Plasma Hepatitis E Virus Kinetics in Solid Organ Transplant Patients Receiving Ribavirin
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Ethical Statement
2.2. Biochemical Measurements
2.3. Detection of Anti-HEV IgM and IgG
2.4. HEV RNA Measurements
2.5. Genotype Determination
2.6. Determination of G1634R Polymorphism by Direct Nucleotide Sequencing
2.7. Measurement of Ribavirin Levels
2.8. Characterization of Plasma Viral Kinetic Patterns
2.9. Statistical Analysis
3. Results
3.1. Baseline Characteristics
3.2. Response to Ribavirin Therapy
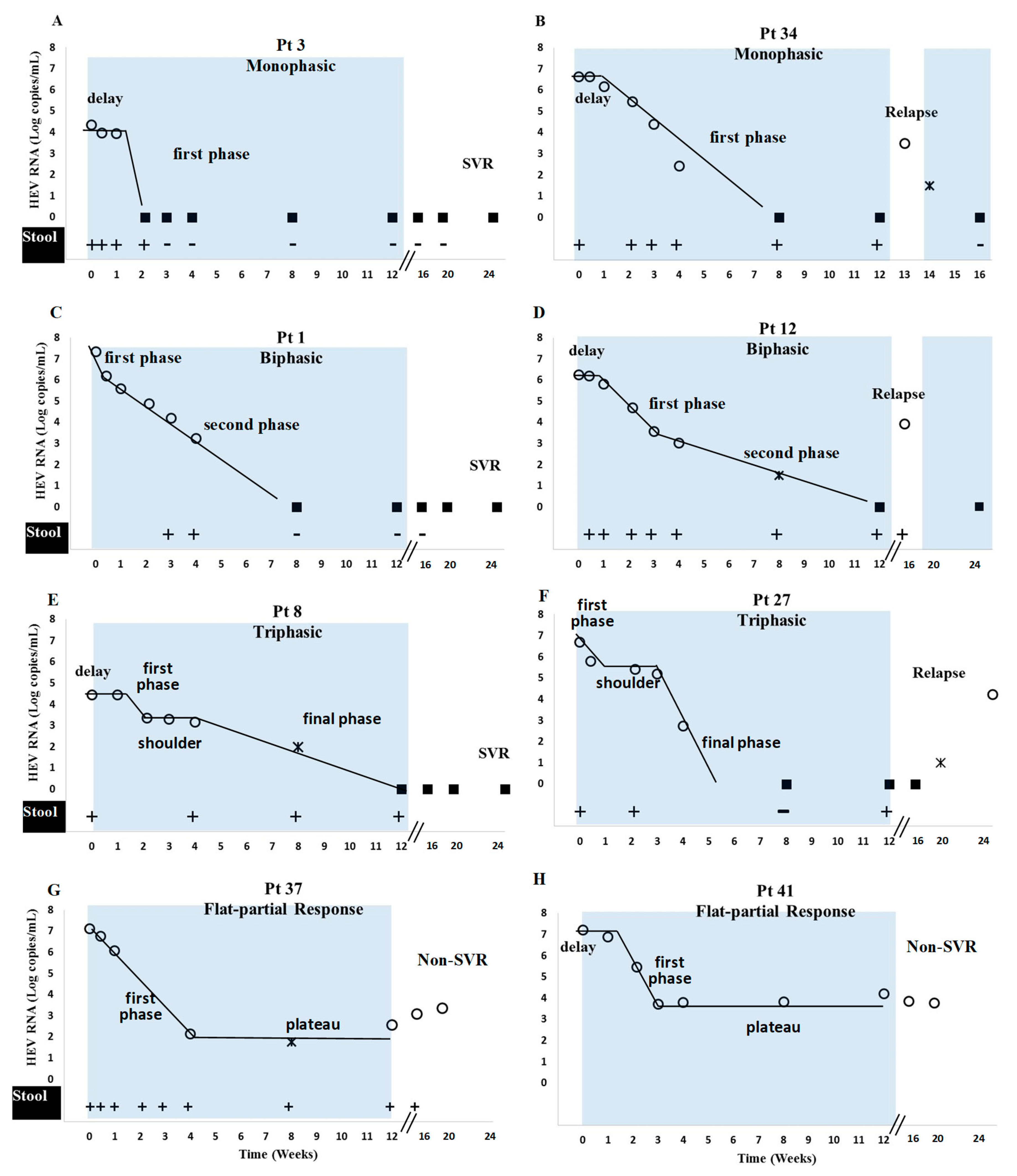
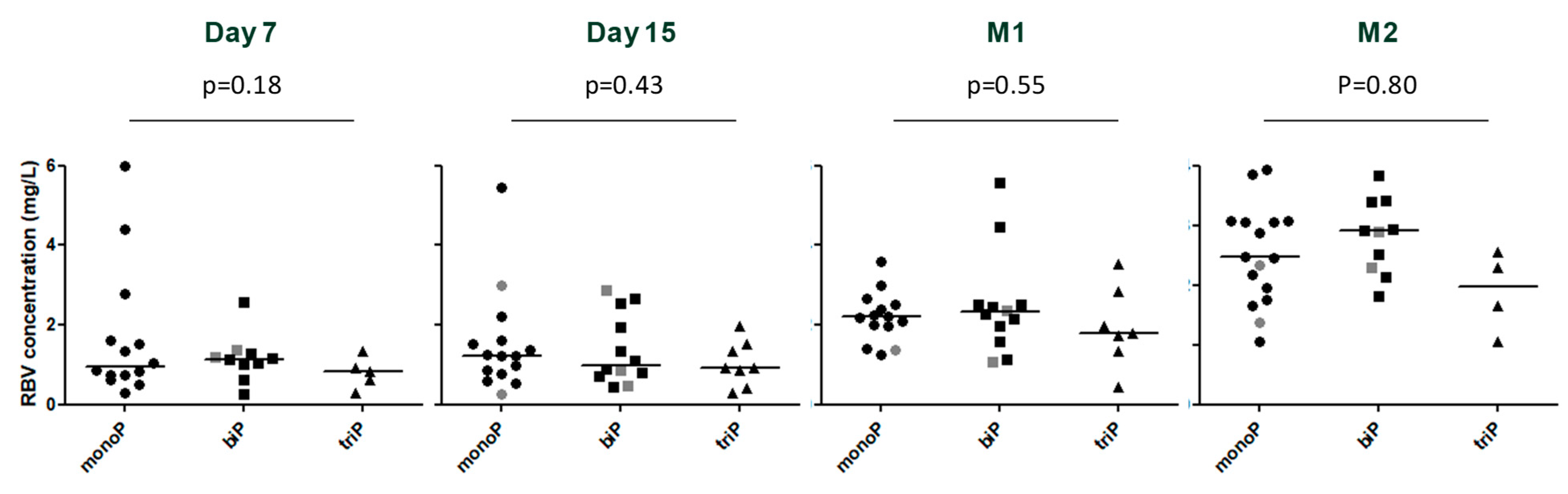
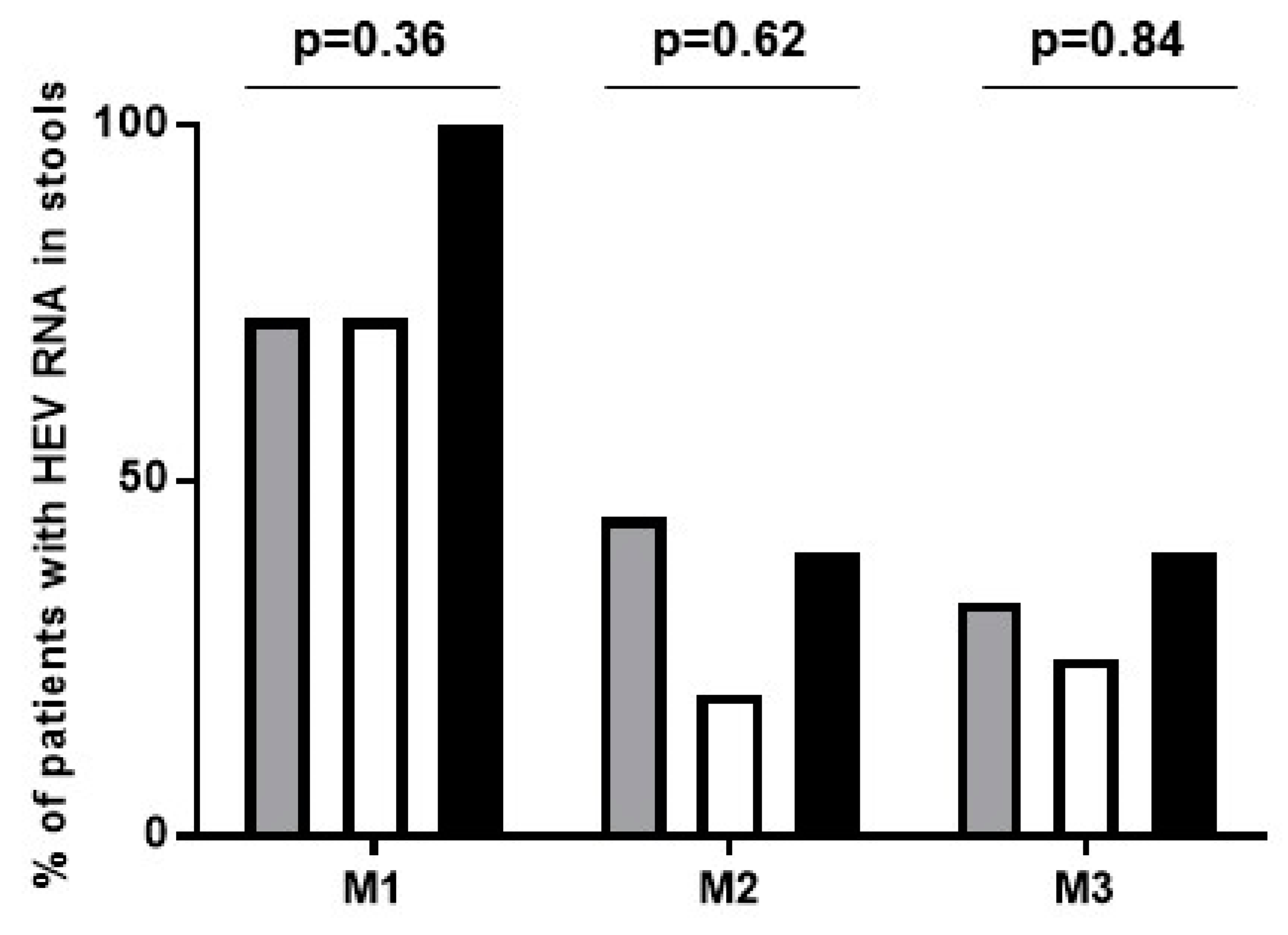
3.3. Viral Kinetics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rein, D.B.; Stevens, G.A.; Theaker, J.; Wittenborn, J.S.; Wiersma, S.T. The global burden of hepatitis E virus genotypes 1 and 2 in 2005. Hepatology 2012, 55, 988–997. [Google Scholar] [CrossRef]
- European Association for the Study of the Liver. EASL Clinical Practice Guidelines on hepatitis E virus infection. J. Hepatol. 2018. [Google Scholar] [CrossRef]
- Meng, X.J.; Purcell, R.H.; Halbur, P.G.; Lehman, J.R.; Webb, D.M.; Tsareva, T.S.; Haynes, J.S.; Thacker, B.J.; Emerson, S.U. A novel virus in swine is closely related to the human hepatitis E virus. Proc. Natl. Acad. Sci. USA 1997, 94, 9860–9865. [Google Scholar] [CrossRef]
- Kamar, N.; Izopet, J.; Pavio, N.; Aggarwal, R.; Labrique, A.; Wedemeyer, H.; Dalton, H.R. Hepatitis E virus infection. Nat. Rev. Dis. Primers 2017, 3, 17086. [Google Scholar] [CrossRef]
- Kamar, N.; Mansuy, J.M.; Cointault, O.; Selves, J.; Abravanel, F.; Danjoux, M.; Otal, P.; Esposito, L.; Durand, D.; Izopet, J.; et al. Hepatitis E virus-related cirrhosis in kidney- and kidney-pancreas-transplant recipients. Am. J. Transplant. 2008, 8, 1744–1748. [Google Scholar] [CrossRef]
- Gerolami, R.; Moal, V.; Colson, P. Chronic hepatitis E with cirrhosis in a kidney-transplant recipient. N. Engl. J. Med. 2008, 358, 859–860. [Google Scholar] [CrossRef]
- Dalton, H.R.; van Eijk, J.J.J.; Cintas, P.; Madden, R.G.; Jones, C.; Webb, G.W.; Norton, B.; Pique, J.; Lutgens, S.; Devooght-Johnson, N.; et al. Hepatitis E virus infection and acute non-traumatic neurological injury: A prospective multicentre study. J. Hepatol. 2017, 67, 925–932. [Google Scholar] [CrossRef]
- Dalton, H.R.; Kamar, N.; van Eijk, J.J.; McLean, B.N.; Cintas, P.; Bendall, R.P.; Jacobs, B.C. Hepatitis E virus and neurological injury. Nat. Rev. Neurol. 2016, 12, 77–85. [Google Scholar] [CrossRef]
- Lhomme, S.; Abravanel, F.; Dubois, M.; Sandres-Saune, K.; Mansuy, J.M.; Rostaing, L.; Kamar, N.; Izopet, J. Characterization of the polyproline region of the hepatitis E virus in immunocompromised patients. J. Virol. 2014, 88, 12017–12025. [Google Scholar] [CrossRef]
- Shukla, P.; Nguyen, H.T.; Torian, U.; Engle, R.E.; Faulk, K.; Dalton, H.R.; Bendall, R.P.; Keane, F.E.; Purcell, R.H.; Emerson, S.U. Cross-species infections of cultured cells by hepatitis E virus and discovery of an infectious virus-host recombinant. Proc. Natl. Acad. Sci. USA 2011, 108, 2438–2443. [Google Scholar] [CrossRef]
- Kamar, N.; Abravanel, F.; Selves, J.; Garrouste, C.; Esposito, L.; Lavayssiere, L.; Cointault, O.; Ribes, D.; Cardeau, I.; Nogier, M.B.; et al. Influence of immunosuppressive therapy on the natural history of genotype 3 hepatitis-E virus infection after organ transplantation. Transplantation 2010, 89, 353–360. [Google Scholar] [CrossRef]
- Kamar, N.; Izopet, J.; Tripon, S.; Bismuth, M.; Hillaire, S.; Dumortier, J.; Radenne, S.; Coilly, A.; Garrigue, V.; D’Alteroche, L.; et al. Ribavirin for chronic hepatitis E virus infection in transplant recipients. N. Engl. J. Med. 2014, 370, 1111–1120. [Google Scholar] [CrossRef]
- Debing, Y.; Gisa, A.; Dallmeier, K.; Pischke, S.; Bremer, B.; Manns, M.; Wedemeyer, H.; Suneetha, P.V.; Neyts, J. A mutation in the hepatitis E virus RNA polymerase promotes its replication and associates with ribavirin treatment failure in organ transplant recipients. Gastroenterology 2014, 147, 1008–1011.e7. [Google Scholar] [CrossRef]
- Lhomme, S.; Kamar, N.; Nicot, F.; Ducos, J.; Bismuth, M.; Garrigue, V.; Petitjean-Lecherbonnier, J.; Ollivier, I.; Alessandri-Gradt, E.; Goria, O.; et al. Mutation in the Hepatitis E Virus Polymerase and Outcome of Ribavirin Therapy. Antimicrob. Agents Chemother. 2015, 60, 1608–1614. [Google Scholar] [CrossRef]
- Abravanel, F.; Lhomme, S.; Rostaing, L.; Kamar, N.; Izopet, J. Protracted fecal shedding of HEV during ribavirin therapy predicts treatment relapse. Clin. Infect. Dis. 2015, 60, 96–99. [Google Scholar] [CrossRef]
- Kamar, N.; Lhomme, S.; Abravanel, F.; Cointault, O.; Esposito, L.; Cardeau-Desangles, I.; Del Bello, A.; Dorr, G.; Lavayssiere, L.; Nogier, M.B.; et al. An Early Viral Response Predicts the Virological Response to Ribavirin in Hepatitis E Virus Organ Transplant Patients. Transplantation 2015, 99, 2124–2131. [Google Scholar] [CrossRef]
- Marion, O.; Lhomme, S.; del Bello, A.; Abravanel, F.; Esposito, L.; Hebral, A.L.; Lavayssiere, L.; Cointault, O.; Ribes, D.; Izopet, J.; et al. Monitoring hepatitis E virus fecal shedding to optimize ribavirin treatment duration in chronically infected transplant patients. J. Hepatol. 2019, 70, 206–209. [Google Scholar] [CrossRef]
- Wu, A. Tietz Clinical Guide to Laboratory Tests, 4th ed.; Elsevier: Amsterdam, The Netherlands; Saunders: Philadelphia, PA, USA, 2006. [Google Scholar]
- Abravanel, F.; Sandres-Saune, K.; Lhomme, S.; Dubois, M.; Mansuy, J.M.; Izopet, J. Genotype 3 diversity and quantification of hepatitis E virus RNA. J. Clin. Microbiol. 2012, 50, 897–902. [Google Scholar] [CrossRef]
- Legrand-Abravanel, F.; Mansuy, J.M.; Dubois, M.; Kamar, N.; Peron, J.M.; Rostaing, L.; Izopet, J. Hepatitis E virus genotype 3 diversity, France. Emerg. Infect. Dis. 2009, 15, 110–114. [Google Scholar] [CrossRef]
- Smith, D.B.; Simmonds, P.; Izopet, J.; Oliveira-Filho, E.F.; Ulrich, R.G.; Johne, R.; Koenig, M.; Jameel, S.; Harrison, T.J.; Meng, X.J.; et al. Proposed reference sequences for hepatitis E virus subtypes. J. Gen. Virol. 2016, 97, 537–542. [Google Scholar] [CrossRef]
- Gandia, P.; Trancart, S.; Nicot, F.; Barange, K.; Alric, L.; Izopet, J.; Seraissol, P.; Lavit, M.; Houin, G. Influence of pre-analytical conditions on plasma ribavirin concentrations. Arzneimittelforschung 2010, 60, 636–639. [Google Scholar] [CrossRef]
- Guedj, J.; Dahari, H.; Pohl, R.T.; Ferenci, P.; Perelson, A.S. Understanding silibinin’s modes of action against HCV using viral kinetic modeling. J. Hepatol. 2012, 56, 1019–1024. [Google Scholar] [CrossRef]
- Kamar, N.; Chatelut, E.; Manolis, E.; Lafont, T.; Izopet, J.; Rostaing, L. Ribavirin pharmacokinetics in renal and liver transplant patients: Evidence that it depends on renal function. Am. J. Kidney Dis. 2004, 43, 140–146. [Google Scholar] [CrossRef]
- Canini, L.; Imamura, M.; Kawakami, Y.; Uprichard, S.L.; Cotler, S.J.; Dahari, H.; Chayama, K. HCV kinetic and modeling analyses project shorter durations to cure under combined therapy with daclatasvir and asunaprevir in chronic HCV-infected patients. PLoS ONE 2017, 12, e0187409. [Google Scholar] [CrossRef]
- Dahari, H.; Canini, L.; Graw, F.; Uprichard, S.L.; Araujo, E.S.; Penaranda, G.; Coquet, E.; Chiche, L.; Riso, A.; Renou, C.; et al. HCV kinetic and modeling analyses indicate similar time to cure among sofosbuvir combination regimens with daclatasvir, simeprevir or ledipasvir. J. Hepatol. 2016, 64, 1232–1239. [Google Scholar] [CrossRef]
- Dahari, H.; Shteingart, S.; Gafanovich, I.; Cotler, S.J.; D’Amato, M.; Pohl, R.T.; Weiss, G.; Ashkenazi, Y.J.; Tichler, T.; Goldin, E.; et al. Sustained virological response with intravenous silibinin: Individualized IFN-free therapy via real-time modelling of HCV kinetics. Liver Int. 2015, 35, 289–294. [Google Scholar] [CrossRef]
- Ferenci, P.; Laferl, H.; Scherzer, T.M.; Gschwantler, M.; Maieron, A.; Brunner, H.; Stauber, R.; Bischof, M.; Bauer, B.; Datz, C.; et al. Peginterferon alfa-2a and ribavirin for 24 weeks in hepatitis C type 1 and 4 patients with rapid virological response. Gastroenterology 2008, 135, 451–458. [Google Scholar] [CrossRef]
- Debing, Y.; Emerson, S.U.; Wang, Y.; Pan, Q.; Balzarini, J.; Dallmeier, K.; Neyts, J. Ribavirin inhibits in vitro hepatitis E virus replication through depletion of cellular GTP pools and is moderately synergistic with alpha interferon. Antimicrob. Agents Chemother. 2014, 58, 267–273. [Google Scholar] [CrossRef]
- Rotman, Y.; Noureddin, M.; Feld, J.J.; Guedj, J.; Witthaus, M.; Han, H.; Park, Y.J.; Park, S.H.; Heller, T.; Ghany, M.G.; et al. Effect of ribavirin on viral kinetics and liver gene expression in chronic hepatitis C. Gut 2014, 63, 161–169. [Google Scholar] [CrossRef]
- Pawlotsky, J.M.; Dahari, H.; Neumann, A.U.; Hezode, C.; Germanidis, G.; Lonjon, I.; Castera, L.; Dhumeaux, D. Antiviral action of ribavirin in chronic hepatitis C. Gastroenterology 2004, 126, 703–714. [Google Scholar] [CrossRef]
- Neumann, A.U.; Lam, N.P.; Dahari, H.; Gretch, D.R.; Wiley, T.E.; Layden, T.J.; Perelson, A.S. Hepatitis C viral dynamics in vivo and the antiviral efficacy of interferon-alpha therapy. Science 1998, 282, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Dahari, H.; Shudo, E.; Ribeiro, R.M.; Perelson, A.S. Modeling complex decay profiles of hepatitis B virus during antiviral therapy. Hepatology 2009, 49, 32–38. [Google Scholar] [CrossRef]
- Guedj, J.; Rotman, Y.; Cotler, S.J.; Koh, C.; Schmid, P.; Albrecht, J.; Haynes-Williams, V.; Liang, T.J.; Hoofnagle, J.H.; Heller, T.; et al. Understanding early serum hepatitis D virus and hepatitis B surface antigen kinetics during pegylated interferon-alpha therapy via mathematical modeling. Hepatology 2014, 60, 1902–1910. [Google Scholar] [CrossRef] [PubMed]
- Koh, C.; Canini, L.; Dahari, H.; Zhao, X.; Uprichard, S.L.; Haynes-Williams, V.; Winters, M.A.; Subramanya, G.; Cooper, S.L.; Pinto, P.; et al. Oral prenylation inhibition with lonafarnib in chronic hepatitis D infection: A proof-of-concept randomised, double-blind, placebo-controlled phase 2A trial. Lancet Infect. Dis. 2015, 15, 1167–1174. [Google Scholar] [CrossRef]
- Wang, Y.; Zhou, X.; Debing, Y.; Chen, K.; Van Der Laan, L.J.; Neyts, J.; Janssen, H.L.; Metselaar, H.J.; Peppelenbosch, M.P.; Pan, Q. Calcineurin inhibitors stimulate and mycophenolic acid inhibits replication of hepatitis E virus. Gastroenterology 2014, 146, 1775–1783. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Wang, Y.; Metselaar, H.J.; Janssen, H.L.; Peppelenbosch, M.P.; Pan, Q. Rapamycin and everolimus facilitate hepatitis E virus replication: Revealing a basal defense mechanism of PI3K-PKB-mTOR pathway. J. Hepatol. 2014, 61, 746–754. [Google Scholar] [CrossRef]
- Todt, D.; Gisa, A.; Radonic, A.; Nitsche, A.; Behrendt, P.; Suneetha, P.V.; Pischke, S.; Bremer, B.; Brown, R.J.; Manns, M.P.; et al. In vivo evidence for ribavirin-induced mutagenesis of the hepatitis E virus genome. Gut 2016, 65, 1733–1743. [Google Scholar] [CrossRef]
- Dahari, H.; Ribeiro, R.M.; Perelson, A.S. Triphasic decline of hepatitis C virus RNA during antiviral therapy. Hepatology 2007, 46, 16–21. [Google Scholar] [CrossRef]
- Dixit, N.M.; Layden-Almer, J.E.; Layden, T.J.; Perelson, A.S. Modelling how ribavirin improves interferon response rates in hepatitis C virus infection. Nature 2004, 432, 922–924. [Google Scholar] [CrossRef]
- Hofmann, W.P.; Polta, A.; Herrmann, E.; Mihm, U.; Kronenberger, B.; Sonntag, T.; Lohmann, V.; Schonberger, B.; Zeuzem, S.; Sarrazin, C. Mutagenic effect of ribavirin on hepatitis C nonstructural 5B quasispecies in vitro and during antiviral therapy. Gastroenterology 2007, 132, 921–930. [Google Scholar] [CrossRef]
- Lutchman, G.; Danehower, S.; Song, B.C.; Liang, T.J.; Hoofnagle, J.H.; Thomson, M.; Ghany, M.G. Mutation rate of the hepatitis C virus NS5B in patients undergoing treatment with ribavirin monotherapy. Gastroenterology 2007, 132, 1757–1766. [Google Scholar] [CrossRef] [PubMed]
- Allweiss, L.; Gass, S.; Giersch, K.; Groth, A.; Kah, J.; Volz, T.; Rapp, G.; Schobel, A.; Lohse, A.W.; Polywka, S.; et al. Human liver chimeric mice as a new model of chronic hepatitis E virus infection and preclinical drug evaluation. J. Hepatol. 2016, 64, 1033–1040. [Google Scholar] [CrossRef] [PubMed]
- Sayed, I.M.; Verhoye, L.; Cocquerel, L.; Abravanel, F.; Foquet, L.; Montpellier, C.; Debing, Y.; Farhoudi, A.; Wychowski, C.; Dubuisson, J.; et al. Study of hepatitis E virus infection of genotype 1 and 3 in mice with humanised liver. Gut 2017, 66, 920–929. [Google Scholar] [CrossRef] [PubMed]
- Thomas, H.; Locarnini, S.; Zuckerman, A. Viral Hepatitis, 4th ed.; Wiley-Blackwell: Hoboken, NJ, USA, 2013. [Google Scholar] [CrossRef]
- Williams, T.P.; Kasorndorkbua, C.; Halbur, P.G.; Haqshenas, G.; Guenette, D.K.; Toth, T.E.; Meng, X.J. Evidence of extrahepatic sites of replication of the hepatitis E virus in a swine model. J. Clin. Microbiol. 2001, 39, 3040–3046. [Google Scholar] [CrossRef] [PubMed]
- Izopet, J.; Dubois, M.; Bertagnoli, S.; Lhomme, S.; Marchandeau, S.; Boucher, S.; Kamar, N.; Abravanel, F.; Guerin, J.L. Hepatitis E virus strains in rabbits and evidence of a closely related strain in humans, France. Emerg. Infect. Dis. 2012, 18, 1274–1281. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Bu, Q.N.; Wang, L.; Han, J.; Du, R.J.; Lei, Y.X.; Ouyang, Y.Q.; Li, J.; Zhu, Y.H.; Lu, F.M.; et al. Transmission of hepatitis E virus from rabbits to cynomolgus macaques. Emerg. Infect. Dis. 2013, 19, 559–565. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Zhao, C.; Huang, W.; Harrison, T.J.; Zhang, H.; Geng, K.; Wang, Y. Detection and assessment of infectivity of hepatitis E virus in urine. J. Hepatol. 2016, 64, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Marion, O.; Capelli, N.; Lhomme, S.; Dubois, M.; Pucelle, M.; Abravanel, F.; Kamar, N.; Izopet, J. Hepatitis E virus genotype 3 and capsid protein in the blood and urine of immunocompromised patients. J. Infect. 2019. [Google Scholar] [CrossRef] [PubMed]
| Variable | Value |
|---|---|
| Age, years (mean, SEM) | 51 ± 14 |
| Sex, Male (%) | 29 (70.7) |
| HEV RNA concentration (log copies/mL) | 6.12 [3.53–7.45] |
| HEV genotype, n (%) | |
| 3f | 25 (60.9) |
| 3c | 12 (29.3) |
| 3e | 2 (4.9) |
| 3 | 1 (2.4) |
| Not determined | 1 (2.4) |
| Type of organ transplant, n (%) | |
| Kidney | 26 (63.4) |
| Liver | 9 (22.0) |
| Heart | 3 (7.3) |
| Kidney/pancreas | 1 (2.4) |
| Lung | 1 (2.4) |
| Liver/kidney | 1 (2.4) |
| Dose of ribavirin (mg/kg/day) | 9.7 [2.7–16.3] |
| Immunosuppressive therapy at the initiation of ribavirin, n (%) | |
| Calcineurin inhibitors | 32 (78.0) |
| Tacrolimus | 31 (75.6) |
| Cyclosporin | 1 (2.4) |
| Antimetabolite | 34 (82.9) |
| Steroids | 35 (85.4) |
| mTOR inhibitors | 11 (26.2) |
| ALT level (IU/L) | 119 [24–1506] |
| AST level (IU/L) | 71 [30–1263] |
| Serum creatinine at the initiation of ribavirin (µmol/L) | 133 [40–360] |
| Lymphocyte count at the initiation of ribavirin (cells/mm3) | 1200 [178–6000] |
| Hemoglobin level at the initiation of ribavirin (g/dL) | 12.6 [9.3–15.6] |
| Platelet count at the initiation of ribavirin (cells/mm3) | 166 [12–1023] |
| Positive test for anti-HEV IgG antibodies at the initiation of ribavirin, n/total (%) | 31/41 (75.6) |
| Positive test for anti-HEV IgM antibodies at the initiation of ribavirin, n/total (%) | 37/41 (90.2) |
| Positive test for HEV RNA at the initiation of ribavirin, n (%) | 41 (100) |
| Interval between transplantation and initiation of ribavirin (months) | 43 [3–268] |
| Interval between immunosuppression decrease and initiation of ribavirin (months) | 4 [0–26] |
| Interval between diagnosis of HEV infection and initiation of ribavirin (months) | 4 [0–47] |
| Patients | Baseline HEV/RNA | Delay | First Phase Slope | First Phase Length | First Phase | Second Phase Slope | Second Phase Plateau Slope | Mean VL at Plateau | Shoulder Phase Slope | Length of Plateau | Mean VL at Plateau | Final Phase Slope | Final Phase | Time to First DNQ | Time to First | Time to First TND in Stool (Days) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| [log10 copies/mL] | [days] | [log10 copies/mL/day] | [days] | t1/2 [days] | [log10 copies/mL/day] | [log10 copies/mL/day] | [log10 copies/mL] | [log10 copies/mL/day] | [days] | [log10 copies/mL] | [log10 copies/mL/day] | t1/2 [days] | [days] | TND [days] | ||
| Monophasic (n = 18) | 5.60 | 3 | 0.14 | 2.21 | 15 | 56 | 60 | |||||||||
| IQR | (1.51) | (7) | (0.13) | (1.35) | (35) | (38) | ||||||||||
| Biphasic (n = 13) | 6.45 | 0.00 | 0.15 | 15.00 | 2.05 | 0.14 | 2.10 | 56 | 56 | 45 | ||||||
| IQR | (0.95) | (3.00) | (0.1) | (4.50) | (1.34) | (0.3) | (4.12) | (56) | (59) | |||||||
| Triphasic (n = 8) | 5.39 | 0 | 0.18 | 7 | 1.71 | 0.01 | 14 | 3.23 | 0.13 | 2.43 | 28 | 56 | 60 | |||
| IQR | (1.19) | (6) | (0.21) | (12) | (1.30) | (0.03) | (6) | (1.34) | (0.24) | (4.06) | (49) | (52) | ||||
| Flat-partial (n = 2) | 6.67 | 3.50 | 0.20 | 21 | 1.50 | 0.01 | 3.02 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lhomme, S.; DebRoy, S.; Kamar, N.; Abravanel, F.; Metsu, D.; Marion, O.; Dimeglio, C.; Cotler, S.J.; Izopet, J.; Dahari, H. Plasma Hepatitis E Virus Kinetics in Solid Organ Transplant Patients Receiving Ribavirin. Viruses 2019, 11, 630. https://doi.org/10.3390/v11070630
Lhomme S, DebRoy S, Kamar N, Abravanel F, Metsu D, Marion O, Dimeglio C, Cotler SJ, Izopet J, Dahari H. Plasma Hepatitis E Virus Kinetics in Solid Organ Transplant Patients Receiving Ribavirin. Viruses. 2019; 11(7):630. https://doi.org/10.3390/v11070630
Chicago/Turabian StyleLhomme, Sebastien, Swati DebRoy, Nassim Kamar, Florence Abravanel, David Metsu, Olivier Marion, Chloé Dimeglio, Scott J. Cotler, Jacques Izopet, and Harel Dahari. 2019. "Plasma Hepatitis E Virus Kinetics in Solid Organ Transplant Patients Receiving Ribavirin" Viruses 11, no. 7: 630. https://doi.org/10.3390/v11070630
APA StyleLhomme, S., DebRoy, S., Kamar, N., Abravanel, F., Metsu, D., Marion, O., Dimeglio, C., Cotler, S. J., Izopet, J., & Dahari, H. (2019). Plasma Hepatitis E Virus Kinetics in Solid Organ Transplant Patients Receiving Ribavirin. Viruses, 11(7), 630. https://doi.org/10.3390/v11070630







