Discovery and Characterization of Bukakata orbivirus (Reoviridae:Orbivirus), a Novel Virus from a Ugandan Bat
Abstract
:1. Introduction
2. Materials and Methods
2.1. Viruses and Cells
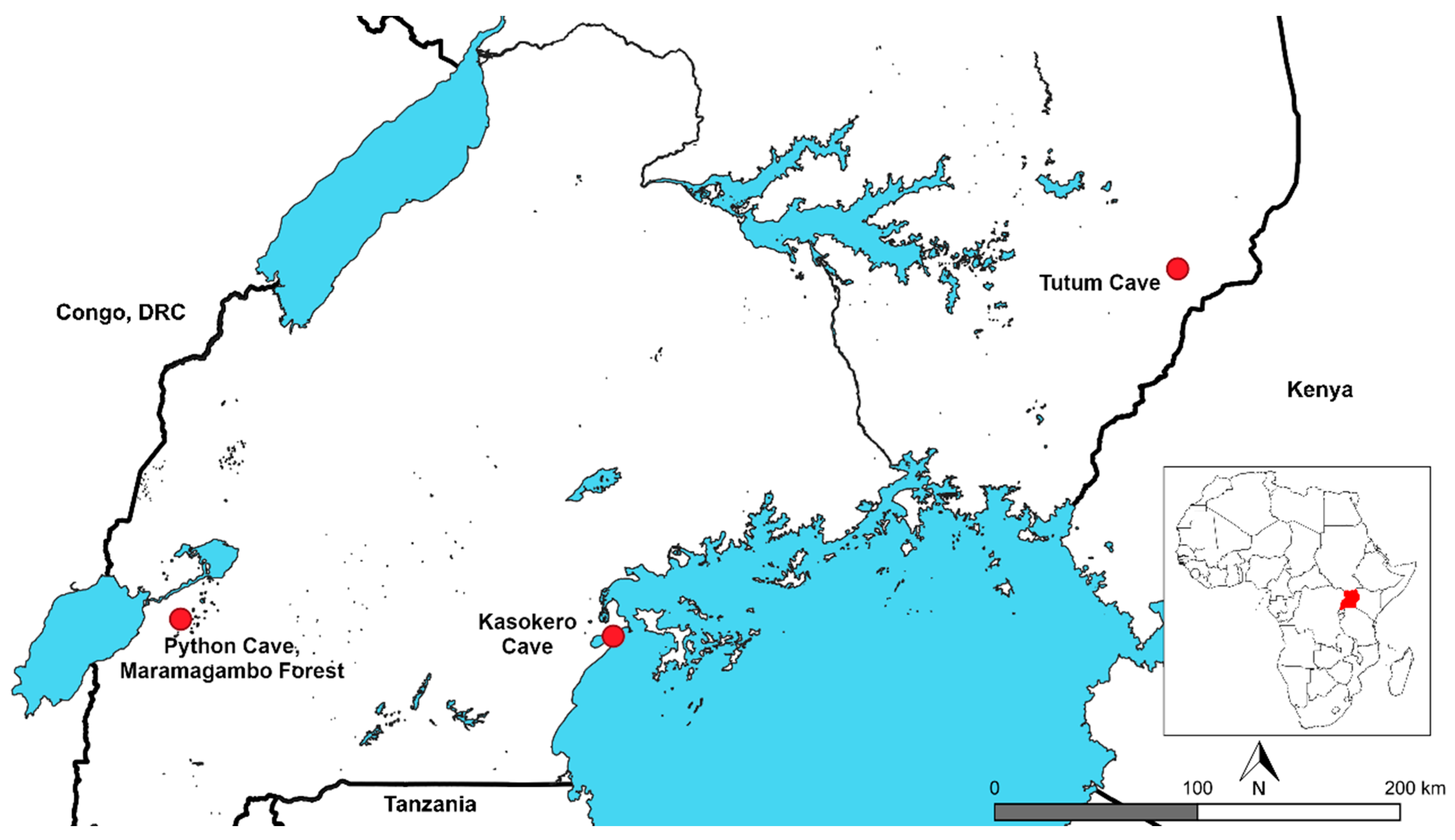
2.2. Bat Capture and Sampling
2.3. Sequencing and Bioinformatics Analysis
2.4. Phylogenetic Analysis
2.5. Multi-Step Growth Curves
2.6. Screening of Archived Field Samples for BUKV
3. Results
3.1. Virus Isolation
3.2. Genome Sequencing
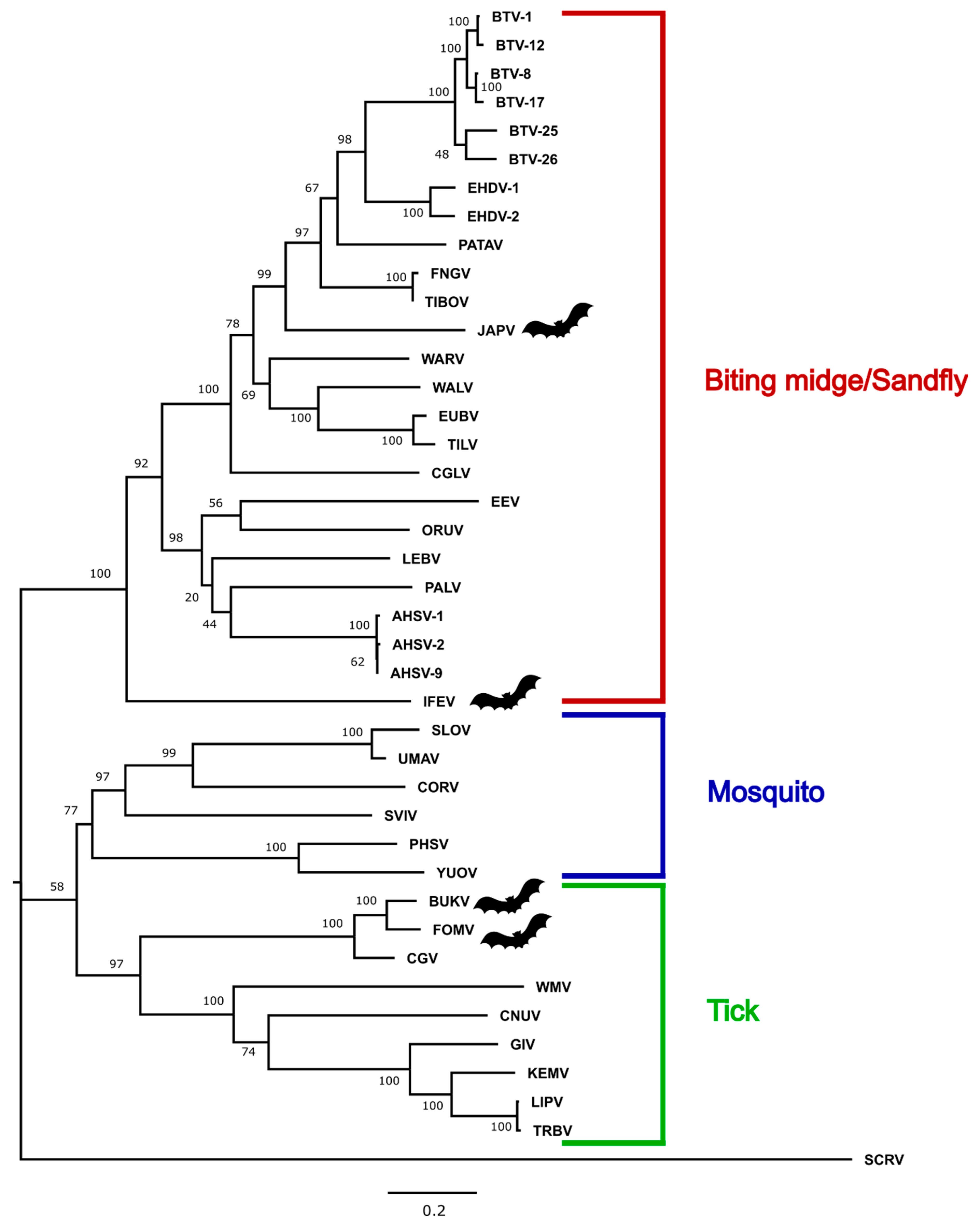
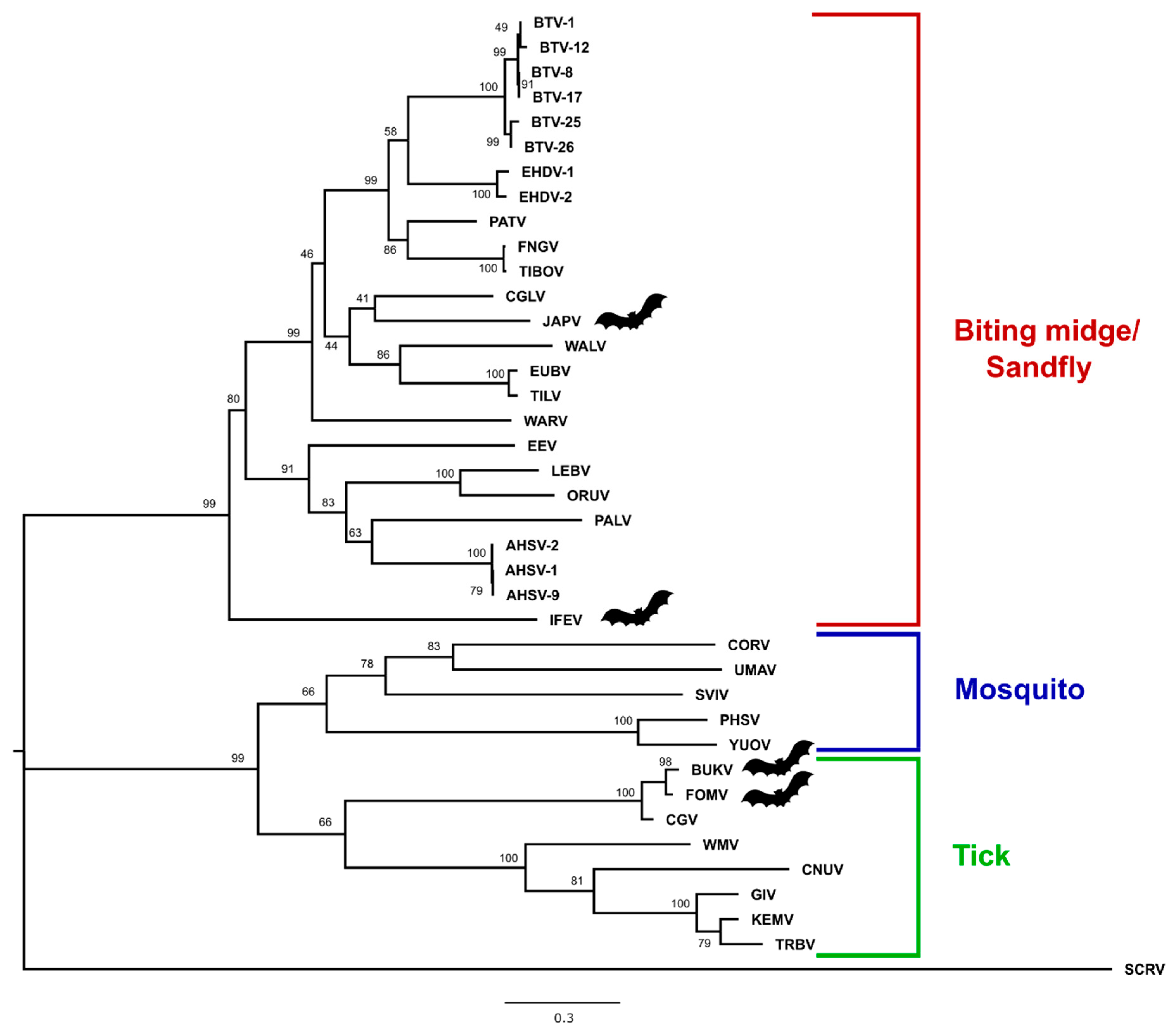
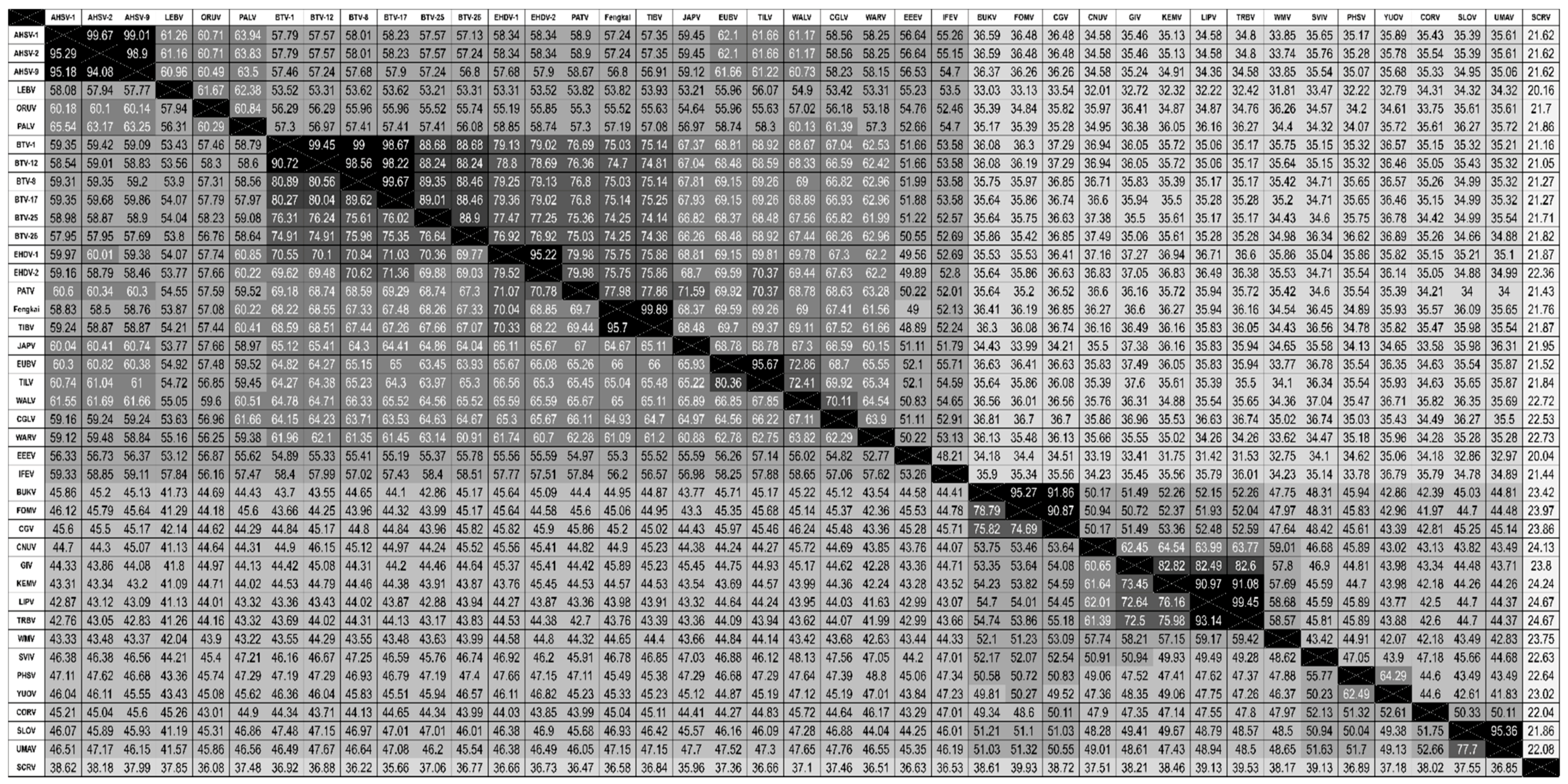
3.3. Phylogenetic Analysis
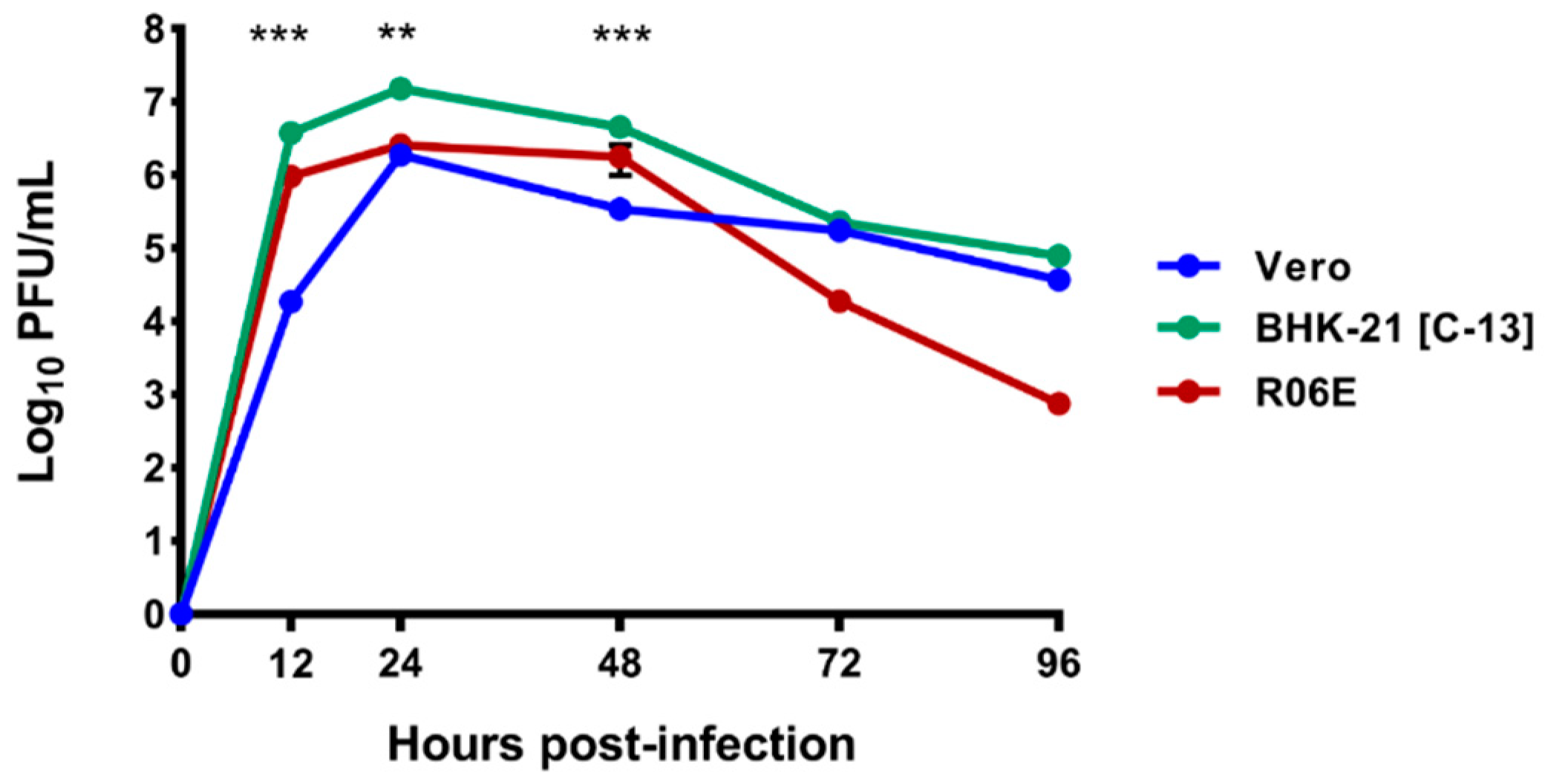
3.4. Growth Curves
3.5. Testing of Additional Bat and Tick Samples
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef] [PubMed]
- Kading, R.C.; Kityo, R.M.; Mossel, E.C.; Borland, E.M.; Nakayiki, T.; Nalikka, B.; Nyakarahuka, L.; Ledermann, J.P.; Panella, N.A.; Gilbert, A.T. Neutralizing antibodies against flaviviruses, Babanki virus, and Rift Valley fever virus in Ugandan bats. Infect. Ecol. Epidemiol. 2018, 8, 1439215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Attoui, H.; Mertens, P.; Becnel, J.; Belaganahalli, S.; Bergoin, M.; Brussaard, C.; Chappell, J.; Ciarlet, M.; del Vas, M.; Dermody, T. Orbiviruses, Reoviridae. Virus Taxon. Ninth Rep. Int. Comm. Taxon. Viruses 2011, 592, 603. [Google Scholar]
- Harrison, J.; Warrilow, D.; McLean, B.; Watterson, D.; O’Brien, C.; Colmant, A.; Johansen, C.; Barnard, R.; Hall-Mendelin, S.; Davis, S. A new orbivirus isolated from mosquitoes in North-Western Australia shows antigenic and genetic similarity to Corriparta virus but does not replicate in vertebrate cells. Viruses 2016, 8, 141. [Google Scholar] [CrossRef] [PubMed]
- Libikova, H.; Heinz, F.; Ujhazyova, D.; Stünzner, D. Orbiviruses of the Kemerovo complex and neurological diseases. Med. Microbiol. Immunol. 1978, 166, 255–263. [Google Scholar] [CrossRef] [PubMed]
- Malkova, D.; Holubova, J.; Kolman, J.; Marhoul, Z.; Hanzal, F.; Kulkova, H.; Markvart, K.; Simkova, L. Antibodies against some arboviruses in persons with various neuropathies. Acta Virol. 1980, 24, 298. [Google Scholar] [PubMed]
- Mohd Jaafar, F.; Belhouchet, M.; Belaganahalli, M.; Tesh, R.B.; Mertens, P.P.C.; Attoui, H. Full-Genome Characterisation of Orungo, Lebombo and Changuinola Viruses Provides Evidence for Co-Evolution of Orbiviruses with Their Arthropod Vectors. PLoS ONE 2014, 9, e86392. [Google Scholar] [CrossRef] [PubMed]
- MacLachlan, N.J.; Guthrie, A.J. Re-emergence of bluetongue, African horse sickness, and other orbivirus diseases. Vet. Res. 2010, 41, 35. [Google Scholar] [CrossRef] [PubMed]
- Schnagl, R.D.; Holmes, I.H. Electron microscopy of Japanaut and Tilligerry viruses: Two proposed members of the orbivirus group. Aust. J. Biol. Sci. 1975, 28, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Arbovirus Catalog: Japanaut. Available online: https://wwwn.cdc.gov/arbocat/VirusDetails.aspx?ID=207 (accessed on 19 September 2017).
- Miura, T.; Kitaoka, M. Viruses isolated from bats in Japan. Arch. Virol. 1977, 53, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Krishnamurthy, S.; Cai, Z.; Popov, V.L.; Travassos da Rosa, A.P.; Guzman, H.; Cao, S.; Virgin, H.W.; Tesh, R.B.; Wang, D. Identification of Novel Viruses Using VirusHunter—An Automated Data Analysis Pipeline. PLoS ONE 2013, 8, e78470. [Google Scholar] [CrossRef] [PubMed]
- Kemp, G.; Le, G.G.; Karabatsos, N.; Rickenbach, A.; Cropp, C. IFE: A new African orbivirus isolated from Eidolon helvum bats captured in Nigeria, Cameroon and the Central African Republic. Bull. de la Soc. de Pathol. Exot. et de Ses Fil. 1988, 81, 40–48. [Google Scholar]
- Ezeifeka, G.; Umoh, J.; Ezeokoli, C.; Ezealor, A. Prevalence of Ife virus infection in wild rodents and birds from Zaria, Nigeria. J. Wildl. Dis. 1987, 23, 663–665. [Google Scholar] [CrossRef] [PubMed]
- Ezeifeka, G.; Umoh, J.; Ezeokoli, C.; Gomwalk, N. Serological evidence of Ife virus infection in Nigerian indigenous domestic ruminants. Trop. Anim. Health Prod. 1989, 21, 55–57. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Arbovirus Catalog: Ife. Available online: https://wwwn.cdc.gov/arbocat/VirusDetails.aspx?ID=187&SID=4 (accessed on 28 September 2017).
- Butenko, A. Arbovirus circulation in the Republic of Guinea. Med. Parazitol. i Parazit. Bolezn. 1996, 2, 40–45. [Google Scholar]
- Konstantinov, O.; Diallo, S.; Inapogi, A.; Ba, A.; Kamara, S. The mammals of Guinea as reservoirs and carriers of arboviruses. Med. Parazitol. i Parazit. Bolezn. 2006, 1, 34–39. [Google Scholar]
- Boiro, I.; Fidarov, F.; Lomonossov, N.; Linev, M.; Bachkirsov, V.; Inapogui, A. Isolation of the Fomédé virus from Chiroptera, Nycteris nana, in the Republic of Guinea. Bull. de la Soc. de Pathol. Exot. et de Ses Fil. 1986, 79, 180–182. [Google Scholar]
- Justines, G.; Kuns, M. Matucare virus, a new agent recovered from Ornithodoros (Alectorobius) boliviensis. Am. J. Trop. Med. Hyg. 1970, 19, 697–702. [Google Scholar] [CrossRef] [PubMed]
- Attoui, H.; Mendez-Lopez, M.R.; Rao, S.; Hurtado-Alendes, A.; Lizaraso-Caparo, F.; Jaafar, F.M.; Samuel, A.R.; Belhouchet, M.; Pritchard, L.I.; Melville, L.; et al. Peruvian horse sickness virus and Yunnan orbivirus, isolated from vertebrates and mosquitoes in Peru and Australia. Virology 2009, 394, 298–310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jordan, I.; Horn, D.; Oehmke, S.; Leendertz, F.H.; Sandig, V. Cell lines from the Egyptian fruit bat are permissive for modified vaccinia Ankara. Virus Res. 2009, 145, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, N.V.; Zemlak, T.S.; Hanner, R.H.; Hebert, P.D. Universal primer cocktails for fish DNA barcoding. Mol. Ecol. Notes 2007, 7, 544–548. [Google Scholar] [CrossRef]
- Messing, J. New M13 vectors for cloning. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 1983; Volume 101, pp. 20–78. [Google Scholar]
- Towner, J.S.; Amman, B.R.; Sealy, T.K.; Carroll, S.A.R.; Comer, J.A.; Kemp, A.; Swanepoel, R.; Paddock, C.D.; Balinandi, S.; Khristova, M.L. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Pathog. 2009, 5, e1000536. [Google Scholar] [CrossRef] [PubMed]
- Miller, B.R.; Mitchell, C.J.; Ballinger, M.E. Replication, tissue tropisms and transmission of yellow fever virus in Aedes albopictus. Trans. R. Soc. Trop. Med. Hyg. 1989, 83, 252–255. [Google Scholar] [CrossRef]
- Kading, R.C.; Gilbert, A.T.; Mossel, E.C.; Crabtree, M.B.; Kuzmin, I.V.; Niezgoda, M.; Agwanda, B.; Markotter, W.; Weil, M.R.; Montgomery, J.M. Isolation and molecular characterization of Fikirini rhabdovirus, a novel virus from a Kenyan bat. J. Gen. Virol. 2013, 94, 2393–2398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Belhouchet, M.; Jaafar, F.M.; Tesh, R.; Grimes, J.; Maan, S.; Mertens, P.P.; Attoui, H. Complete sequence of Great Island virus and comparison with the T2 and outer-capsid proteins of Kemerovo, Lipovnik and Tribec viruses (genus Orbivirus, family Reoviridae). J. Gen. Virol. 2010, 91, 2985–2993. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belaganahalli, M.N.; Maan, S.; Maan, N.S.; Tesh, R.; Attoui, H.; Mertens, P.P. Umatilla virus genome sequencing and phylogenetic analysis: Identification of stretch lagoon orbivirus as a new member of the Umatilla virus species. PLoS ONE 2011, 6, e23605. [Google Scholar] [CrossRef] [PubMed]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Le, S.Q.; Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 2008, 25, 1307–1320. [Google Scholar] [CrossRef] [PubMed]
- Beaty, B.; Calisher, C.; Shope, R.; Schmidt, N.; Emmons, R. Arboviruses. In Diagnostic Procedures for Viral, Rickettsial and Chlamydial Infections, 6th ed.; American Public Health Association: Washington, DC, USA, 1989; pp. 797–855. [Google Scholar]
- Schuh, A.J.; Amman, B.R.; Apanaskevich, D.A.; Sealy, T.K.; Nichol, S.T.; Towner, J.S. No evidence for the involvement of the argasid tick Ornithodoros faini in the enzootic maintenance of marburgvirus within Egyptian rousette bats (Rousettus aegyptiacus). Parasites Vectors 2016, 9, 128. [Google Scholar] [CrossRef] [PubMed]
- Belhouchet, M.; Mohd Jaafar, F.; Firth, A.E.; Grimes, J.M.; Mertens, P.P.C.; Attoui, H. Detection of a fourth orbivirus non-structural protein. PLoS ONE 2011, 6, e25697. [Google Scholar] [CrossRef] [PubMed]
- Van Dijk, A.A.; Huismans, H. In vitro transcription and translation of bluetongue virus mRNA. J. Gen. Virol. 1988, 69, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Zeller, H.; Karabatsos, N.; Calisher, C.; Digoutte, J.-P.; Cropp, C.; Murphy, F.; Shope, R. Electron microscopic and antigenic studies of uncharacterized viruses. III. Evidence suggesting the placement of viruses in the familyReoviridae. Arch. Virol. 1989, 109, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Karabatsos, N. International Catalogue of Arthropod-Borne Viruses, 3rd ed.; American Society of Tropical Medicine and Hygiene: San Antonio, TX, USA, 1985. [Google Scholar]
- Centers for Disease Control and Prevention. Arbovirus Catalog: Chobar Gorge. Available online: https://wwwn.cdc.gov/arbocat/VirusDetails.aspx?ID=111&SID=5 (accessed on 29 September 2017).
- Attoui, H.; Stirling, J.M.; Munderloh, U.G.; Billoir, F.; Brookes, S.M.; Burroughs, J.N.; de Micco, P.; Mertens, P.P.; de Lamballerie, X. Complete sequence characterization of the genome of the St Croix River virus, a new orbivirus isolated from cells of Ixodes scapularis. J. Gen. Virol. 2001, 82, 795–804. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shaw, A.E.; Ratinier, M.; Nunes, S.F.; Nomikou, K.; Caporale, M.; Golder, M.; Allan, K.; Hamers, C.; Hudelet, P.; Zientara, S. Reassortment between two serologically unrelated bluetongue virus strains is flexible and can involve any genome segment. J. Virol. 2013, 87, 543–557. [Google Scholar] [CrossRef] [PubMed]
- Nuttall, P.A.; Moss, S.R. Genetic reassortment indicates a new grouping for tick-borne orbiviruses. Virology 1989, 171, 156–161. [Google Scholar] [CrossRef]
- Hölzer, M.; Krähling, V.; Amman, F.; Barth, E.; Bernhart, S.H.; Carmelo, V.A.; Collatz, M.; Doose, G.; Eggenhofer, F.; Ewald, J. Differential transcriptional responses to Ebola and Marburg virus infection in bat and human cells. Sci. Rep. 2016, 6, 34589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perry, A.K.; Gang, C.; Zheng, D.; Hong, T.; Cheng, G. The host type I interferon response to viral and bacterial infections. Cell Res. 2005, 15, 407. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Tachedjian, M.; Wynne, J.W.; Boyd, V.; Cui, J.; Smith, I.; Cowled, C.; Ng, J.H.; Mok, L.; Michalski, W.P. Contraction of the type I IFN locus and unusual constitutive expression of IFN-α in bats. Proc. Natl. Acad. Sci. USA 2016, 113, 2696–2701. [Google Scholar] [CrossRef] [PubMed]
- Pavlovich, S.S.; Lovett, S.P.; Koroleva, G.; Guito, J.C.; Arnold, C.E.; Nagle, E.R.; Kulcsar, K.; Lee, A.; Thibaud-Nissen, F.; Hume, A.J. The Egyptian rousette genome reveals unexpected features of bat antiviral immunity. Cell 2018, 173, 1098–1110.e1018. [Google Scholar] [CrossRef] [PubMed]

| Location | Latitude | Longitude | n | Tissue Type | Date Collected |
|---|---|---|---|---|---|
| Python Cave, Maramagambo Forest | −0.26667 | 30.05000 | 71 | Liver and spleen | Nov 09 |
| Kasokero cave | −0.34214 | 31.96627 | 56 | Spleen | Jan 13 |
| Tutum cave | 1.28333 | 34.46667 | 45 | Spleen | Feb 12 |
| Seg. | Protein Encoded | ORF Length (nt) | Pred. Protein Length | %GC Content (ORF) | Top Blastx (ORF) [Nearest Virus 1], Accession, % Pairwise Identity | Accession # |
|---|---|---|---|---|---|---|
| 1 | VP1 | 3855 | 1285 | 51.9 | VP1 (Pol) [CGV], YP_009158901, 83.6% | MK359215 |
| 2 | VP2 (T2) | 2730 | 2730 | 52.3 | VP2 (T2) [CGV], YP_009158902, 91.7% | MK359216 |
| 3 | VP3 (Cap) | N/A 2 | N/A 2 | N/A 2 | VP3 (Cap) [CGV], YP)009158903, 74.5% | MK359217 |
| 4 | VP4 (OC1) | 1761 | 587 | 50.9 | VP4 (OC1) [CGV], YP_009158904, 60.1% | MK359218 |
| 5 | NS1 (TuP) | 1575 | 525 | 55.3 | NS1 (TuP) [CGV], YP_009158905, 80.2% | MK359219 |
| 6 | VP5 (OC2) | 1608 | 536 | 53.7 | VP5 (OC2) [CGV], YP_009158906, 79.8% | MK359220 |
| 7 | VP7 (T13) | 1068 | 356 | 54.1 | VP7 (T13) [CGV], YP_009158907, 88.2% | MK359222 |
| 8 | NS2 (ViP) | 1110 | 370 | 54.6 | NS2 (ViP) [CGV], YP_009158908, 76.7% | MK359221 |
| 9 | VP6 (Hel) | 1041 | 347 | 53.2 | VP6 (Hel) [CGV], YP_009158909, 54.3% | MK359223 |
| NS4 | 717 | 239 | 54.4 | VP6 (Hel) [CGV], YP_009158909, 54.3% | ||
| 10 | NS3 | 621 | 207 | 52.3 | NS3 [CGV], YP_009158911, 90.3% | MK359224 |
| NS3a | 579 | 193 | 52.0 | NS3a [CGV], YP_009158912, 90.1% |
| Seg. | Protein Encoded | ORF Length (nt) | Pred. Protein Length | %GC Content (ORF) | Top Blastx (ORF) [Nearest Virus 1], Accession, % Pairwise Identity | Accession # |
|---|---|---|---|---|---|---|
| 1 | VP1 | 3855 | 1285 | 52.7 | VP1 (Pol) [CGV], YP_009158901, 82.8% | MK359225 |
| 2 | VP2 (T2) | 2730 | 910 | 50.9 | VP2 (T2) [CGV], YP_009158902, 90.9% | MK359226 |
| 3 | VP3 (Cap) | 1908 | 636 | 53.7 | VP3 (Cap) [CGV], YP_009158903, 71.4% | MK359227 |
| 4 | VP4 (OC1) | 1761 | 587 | 50.3 | VP4 (OC1) [CGV], YP_009158904, 69.3% | MK359228 |
| 5 | NS1 (TuP) | 1575 | 525 | 55.7 | NS1 (TuP) [CGV], YP_009158905, 78.6% | MK359229 |
| 6 | VP5 (OC2) | 1608 | 536 | 54.7 | VP5 (OC2) [CGV], YP_009158906, 83.2% | MK359230 |
| 7 | VP7 (T13) | 1068 | 356 | 53.2 | VP7 (T13) [CGV], YP_009158907, 89.0% | MK359232 |
| 8 | NS2 (ViP) | 1110 | 370 | 56.0 | NS2 (ViP) [CGV], YP_009158908, 77.2% | MK359231 |
| 9 | VP6 (Hel) | 1041 | 347 | 55.1 | N/A 2 | MK359233 |
| NS4 | 717 | 239 | 55.2 | NS4 [CGV], YP_009158910, 60.2% | ||
| 10 | NS3 | 621 | 207 | 53.9 | NS3 [CGV], YP_009158911, 88.8% | MK359234 |
| NS3a | 570 | 190 | 53.5 | NS3a [CGV], YP_009158912, 89.4% |
| Seg. | Protein Encoded | ORF Length (nt) | Pred. Protein Length | %GC Content (ORF) | Top Blastx (ORF) [Nearest Virus 1], Accession, % Pairwise Identity | Accession # |
|---|---|---|---|---|---|---|
| 1 | VP1 | 3900 | 1300 | 38.3 | VP1 [TIBOV], APT68074, 66.3% | MK359235 |
| 2 | VP2 (OC1) | 3636 | 1212 | 39.2 | VP2 (OC1) [BTV-2], CAO79540, 22.1% | MK359236 |
| 3 | VP3 (T2) | 2703 | 901 | 40.9 | VP3 (T2) [PATV], AFH41521, 72.0% | MK359237 |
| 4 | VP4 (Cap) | 1932 | 644 | 40.7 | VP4 (Cap) [BTV-12], ASV51737, 57.6% | MK359238 |
| 5 | NS1 (TuP) | 1662 | 554 | 44.8 | NS1 (TuP) [EHDV-7], AIY25176, 30.3% | MK359239 |
| 6 | VP5 (OC2) | 1590 | 530 | 43.1 | VP5 (OC2) [CGLV], AGZ91948, 54.0% | MK359240 |
| 7 | VP7 (T13) | 1053 | 351 | 44.5 | VP7 (T13) [CGLV], YP_008719923, 57.1% | MK359241 |
| 8 | NS2 (ViP) | 1011 | 337 | 43.8 | NS2 (ViP) [CGLV], AGZ91980, 42.6% | MK359242 |
| 9 | VP6 (Hel) | 804 | 268 | 44.7 | VP6 (Hel) [WALV], YP_008658421, 32.7% | MK359243 |
| NS4 | 234 | 78 | 48.7 | N/A 2 | ||
| 10 | NS3 | 786 | 262 | 43.6 | NS3 [TILV], AFH41508, 39.1% | MK359244 |
| NS3a | 642 | 214 | 44.5 | NS3 [TILV], AFH41508, 39.1% |
| Seg. | Protein Encoded | ORF Length (nt) | Pred. Protein Length | %GC (ORF) | Top Blastx (ORF) [Nearest Virus 1], Accession, % Pairwise Identity | Accession # |
|---|---|---|---|---|---|---|
| 1 | VP1 | 3900 | 1300 | 39.7 | VP1 [Heramatsu], AGZ62525, 62.0% | MK359245 |
| 2 | VP3 (T2) | 2685 | 895 | 42.6 | VP3 (T2) [Heramatsu], AGZ62528, 61.9% | MK359246 |
| 3 | VP2 (OC1) | 2523 | 841 | 41.7 | VP2 (OC1) [Heramatsu], AGZ62527, 29.1% | MK359247 |
| 4 | VP4 (Cap) | 1902 | 634 | 42.0 | VP4 (Cap) partial [Heramatsu], AGZ62529, 52.7% | MK359248 |
| 5 | NS1 (TuP) | 1581 | 527 | 44.7 | NS1 (TuP) [LEBV], YP_009507714, 32.0% | MK359249 |
| 6 | VP5 (OC2) | 1569 | 523 | 43.0 | VP5 (OC2) [CGLV], AGZ91955, 46.8% | MK359250 |
| 7 | VP7 (T13) | 1047 | 349 | 46.2 | VP7 (T13) [WALV], YP_008658420, 41.7% | MK359251 |
| 8 | NS2 (ViP) | 990 | 330 | 44.1 | NS2 (ViP) [Heramatsu], AGZ62533, 40.5% | MK359252 |
| 9 | VP6 (Hel) | 768 | 256 | 47.5 | VP6 (Hel) partial [Heramatsu], AGZ62534, 34.7% | MK359253 |
| NS4 | 240 | 80 | 53.8 | N/A 2 | ||
| 10 | NS3 | 612 | 204 | 46.6 | NS3 [Heramatsu], AGZ62526, 47.1% | MK359254 |
| NS3a | 561 | 187 | 46.3 | NS3 [Heramatsu], AGZ62526, 47.1% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fagre, A.C.; Lee, J.S.; Kityo, R.M.; Bergren, N.A.; Mossel, E.C.; Nakayiki, T.; Nalikka, B.; Nyakarahuka, L.; Gilbert, A.T.; Peterhans, J.K.; et al. Discovery and Characterization of Bukakata orbivirus (Reoviridae:Orbivirus), a Novel Virus from a Ugandan Bat. Viruses 2019, 11, 209. https://doi.org/10.3390/v11030209
Fagre AC, Lee JS, Kityo RM, Bergren NA, Mossel EC, Nakayiki T, Nalikka B, Nyakarahuka L, Gilbert AT, Peterhans JK, et al. Discovery and Characterization of Bukakata orbivirus (Reoviridae:Orbivirus), a Novel Virus from a Ugandan Bat. Viruses. 2019; 11(3):209. https://doi.org/10.3390/v11030209
Chicago/Turabian StyleFagre, Anna C., Justin S. Lee, Robert M. Kityo, Nicholas A. Bergren, Eric C. Mossel, Teddy Nakayiki, Betty Nalikka, Luke Nyakarahuka, Amy T. Gilbert, Julian Kerbis Peterhans, and et al. 2019. "Discovery and Characterization of Bukakata orbivirus (Reoviridae:Orbivirus), a Novel Virus from a Ugandan Bat" Viruses 11, no. 3: 209. https://doi.org/10.3390/v11030209
APA StyleFagre, A. C., Lee, J. S., Kityo, R. M., Bergren, N. A., Mossel, E. C., Nakayiki, T., Nalikka, B., Nyakarahuka, L., Gilbert, A. T., Peterhans, J. K., Crabtree, M. B., Towner, J. S., Amman, B. R., Sealy, T. K., Schuh, A. J., Nichol, S. T., Lutwama, J. J., Miller, B. R., & Kading, R. C. (2019). Discovery and Characterization of Bukakata orbivirus (Reoviridae:Orbivirus), a Novel Virus from a Ugandan Bat. Viruses, 11(3), 209. https://doi.org/10.3390/v11030209







