, F.A.J. Disease-Regulated Local IL-10 Gene Therapy Diminishes Synovitis and Cartilage Proteoglycan Depletion in Experimental Arthritis. Ann. Rheum. Dis. 2015, 74, 2084–2091. [Google Scholar] [CrossRef] [PubMed]
- Chassin, H.; Geering, B.; Schukur, L.; Ausländer, D.; Lang, B.; Fussenegger, M. Sensing and Responding to Allergic Response Cytokines Through a Genetically Encoded Circuit. Nat. Commun. 2017, 8, 1101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schukur, L.; Fussenegger, M. Engineering of Synthetic Gene Circuits for (Re-)balancing Physiological Processes in Chronic Diseases. Wiley Interdiscip. Rev. Syst. Biol. Med. 2016, 8, 402–422. [Google Scholar] [CrossRef] [PubMed]
- Schukur, L.; Geering, B.; Charpin-El Hamri, G.; Fussenegger, M. Implantable Synthetic Cytokine Converter Cells with AND-Gate Logic Treat Experimental Psoriasis. Sci. Transl. Med. 2015, 7, 318ra201. [Google Scholar] [CrossRef] [PubMed]
- Qudrat, A.; Mosabbir, A.A.; Truong, K. Engineered Proteins Program Mammalian Cells to Target Inflammatory Disease Sites. Cell Chem. Biol. 2017, 24, 703–711.e2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smole, A.; Lainšček, D.; Bezeljak, U.; Horvat, S.; Jerala, R. A Synthetic Mammalian Therapeutic Gene Circuit for Sensing and Suppressing Inflammation. Mol. Ther. 2017, 25, 102–119. [Google Scholar] [CrossRef] [Green Version]
- Schwarz, K.A.; Leonard, J.N. Engineering Cell-Based Therapies to Interface Robustly with Host Physiology. Adv. Drug Deliv. Rev. 2016, 105, 55–65. [Google Scholar] [CrossRef] [Green Version]
- Roybal, K.T.; Lim, W.A. Synthetic Immunology: Hacking Immune Cells to Expand Their Therapeutic Capabilities. Annu. Rev. Immunol. 2017, 35, 229–253. [Google Scholar] [CrossRef] [Green Version]
- Scheller, L.; Strittmatter, T.; Fuchs, D.; Bojar, D.; Fussenegger, M. Generalized Extracellular Molecule Sensor Platform for Programming Cellular Behavior. Nat. Chem. Biol. 2018, 14, 723–729. [Google Scholar] [CrossRef]
- Kojima, R.; Aubel, D.; Fussenegger, M. Building Sophisticated Sensors of Extracellular Cues That Enable Mammalian Cells to Work as “Doctors” in the Body. Cell. Mol. Life Sci. 2020, 77, 3567–3581. [Google Scholar] [CrossRef] [Green Version]
- Morsut, L.; Roybal, K.T.; Xiong, X.; Gordley, R.M.; Coyle, S.M.; Thomson, M.; Lim, W.A. Engineering Customized Cell Sensing and Response Behaviors Using Synthetic Notch Receptors. Cell 2016, 164, 780–791. [Google Scholar] [CrossRef] [Green Version]
- Schwarz, K.A.; Daringer, N.M.; Dolberg, T.B.; Leonard, J.N. Rewiring Human Cellular Input–Output Using Modular Extracellular Sensors. Nat. Chem. Biol. 2017, 13, 202–209. [Google Scholar] [CrossRef]
- Daringer, N.M.; Dudek, R.M.; Schwarz, K.A.; Leonard, J.N. Modular Extracellular Sensor Architecture for Engineering Mammalian Cell-Based Devices. ACS Synth. Biol. 2014, 3, 892–902. [Google Scholar] [CrossRef]
- Higashikuni, Y.; Chen, W.C.; Lu, T.K. Advancing Therapeutic Applications of Synthetic Gene Circuits. Curr. Opin. Biotechnol. 2017, 47, 133–141. [Google Scholar] [CrossRef]
- Roybal, K.T.; Rupp, L.J.; Morsut, L.; Walker, W.J.; McNally, K.A.; Park, J.S.; Lim, W.A. Precision Tumor Recognition by T Cells with Combinatorial Antigen-Sensing Circuits. Cell 2016, 164, 770–779. [Google Scholar] [CrossRef] [Green Version]
- Aubrey, B.J.; Kelly, G.L.; Kueh, A.J.; Brennan, M.S.; O’Connor, L.; Milla, L.; Wilcox, S.; Tai, L.; Strasser, A.; Herold, M.J. An Inducible Lentiviral Guide RNA Platform Enables the Identification of Tumor-Essential Genes and Tumor-Promoting Mutations In Vivo. Cell Rep. 2015, 10, 1422–1432. [Google Scholar] [CrossRef] [Green Version]
- Wiznerowicz, M.; Trono, D. Conditional Suppression of Cellular Genes: Lentivirus Vector-Mediated Drug-Inducible RNA Interference. J. Virol. 2003, 77, 8957–8961. [Google Scholar] [CrossRef] [Green Version]
- Christel, C.J.; Schmied, P.; Jagusch, V.; Schrödel, S.; Thirion, C.; Schmitt, K.; Salomon, M. Versatile Viral Vector Strategies for Postscreening Target Validation and RNAi ON-Target Control. J. Biomol. Screen. 2015, 20, 976–984. [Google Scholar] [CrossRef] [Green Version]
- Peng, Y.; Yang, T.; Tang, X.; Chen, F.; Wang, S. Construction of an Inducible CRISPR/Cas9 System for CXCR4 Gene and Demonstration of Its Effects on MKN-45 Cells. Cell Biochem. Biophys. 2020, 78, 23–30. [Google Scholar] [CrossRef]
- Meerbrey, K.L.; Hu, G.; Kessler, J.D.; Roarty, K.; Li, M.Z.; Fang, J.E.; Herschkowitz, J.I.; Burrows, A.E.; Ciccia, A.; Sun, T.; et al. The pINDUCER Lentiviral Toolkit for Inducible RNA Interference In Vitro and In Vivo. Proc. Natl. Acad. Sci. USA 2011, 108, 3665–3670. [Google Scholar] [CrossRef] [Green Version]
- He, J.; Huang, L.; Qiu, H.; Li, J.; Luo, L.; Li, Y.; Tian, S.; Kang, K.; Luo, J.; Liu, L.; et al. A New Design of a Lentiviral shRNA Vector with Inducible Co-expression of ARGONAUTE 2 for Enhancing Gene Silencing Efficiency. Cell Biosci. 2015, 5, 67. [Google Scholar] [CrossRef] [Green Version]
- Kaspar, P.; Prochazka, J.; Efenberkova, M.; Juhasz, A.; Novosadova, V.; Sedlacek, R. c-Myb Regulates Tumorigenic Potential of Embryonal Rhabdomyosarcoma Cells. Sci. Rep. 2019, 9, 6342. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Liu, X.; Wang, H.; Zhou, Q.; Liang, Y.; Sui, A.; Yao, R.; Zhao, B.; Sun, M. Lentiviral Vector-Mediated Doxycycline-Inducible USP39 shRNA or cDNA Expression in Triple-Negative Breast Cancer Cells. Oncol. Rep. 2015, 33, 2477–2483. [Google Scholar] [CrossRef] [Green Version]
- Merentie, M.; Rissanen, R.; Lottonen-Raikaslehto, L.; Huusko, J.; Gurzeler, E.; Turunen, M.P.; Holappa, L.; Mäkinen, P.; Ylä-Herttuala, S. Doxycycline Modulates VEGF-A Expression: Failure of Doxycycline-Inducible Lentivirus shRNA Vector to Knockdown VEGF-A Expression in Transgenic Mice. PLoS ONE 2018, 13, e0190981. [Google Scholar] [CrossRef]
- Wiederschain, D.; Susan, W.; Chen, L.; Loo, A.; Yang, G.; Huang, A.; Chen, Y.; Caponigro, G.; Yao, Y.; Lengauer, C.; et al. Single-Vector Inducible Lentiviral RNAi System for Oncology Target Validation. Cell Cycle 2009, 8, 498–504. [Google Scholar] [CrossRef]
- Brown, C.Y.; Sadlon, T.; Gargett, T.; Melville, E.; Zhang, R.; Drabsch, Y.; Ling, M.; Strathdee, C.A.; Gonda, T.J.; Barry, S.C. Robust, Reversible Gene Knockdown Using a Single Lentiviral Short Hairpin RNA Vector. Hum. Gene Ther. 2010, 21, 1005–1017. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Zhang, Y.; Pan, R.; Chen, M.; Wang, X.; Kong, E.; Yu, W.; Sun, Y.; Wu, F. Lentiviral-Mediated Inducible Silencing of TLR4 Attenuates Neuropathic Pain in a Rat Model of Chronic Constriction Injury. Mol. Med. Rep. 2018, 18, 5545–5551. [Google Scholar] [CrossRef]
- Wang, Y.; Yu, J.; Liu, L.; Li, W.; Duan, X.; Peng, Y.; Zeng, S.; Ouyang, Q.; Lu, G.; Lin, G.; et al. Generation of a Human Embryonic Stem Cell line, NERCe003-A-1, with Lentivirus Vector-Mediated Inducible CTNNB1 Overexpression. Stem Cell Res. 2018, 28, 61–65. [Google Scholar] [CrossRef]
- Bahi, A.; Dreyer, J.-L. Lentiviral-Mediated Let-7d microRNA Overexpression Induced Anxiolytic- and Anti-depressant-like Behaviors and Impaired Dopamine D3 Receptor Expression. Eur. Neuropsychopharmacol. 2018, 28, 1394–1404. [Google Scholar] [CrossRef] [Green Version]
- Shuen, W.H.; Kan, R.; Yu, Z.; Lung, H.L.; Lung, M.L. Novel Lentiviral-Inducible Transgene Expression Systems and Versatile Single-Plasmid Reporters for In Vitro and In Vivo Cancer Biology Studies. Cancer Gene Ther. 2015, 22, 207–214. [Google Scholar] [CrossRef]
- Belian, E.; Noseda, M.; Abreu Paiva, M.S.; Leja, T.; Sampson, R.; Schneider, M.D. Forward Programming of Cardiac Stem Cells by Homogeneous Transduction with MYOCD plus TBX5. PLoS ONE 2015, 10, e0125384. [Google Scholar] [CrossRef] [PubMed]
- Ballester, M.; Bolonio, M.; Santamaria, R.; Castell, J.V.; Ribes-Koninckx, C.; Bort, R. Direct Conversion of Human Fibroblast to Hepatocytes Using a Single Inducible Polycistronic Vector. Stem Cell Res. Ther. 2019, 10, 317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stahlhut, M.; Schwarzer, A.; Eder, M.; Yang, M.; Li, Z.; Morgan, M.; Schambach, A.; Kustikova, O.S. Lentiviral Vector System for Coordinated Constitutive and Drug Controlled Tetracycline-Regulated Gene Co-expression. Biomaterials 2015, 63, 189–201. [Google Scholar] [CrossRef] [PubMed]
- Tan, G.C.; Wong, Y.P.; Cui, W.; Dibb, N. Construction of a Doxycycline Inducible Lentivirus That Expresses Stem Cell-Specific miR-302 Cluster. Malays. J. Pathol. 2020, 42, 91–97. [Google Scholar]
- Yamaguchi, T.; Hamanaka, S.; Kamiya, A.; Okabe, M.; Kawarai, M.; Wakiyama, Y.; Umino, A.; Hayama, T.; Sato, H.; Lee, Y.-S.; et al. Development of an All-in-One Inducible Lentiviral Vector for Gene Specific Analysis of Reprogramming. PLoS ONE 2012, 7, e41007. [Google Scholar] [CrossRef] [Green Version]
- Sommer, C.A.; Stadtfeld, M.; Murphy, G.J.; Hochedlinger, K.; Kotton, D.N.; Mostoslavsky, G. Induced Pluripotent Stem Cell Generation Using a Single Lentiviral Stem Cell Cassette. Stem Cells 2009, 27, 543–549. [Google Scholar] [CrossRef] [Green Version]
- López-Ornelas, A.; Mejía-Castillo, T.; Vergara, P.; Segovia, J. Lentiviral Transfer of an Inducible Transgene Expressing a Soluble Form of Gas1 Causes Glioma Cell Arrest, Apoptosis and Inhibits Tumor Growth. Cancer Gene Ther. 2011, 18, 87–99. [Google Scholar] [CrossRef] [Green Version]
- Varga, E.; Nemes, C.; Davis, R.P.; Ujhelly, O.; Klincumhom, N.; Polgar, Z.; Muenthaisong, S.; Pirity, M.K.; Dinnyes, A. Generation of Transgene-Free Mouse Induced Pluripotent Stem Cells Using an Excisable Lentiviral System. Exp. Cell Res. 2014, 322, 335–344. [Google Scholar] [CrossRef]
- Vigna, E.; Amendola, M.; Benedicenti, F.; Simmons, A.D.; Follenzi, A.; Naldini, L. Efficient Tet-Dependent Expression of Human Factor IX In Vivo by a New Self-Regulating Lentiviral Vector. Mol. Ther. 2005, 11, 763–775. [Google Scholar] [CrossRef]
- Shi, Q.; Tian, X.; Zhao, Y.; Luo, H.; Tian, Y.; Luo, A. Anti-Arthritic Effects of FasL Gene Transferred Intra-Articularly by an Inducible Lentiviral Vector Containing Improved Tet-On System. Rheumatol. Int. 2014, 34, 51–57. [Google Scholar] [CrossRef]
- Ge, G.; Chen, C.; Guderyon, M.J.; Liu, J.; He, Z.; Yu, Y.; Clark, R.A.; Li, S. Regulatable Lentiviral Hematopoietic Stem Cell Gene Therapy in a Mouse Model of Parkinson’s Disease. Stem Cells Dev. 2018, 27, 995–1005. [Google Scholar] [CrossRef] [PubMed]
- Lachmann, N.; Brennig, S.; Pfaff, N.; Schermeier, H.; Dahlmann, J.; Phaltane, R.; Gruh, I.; Modlich, U.; Schambach, A.; Baum, C.; et al. Efficient In Vivo Regulation of Cytidine Deaminase Expression in the Haematopoietic System Using a Doxycycline-Inducible Lentiviral Vector System. Gene Ther. 2013, 20, 298–307. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lachmann, N.; Brennig, S.; Hillje, R.; Schermeier, H.; Phaltane, R.; Dahlmann, J.; Gruh, I.; Heinz, N.; Schiedlmeier, B.; Baum, C.; et al. Tightly Regulated ‘All-in-One’ Lentiviral Vectors for Protection of Human Hematopoietic Cells from Anticancer Chemotherapy. Gene Ther. 2015, 22, 883–892. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, F.; Okawa, H.; Kamano, Y.; Niibe, K.; Kayashima, H.; Osathanon, T.; Pavasant, P.; Saeki, M.; Yatani, H.; Egusa, H. Controlled Osteogenic Differentiation of Mouse Mesenchymal Stem Cells by Tetracycline-Controlled Transcriptional Activation of Amelogenin. PLoS ONE 2015, 10, e0145677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wübbenhorst, D.; Biol, D.; Dumler, K.; Ing, D.; Wagner, B.; Wexel, G.; Imhoff, A.; Gansbacher, B.; Vogt, S.; Anton, M. Tet-Regulated BMP-2 Gene Expression in Lentivirally Transduced Primary Rabbit Chondrocytes for Treatment of Cartilage Defects. Arthritis Rheum. 2010, 62, 2037–2046. [Google Scholar] [CrossRef]
- Alaee, F.; Sugiyama, O.; Virk, M.S.; Tang, H.; Drissi, H.; Lichtler, A.C.; Lieberman, J.R. Suicide Gene Approach Using a Dual-Expression Lentiviral Vector to Enhance the Safety of Ex Vivo Gene Therapy for Bone Repair. Gene Ther. 2014, 21, 139–147. [Google Scholar] [CrossRef] [Green Version]
- Dossa, R.G.; Cunningham, T.; Sommermeyer, D.; Medina-Rodriguez, I.; Biernacki, M.A.; Foster, K.; Bleakley, M. Development of T-Cell Immunotherapy for Hematopoietic Stem Cell Transplantation Recipients at Risk of Leukemia Relapse. Blood 2018, 131, 108–120. [Google Scholar] [CrossRef]
- Di Stasi, A.; Tey, S.-K.; Dotti, G.; Fujita, Y.; Kennedy-Nasser, A.; Martinez, C.; Straathof, K.; Liu, E.; Durett, A.G.; Grilley, B.; et al. Inducible Apoptosis as a Safety Switch for Adoptive Cell Therapy. N. Engl. J. Med. 2011, 365, 1673–1683. [Google Scholar] [CrossRef] [Green Version]
- Zhang, R.-Y.; Wei, D.; Liu, Z.-K.; Yong, Y.-L.; Wei, W.; Zhang, Z.-Y.; Lv, J.-J.; Zhang, Z.; Chen, Z.-N.; Bian, H. Doxycycline Inducible Chimeric Antigen Receptor T Cells Targeting CD147 for Hepatocellular Carcinoma Therapy. Front. Cell Dev. Biol. 2019, 7, 233. [Google Scholar] [CrossRef] [Green Version]
- Gu, X.; He, D.; Li, C.; Wang, H.; Yang, G. Development of Inducible CD19-CAR T Cells with a Tet-On System for Controlled Activity and Enhanced Clinical Safety. IJMS 2018, 19, 3455. [Google Scholar] [CrossRef] [Green Version]
- Kulemzin, S.V.; Matvienko, D.A.; Sabirov, A.H.; Sokratyan, A.M.; Chernikova, D.S.; Belovezhets, T.N.; Chikaev, A.N.; Taranin, A.V.; Gorchakov, A.A. Design and Analysis of Stably Integrated Reporters for Inducible Transgene Expression in Human T Cells and CAR NK-Cell Lines. BMC Med. Genom. 2019, 12, 44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chmielewski, M.; Abken, H. TRUCKS, the Fourth-Generation CAR T Cells: Current Developments and Clinical Translation. Adv. Cell Gene Ther. 2020, 3. [Google Scholar] [CrossRef]
- Chilov, D.; Fussenegger, M. Toward Construction of a Self-Sustained Clock-Like Expression System Based on the Mammalian Circadian Clock. Biotechnol. Bioeng. 2004, 87, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Fusil, F.; Calattini, S.; Amirache, F.; Mancip, J.; Costa, C.; Robbins, J.B.; Douam, F.; Lavillette, D.; Law, M.; Defrance, T.; et al. A Lentiviral Vector Allowing Physiologically Regulated Membrane-Anchored and Secreted Antibody Expression Depending on B-Cell Maturation Status. Mol. Ther. 2015, 23, 1734–1747. [Google Scholar] [CrossRef] [PubMed]
- Berens, C.; Suess, B. Riboswitch Engineering—Making the All-Important Second and Third Steps. Curr. Opin. Biotechnol. 2015, 31, 10–15. [Google Scholar] [CrossRef]
- Michel, G.; Yu, Y.; Chang, T.; Yee, J.-K. Site-Specific Gene Insertion Mediated by a Cre-loxP-Carrying Lentiviral Vector. Mol. Ther. 2010, 18, 1814–1821. [Google Scholar] [CrossRef]
- Mangeot, P.E.; Risson, V.; Fusil, F.; Marnef, A.; Laurent, E.; Blin, J.; Mournetas, V.; Massouridès, E.; Sohier, T.J.M.; Corbin, A.; et al. Genome Editing in Primary Cells and In Vivo Using Viral-Derived Nanoblades Loaded with Cas9-sgRNA Ribonucleoproteins. Nat. Commun. 2019, 10, 45. [Google Scholar] [CrossRef] [Green Version]
- Folcher, M.; Oesterle, S.; Zwicky, K.; Thekkottil, T.; Heymoz, J.; Hohmann, M.; Christen, M.; Daoud El-Baba, M.; Buchmann, P.; Fussenegger, M. Mind-Controlled Transgene Expression by a Wireless-Powered Optogenetic Designer Cell Implant. Nat. Commun. 2014, 5, 5392. [Google Scholar] [CrossRef] [Green Version]
- Shao, J.; Xue, S.; Yu, G.; Yu, Y.; Yang, X.; Bai, Y.; Zhu, S.; Yang, L.; Yin, J.; Wang, Y.; et al. Smartphone-Controlled Optogenetically Engineered Cells Enable Semiautomatic Glucose Homeostasis in Diabetic Mice. Sci. Transl. Med. 2017, 9, eaal2298. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Page, A.; Fusil, F.; Cosset, F.-L. Toward Tightly Tuned Gene Expression Following Lentiviral Vector Transduction. Viruses 2020, 12, 1427. https://doi.org/10.3390/v12121427
Page A, Fusil F, Cosset F-L. Toward Tightly Tuned Gene Expression Following Lentiviral Vector Transduction. Viruses. 2020; 12(12):1427. https://doi.org/10.3390/v12121427
Chicago/Turabian StylePage, Audrey, Floriane Fusil, and François-Loïc Cosset. 2020. "Toward Tightly Tuned Gene Expression Following Lentiviral Vector Transduction" Viruses 12, no. 12: 1427. https://doi.org/10.3390/v12121427
APA StylePage, A., Fusil, F., & Cosset, F.-L. (2020). Toward Tightly Tuned Gene Expression Following Lentiviral Vector Transduction. Viruses, 12(12), 1427. https://doi.org/10.3390/v12121427