The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements
Abstract
:1. Introduction
2. Materials and Methods
2.1. Constructs
2.2. Agro-Infiltration
2.3. Protein Extraction and Western Blotting
2.4. Confocal Microscopy
3. Results
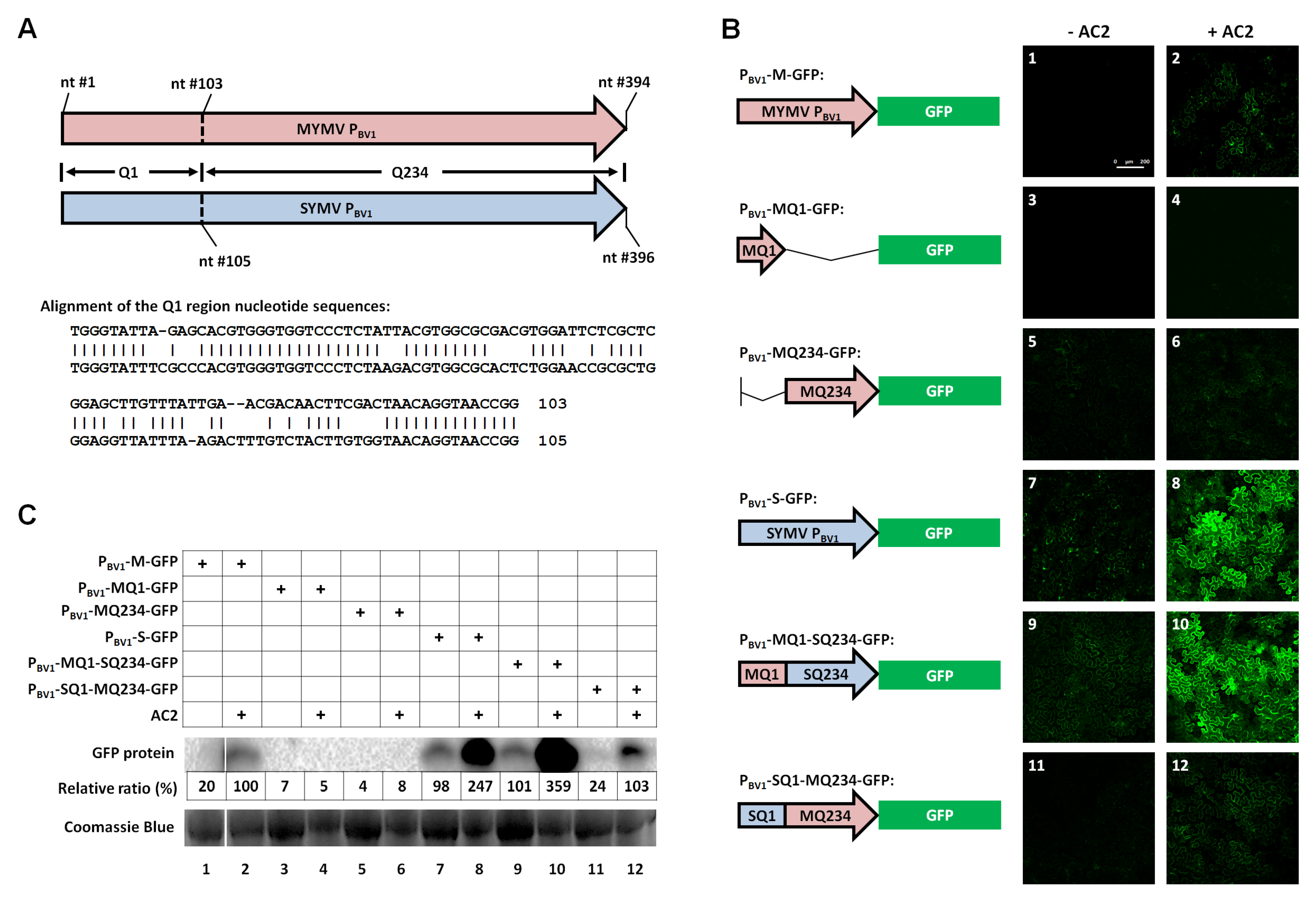
3.1. The First 100 nt of MYMV PBV1 Contains AC2-Responsive DNA Element(s)
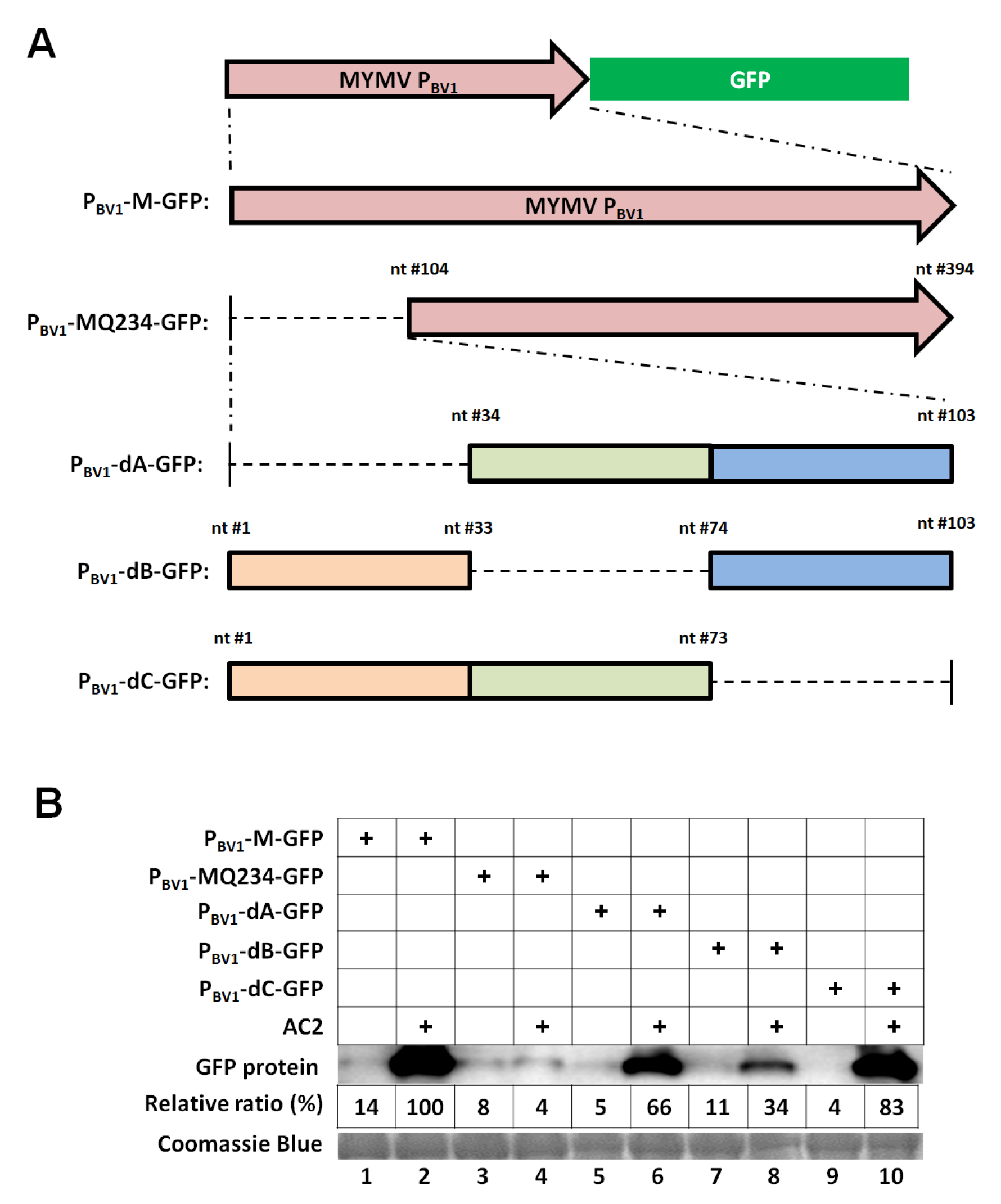
3.2. The First 73 nt of the MYMV PBV1 is Sufficient to Accommodate Specific AC2 Activation
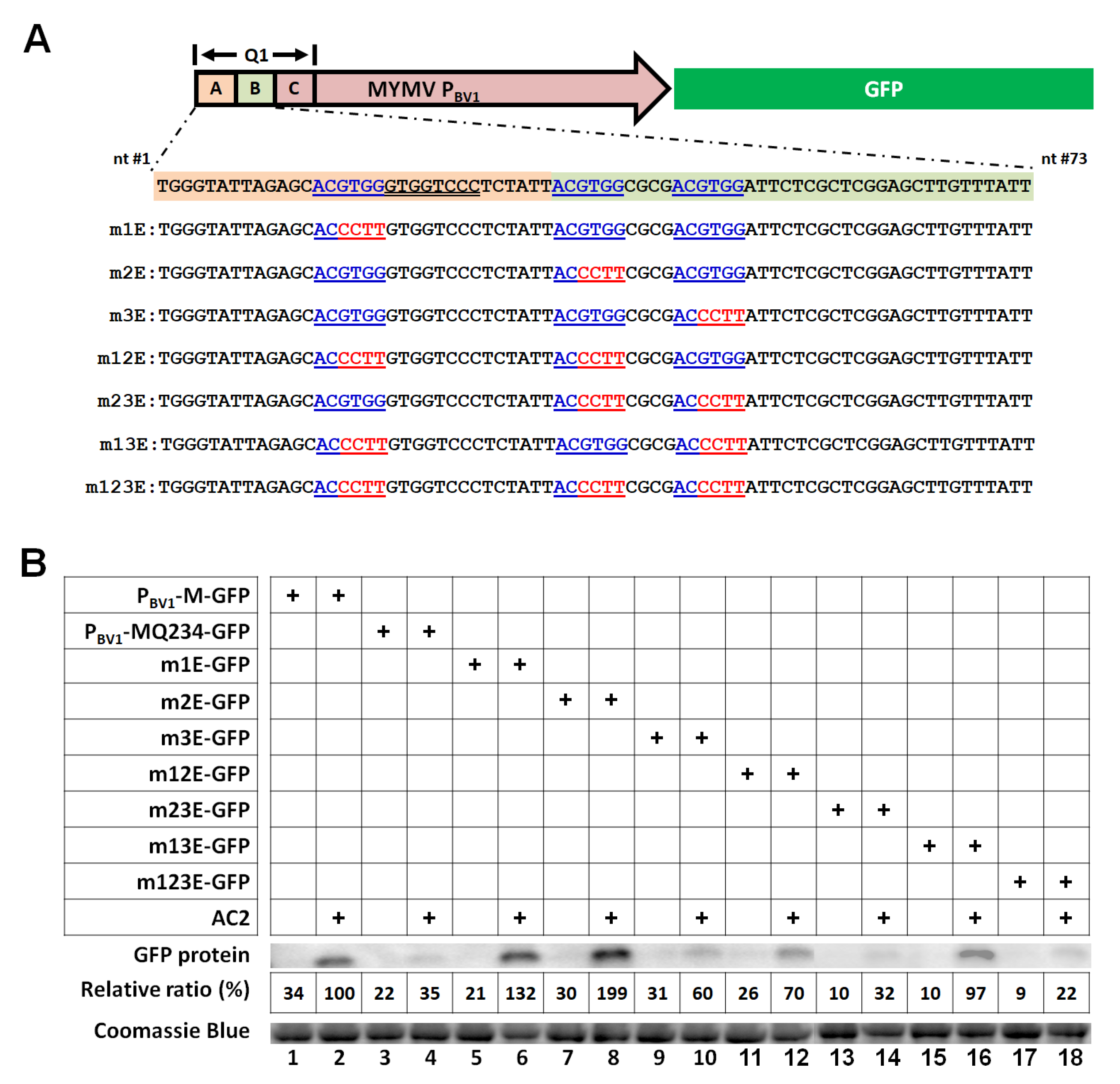
3.3. Three ABREs within the First 73 nt of the MYMV PBV1 Cooperatively Mediate Transcriptional Activation by AC2
3.4. BV1 Promoters of Many Old World Bipartite Begomoviruses Contain ABREs and Other ABA-Responsive Promoter Motifs
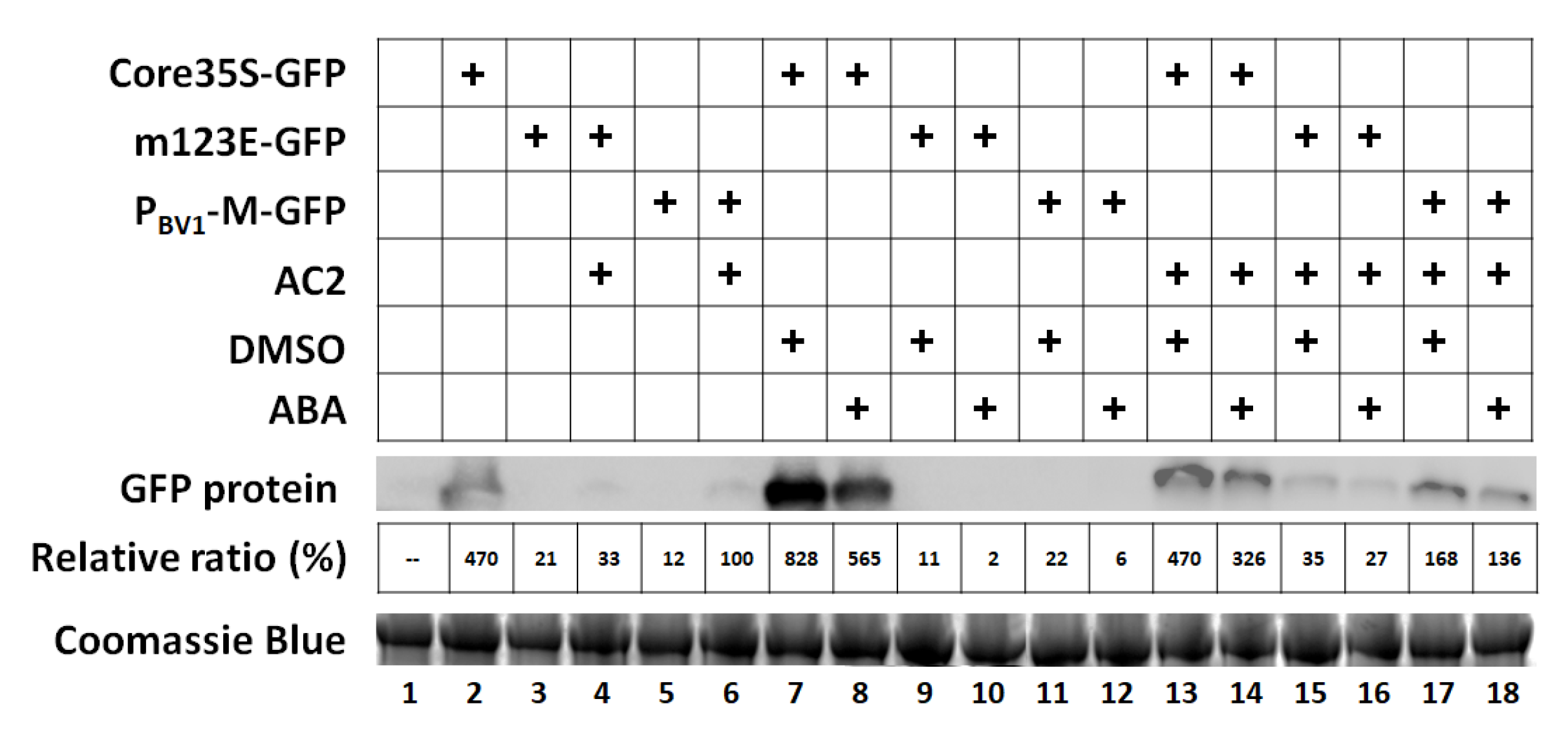
3.5. Exogenous ABA Has No Effect on PBV1 Promoter Activity or AC2-Mediated Transcriptional Activation
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fondong, V. Geminivirus protein structure and function. Mol. Plant Pathol. 2013, 14, 635–649. [Google Scholar] [CrossRef]
- Hanley-Bowdoin, L.; Bejarano, E.R.; Robertson, D.; Mansoor, S. Geminiviruses: Masters at redirecting and reprogramming plant processes. Nat. Rev. Genet. 2013, 11, 777–788. [Google Scholar] [CrossRef]
- Aguilar, E.; Gomez, B.G.; Lozano-Durán, R. Recent advances on the plant manipulation by geminiviruses. Curr. Opin. Plant Biol. 2020, 56, 56–64. [Google Scholar] [CrossRef]
- Guerrero, J.; Regedanz, E.; Lu, L.; Ruan, J.; Bisaro, D.M.; Sunter, G. Manipulation of the Plant Host by the Geminivirus AC2/C2 Protein, a Central Player in the Infection Cycle. Front. Plant Sci. 2020, 11, 591. [Google Scholar] [CrossRef] [PubMed]
- Chikoti, P.C.; Mulenga, R.M.; Tembo, M.; Sseruwagi, P. Cassava mosaic disease: A review of a threat to cassava production in Zambia. J. Plant Pathol. 2019, 101, 467–477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prasad, A.; Sharma, N.; Hari-Gowthem, G.; Muthamilarasan, M.; Prasad, M. Tomato Yellow Leaf Curl Virus: Impact, Challenges, and Management. Trends Plant Sci. 2020, 25, 897–911. [Google Scholar] [CrossRef] [PubMed]
- Varsani, A.; Roumagnac, P.; Fuchs, M.; Navas-Castillo, J.; Moriones, E.; Idris, A.; Briddon, R.W.; Rivera-Bustamante, R.; Zerbini, F.M.; Martin, D.P. Capulavirus and Grablovirus: Two new genera in the family Geminiviridae. Arch. Virol. 2017, 162, 1819–1831. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Briddon, R.W.; Patil, B.L.; Bagewadi, B.; Nawaz-Ul-Rehman, M.S.; Fauquet, C.M. Distinct evolutionary histories of the DNA-A and DNA-B components of bipartite begomoviruses. BMC Evol. Biol. 2010, 10, 97. [Google Scholar] [CrossRef] [Green Version]
- Fontes, E.P.B.; Gladfelter, H.J.; Schaffer, R.L.; Petty, T.D.; Hanley-Bowdoin, L. Geminivirus replication origins have a modular organization. Plant Cell 1994, 6, 405–416. [Google Scholar] [CrossRef] [Green Version]
- Jupin, I.; Héricourt, F.; Benz, B.; Gronenborn, B. DNA replication specificity of TYLCV geminivirus is mediated by the amino-terminal 116 amino acids of the Rep protein. FEBS Lett. 1995, 362, 116–120. [Google Scholar] [CrossRef] [Green Version]
- Fondong, V.N. The Ever-Expanding Role of C4/AC4 in Geminivirus Infection: Punching above Its Weight? Mol. Plant 2019, 12, 145–147. [Google Scholar] [CrossRef] [Green Version]
- Mei, Y.; Zhang, F.; Wang, M.; Li, F.; Wang, Y.; Zhou, X. Divergent Symptoms Caused by Geminivirus-Encoded C4 Proteins Correlate with Their Ability to Bind NbSKη. J. Virol. 2020, 94, 01307-20. [Google Scholar] [CrossRef]
- Haley, A.; Zhan, X.; Richardson, K.; Head, K.; Morris, B. Regulation of the activities of African cassava mosaic virus promoters by the AC1, AC2, and AC3 gene products. Virology 1992, 188, 905–909. [Google Scholar] [CrossRef]
- Sunter, G.; Bisaro, D.M. Transactivation of geminivirus AR1 and BR1 gene expression by the viral AL2 gene product occurs at the level of transcription. Plant Cell 1992, 4, 1321–1331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Noris, E.; Jupin, I.; Accotto, G.P.; Gronenborn, B. DNA-Binding Activity of the C2 Protein of Tomato Yellow Leaf Curl Geminivirus. Virology 1996, 217, 607–612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sung, Y.; Coutts, R. Potato yellow mosaic geminivirus AC2 protein is a sequence non-specific DNA binding protein. FEBS Lett. 1996, 383, 51–54. [Google Scholar] [CrossRef] [Green Version]
- Hartitz, M.D.; Sunter, G.; Bisaro, D.M. The Tomato Golden Mosaic Virus Transactivator (TrAP) is a Single-Stranded DNA and Zinc-Binding Phosphoprotein with an Acidic Activation Domain. Virology 1999, 263, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Lacatus, G.; Sunter, G. The Arabidopsis PEAPOD2 transcription factor interacts with geminivirus AL2 protein and the coat protein promoter. Virology 2009, 392, 196–202. [Google Scholar] [CrossRef] [Green Version]
- Berger, M.R.; Sunter, G. Identification of sequences required for AL2-mediated activation of the tomato golden mosaic virus-yellow vein BR1 promoter. J. Gen. Virol. 2013, 94, 1398–1406. [Google Scholar] [CrossRef] [Green Version]
- Karthikeyan, A.S.; Vanitharani, R.; Balaji, V.; Anuradha, S.; Thillaichidambaram, P.; Shivaprasad, P.V.; Parameswari, C.; Balamani, V.; Saminathan, M.; Veluthambi, K. Analysis of an isolate of Mungbean yellow mosaic virus (MYMV) with a highly variable DNA B component. Arch. Virol. 2004, 149. [Google Scholar] [CrossRef]
- Shivaprasad, P.V.; Akbergenov, R.; Trinks, D.; Rajeswaran, R.; Veluthambi, K.; Hohn, T.; Pooggin, M.M. Promoters, Transcripts, and Regulatory Proteins of Mungbean Yellow Mosaic Geminivirus. J. Virol. 2005, 79, 8149–8163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trinks, D.; Rajeswaran, R.; Shivaprasad, P.V.; Akbergenov, R.; Oakeley, E.J.; Veluthambi, K.; Hohn, T.; Pooggin, M.M. Suppression of RNA Silencing by a Geminivirus Nuclear Protein, AC2, Correlates with Transactivation of Host Genes. J. Virol. 2005, 79, 2517–2527. [Google Scholar] [CrossRef] [Green Version]
- Choi, H.-I.; Hong, J.-H.; Ha, J.-O.; Kang, J.-Y.; Kim, S.Y. ABFs, a Family of ABA-responsive Element Binding Factors. J. Biol. Chem. 2000, 275, 1723–1730. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.-K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Narusaka, Y.; Nakashima, K.; Shinwari, Z.K.; Sakuma, Y.; Furihata, T.; Abe, H.; Narusaka, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J. 2003, 34, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.J.; Casaretto, J.A.; Zhang, P.; Ho, T.-H.D. Functional definition of ABA-response complexes: The promoter units necessary and sufficient for ABA induction of gene expression in barley (Hordeum vulgare L.). Plant Mol. Biol. 2004, 54, 111–124. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Todaka, D.; Mizoi, J.; Yoshida, T.; Kidokoro, S.; Matsukura, S.; Takasaki, H.; Sakurai, T.; Yamamoto, Y.Y.; Yoshiwara, K.; et al. Identification of Cis-Acting Promoter Elements in Cold- and Dehydration-Induced Transcriptional Pathways in Arabidopsis, Rice, and Soybean. DNA Res. 2012, 19, 37–49. [Google Scholar] [CrossRef]
- Zhang, S.; Sun, R.; Guo, Q.; Zhang, X.-F.; Qu, F. Repression of turnip crinkle virus replication by its replication protein p88. Virology 2019, 526, 165–172. [Google Scholar] [CrossRef]
- Sun, R.; Zhang, S.; Zheng, L.; Qu, F. Translation-Independent Roles of RNA Secondary Structures within the Replication Protein Coding Region of Turnip Crinkle Virus. Viruses 2020, 12, 350. [Google Scholar] [CrossRef] [Green Version]
- Qu, F.; Ren, T.; Morris, T.J. The Coat Protein of Turnip Crinkle Virus Suppresses Posttranscriptional Gene Silencing at an Early Initiation Step. J. Virol. 2003, 77, 511–522. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.-F.; Guo, J.; Zhang, X.; Meulia, T.; Paul, P.; Madden, L.V.; Li, Y.-Y.; Qu, F. Random Plant Viral Variants Attain Temporal Advantages During Systemic Infections and in Turn Resist other Variants of the Same Virus. Sci. Rep. 2015, 5, 15346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.-F.; Sun, R.; Guo, Q.; Zhang, S.; Meulia, T.; Halfmann, R.; Li, D.; Qu, F. A self-perpetuating repressive state of a viral replication protein blocks superinfection by the same virus. PLoS Pathog. 2017, 13, e1006253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Girish, K.; Usha, R. Molecular characterization of two soybean-infecting begomoviruses from India and evidence for recombination among legume-infecting begomoviruses from South-East Asia. Virus Res. 2005, 108, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Qu, F.; Morris, T.J. Efficient Infection of Nicotiana benthamiana by Tomato bushy stunt virus Is Facilitated by the Coat Protein and Maintained by p19 Through Suppression of Gene Silencing. Mol. Plant Microbe Interact. 2002, 15, 193–202. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, M.; Ketterling, M.G.; Mccarty, D.R. Quantitative Statistical Analysis of cis-Regulatory Sequences in ABA/VP1- and CBF/DREB1-Regulated Genes of Arabidopsis. Plant Physiol. 2005, 139, 437–447. [Google Scholar] [CrossRef] [Green Version]
- Chung, S.; Parish, R.W. Combinatorial interactions of multiple cis-elements regulating the induction of the Arabidopsis XERO2 dehydrin gene by abscisic acid and cold. Plant J. 2007, 54, 15–29. [Google Scholar] [CrossRef]
- Ezer, D.; Shepherd, S.J.; Brestovitsky, A.; Dickinson, P.; Cortijo, S.; Charoensawan, V.; Box, M.S.; Biswas, S.; Jaeger, K.E.; Wigge, P.A. The G-Box Transcriptional Regulatory Code in Arabidopsis. Plant Physiol. 2017, 175, 628–640. [Google Scholar] [CrossRef] [Green Version]
- Niu, X.; Helentjaris, T.; Bate, N.J. Maize ABI4 Binds Coupling Element1 in Abscisic Acid and Sugar Response Genes. Plant Cell 2002, 14, 2565–2575. [Google Scholar] [CrossRef] [Green Version]
- Timmerhaus, G.; Hanke, S.; Buchta, K.; Rensing, S.A. Prediction and Validation of Promoters Involved in the Abscisic Acid Response in Physcomitrella patens. Mol. Plant 2011, 4, 713–729. [Google Scholar] [CrossRef]
- Tian, R.; Wang, F.; Zheng, Q.; Niza, V.M.A.G.E.; Downie, A.B.; Perry, S.E. Direct and indirect targets of the arabidopsis seed transcription factor ABSCISIC ACID INSENSITIVE3. Plant J. 2020, 103, 1679–1694. [Google Scholar] [CrossRef]
- Argüello-Astorga, G.; Guevara-González, R.; Herrera-Estrella, L.; Rivera-Bustamante, R. Geminivirus Replication Origins Have a Group-Specific Organization of Iterative Elements: A Model for Replication. Virology 1994, 203, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Medrano, R.; Guevara-Gonzalez, R.; Argüello-Astorga, G.; Monsalve-Fonnegra, Z.; Herrera-Estrella, L.R.; Rivera-Bustamante, R.; Rivera-Bustamante, R.F. Identification of a Sequence Element Involved in AC2-Mediated Transactivation of the Pepper Huasteco Virus Coat Protein Gene. Virology 1999, 253, 162–169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eagle, P.A.; Hanley-Bowdoin, L. Cis elements that contribute to geminivirus transcriptional regulation and the efficiency of DNA replication. J. Virol. 1997, 71, 6947–6955. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miozzi, L.; Napoli, C.; Sardo, L.; Accotto, G.P. Transcriptomics of the Interaction between the Monopartite Phloem-Limited Geminivirus Tomato Yellow Leaf Curl Sardinia Virus and Solanum lycopersicum Highlights a Role for Plant Hormones, Autophagy and Plant Immune System Fine Tuning during Infection. PLoS ONE 2014, 9, e89951. [Google Scholar] [CrossRef] [Green Version]
- Alazem, M.; He, M.-H.; Moffett, P.; Lin, N.-S. Abscisic Acid Induces Resistance against Bamboo Mosaic Virus through Argonaute2 and 3. Plant Physiol. 2017, 174, 339–355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Virus_DNA-B Accession | DNA-B nt #26-175 (150 nt), Conserved Promoter Elements Denoted |
|---|---|
| MYMV-Vigna [KA22] _AJ132574.1 | TGGGTATTAGAGCACGTGGGTGGTCCCTCTATTACGTGGCGCGACGTGGATTCTCGCTCGGAGCTTGTTTATTGAACGACAACTTCGACTAACAGGTAACCGGTTTGGTTACCTTTCGTACATGGACAAATTTGTCTTTTCCTCAAAAAG |
| MYMV-Vigna [KA34] _AJ439057.1 | TGGGTATTAGAGCACGTGGGTGGTCCCTCTATTACGTGGCGCGACGTGGATTCTCGCTCGGAGGTTGTTTATTGAACGACAACTTGGACTAACAGGTAACCGGTTTGGTTACCTTTCGTACATGGACAAATTTGTCTTTTCCTCAAAAAG |
| SYMV_AJ582267.1 | TGGGTATTTCGCCCACGTGGGTGGTCCCTCTAAGACGTGGCGCACTCTGGAACCGCGCTGGGAGGTTATTTAAGACTTTGTCTACTTGTGGTAACAGGTAACCGGCTAAGTGACCGGTTGGAAAGCGTGCCTTTCCCACCCCTGGCAAAT |
| MYMIV_AY271894.1 | TCGGTTTTAGAGCACGTGGGTGGTCCCTCTAGTACGTGGCGCGCTCTGGAGTCTCGCTCGAAGCTTGTTTATCGAACGACTACTTAGAGTTACAGGTAACCGGATAGGTGACCGATCGTACATGGACAAATTTGTCTTTTCCTCAAAAAG |
| HYMV_AJ627905.1 | GACCTGGATCACTCACGTGGGTGGTCCCGCTCATCGTGGCGCAACGAGGAGTCTCGTTGGGAGTTTTTATAATGGGTTACTACTTGGGAGAAGTGGGGAAACCGGAAGCTACCGCCGCAAATCGAGAAATTTGCTTTTAATGCGAAATTA |
| SLCMV_AJ579308.1 | GTGGTGGCCCCCCCCACGTGGGGATGTCCCCCTCTCACAACGCTCACTAGAAGGTTCAACATGTTGGTGGCCCCACGATGTTGTTTATAACGTCTATAACGTTTGAGACTCGAAGCTTGTGATGCCACGTATGCGTTATTAGTACTTCGT |
| ICMV_AJ314740.1 | GTGGTGGACCCCTCCCCCACGTGGAGATGTCCCCCACTCAGAACGCTCACTAGGAGGTTCAACATGTTGGTGGCCCCACGATGTTGTTTATAACGTCTATAACGTCTGAGACTCCAAGCTTGTGATGCCACGTATGTGTTATTTGTACTT |
| ClGMV_DQ641693.1 | TATAGTATGTGTCATGTGTGGGTCCCATGTCACGTGATTTTGTACCTTTCTGTCCCCCAGCTAGCCGACAAAGTGGGACCCACTCACGTGATTAGCACTTGCCTTGTACCTTCCTTCTGTGTACGTTCTGGTCAAAAAGACTAGTATACC |
| KuMV_DQ641691.1 | TGCGGTGTGGTCCCCCCGCCCATGTGCTTTCAATCTCGTCCGCACGTTTTTTTTAGTCTTCGCGCGTGGTGTGAATGCGCCGTAATGCGCTTGCCGCAAATAACGTTATAACTGCTTTAATTTGAATTTCTTTAAATTATGTGGAATACC |
| SLCCV_AM260207.1 | CTCTAAATCTGGCCGTCTATTTTCTATAAATGCGCATCAAACGAGTGCGATGAACAACACGTGTCCTATGTGTCATCCTTTACTAAAGGAAAGGATTGTACGATATCGAAATTTGCACATGGTGGACCCCAGTGACTAGATATGCAATAT |
| SLCPHV_AB085794.1 | CTTCAAATCTGTCCGTCTATTTTGTACAAAAGCTTATCAAGGTGCTGCGATGCACAACACGTGTCCTCTATTACATCCTTTACTAAAGTAAAGGTTTGTACGATTTCCAAGTGCCGCACATGGTGGTCCCCAGTGACTAGATATGCAATA |
| LYMV_AF509740.1 | TCTCCCCACAGTCGTTGATTTGTTCTAAAGGCGCGCACGATTGCATGTTTTCGCAACACGTGTCCTCTTATGATTACTTTATGAAAATAAAGTTAGCTGTGTTCTGAAATTTCGCACATGGTGGTCCCCCAGATACTAGATAGAAATATC |
| ToLCNDV_AJ875158.1 | CCCTATCCTGACCGTTGCTGTGTAATCATTGCACCAAGTTAGTCATCCGATTTGCAACACGTGTATCCCACTAACAGACTTTATGCAAATAAATGTCCGATATCTGTGTCGACAATGCTTGTGTGTTCCCCTTATATCTTGTCGTACCCC |
| ToLCuGuV_AY190291.1 | CCTTATCTTGACCGTTGCTGCGTAATCATTGCACCAAGTTACTCATCCGATTTGCAACACGTGTATCCCACTAGCAGACTTTATGCAATTAAATGTCTGATATCTGTGTGTACAATGCATATGTGTTCCCCTTATATCTTGTCGTACCCC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, R.; Han, J.; Zheng, L.; Qu, F. The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements. Viruses 2020, 12, 1403. https://doi.org/10.3390/v12121403
Sun R, Han J, Zheng L, Qu F. The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements. Viruses. 2020; 12(12):1403. https://doi.org/10.3390/v12121403
Chicago/Turabian StyleSun, Rong, Junping Han, Limin Zheng, and Feng Qu. 2020. "The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements" Viruses 12, no. 12: 1403. https://doi.org/10.3390/v12121403
APA StyleSun, R., Han, J., Zheng, L., & Qu, F. (2020). The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements. Viruses, 12(12), 1403. https://doi.org/10.3390/v12121403







