Diversity of tRNA Clusters in the Chloroviruses
Abstract
1. Introduction
2. Materials and Methods
2.1. Genomic Sequence Data
2.2. Identification and Localization of the tRNA Genes
2.3. CUB in Chloroviruses and Their Host
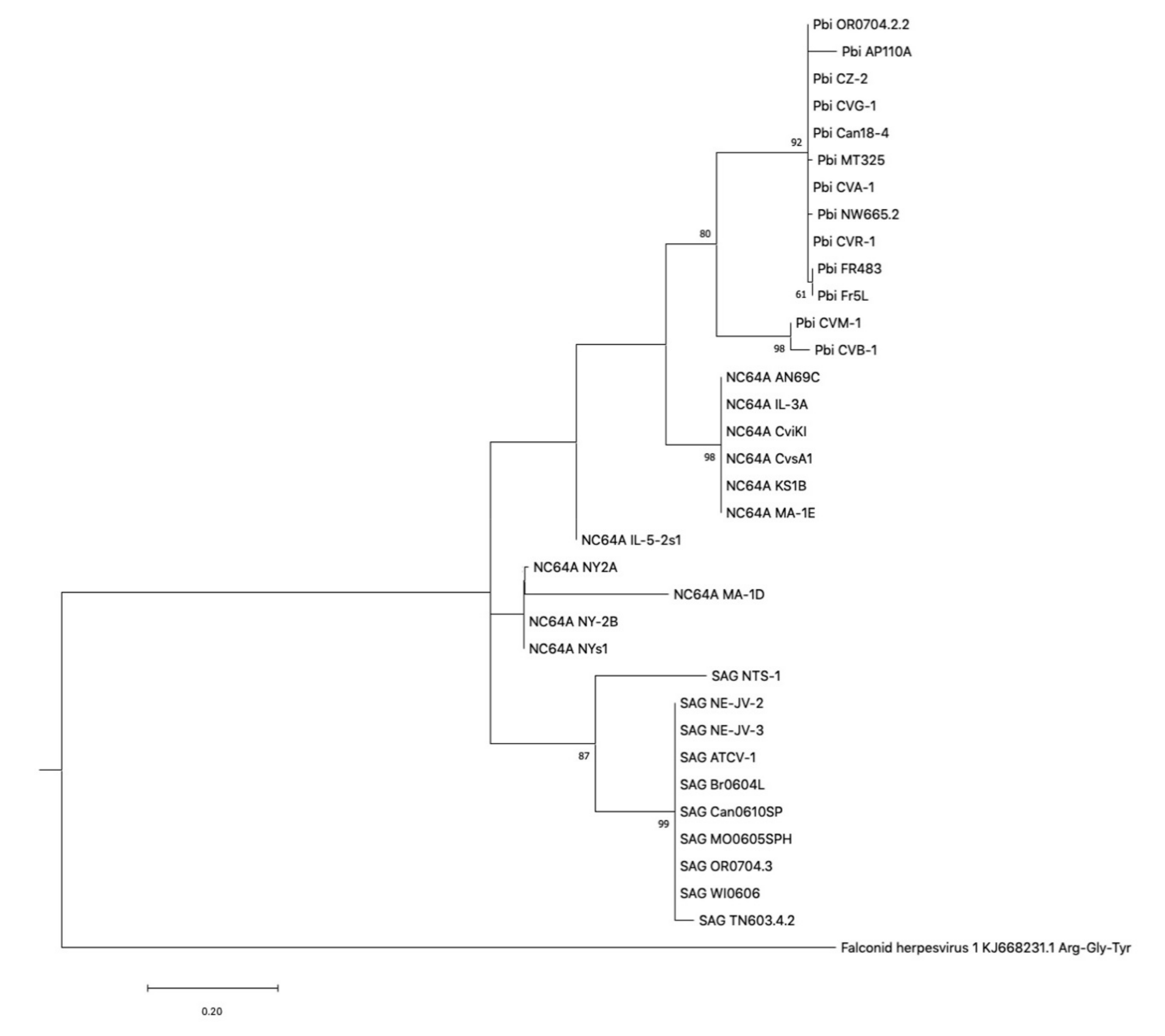
2.4. Phylogenetic Tree Construction of tRNA Genes
3. Results and Discussion
3.1. Chlorovirus tRNA Gene Clusters
3.2. Genome Location of the Chlorovirus tRNA Clusters
3.3. Potential tRNA Transcription Promoters
3.4. Presence of CUB Differences in Host and Viruses
3.5. Not All Chlorovirus tRNAs Assist in Overcoming CUB
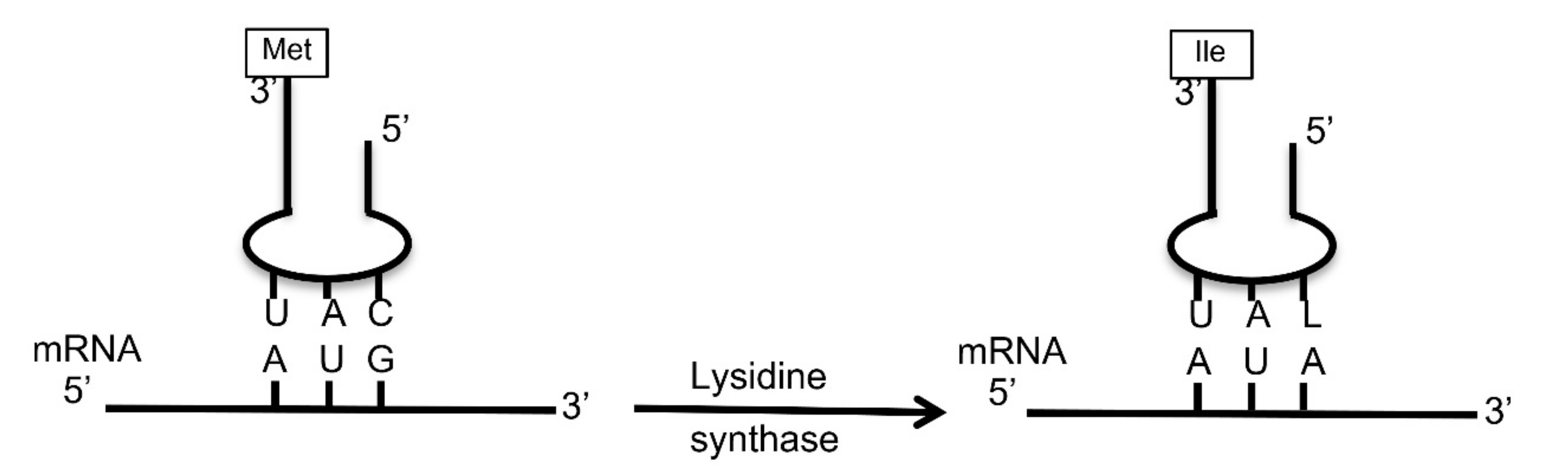
3.6. Chloroviruses Encode a Putative tRNA Isoleucine Lysidine Synthase (TilS)
3.7. Acquisition of Chlorovirus tRNA Clusters
3.8. Generation of Chlorovirus tRNAs
3.9. tRNA Gene Clusters in other Phycodnavirues
3.10. tRNA Genes and Gene Clusters in other Large DNA Viruses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Morgado, S.M.; Vicente, A.C.P. Global in-silico scenario of tRNA genes and their organization in virus genomes. Viruses 2019, 11, 180. [Google Scholar] [CrossRef] [PubMed]
- Abe, T.; Inokuchi, H.; Yamada, Y.; Muto, A.; Iwasaki, Y.; Ikemura, T. tRNADB-CE: tRNA gene database well-timed in the era of big sequence data. Front. Genet. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Michely, S.; Toulza, E.; Subirana, L.; John, U.; Cognat, V.; Maréchal-Drouard, L.; Grimsley, N.; Moreau, H.; Piganeau, G. Evolution of codon usage in the smallest photosynthetic eukaryotes and their giant viruses. Genome Biol. Evol. 2013, 5, 848–859. [Google Scholar] [CrossRef] [PubMed]
- Pagarete, A.; Lanzén, A.; Puntervoll, P.; Sandaa, R.A.; Larsen, A.; Larsen, J.B.; Allen, M.J.; Bratbak, G. Genomic sequence and analysis of EhV-99B1, a new coccolithovirus from the Norwegian fjords. Intervirology 2013, 56, 60–66. [Google Scholar] [CrossRef] [PubMed]
- Santini, S.; Jeudy, S.; Bartoli, J.; Poirot, O.; Lescot, M.; Abergel, C.; Barbe, V.; Wommack, K.E.; Noordeloos, A.A.M.; Brussaard, C.P.D.; et al. Genome of Phaeocystis globosa virus PgV-16T highlights the common ancestry of the largest known DNA viruses infecting eukaryotes. Proc. Natl. Acad. Sci. USA 2013, 110, 10800–10805. [Google Scholar] [CrossRef] [PubMed]
- Derelle, E.; Monier, A.; Cooke, R.; Worden, A.Z.; Grimsley, N.H.; Moreau, H. Diversity of viruses infecting the green microalga Ostreococcus lucimarinus. J. Virol. 2015, 89, 5812–5821. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lu, Z.; Sun, L.; Ropp, S.; Kutish, G.F.; Rock, D.L.; Van Etten, J.L. Analysis of 74 kb of DNA located at the right end of the 330-kb chlorella virus PBCV-1 genome. Virology 1997, 237, 360–377. [Google Scholar] [CrossRef]
- Nishida, K.; Kawasaki, T.; Fujie, M.; Usami, S.; Yamada, T. Aminoacylation of tRNAs encoded by chlorella virus CVK2. Virology 1999, 263, 220–229. [Google Scholar] [CrossRef]
- Lee, D.Y.; Graves, M.V.; Van Etten, J.L.; Choi, T.J. Functional implication of the tRNA genes encoded in the chlorella virus PBCV-1 genome. Plant. Pathol. J. 2005, 21, 334–342. [Google Scholar] [CrossRef]
- Fitzgerald, L.A.; Graves, M.V.; Li, X.; Feldblyum, T.; Nierman, W.C.; Van Etten, J.L. Sequence and annotation of the 369-kb NY-2A and the 345-kb AR158 viruses that infect Chlorella NC64A. Virology 2007, 358, 472–484. [Google Scholar] [CrossRef]
- Fitzgerald, L.A.; Graves, M.V.; Li, X.; Hartigan, J.; Pfitzner, A.J.; Hoffart, E.; Van Etten, J.L. Sequence and annotation of the 288-kb ATCV-1 virus that infects an endosymbiotic chlorella strain of the heliozoon Acanthocystis turfacea. Virology 2007, 362, 350–361. [Google Scholar] [CrossRef]
- Fitzgerald, L.A.; Graves, M.V.; Li, X.; Feldblyum, T.; Hartigan, J.; Van Etten, J.L. Sequence and annotation of the 314-kb MT325 and the 321-kb FR483 viruses that infect Chlorella Pbi. Virology 2007, 358, 459–471. [Google Scholar] [CrossRef] [PubMed]
- Jeanniard, A.; Dunigan, D.D.; Gurnon, J.R.; Agarkova, I.V.; Kang, M.; Vitek, J.; Duncan, G.; McClung, O.W.; Larsen, M.; Claverie, J.-M.; et al. Towards defining the chloroviruses: A genomic journey through a genus of large DNA viruses. BMC Genom. 2013, 14, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Van Etten, J.L.; Agarkova, I.V.; Dunigan, D.D. Chloroviruses. Viruses 2020, 12, 20. [Google Scholar] [CrossRef]
- Blanc, G.; Duncan, G.; Agarkova, I.; Borodovsky, M.; Gurnon, J.; Kuo, A.; Lindquist, E.; Lucas, S.; Pangilinan, J.; Polle, J.; et al. The Chlorella variabilis NC64A genome reveals adaptation to photosymbiosis, coevolution with viruses, and cryptic sex. Plant. Cell 2010, 22, 2943–2955. [Google Scholar] [CrossRef] [PubMed]
- Arriola, M.B.; Velmurugan, N.; Zhang, Y.; Plunkett, M.H.; Hondzo, H.; Barney, B.M. Genome sequences of Chlorella sorokiniana UTEX 1602 and Micractinium conductrix SAG 241.80: Implications to maltose excretion by a green alga. Plant J. 2018, 93, 566–586. [Google Scholar] [CrossRef] [PubMed]
- Bermudez-Santana, C.; Attolini, C.S.O.; Kirsten, T.; Engelhardt, J.; Prohaska, S.J.; Steigele, S.; Stadler, P.F. Genomic organization of eukaryotic tRNAs. BMC Genom. 2010, 11, 270. [Google Scholar] [CrossRef]
- Morgado, S.M.; Vicente, A.C.P. Beyond the limits: tRNA array units in Mycobacterium genomes. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Morgado, S.M.; Vicente, A.C.P. Exploring tRNA gene cluster in archaea. Mem. Inst. 2019, 114. [Google Scholar] [CrossRef]
- Fan, W.; Guo, W.; Van Etten, J.L.; Mower, J.P. Multiple origins of endosymbionts in Chlorellaceae with no reductive effects on the plastid or mitochondrial genomes. Sci. Rep. 2017, 7, 10101. [Google Scholar] [CrossRef]
- Friedrich, A.; Jung, P.P.; Hou, J.; Neuvéglise, C.; Schacherer, J. Comparative mitochondrial genomics within and among yeast species of the Lachancea genus. PLoS ONE 2012, 7, e47834. [Google Scholar] [CrossRef] [PubMed]
- Dunigan, D.D.; Cerny, R.L.; Bauman, A.T.; Roach, J.C.; Lane, L.C.; Agarkova, I.V.; Wulser, K.; Yanai-Balser, G.M.; Gurnon, J.R.; Vitek, J.C.; et al. Paramecium bursaria Chlorella virus 1 proteome reveals novel architectural and regulatory features of a giant virus. J. Virol. 2012, 86, 8821–8834. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Lowe, T.M.; Chan, P.P. tRNAscan-SE On-line: Integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 2016, 44, W54–W57. [Google Scholar] [CrossRef]
- Stecher, G.; Tamura, K.; Kumar, S. Molecular evolutionary genetics analysis (MEGA) for macOS. Mol. Biol. Evol. 2020, 37, 1237–1239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Calin-Jageman, I.; Gurnon, J.R.; Choi, T.J.; Adams, B.; Nicholson, A.W.; Van Etten, J.L. Characterization of a chlorella virus PBCV-1 encoded ribonuclease III. Virology 2003, 317, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Rak, R.; Dahan, O.; Pilpel, Y. Repertoires of tRNAs: The couplers of genomics and proteomics. Annu. Rev. Cell Dev. Biol. 2018, 34, 239–264. [Google Scholar] [CrossRef]
- Diebel, K.W.; Claypool, D.J.; van Dyk, L.F. A conserved RNA polymerase III promoter required for gammaherpesvirus TMER transcription and microRNA processing. Gene 2014, 544, 8–18. [Google Scholar] [CrossRef]
- Arimbasseri, A.G.; Maraia, R.J. RNA polymerase III advances: Structural and tRNA functional views. Trends Biochem. Sci. 2016, 41, 546–559. [Google Scholar] [CrossRef]
- Tatosyan, K.A.; Stasenko, D.V.; Koval, A.P.; Gogolevskaya, I.K.; Kramerov, D.A. TATA-like boxes in RNA polymerase III promoters: Requirements for nucleotide sequences. Int. J. Mol. Sci. 2020, 21, 3706. [Google Scholar] [CrossRef]
- Crick, F.H.C. The origin of the genetic code. J. Mol. Biol. 1968, 38, 367–379. [Google Scholar] [CrossRef]
- Nakanishi, K.; Fukai, S.; Ikeuchi, Y.; Soma, A.; Sekine, Y.; Suzuki, T.; Nureki, O. Structural basis for lysidine formation by ATP pyrophosphatase accompanied by a lysine-specific loop and a tRNA-recognition domain. Proc. Natl. Acad. Sci. USA 2005, 102, 7487–7492. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Miyauchi, K. Discovery and characterization of tRNAIle lysidine synthetase (TilS). FEBS Lett. 2010, 584, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Blanc, G.; Mozar, M.; Agarkova, I.V.; Gurnon, J.R.; Yanai-Balser, G.M.; Rowe, J.M.; Xia, Y.; Riethoven, J.-J.; Dunigan, D.D.; Van Etten, J.L. Deep RNA sequencing reveals hidden features and dynamics of early gene transcription in Paramecium bursaria chlorella virus 1. PLoS ONE 2014, 9, e90989. [Google Scholar] [CrossRef] [PubMed]
- Belfield, G.P.; Tuite, M.F. Translation elongation factor 3: A fungus-specific translation factor? Mol. Microbiol. 1993, 9, 411–418. [Google Scholar] [CrossRef] [PubMed]
- Belfield, G.P.; Ross-Smith, N.J.; Tuite, M.F. Translation elongation factor-3 (EF-3): An evolving eukaryotic ribosomal protein? J. Mol. Evol. 1995, 41, 376–387. [Google Scholar] [CrossRef]
- Mateyak, M.K.; Pupek, J.K.; Garino, A.E.; Knapp, M.C.; Colmer, S.F.; Kinzy, T.G.; Dunaway, S. Demonstration of translation elongation factor 3 activity from a non-fungal species, Phytophthora infestans. PLoS ONE 2018, 13, e0190524. [Google Scholar] [CrossRef]
- Velandia-Huerto, C.A.; Berkemer, S.J.; Hoffmann, A.; Retzlaff, N.; Romero Marroquín, L.C.; Hernández-Rosales, M.; Stadler, P.F.; Bermúdez-Santana, C.I. Orthologs, turn-over, and remolding of tRNAs in primates and fruit flies. BMC Genom. 2016, 17, 617. [Google Scholar] [CrossRef]
- Pope, W.H.; Anders, K.R.; Baird, M.; Bowman, C.A.; Boyle, M.M.; Broussard, G.W.; Chow, T.; Clase, K.L.; Cooper, S.; Cornely, K.A.; et al. Cluster M mycobacteriophages Bongo, PegLeg, and Rey with unusually large repertoires of tRNA isotypes. J. Virol. 2014, 88, 2461–2480. [Google Scholar] [CrossRef][Green Version]
- Koonin, E.V.; Yutin, N. Multiple evolutionary origins of giant viruses. F1000Research 2018, 7, 1840. [Google Scholar] [CrossRef]
- Koonin, E.V.; Yutin, N. Evolution of the large nucleocytoplasmic DNA viruses of eukaryotes and convergent origins of viral gigantism. Adv. Virus Res. 2019, 103, 167–202. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Aravind, L.; Koonin, E.V. Common origin of four diverse families of large eukaryotic DNA viruses. J. Virol. 2001, 75, 11720–11734. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Balaji, S.; Koonin, E.V.; Aravind, L. Evolutionary genomics of nucleo-cytoplasmic large DNA viruses. Virus Res. 2006, 117, 156–184. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Yutin, N. Origin and evolution of eukaryotic large nucleo-cytoplasmic DNA viruses. Intervirology 2010, 53, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Lyons, S.M.; Fay, M.M.; Ivanov, P. The role of RNA modifications in the regulation of tRNA cleavage. FEBS Lett. 2018, 592, 2828–2844. [Google Scholar] [CrossRef] [PubMed]
| Leu-1 UUG | Ile AUA | Asn-1 AAC | Leu-2 UUA | Arg-1 AGA | Asn-2 AAC | Gly GGA | Asn-3 AAC | Lys-1 AAG | Gln CAG | Lys-2 AAG | Tyr UAC a | Lys-3 AAA | Lys-4 AAG | Arg-2 AGA | Asp GAC | Val GUU | Total tRNAs | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MA-1E | 25 | 23 | 24 | 3 | 23 | 3 | 27 | 3 | 22 | 23 | 23 | 30 | 1 | 14 | ||||||||||||||||||||
| CvsA1 | 25 | 23 | 24 | 3 | 23 | 3 | 27 | 3 | 22 | 23 | 23 | 30 | 1 | 14 | ||||||||||||||||||||
| CviK1 | 25 | 23 | 24 | 3 | 23 | 3 | 27 | 3 | 22 | 23 | 23 | 30 | 1 | 14 | ||||||||||||||||||||
| KS1B | 3 | 23 | 109 | 3 | 23 | 3 | 27 | 3 | 22 | 23 | 23 | 30 | 1 | 12 | ||||||||||||||||||||
| PBCV-1 | 25 | 23 | 23 | 3 | b | 24 | 3 | 24 | 22 | 23 | 33 | 11 | ||||||||||||||||||||||
| IL-3A | 25 | 23 | 24 | 3 | 23 | 3 | 25 | c | 3 | 84 | 22 | 23 | 10 | |||||||||||||||||||||
| MA-1D | 25 | 23 | 24 | 5 | 15 | 3 | 25 | 3 | 22 | 23 | 33 | 12 | ||||||||||||||||||||||
| NE-JV-4 | 25 | 23 | 24 | 3 | 23 | 3 | 72 | c | 3 | 23 | 33 | 11 | ||||||||||||||||||||||
| AN69C | 25 | 23 | 24 | 3 | 23 | 3 | 12 | 22 | 23 | 10 | ||||||||||||||||||||||||
| NY-2B | 25 | 23 | 23 | 3 | 23 | 22 | 2 | 25 | 8 | |||||||||||||||||||||||||
| IL-5-2s1 | 25 | 23 | 23 | 3 | 23 | 22 | 2 | 25 | 8 | |||||||||||||||||||||||||
| NY-2A | 142 | 23 | 3 | 24 | 22 | 2 | 25 | 8 | ||||||||||||||||||||||||||
| NYs-1 | 25 | 25 | 229 | 3 | 1058 | 2 | 25 | 8 | ||||||||||||||||||||||||||
| AR158 | 25 | 25 | 51 | 23 | 3 | 24 | 7 | |||||||||||||||||||||||||||
| SAG Viruses | Codons and Cognate Amino Acids Recognized by SAG Virus tRNAs | |||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ile-1 AUA | Ser AGU | Arg AGA | Asn-1 AAC | Gly GGA | Ile-2 AUU | Asn-2 AAC | Met AUG | Asp-1 GAC | Val-1 GUU | Val-2 GUU | Asn-3 AAC | Tyr UAC a | Lys AAG | Asn-4 AAC | Asp-2 GAC | Leu-1 UUA | Asn-5 AAC | Leu-2 UUG | Thr ACU b | Total tRNAs | ||||||||||||||||||||
| Can0610SP | 4 | 25 | 22 | 22 | 25 | 23 | 22 | 22 | 2 | 21 | 4 | 36k | 13 | |||||||||||||||||||||||||||
| OR0704.3 | 4 | 25 | 22 | 22 | 25 | 23 | 22 | 22 | 2 | 21 | 148 | 32k | 13 | |||||||||||||||||||||||||||
| NE-JV-2 | 22 | 4 | 25 | 22 | 22 | 22 | 23 | 22 | 2 | 21 | 148 | 29k | 13 | |||||||||||||||||||||||||||
| NE-JV-3 | 4 | 25 | 22 | 22 | 22 | 23 | 22 | 2 | 21 | 148 | 31k | 12 | ||||||||||||||||||||||||||||
| ATCV-1 | 5 | 25 | 22 | 23 | 22 | 22 | 22 | 2 | 22 | 148 | c | 31k | 11 | |||||||||||||||||||||||||||
| WI0606 | 4 | 25 | 22 | 23 | 22 | 22 | 2 | 22 | 148 | 31k | 11 | |||||||||||||||||||||||||||||
| MO0605SPH | 4 | 25 | 22 | 23 | 22 | 22 | 2 | 22 | 148 | 30k | 11 | |||||||||||||||||||||||||||||
| GM0701.1 | 25 | 1 | 22 | 24 | 23 | 148 | 77 | 4 | 35k | 10 | ||||||||||||||||||||||||||||||
| Br0604L | 4 | 25 | 22 | 25 | 23 | 2 | 242 | 32k | 9 | |||||||||||||||||||||||||||||||
| TN603.4.2 | 25 | 22 | 24 | 23 | 2 | 22 | 227 | 32k | 9 | |||||||||||||||||||||||||||||||
| Canal-1 | 158 | 5 | 25 | 22 | 65 | 23 | 24 | 54 | 29k | 9 | ||||||||||||||||||||||||||||||
| MN0810.1 | 23 | 24 | 1 | 22 | 22 | 22 | 4 | 35k | 9 | |||||||||||||||||||||||||||||||
| NTS-1 | 23 | 25 | 22 | 21 | 23 | 33k | 7 | |||||||||||||||||||||||||||||||||
| Pbi Viruses | Codons and Cognate Amino Acids Recognized by Pbi Virus tRNAs | |||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ile AUA | Leu UUA | Phe UUC | Arg AGA | Gly GGA | Asn-1 AAC | Tyr-1 UAC a | Lys-1 AAG | Asn-2 AAC | Asn-3 AAC | Asn-4 AAC | Tyr-2 UAC a | Lys-2 AAG | Thr-1 ACG | Thr-2 ACG | Total tRNAs | |||||||||||||||
| Fr5L | 23 | 3 | 22 | 22 | 2 | 25 | 22 | 22 | 2 | 245 | 11 | |||||||||||||||||||
| CZ-2 | 23 | 3 | 22 | 23 | 23 | 22 | 22 | 2 | 243 | 10 | ||||||||||||||||||||
| MT325 | 24 | 24 | 23 | 3 | 23 | 22 | 22 | 2 | 161 | 10 | ||||||||||||||||||||
| Can18-4 | 24 | 24 | 23 | 3 | 23 | 22 | 22 | 2 | 1041 | 10 | ||||||||||||||||||||
| CVB-1 | 24 | 24 | 26 | 985 | 24 | 23 | 22 | 2 | 161 | 10 | ||||||||||||||||||||
| FR483 | 22 | 24 | 3 | 23 | 23 | 22 | 2 | 161 | 9 | |||||||||||||||||||||
| CVG-1 | 132 | 23 | 3 | 23 | 23 | 22 | 2 | 161 | 9 | |||||||||||||||||||||
| CVR-1 | 132 | 23 | 3 | 23 | 23 | 22 | 2 | 159 | 9 | |||||||||||||||||||||
| CVA-1 | 132 | 23 | 3 | 23 | 23 | 22 | 2 | 159 | 9 | |||||||||||||||||||||
| AP110A | 132 | 23 | 3 | 23 | 23 | 23 | 2 | 159 | 9 | |||||||||||||||||||||
| CVM-1 | 132 | 23 | 3 | 24 | 96 | 22 | 2 | 161 | 9 | |||||||||||||||||||||
| NW665.2 | 1142 | 3 | 23 | 23 | 22 | 2 | 161 | 8 | ||||||||||||||||||||||
| OR0704.2.2 | 3 | 23 | 22 | 22 | 2 | 159 | 7 | |||||||||||||||||||||||
| NE-JV-1 | 132 | 1416 | 3 | |||||||||||||||||||||||||||
| tRNA | Codon | NC64A | SAG | Pbi |
|---|---|---|---|---|
| Ile-1 | AUA | c | c | c |
| Leu-1 | UUA | c | c | c |
| Asn-1 | AAC | c | c | c |
| Gly-1 | GGA | c | c | c |
| Lys-1 | AAG | c | c | c |
| Tyr-1 | UAC | c | c | c |
| Arg-1 | AGA | c | c | c |
| Asp-1 | GAC | d | d | |
| Val-1 | GUU | d | d | |
| Leu-2 | UUG | d | d | |
| Gln-1 | CAG | u | ||
| Lys-2 | AAA | u | ||
| Ser-1 | AGU | u | ||
| Ile-2 | AUU | u | ||
| Met-1 | AUG | u | ||
| Thr-1 | ACU | u | ||
| Phe-1 | UUC | u | ||
| Thr-1 | ACG | u | ||
| Thr-2 | ACG | u |
| Codon AA | PBCV-1 | AN69C | C. variabilis | Ratio 1 | Codon AA | PBCV-1 | AN69C | C. variabilis | Ratio 1 |
|---|---|---|---|---|---|---|---|---|---|
| GCA A | 1.92 | 1.15 | 2.42 | 0.63 | AAC N | 2.62 | 2.28 | 1.48 | 1.66 |
| GCC A | 0.86 | 0.64 | 5.54 | 0.14 | AAU N | 3.16 | 2.78 | 0.30 | 9.9 |
| GCG A | 1.29 | 0.84 | 5.46 | 0.20 | CCA P | 1.42 | 1.24 | 1.08 | 1.23 |
| GCU A | 1.30 | 0.82 | 1.72 | 0.62 | CCC P | 1.07 | 0.91 | 2.55 | 0.39 |
| UGC C | 0.67 | 1.08 | 1.77 | 0.49 | CCG P | 0.96 | 0.97 | 2.30 | 0.42 |
| UGU C | 1.23 | 1.88 | 0.32 | 4.86 | CCU P | 1.33 | 0.89 | 0.96 | 1.16 |
| GAC D | 1.97 | 1.31 | 3.04 | 0.54 | CAA Q | 1.84 | 2.09 | 0.59 | 3.33 |
| GAU D | 3.02 | 1.91 | 1.06 | 4.56 | CAG Q | 0.87 | 1.06 | 4.93 | 0.2 |
| GAA E | 3.69 | 2.47 | 0.54 | 5.70 | AGA R | 1.43 | 1.71 | 0.25 | 6.28 |
| GAG E | 1.26 | 1.08 | 4.92 | 0.24 | AGG R | 0.68 | 0.84 | 0.92 | 0.83 |
| UUC F | 2.53 | 2.40 | 1.55 | 1.59 | CGA R | 0.66 | 1.48 | 0.44 | 2.43 |
| UUU F | 2.94 | 3.49 | 0.95 | 3.38 | CGC R | 0.66 | 0.84 | 3.22 | 0.23 |
| GGA G | 1.71 | 1.28 | 0.76 | 1.97 | CGG R | 0.45 | 0.94 | 2.11 | 0.32 |
| GGC G | 0.61 | 0.61 | 5.88 | 0.10 | CGU R | 1.05 | 1.38 | 0.44 | 2.76 |
| GGG G | 1.12 | 0.92 | 2.07 | 0.52 | AGC S | 0.71 | 0.82 | 3.03 | 0.25 |
| GGU G | 2.10 | 1.42 | 0.67 | 2.63 | AGU S | 1.26 | 1.24 | 0.27 | 4.63 |
| CAC H | 0.89 | 1.21 | 1.89 | 0.56 | UCA S | 1.48 | 1.78 | 0.43 | 3.79 |
| CAU H | 1.26 | 1.87 | 0.52 | 3.01 | UCC S | 0.98 | 1.34 | 1.33 | 0.87 |
| AUA I | 2.44 | 2.64 | 0.20 | 12.70 | UCG S | 1.10 | 1.38 | 1.05 | 1.18 |
| AUC I | 2.00 | 1.80 | 1.68 | 1.13 | UCU S | 1.88 | 1.69 | 0.51 | 3.5 |
| AUU I | 2.87 | 2.88 | 0.44 | 6.53 | ACA T | 2.01 | 1.87 | 0.61 | 3.18 |
| AAA K | 4.70 | 3.76 | 0.25 | 16.92 | ACC T | 1.32 | 1.38 | 1.98 | 0.68 |
| AAG K | 2.48 | 1.91 | 2.53 | 0.87 | ACG T | 1.62 | 1.57 | 1.29 | 1.24 |
| CUA L | 0.85 | 0.93 | 0.27 | 3.30 | ACU T | 1.57 | 1.31 | 0.39 | 3.69 |
| CUC L | 1.26 | 1.09 | 1.47 | 0.80 | GUA V | 1.80 | 1.54 | 0.25 | 6.68 |
| CUG L | 0.85 | 1.08 | 6.85 | 0.14 | GUC V | 1.35 | 1.25 | 1.16 | 1.12 |
| CUU L | 1.65 | 1.67 | 0.50 | 2.00 | GUG V | 1.57 | 1.37 | 4.23 | 0.35 |
| UUA L | 1.39 | 1.81 | 0.06 | 26.67 | GUU V | 2.34 | 2.28 | 0.40 | 5.78 |
| UUG L | 1.76 | 2.06 | 0.56 | 3.41 | UGG W | 1.09 | 1.24 | 1.61 | 0.72 |
| AUG M | 2.76 | 1.97 | 1.84 | 2.04 | UAC Y | 1.52 | 1.46 | 1.42 | 1.05 |
| UAU Y | 2.24 | 2.66 | 0.38 | 6.45 |
| Phycodnaviridae Viruses | Accession Number | Total tRNA Genes | tRNA Genes in Cluster b |
|---|---|---|---|
| Bathycoccus sp. RCC1105 virus BpV1 | HM004432.1 | 2 | 2 |
| Ectocarpus siliculosus virus 1 | AF204951.2 | 0 | 0 |
| Emiliania huxleyi virus 86 isolate EhV86 | AJ890364.1 | 4 | 2 |
| Heterosigma akashiwo virus 01 | NC_038553 | 3 | 2 |
| Micromonas pusilla virus 12T | NC_020864.1 | 6 | 5 |
| Micromonas sp. RCC1109 virus MpV1 | HM004429.1 | 6 | 3 |
| Only Syngen Nebraska virus 5 | KX857749.1 | 14 | 14 |
| Ostreococcus lucimarinus virus OlV1 | HM004431.1 | 4 | 4 |
| Ostreococcus lucimarinus virus 2 isolate Olv2 | KP874736.1 | 5 | 4 |
| Ostreococcus lucimarinus virus 7 isolate OlV7 | KP874737.1 | 4 | 4 |
| Ostreococcus mediterraneus virus 1 isolate OmV1 | KP874735.1 | 3 | 3 |
| Ostreococcus tauri virus 2 | FN600414.1 | 3 | 3 |
| Ostreococcus tauri virus OtV5 | EU304328.2 | 3 | 3 |
| Yellowstone lake phycodnavirus 1 | LC015647.1 | 7 | 2 and 5 c |
| Yellowstone lake phycodnavirus 2 | LC015648.1 | 4 | 4 |
| Yellowstone lake phycodnavirus 3 | LC015649.1 | 4 | 3 |
| Ascoviridae | Accession Number | Total tRNA Genes | tRNA Genes in Cluster c |
|---|---|---|---|
| Heliothis virescens ascovirus 3f isolate LD135790 | KJ755191.1 | 1 | 0 |
| Heliothis virescens ascovirus 3g | JX491653.1 | 1 | 0 |
| Trichoplusia ni ascovirus 2c | DQ517337.1 | 3 | 0 |
| Asfarviridae | |||
| African swine fever virus strain Ken06.Bus | NC_044946 | 0 | 0 |
| African swine fever virus strain BA71V | NC_001659 | 0 | 0 |
| African swine fever virus isolate ASFV Belgium 2018/1 | LR536725.1 | 0 | 0 |
| Faustoviruses | |||
| Faustovirus strain E9 | MT335755 | 0 | 0 |
| Faustovirus strain D3 | KU556803 | 0 | 0 |
| Faustovirus strain E24 | KU702948.1 | 0 | 0 |
| Iridoviridae | |||
| Aedes taeniorhynchus iridescent virus, | DQ643392.1 | 1 | 0 |
| Lymphocystis disease virus—isolate China | AY380826.1 | 1 | 0 |
| Scale drop disease virus isolate C4575 | KR139659.1 | 1 | 0 |
| Singapore grouper iridovirus | AY521625.1 | 1 | 0 |
| Kaumeobavirus | |||
| Kaumeobavirus isolate Sc | NC_034249.1 | 0 | 0 |
| Kaumoebavirus isolate Sc | KX552040.1 | 0 | 0 |
| Marseilliviridae | |||
| Brazilian marseillevirus strain BH2014 | NC_029692.1 | 0 | 0 |
| Golden Marseillevirus | NC_031465.1 | 0 | 0 |
| Marseillevirus strain T19 | NC_013756 | 0 | 0 |
| Mimiviridae | |||
| Acanthamoeba polyphaga mimivirus | HQ336222.2 | 6 | 2 |
| Acanthamoeba polyphaga moumouvirus | JX962719.1 | 2 | 2 |
| Aureococcus anophagefferens virus isolate BtV-01 | KJ645900.1 | 7 | 5 |
| Cafeteria roenbergensis virus BV-PW1 | GU244497.1 | 22 | 22 |
| Chrysochromulina ericina virus isolate CeV-01B | KT820662.1 | 12 | 3 and 7 d |
| Megavirus chiliensis | JN258408.1 | 3 | 0 |
| Mimivirus terra2 | KF527228.1 | 6 | 2 |
| Phaeocystis globosa virus strain 16T | KC662249.1 | 8 | 7 |
| Tetraselmis virus 1 | KY322437.1 | 10 | 10 |
| Molliviruses | |||
| Mollivirus kamchatka strain Kronotsky | MN812837.1 | 0 | 0 |
| Mollivirus sibericum isolate P1084-T | NC_027867.1 | 2 | 0 |
| Mollivirus sibericum isolate P1084-T | KR921745.1 | 3 | 0 |
| Orpheovirus | |||
| Orpheovirus IHUMI-LCC2 | NC_036594.1 | 0 | 0 |
| Pacmanvirus | |||
| Pacmanvirus A23 | NC_034383.1 | 0 | 0 |
| Pandoraviruses | |||
| Pandoravirus macleodensis | NC_037665 | 1 | 0 |
| Pandoravirus neocaledonia | NC_037666 | 3 | 0 |
| Pandoravirus quercus | NC_037667 | 1 | 0 |
| Pithoviridae | |||
| Brazilian cedratvirus IHUMI strain IHUMI-27.7 | LT994651 | 0 | 0 |
| Cedratvirus kamchatka isolate P4 | MN873693.1 | 0 | 0 |
| Pithovirus sp. LC8 | LT598836 | 0 | 0 |
| Poxviridae | |||
| Parapoxvirus red deer/HL953 strain HL953 | KM502564.1 | 1 | 0 |
| Squirrelpox virus Berlin_2015 (unverified) | MF503315.1 | 1 | 0 |
| Baculoviridae | |||
| Anticarsia gemmatalis multicapsid nucleopolyhedrovirus isolate AgMNPV-37 | KR815466.1 | 1 | 0 |
| Anticarsia gemmatalis nucleopolyhedrovirus | DQ813662.2 | 1 | 0 |
| Autographa californica nucleopolyhedrovirus clone C6 | L22858.1 | 1 | 0 |
| Neodiprion lecontei NPV | AY349019.1 | 1 | 0 |
| Antheraea pernyi nucleopolyhedrovirus | DQ486030.3 | 1 | 0 |
| Helicoverpa armigera granulovirus | EU255577.1 | 1 | 0 |
| Mamestra brassicae MNPV strain K1 | JQ798165.1 | 1 | 0 |
| Mamestra configurata NPV-A strain 90/2 | U59461.2 | 1 | 0 |
| Xestia c-nigrum granulovirus | AF162221.1 | 1 | 0 |
| Herpesviridae | |||
| Bovine herpesvirus type 1.1 | AJ004801.1 | 2 | 0 |
| Cercopithecine herpesvirus 16 strain X313 | DQ149153.1 | 2 | 0 |
| Columbid alphaherpesvirus 1 strain HLJ | KX589235.1 | 19 | 0 |
| Falconid herpesvirus 1 strain S-18 | KJ668231.1 | 19 | 0 |
| Macropodid herpesvirus 1 isolate MaHV1.3076/08 | KT594769.1 | 1 | 0 |
| Murine herpesvirus 68 strain WUMS | U97553.2 | 7 | 4 and 2 e |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duncan, G.A.; Dunigan, D.D.; Van Etten, J.L. Diversity of tRNA Clusters in the Chloroviruses. Viruses 2020, 12, 1173. https://doi.org/10.3390/v12101173
Duncan GA, Dunigan DD, Van Etten JL. Diversity of tRNA Clusters in the Chloroviruses. Viruses. 2020; 12(10):1173. https://doi.org/10.3390/v12101173
Chicago/Turabian StyleDuncan, Garry A., David D. Dunigan, and James L. Van Etten. 2020. "Diversity of tRNA Clusters in the Chloroviruses" Viruses 12, no. 10: 1173. https://doi.org/10.3390/v12101173
APA StyleDuncan, G. A., Dunigan, D. D., & Van Etten, J. L. (2020). Diversity of tRNA Clusters in the Chloroviruses. Viruses, 12(10), 1173. https://doi.org/10.3390/v12101173






