BST2/Tetherin Overexpression Modulates Morbillivirus Glycoprotein Production to Inhibit Cell–Cell Fusion
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells
2.2. Viruses and Viral RNA Quantification
2.3. Plasmids
2.4. Infections
2.5. Pseudotyped Viruses
2.6. Fusion Assays
2.7. Incucyte Fluorometric Quantification
2.8. Protein Labeling and Quantification
2.9. Phylogenetic Analysis
2.10. Statistical Analysis and Data Handling
3. Results
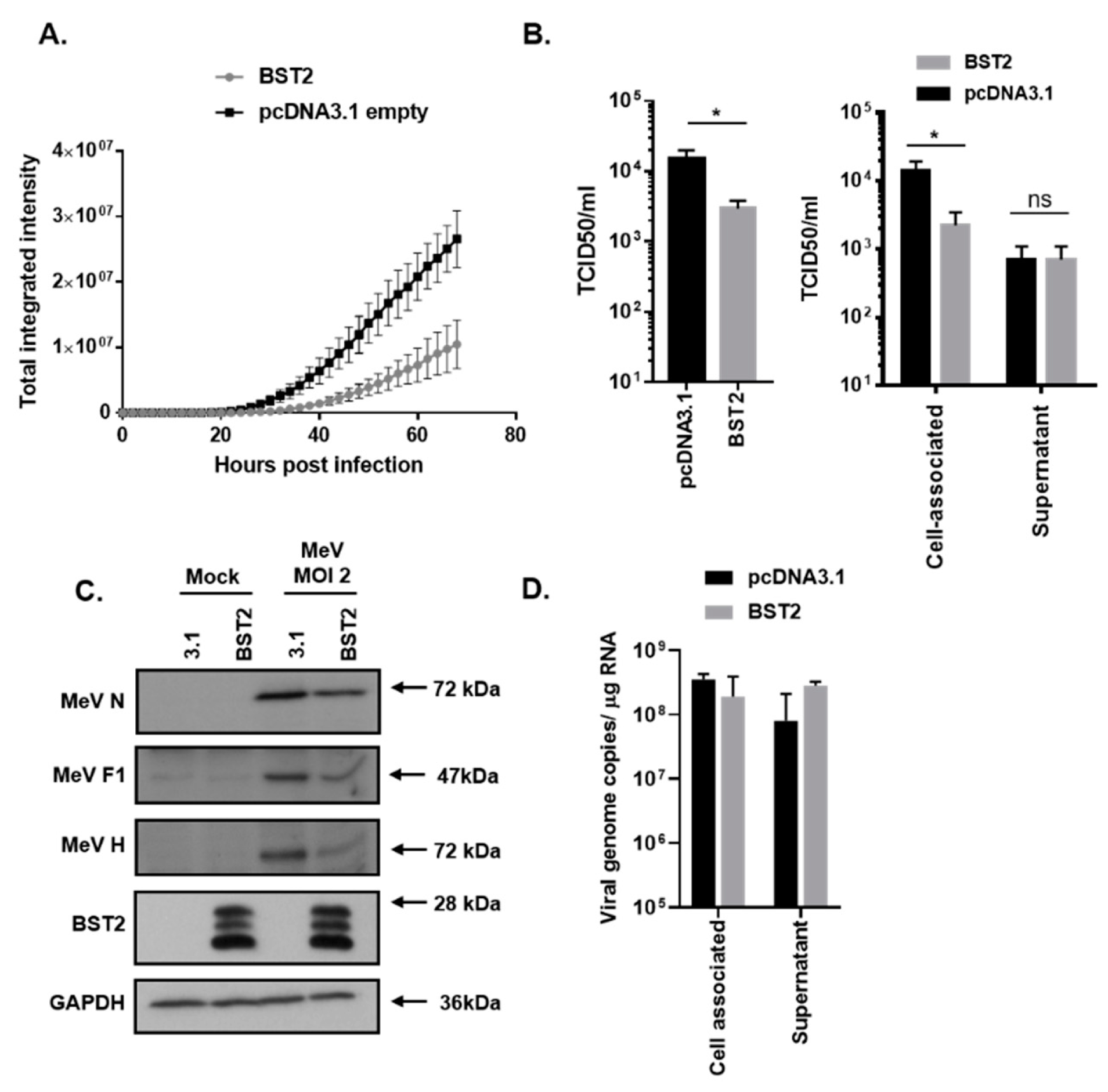
3.1. MeV Replication is Sensitive to BST2 Overexpression
3.2. MeV Entry is Not Inhibited by BST2
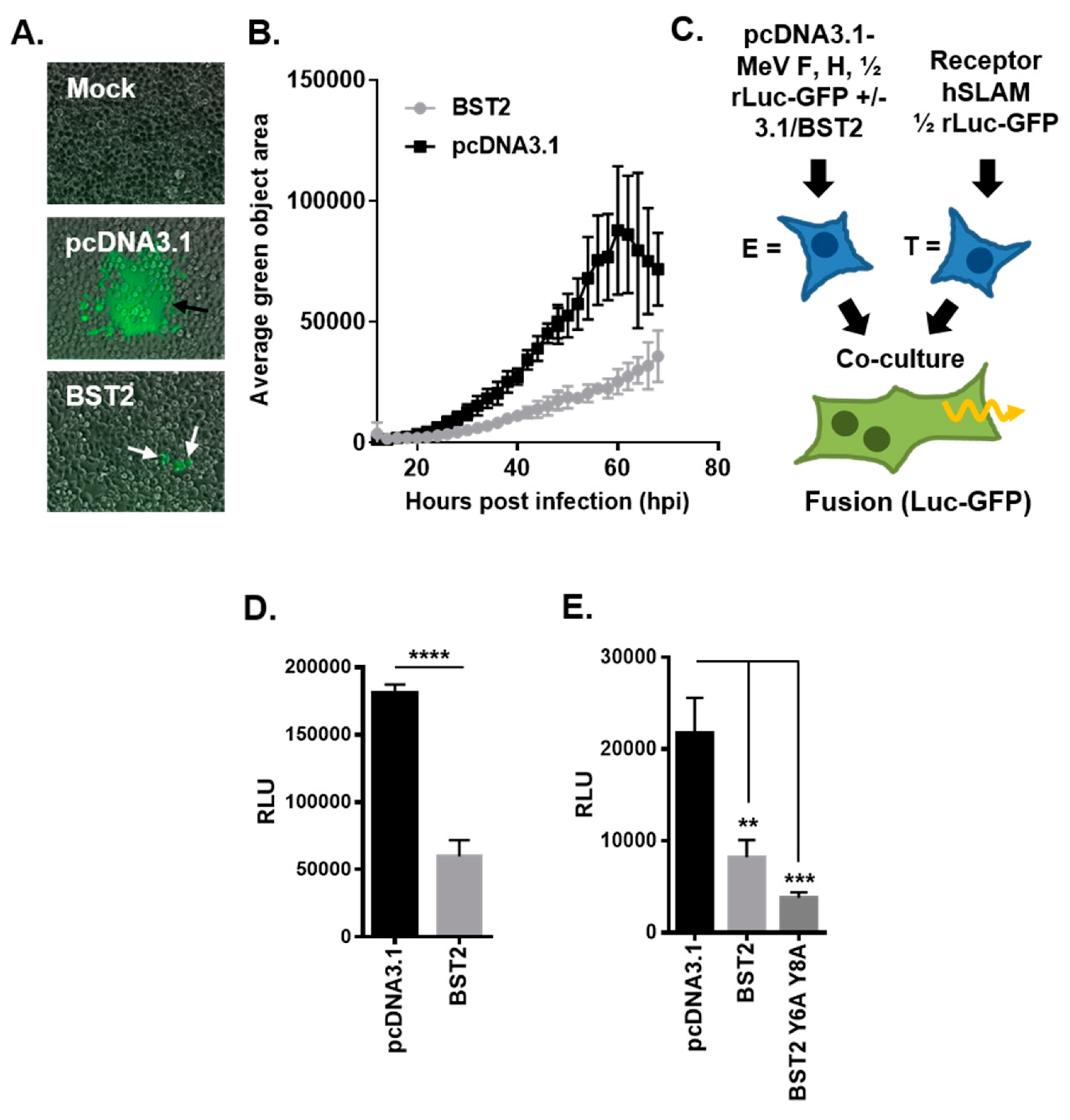
3.3. MeV Cell–Cell Fusion is Inhibited by BST2
3.4. The MeV Haemagglutinin Glycoprotein is Targeted by BST2
3.5. The BST2 GPI Anchor is Required for Restriction of MeV Replication
3.6. Morbilliviruses are Broadly Targeted by Mammalian BST2 Proteins
3.7. H Distribution is Modified in BST2-Transfected PPRV-Infected Cells
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Orgnization. Measles (Fact Sheet No 286). Available online: www.who.int/mediacentre/factsheets/fs286/en/ (accessed on 24 July 2019).
- El Najjar, F.; Schmitt, A.P.; Dutch, R.E. Paramyxovirus glycoprotein incorporation, assembly and budding: A three way dance for infectious particle production. Viruses 2014, 6, 3019–3054. [Google Scholar] [CrossRef] [PubMed]
- Muhlebach, M.D.; Mateo, M.; Sinn, P.L.; Prufer, S.; Uhlig, K.M.; Leonard, V.H.; Navaratnarajah, C.K.; Frenzke, M.; Wong, X.X.; Sawatsky, B.; et al. Adherens junction protein nectin-4 is the epithelial receptor for measles virus. Nature 2011, 480, 530–533. [Google Scholar] [CrossRef] [PubMed]
- Ludlow, M.; McQuaid, S.; Milner, D.; de Swart, R.L.; Duprex, W.P. Pathological consequences of systemic measles virus infection. J. Pathol. 2015, 235, 253–265. [Google Scholar] [CrossRef]
- White, R.G.; Boyd, J.F. The effect of measles on the thymus and other lymphoid tissues. Clin. Exp. Immunol. 1973, 13, 343–357. [Google Scholar] [PubMed]
- Podbilewicz, B. Virus and cell fusion mechanisms. Annu. Rev. Cell Dev. Biol. 2014, 30, 111–139. [Google Scholar] [CrossRef] [PubMed]
- Gotoh, B.; Komatsu, T.; Takeuchi, K.; Yokoo, J. Paramyxovirus accessory proteins as interferon antagonists. Microbiol. Immunol. 2001, 45, 787–800. [Google Scholar] [CrossRef] [PubMed]
- Zilliox, M.J.; Moss, W.J.; Griffin, D.E. Gene expression changes in peripheral blood mononuclear cells during measles virus infection. Clin. Vaccine Immunol.: Cvi. 2007, 14, 918–923. [Google Scholar] [CrossRef] [PubMed]
- Holmgren, A.M.; Miller, K.D.; Cavanaugh, S.E.; Rall, G.F. Bst2/Tetherin Is Induced in Neurons by Type I Interferon and Viral Infection but Is Dispensable for Protection against Neurotropic Viral Challenge. J. Virol. 2015, 89, 11011–11018. [Google Scholar] [CrossRef] [PubMed]
- Zilliox, M.J.; Parmigiani, G.; Griffin, D.E. Gene expression patterns in dendritic cells infected with measles virus compared with other pathogens. Proc. Natl. Acad. Sci. USA 2006, 103, 3363–3368. [Google Scholar] [CrossRef] [PubMed]
- Neil, S.J. The antiviral activities of tetherin. Curr. Top. Microbiol. Immunol. 2013, 371, 67–104. [Google Scholar] [PubMed]
- Neil, S.J.; Zang, T.; Bieniasz, P.D. Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu. Nature 2008, 451, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Giese, S.; Marsh, M. Tetherin can restrict cell-free and cell-cell transmission of HIV from primary macrophages to T cells. PLoS Pathog. 2014, 10, e1004189. [Google Scholar] [CrossRef] [PubMed]
- Jia, X.; Weber, E.; Tokarev, A.; Lewinski, M.; Rizk, M.; Suarez, M.; Guatelli, J.; Xiong, Y. Structural basis of HIV-1 Vpu-mediated BST2 antagonism via hijacking of the clathrin adaptor protein complex 1. eLife 2014, 3, e02362. [Google Scholar] [CrossRef]
- Gustin, J.K.; Bai, Y.; Moses, A.V.; Douglas, J.L. Ebola Virus Glycoprotein Promotes Enhanced Viral Egress by Preventing Ebola VP40 From Associating With the Host Restriction Factor BST2/Tetherin. J. Infect. Dis. 2015, 212, S181–S190. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.B.; Qu, X.W.; Jiang, D.; Zhao, X.L.; Han, J.C.; Wei, L. BST2/Tetherin inhibits hepatitis C virus production in human hepatoma cells. Antivir. Res. 2013, 98, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.B.; Han, J.C.; Cong, X.; Wei, L. BST2/tetherin inhibits dengue virus release from human hepatoma cells. PLoS ONE 2012, 7, e51033. [Google Scholar] [CrossRef] [PubMed]
- Zenner, H.L.; Mauricio, R.; Banting, G.; Crump, C.M. Herpes simplex virus 1 counteracts tetherin restriction via its virion host shutoff activity. J. Virol. 2013, 87, 13115–13123. [Google Scholar] [CrossRef] [PubMed]
- Galao, R.P.; Pickering, S.; Curnock, R.; Neil, S.J. Retroviral retention activates a Syk-dependent HemITAM in human tetherin. Cell Host Microbe 2014, 16, 291–303. [Google Scholar] [CrossRef]
- Hotter, D.; Sauter, D.; Kirchhoff, F. Emerging role of the host restriction factor tetherin in viral immune sensing. J. Mol. Biol. 2013, 425, 4956–4964. [Google Scholar] [CrossRef]
- Birch, J.; Juleff, N.; Heaton, M.P.; Kalbfleisch, T.; Kijas, J.; Bailey, D. Characterization of ovine Nectin-4, a novel peste des petits ruminants virus receptor. J. Virol. 2013, 87, 4756–4761. [Google Scholar] [CrossRef]
- Hashimoto, K.; Ono, N.; Tatsuo, H.; Minagawa, H.; Takeda, M.; Takeuchi, K.; Yanagi, Y. SLAM (CD150)-independent measles virus entry as revealed by recombinant virus expressing green fluorescent protein. J. Virol. 2002, 76, 6743–6749. [Google Scholar] [CrossRef] [PubMed]
- Frecha, C.; Costa, C.; Negre, D.; Gauthier, E.; Russell, S.J.; Cosset, F.L.; Verhoeyen, E. Stable transduction of quiescent T cells without induction of cycle progression by a novel lentiviral vector pseudotyped with measles virus glycoproteins. Blood 2008, 112, 4843–4852. [Google Scholar] [CrossRef] [PubMed]
- Moll, M.; Klenk, H.D.; Maisner, A. Importance of the cytoplasmic tails of the measles virus glycoproteins for fusogenic activity and the generation of recombinant measles viruses. J. Virol. 2002, 76, 7174–7186. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, H.; Meng, F.; Kondo, N.; Iwamoto, A.; Matsuda, Z. Generation of a dual-functional split-reporter protein for monitoring membrane fusion using self-associating split GFP. Protein Eng. Des. Sel.: Peds 2012, 25, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Cathomen, T.; Naim, H.Y.; Cattaneo, R. Measles viruses with altered envelope protein cytoplasmic tails gain cell fusion competence. J. Virol. 1998, 72, 1224–1234. [Google Scholar] [PubMed]
- Galao, R.P.; Le Tortorec, A.; Pickering, S.; Kueck, T.; Neil, S.J. Innate sensing of HIV-1 assembly by Tetherin induces NFkappaB-dependent proinflammatory responses. Cell Host Microbe 2012, 12, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Murphy, L.; Varela, M.; Desloire, S.; Ftaich, N.; Murgia, C.; Golder, M.; Neil, S.; Spencer, T.E.; Wootton, S.K.; Lavillette, D.; et al. The sheep tetherin paralog oBST2B blocks envelope glycoprotein incorporation into nascent retroviral virions. J. Virol. 2015, 89, 535–544. [Google Scholar] [CrossRef]
- Gnirss, K.; Fiedler, M.; Kramer-Kuhl, A.; Bolduan, S.; Mittler, E.; Becker, S.; Schindler, M.; Pohlmann, S. Analysis of determinants in filovirus glycoproteins required for tetherin antagonism. Viruses 2014, 6, 1654–1671. [Google Scholar] [CrossRef]
- Vande Burgt, N.H.; Kaletsky, R.L.; Bates, P. Requirements within the Ebola Viral Glycoprotein for Tetherin Antagonism. Viruses 2015, 7, 5587–5602. [Google Scholar] [CrossRef]
- Hauser, H.; Lopez, L.A.; Yang, S.J.; Oldenburg, J.E.; Exline, C.M.; Guatelli, J.C.; Cannon, P.M. HIV-1 Vpu and HIV-2 Env counteract BST-2/tetherin by sequestration in a perinuclear compartment. Retrovirology 2010, 7, 51. [Google Scholar] [CrossRef]
- Blondeau, C.; Pelchen-Matthews, A.; Mlcochova, P.; Marsh, M.; Milne, R.S.; Towers, G.J. Tetherin restricts herpes simplex virus 1 and is antagonized by glycoprotein M. J. Virol. 2013, 87, 13124–13133. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Luo, S.; He, S.; Zhang, M.; Wang, P.; Li, C.; Huang, W.; Hu, B.; Griffin, G.E.; Shattock, R.J.; et al. Tetherin restricts HSV-2 release and is counteracted by multiple viral glycoproteins. Virology 2015, 475, 96–109. [Google Scholar] [CrossRef] [PubMed]
- Narkpuk, J.; Wanitchang, A.; Kramyu, J.; Frantz, P.N.; Jongkaewwattana, A.; Teeravechyan, S. An unconventional BST-2 function: Down-regulation of transient protein expression. Biochem. Biophys. Res. Commun. 2014, 450, 1469–1474. [Google Scholar] [CrossRef] [PubMed]
- Goncalves-Carneiro, D.; McKeating, J.A.; Bailey, D. The measles virus receptor SLAMF1 can mediate particle endocytosis. J. Virol. 2017, 91, e02255-16. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S.; Ono, N.; Takeda, M.; Takeuchi, K.; Yanagi, Y. Dissection of measles virus V protein in relation to its ability to block alpha/beta interferon signal transduction. J. Gen. Virol. 2004, 85, 2991–2999. [Google Scholar] [CrossRef] [PubMed]
- Takaki, H.; Watanabe, Y.; Shingai, M.; Oshiumi, H.; Matsumoto, M.; Seya, T. Strain-to-strain difference of V protein of measles virus affects MDA5-mediated IFN-beta-inducing potential. Mol. Immunol. 2011, 48, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Ohta, K.; Goto, H.; Yumine, N.; Nishio, M. Human parainfluenza virus type 2 V protein inhibits and antagonizes tetherin. J. Gen. Virol. 2016, 97, 561–570. [Google Scholar] [CrossRef]
- Nishio, M.; Garcin, D.; Simonet, V.; Kolakofsky, D. The carboxyl segment of the mumps virus V protein associates with Stat proteins in vitro via a tryptophan-rich motif. Virology 2002, 300, 92–99. [Google Scholar] [CrossRef]
- Gnirss, K.; Zmora, P.; Blazejewska, P.; Winkler, M.; Lins, A.; Nehlmeier, I.; Gartner, S.; Moldenhauer, A.S.; Hofmann-Winkler, H.; Wolff, T.; et al. Tetherin Sensitivity of Influenza A Viruses Is Strain Specific: Role of Hemagglutinin and Neuraminidase. J. Virol. 2015, 89, 9178–9188. [Google Scholar] [CrossRef]
- Arnaud, F.; Black, S.G.; Murphy, L.; Griffiths, D.J.; Neil, S.J.; Spencer, T.E.; Palmarini, M. Interplay between ovine bone marrow stromal cell antigen 2/tetherin and endogenous retroviruses. J. Virol. 2010, 84, 4415–4425. [Google Scholar] [CrossRef]
- Weinelt, J.; Neil, S.J. Differential sensitivities of tetherin isoforms to counteraction by primate lentiviruses. J. Virol. 2014, 88, 5845–5858. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kelly, J.T.; Human, S.; Alderman, J.; Jobe, F.; Logan, L.; Rix, T.; Gonçalves-Carneiro, D.; Leung, C.; Thakur, N.; Birch, J.; et al. BST2/Tetherin Overexpression Modulates Morbillivirus Glycoprotein Production to Inhibit Cell–Cell Fusion. Viruses 2019, 11, 692. https://doi.org/10.3390/v11080692
Kelly JT, Human S, Alderman J, Jobe F, Logan L, Rix T, Gonçalves-Carneiro D, Leung C, Thakur N, Birch J, et al. BST2/Tetherin Overexpression Modulates Morbillivirus Glycoprotein Production to Inhibit Cell–Cell Fusion. Viruses. 2019; 11(8):692. https://doi.org/10.3390/v11080692
Chicago/Turabian StyleKelly, James T., Stacey Human, Joseph Alderman, Fatoumatta Jobe, Leanne Logan, Thomas Rix, Daniel Gonçalves-Carneiro, Corwin Leung, Nazia Thakur, Jamie Birch, and et al. 2019. "BST2/Tetherin Overexpression Modulates Morbillivirus Glycoprotein Production to Inhibit Cell–Cell Fusion" Viruses 11, no. 8: 692. https://doi.org/10.3390/v11080692
APA StyleKelly, J. T., Human, S., Alderman, J., Jobe, F., Logan, L., Rix, T., Gonçalves-Carneiro, D., Leung, C., Thakur, N., Birch, J., & Bailey, D. (2019). BST2/Tetherin Overexpression Modulates Morbillivirus Glycoprotein Production to Inhibit Cell–Cell Fusion. Viruses, 11(8), 692. https://doi.org/10.3390/v11080692




