High-Throughput Sequencing Assists Studies in Genomic Variability and Epidemiology of Little Cherry Virus 1 and 2 infecting Prunus spp. in Belgium
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material Collection and Samples Preparation
2.2. RT-PCR and Sequencing
2.3. Sequences and Phylogenetic Analysis
2.4. Total RNA Sequencing and Bioinformatics Analysis
3. Results
3.1. Incidence and Etiology of LChV-1 and LChV-2 in Prunus Trees Grown in Belgium
3.2. Diversity and Comparison of LChV-1 and LChV-2 Sequences Encoding RdRp and CP Gene
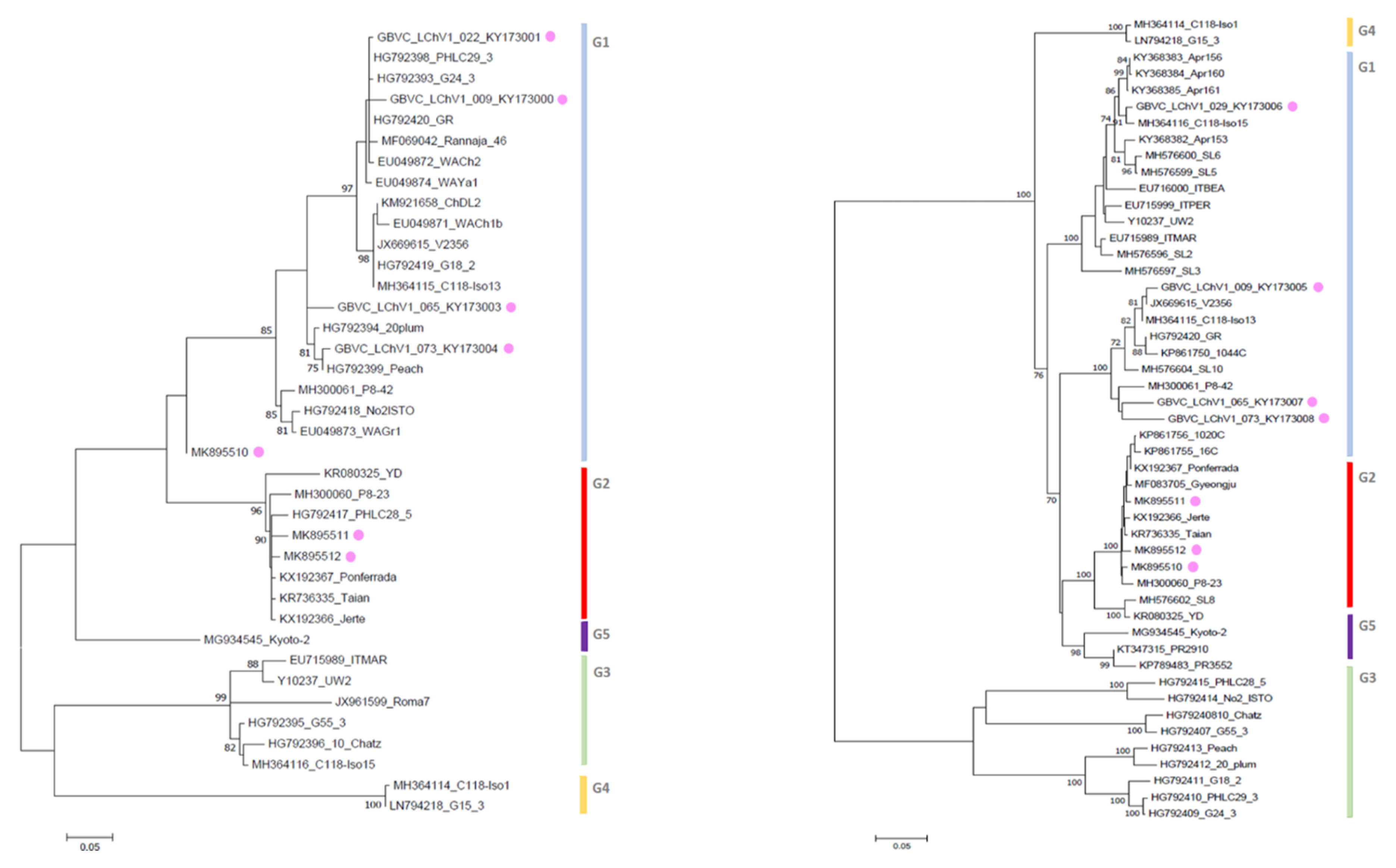
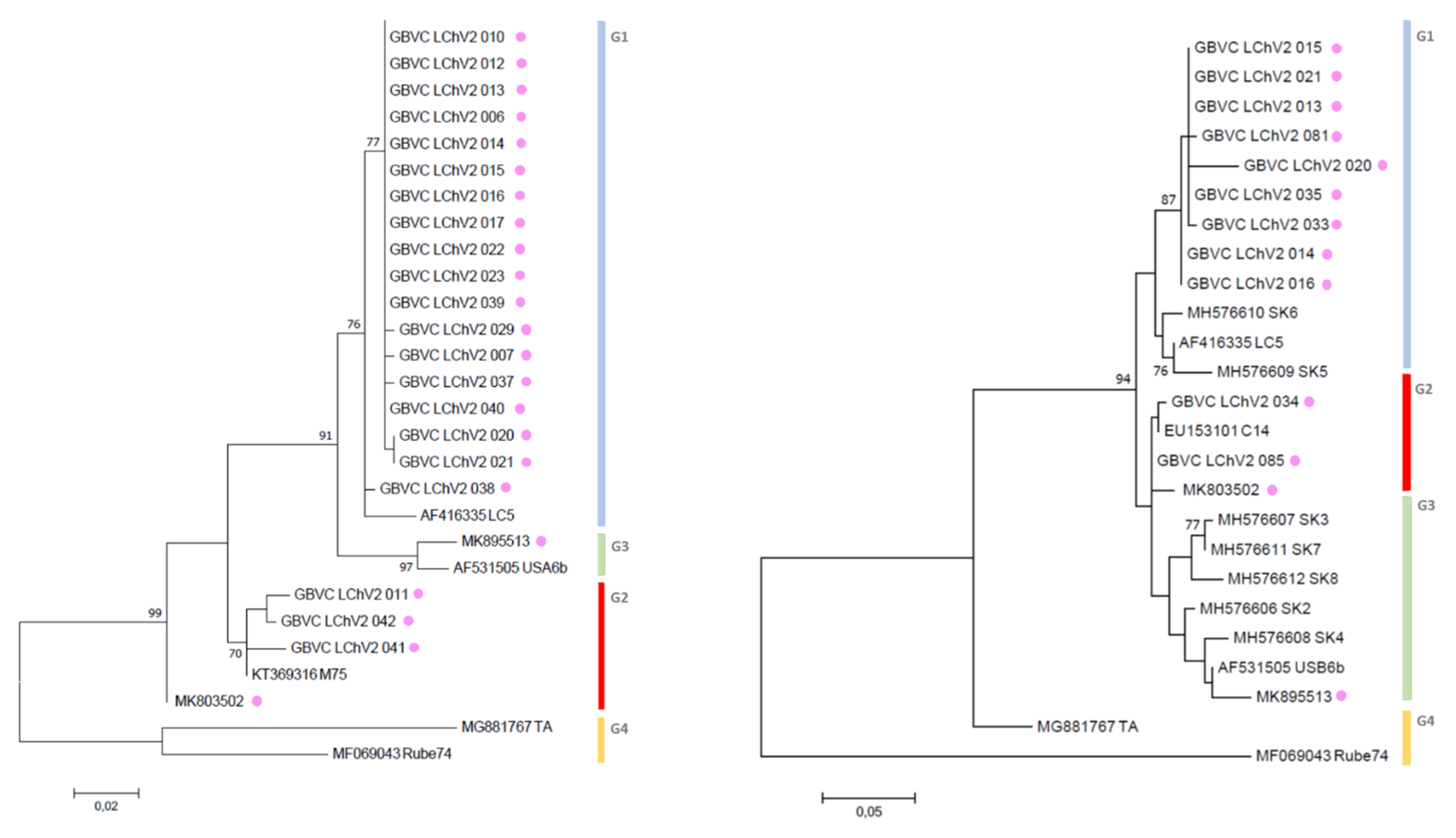
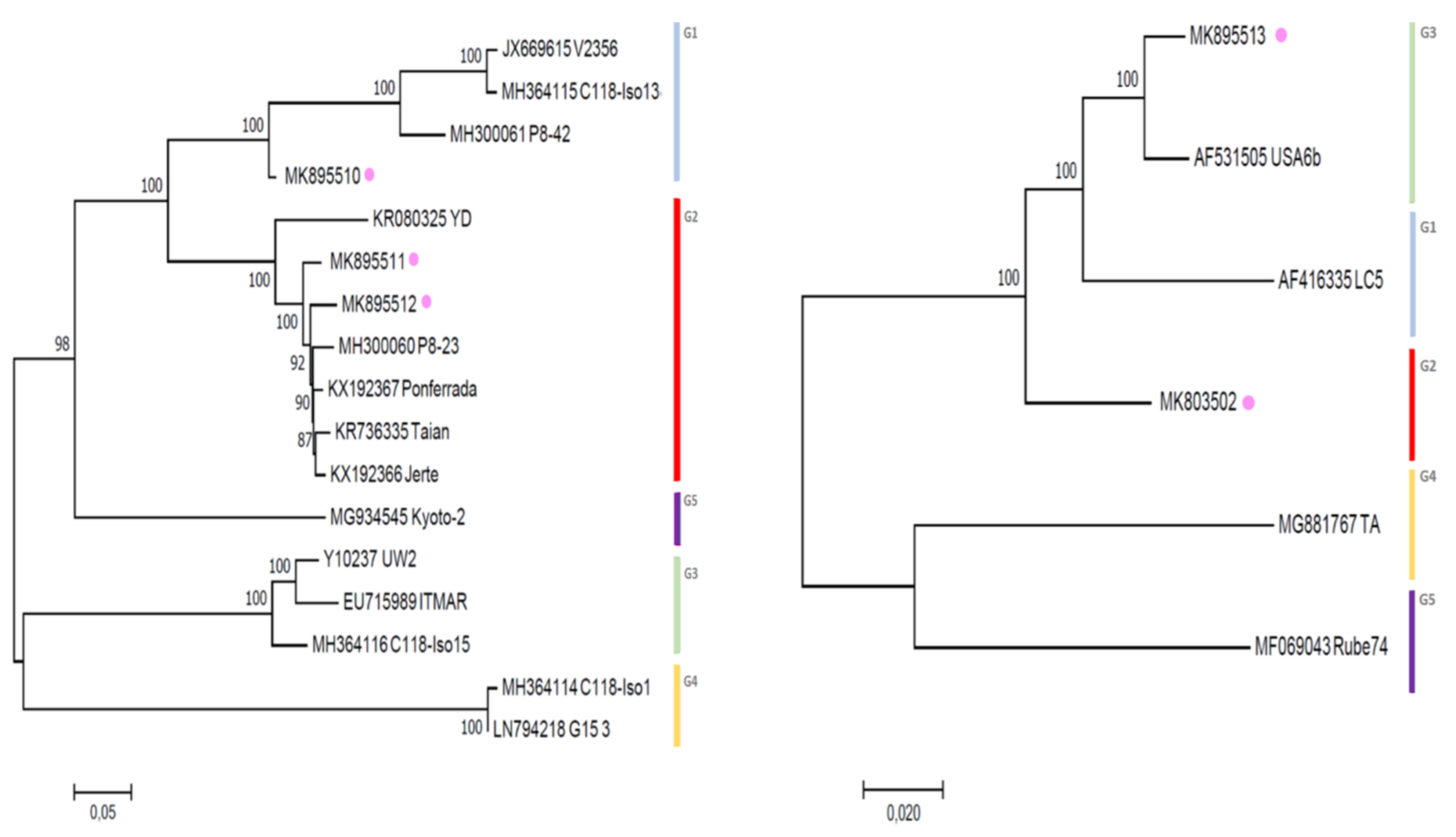
3.3. Phylogenetic Inference Reveals the Diversity of LChV-1 and LChV-2 in Belgium
3.4. Total RNA HTS also Revealed the Presence of CVA, Prune dwarf viruses (PDV) and PrVF in Belgium
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Al Rwahnih, M.; Dolja, V.V.; Daubert, S.; Koonin, E.V.; Rowhani, A. Genomic and biological analysis of Grapevine leafroll-associated virus 7 reveals a possible new genus within the family Closteroviridae. Virus Res. 2012, 163, 302–309. [Google Scholar] [CrossRef] [PubMed]
- Katsiani, A.T.; Maliogka, V.I.; Amoutzias, G.D.; Efthimiou, K.E.; Katis, N.I. Insights into the genetic diversity and evolution of Little cherry virus 1. Plant Pathol. 2015, 64, 817–824. [Google Scholar] [CrossRef]
- Jelkmann, W.; Fechter, B.; Agranovsky, A.A. Complete genome structure and phylogenetic analysis of Little cherry virus, a mealybug- transmissible Closterovirus. J. Gen. Virol. 1997, 78, 2067–2071. [Google Scholar] [CrossRef] [PubMed]
- Rott, M.E.; Jelkmann, W. Detection and partial characterization of a second Closterovirus associated with little cherry disease, Little cherry virus-2. Phytopathology 2001, 91, 261–267. [Google Scholar] [CrossRef] [PubMed]
- FAO. Food and Agriculture organization of the United Nations 2013. Available online: http://www.fao.org/statistics (accessed on 17 January 2019).
- Vavilov, N.I. The origin, variation and breeding of cultivated plants. Chronica Botanica 1951, 72, 482. [Google Scholar] [CrossRef]
- Webster, A.D.; Looney, N.E. The Taxonomic Classification of Sweet and Sour Cherries and a Brief History of Their Cultivation. In Cherries: Crop Physiology, Production and Uses, 1st ed.; CABI: Wallingford, CT, USA, 1996; pp. 3–24. ISBN 9780851989365. [Google Scholar]
- Jelkmann, W.; Eastwell, K.C. Little cherry viruses 1 and 2. In Virus and virus-like diseases of pome fruits and stone fruits; Barba, M., Candresse, T., Hadidi, A., Jelkmann, W., Eds.; APS Press: St. Paul, MN, USA, 2011; ISBN 978-0-89054-396-2. [Google Scholar]
- Raine, J.; McMullen, R.; Forbes, A. Transmission of the agent causing little cherry disease by the apple mealybug Phenacoccus aceris and the dodder Cuscuta lupuliformis. Can. J. Plant Pathol. 1986, 8, 6–11. [Google Scholar] [CrossRef]
- Isogai, M.; Aoyagi, J.; Nakagawa, M.; Kubodera, Y.; Satoh, K.; Katoh, T.; Inamori, M.; Yamashita, K.; Yoshikawa, N. Molecular detection of five cherry viruses from sweet cherry trees in Japan. J. Gen. Plant Pathol. 2004, 70, 288–291. [Google Scholar] [CrossRef]
- Komorowska, B.; Cieslinska, M. First Report of Cherry virus A and Little cherry virus 1 in Poland. Plant Dis. 2004, 88, 909. [Google Scholar] [CrossRef]
- Marais, A.; Faure, C.; Theil, S.; Svanella-Dumas, L.; Brans, Y.; Maurice, I.; Blin, V.; Candresse, T. First Report of Little cherry virus 1 on Plum in France. Plant Dis. 2016, 100, 2544. [Google Scholar] [CrossRef]
- Matic, S.; Myrta, A.; Minafra, A. First report of Little cherry virus 1 in cherry, plum, almond and peach in Italy. J. Plant Pathol. 2007, 8, S69–S76. [Google Scholar]
- Komorowska, B.; Cieslinska, M. First Report of Little cherry virus 2 from Sweet Cherry in Poland. Plant Dis. 2008, 92, 1366. [Google Scholar] [CrossRef] [PubMed]
- Rao, W.L.; Li, F.; Zuo, R.J.; Li, R. First report of Little cherry virus 2 in Flowering and Sweet Cherry trees in China. Plant Dis. 2011, 95, 1484. [Google Scholar] [CrossRef]
- Ludvikova, H.; Sucha, J. First Report of Little Cherry Disease from Sweet Cherry (Prunus avium) and Sour Cherry (P. cerasus) in the Czech Republic. Plant Dis. 2011, 95, 1197. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.G.; Gao, R.; Chen, R.R.; Chen, B.; Wu, Z.X.; Zhang, Z.X.; Li, S.F. First Report of Little cherry virus 1 in Sweet Cherry trees in China. Plant Dis. 2015, 99, 1191. [Google Scholar] [CrossRef]
- Voncina, D.; Simon, S.; Razov, J.; Leong, L. First Report of Little cherry virus 2 on Prunus cerasus var. Marasca in Croatia. J. Plant Pathol. 2016, 98, 171–185. [Google Scholar] [CrossRef]
- Ruiz-Garcia, A.B.; Martinez, C.; Santiago, R.; Garcia, M.T.; de Prado, N.; Olmos, A. First report of Little cherry virus 1 (LChV-1) in Sweet Cherry in Spain. Plant Dis. 2016, 100, 2340. [Google Scholar] [CrossRef]
- Safarova, D.; Faure, C.; Candresse, T.; Navratil, M.; Necas, T.; Marais, A. First Report of Little cherry virus 1 infecting Apricot in the Czech Republic. Plant Dis. 2017, 101, 845. [Google Scholar] [CrossRef]
- Fiore, N.; Fernandez, C.; Quiroga, N.; Pino, A.M.; Rivera, L.; Sagredo, K.; Zamorano, A. First report of Little cherry virus 1 in Chile. Plant Dis. 2018, 102, 3. [Google Scholar] [CrossRef]
- Bajet, N.B.; Unruh, T.R.; Druffel, K.L.; Eastwell, K.C. Occurrence of two little cherry viruses in sweet cherry in Washington State. Plant Dis. 2008, 92, 234–238. [Google Scholar] [CrossRef]
- Glasa, M.; Benediková, D.; Predajňa, L. First report of Little cherry virus 1 in Slovakia. J. Plant Pathol. 2015, 97, 542. [Google Scholar] [CrossRef]
- Serce, C.U.; Gazel, M.; Caglayan, K. First report of little cherry virus 1 in Turkey. J. Plant Pathol. 2011, 93, 66. [Google Scholar]
- Legrand, G.; Gilles, G.; Verhoyen, M. Little cherry disease in ornamental cherry trees in Belgium [description, diagnosis, viral disease, epidemiology, transmission by Macrosteles fascifrons, Phenacoccus aceris]. Fruit Belge 1982, 50, 125–131. [Google Scholar]
- Tahzima, R.; Foucart, Y.; Peusens, P.; Beliën, T.; Massart, S.; De Jonghe, K. First Report of Little cherry virus 1 affecting European Plum Prunus domestica (L.) in Belgium. Plant Dis. 2017, 101, 1557. [Google Scholar] [CrossRef]
- Matic, S.; Minafra, A.; Sanchez-Navarro, J.A.; Pallas, V.; Myrta, A.; Martelli, G.P. Kwanzan stunting syndrome: Detection and molecular characterization of an Italian isolate of Little cherry virus 1. Virus Res. 2009, 143, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhu, D.; Tan, Y.; Zong, X.; Wei, H.; Hammond, R.W.; Liu, Q. Complete nucleotide sequence of Little cherry virus 1 infecting sweet cherry in China. Arch. Virol. 2016, 161, 749–753. [Google Scholar] [CrossRef]
- Eastwell, K.C.; Bernardy, M.; Li, T.S. Comparison between woody indexing and a rapid hybridization assay for the diagnosis of little cherry disease in cherry trees. Ann. Appl. Biol. 1996, 128, 269–277. [Google Scholar] [CrossRef]
- Eastwell, K.C.; Bernardy, M.G. Partial characterization of a Closterovirus associated with apple mealybug-transmitted little cherry disease in North America. Phytopathology 2001, 91, 268–273. [Google Scholar] [CrossRef]
- Jelkmann, W.; Leible, S.; Rott, M. Little Cherry Closteroviruses -1 and -2, their genetic variability and detection by real-time-PCR. Acta Hortic. 2008, 781, 321–330. [Google Scholar] [CrossRef]
- Theilmann, J.; Mozafari, J.; Reade, R.; Wu, Z.; Xie, W.; Jesperson, G.; Bernardy, M.; Eastwell, K.C.; Rochon, D. Partial nucleotide sequence and genome organization of a Canadian isolate of Little cherry virus and development of an enzyme-linked immunosorbent assay-based diagnostic test. Phytopathology 2002, 92, 87–98. [Google Scholar] [CrossRef]
- Theilmann, J.; Orban, S.; Rocon, D. High Sequence Variability Among Little cherry virus Isolates Occurring in British Columbia. Plant Dis. 2004, 88, 1092–1098. [Google Scholar] [CrossRef]
- Candresse, T.; Marais, A.; Faure, C.; Gentit, P. Association of Little cherry virus 1 (LChV-1) with the Shirofugen Stunt Disease and Characterization of the Genome of a Divergent LChV-1 Isolate. Phytopathology 2013, 103, 293–298. [Google Scholar] [CrossRef] [PubMed]
- Zong, X.; Wang, W.; Wei, H.; Wang, J.; Yan, X.; Hammond, R.W.; Liu, Q. Incidence of sweet cherry viruses in Shandong province, China and a case study on multiple infection with five viruses. J. Plant Pathol. 2015, 97, 61–68. [Google Scholar] [CrossRef]
- Katsiani, A.; Maliogka, V.I.; Katis, N.I.; Svanella-Dumas, L.; Olmos, A.; Ruis-Garcia, A.B.; Marais, A.; Faure, C.; Theil, S.; Lotos, L.; et al. High-throughput sequencing reveals further diversity of Little cherry virus 1 with implication for diagnostics. Viruses 2018, 10, 385. [Google Scholar] [CrossRef] [PubMed]
- EFSA. Pest categorization of Little cherry pathogen (non-EU isolates). EFSA J. 2017, 15, 4926. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; MCWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. CLUSTALW and CLUSTALX version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Gao, S.; Padmanabhan, C.; Li, R.; Galvez, M.; Gutierrez, D.; Fuentes, S.; Ling, K.S.; Kreuze, J.; Fei, Z. VirusDetect: An automated pipeline for efficient virus discovery using deep sequencing of small RNAs. Virology 2017, 500, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Katsiani, A.; Pappi, P.; Olmos, A.; Efthimiou, K.E.; Maliogka, V.I.; Katis, N.I. Development of a real-time RT-PCR for the universal detection of Little cherry virus 1 and study of the seasonal fluctuation of the viral titer in sweet cherry cultivars. Plant Dis. 2018, 102, 899–904. [Google Scholar] [CrossRef]
- Villamor, D.E.V.; Pillai, S.S.; Eastwell, K.C. High throughput sequencing reveals a novel Fabavirus infecting sweet cherry. Arch. Virol. 2017, 162, 811. [Google Scholar] [CrossRef]
- Koloniuk, I.; Sarkisova, T.; Petrzik, K.; Lenz, O.; Pribylová, J.; Fránová, J.; Špak, J.; Lotos, L.; Beta, C.; Katsiani, A.; et al. Variability studies of two Prunus-infecting fabaviruses with the aid of high-throughput sequencing. Viruses 2018, 10, 204. [Google Scholar] [CrossRef]
- Baráth, D.; Jaksa-Czotter, N.; Molnár, J.; Varga, T.; Balássy, J.; Szabó, L.K.; Kirilla, Z.; Tusnády, G.E.; Preininger, É.; Várallyay, É. Small RNA NGS revealed the presence of Cherry virus A and Little cherry virus 1 on apricots in Hungary. Viruses 2018, 10, 318. [Google Scholar] [CrossRef] [PubMed]
- Tahzima, R.; Foucart, Y.; Peusens, P.; Beliën, T.; Massart, S.; De Jonghe, K. First Report of Little cherry virus 1 infecting Flowering Cherry Cultivars using High-Throughput Sequencing in Belgium. Plant Dis. In preparation.
- Bar-Joseph, M. Closteroviridae: The beginning. Frontiers Microbiol. 2014, 5, 1–2. [Google Scholar] [CrossRef]
- Karasev, A.V. Genetic diversity and evolution of Closteroviruses. Annu. Rev. Phytopathol. 2000, 38, 293–324. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, A.J.; Fargette, D.; Garcia-Arenal, F.; Gibbs, M.J. Time-The Emerging Dimension of Plant Virus Studies. J. Virol. Meth. 2010, 91, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Osman, F.; Al Rwahnih, M.; Golino, D.; Pitman, T.; Cordero, F.; Preece, J.E.; Rowhani, A. Evaluation of the phytosanitary status of the Prunus species in the national clonal germplasm repository in California: Survey of viruses and viroids. J. Plant Pathol. 2012, 94, 249–253. [Google Scholar] [CrossRef]
- Mekuria, T.A.; Smith, T.J.; Beers, E.; Watson, G.W.; Eastwell, K.C. First Report of Transmission of Little cherry virus 2 to Sweet Cherry by Pseudococcus maritimus (Ehrhorn) (Hemiptera: Pseudococcidae). Plant Dis. 2013, 97, 851. [Google Scholar] [CrossRef]
- Whitfield, A.E.; Falk, B.W.; Rotenberg, D. Insect vector-mediated transmission of plant viruses. Virology 2015, 479–480, 278–289. [Google Scholar] [CrossRef]
- Domingo, E.; Sheldon, J.; Perales, C. Viral Quasi-species Evolution. Microbiol. Mol. Biol. R. 2012, 76, 159–216. [Google Scholar] [CrossRef]
- Schneider, W.L.; Roossinck, M.J. Genetic variation in RNA virus quasispecies is controlled by host–virus interactions. J. Virol. 2001, 75, 6566–6571. [Google Scholar] [CrossRef]
- Garcia-Arenal, F.; Fraile, A.; Malpica, J.M. Variability and genetic structure of plant virus populations. Annu. Rev. Phytopathol. 2001, 39, 157–186. [Google Scholar] [CrossRef] [PubMed]
- Maliogka, V.I.; Dovas, C.I.; Katis, N.I. Evolutionary relationships of virus species belonging to a distinct lineage within the Ampelovirus genus. Virus Res. 2008, 135, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Martelli, G.P.; Agranovsky, A.A.; Bar-Joseph, M.; Boscia, D.; Candresse, T.; Coutts, R.H.A.; Dolja, V.V.; Hu, J.S.; Jelkmann, W.; Karasev, A.V.; et al. Family Closteroviridae. In Virus Taxonomy. Ninth Report of the International Committee on Taxonomy of Viruses; King, A., Adams, M.J., Carstens, E.B., Lefkowitz, E., Eds.; Elsevier-Academic Press: Amsterdam, The Netherlands, 2011; pp. 987–1001. ISBN 9780123846846. [Google Scholar]
- Dolja, V.V.; Kreuze, J.F.; Valkonen, J.P. Comparative and functional genomics of Closteroviruses. Virus Res. 2006, 117, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Lyu, M.D.; Li, X.M.; Guo, R.; Li, M.J.; Liu, X.M.; Wang, Q.; Cheng, Y.Q. Prevalence and distribution of Grapevine leafroll-associated virus 7 in China detected by an improved reverse transcription polymerase chain reaction assay. Plant Pathol. 2014, 63, 1168–1176. [Google Scholar] [CrossRef]
- Rowhani, A.; Uyemoto, K.K.; Golino, D.A.; Martelli, G.P. Pathogen testing and certification of Vitis and Prunus species. Annu. Rev. Phytopathol. 2005, 43, 261–278. [Google Scholar] [CrossRef]
- EPPO. Certification scheme for Cherry. EPPO Bull. 2001, 31, 447–461. [Google Scholar] [CrossRef]
- Massart, S.; Olmos, A.; Jijakli, H.; Candresse, T.; Olmos, A.; Jijakli, H.; Candresse, T. Current impact and future directions of high throughput sequencing in plant virus diagnostics. Virus Res. 2014, 188, 90–96. [Google Scholar] [CrossRef]
- Massart, S.; Candresse, T.; Gil, J.; Lacomme, C.; Predajna, L.; Ravnikar, M.; Reynard, J.S.; Rumbou, A.; Saldarelli, P.; Škorić, D.; et al. A framework for the evaluation of biosecurity, commercial, regulatory, and scientific impacts of plant viruses and viroids identified by NGS technologies. Front. Microbiol. 2017, 8, 45. [Google Scholar] [CrossRef]
- Eastwell, K.C.; Bernardy, M.G. Relationship of Cherry virus A to Little cherry virus disease in British Columbia. Acta Hortic. 1998, 472, 305–314. [Google Scholar] [CrossRef]
- Sabanadzovic, S.; Abou Ghanem-Sabanadzovic, N.; Rowhani, A.; Grant, A.; Uyemoto, K. Detection of Cherry virus A, cherry necrotic rusty mottle virus and Little cherry virus 1 in California orchards. Plant Pathol. 2005, 87, 173–177. [Google Scholar] [CrossRef]
- Bhuvaneshwar, K.; Song, L.; Madhavan, S.; Gusev, Y. viGEN: An Open Source Pipeline for the Detection and Quantification of Viral RNA in Human Tumors. Front. Microbiol. 2018, 9, 1172. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tahzima, R.; Foucart, Y.; Peusens, G.; Beliën, T.; Massart, S.; De Jonghe, K. High-Throughput Sequencing Assists Studies in Genomic Variability and Epidemiology of Little Cherry Virus 1 and 2 infecting Prunus spp. in Belgium. Viruses 2019, 11, 592. https://doi.org/10.3390/v11070592
Tahzima R, Foucart Y, Peusens G, Beliën T, Massart S, De Jonghe K. High-Throughput Sequencing Assists Studies in Genomic Variability and Epidemiology of Little Cherry Virus 1 and 2 infecting Prunus spp. in Belgium. Viruses. 2019; 11(7):592. https://doi.org/10.3390/v11070592
Chicago/Turabian StyleTahzima, Rachid, Yoika Foucart, Gertie Peusens, Tim Beliën, Sébastien Massart, and Kris De Jonghe. 2019. "High-Throughput Sequencing Assists Studies in Genomic Variability and Epidemiology of Little Cherry Virus 1 and 2 infecting Prunus spp. in Belgium" Viruses 11, no. 7: 592. https://doi.org/10.3390/v11070592
APA StyleTahzima, R., Foucart, Y., Peusens, G., Beliën, T., Massart, S., & De Jonghe, K. (2019). High-Throughput Sequencing Assists Studies in Genomic Variability and Epidemiology of Little Cherry Virus 1 and 2 infecting Prunus spp. in Belgium. Viruses, 11(7), 592. https://doi.org/10.3390/v11070592





