Low Temperature and Low UV Indexes Correlated with Peaks of Influenza Virus Activity in Northern Europe during 2010–2018
Abstract
1. Introduction
2. Materials and Methods
2.1. The “NorthernFlu” Consortium
2.2. Data Collection
2.3. Statistical Analysis
2.4. Experimental Validation
3. Results
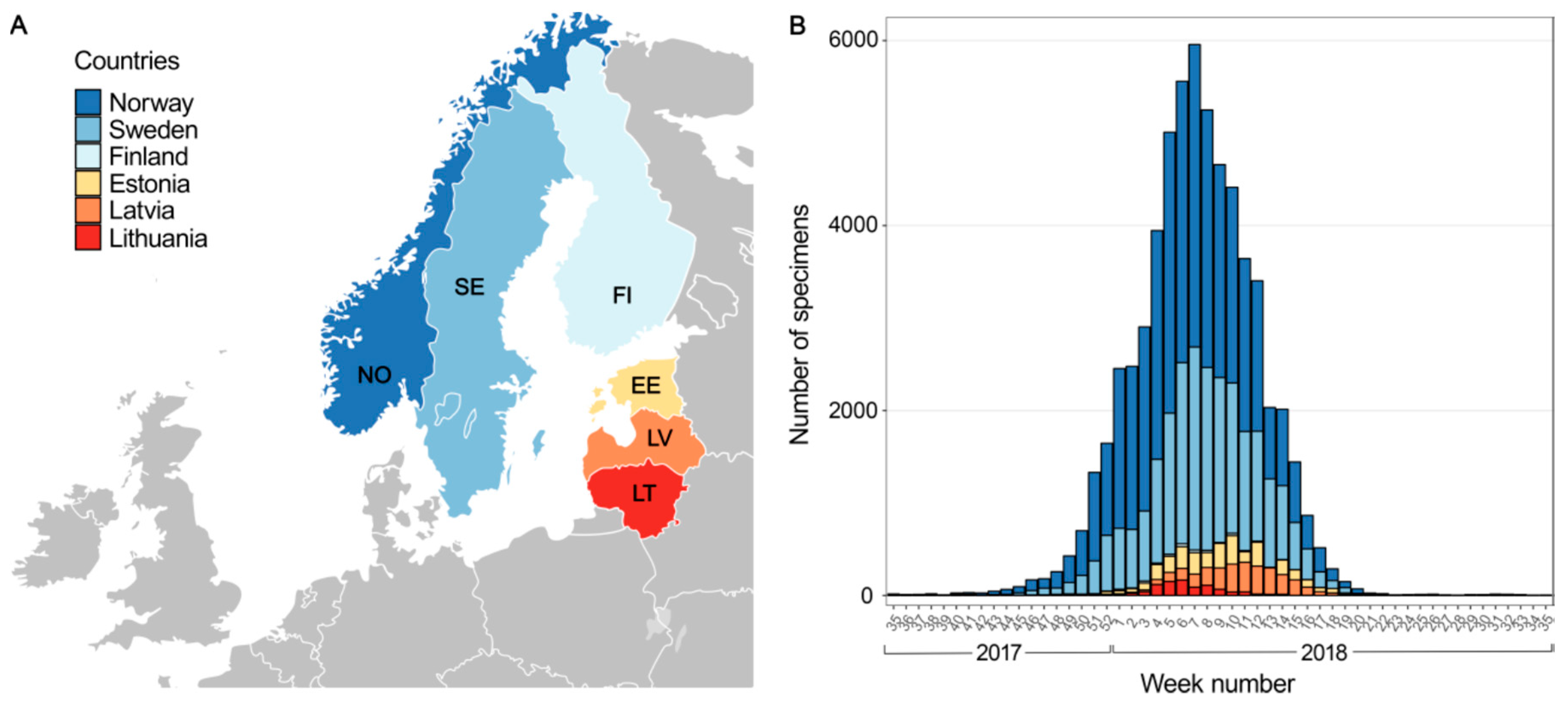
3.1. Influenza Virus Activity during 2017–2018 Season
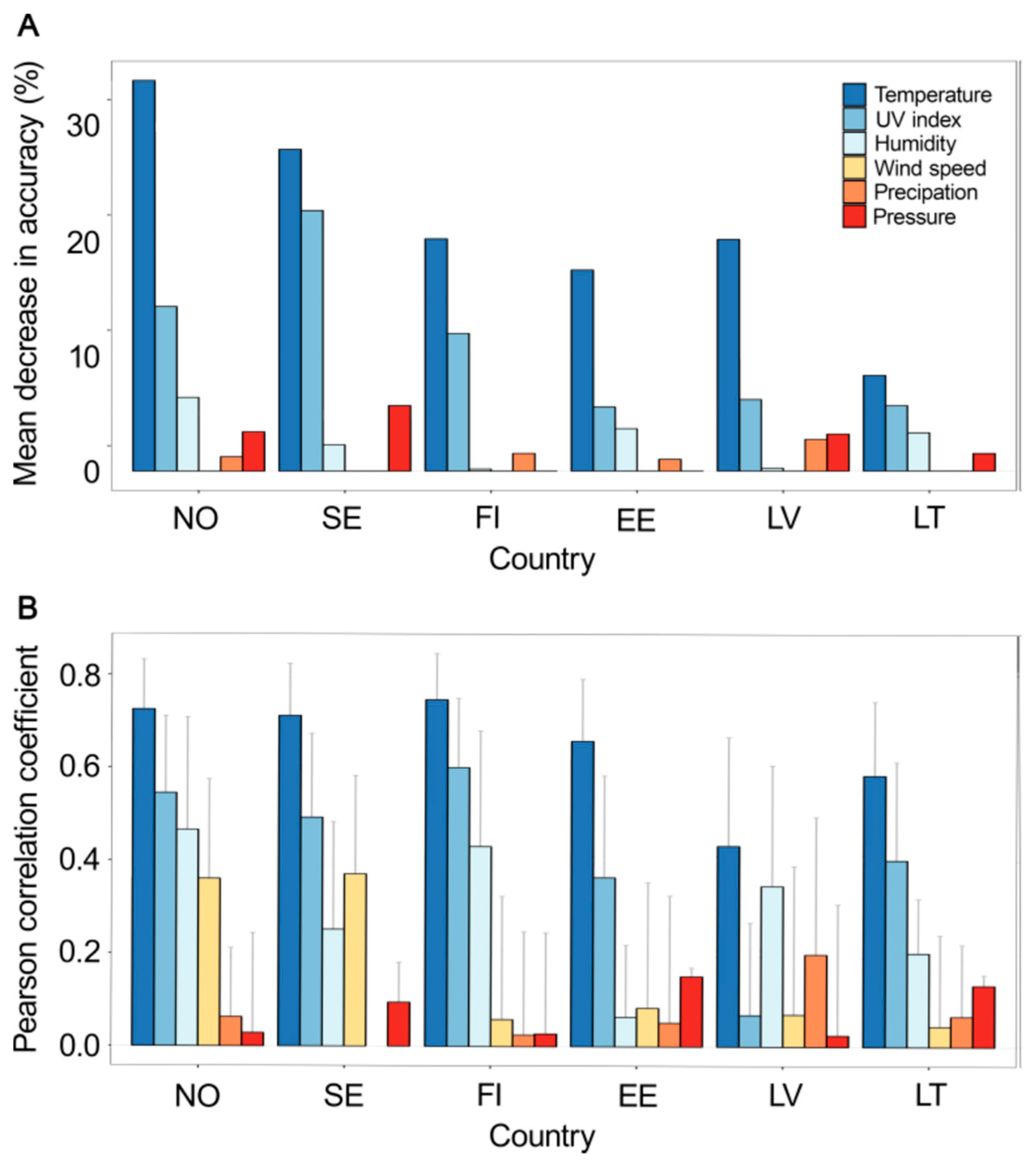
3.2. Temperature and UV Index Are the most Predictive Meteorological Factors of IV Epidemic in 2017–2018 in Northern Europe
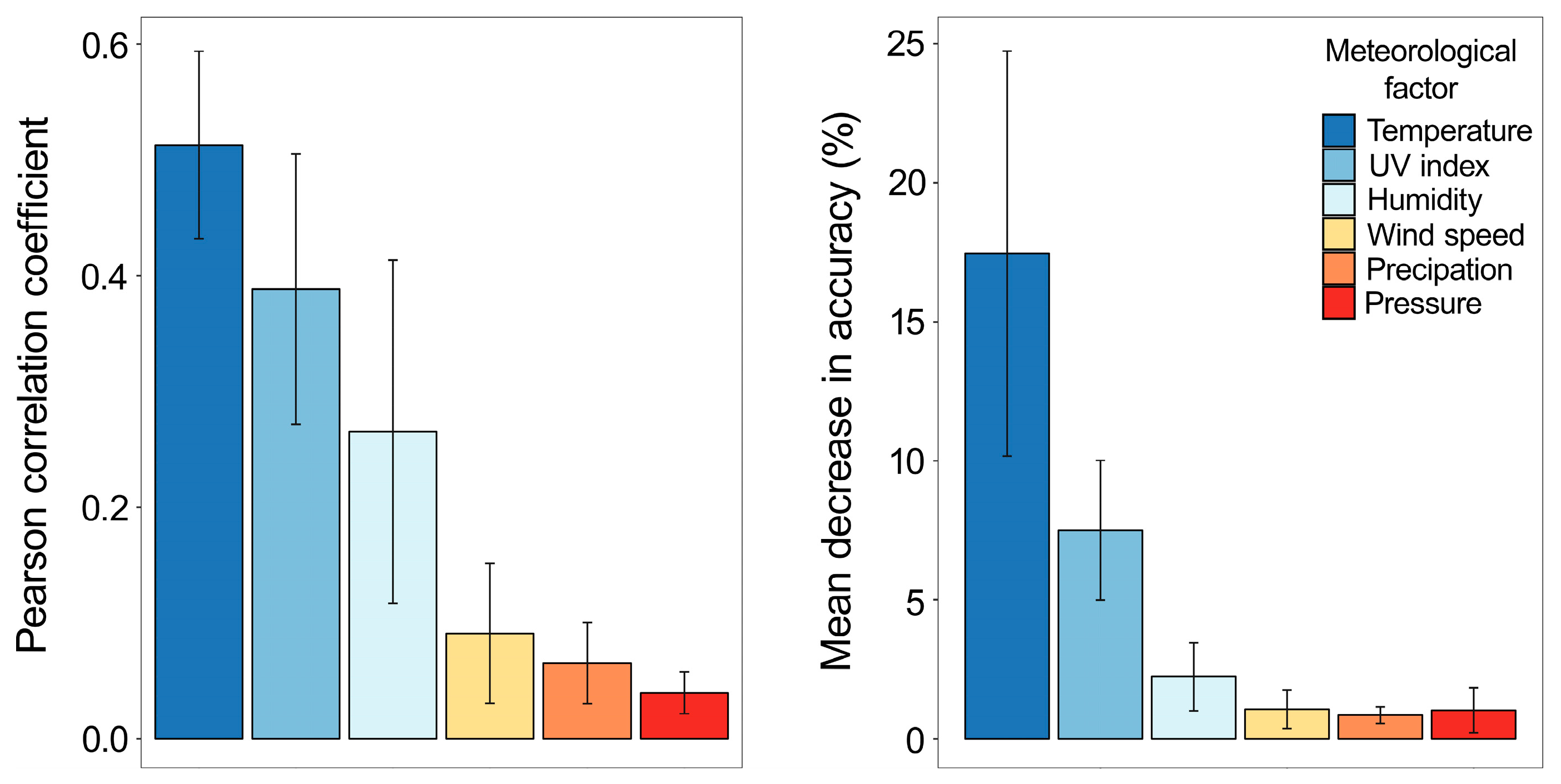
3.3. Temperature and UV Index Are the Most Predictive Meteorological Factors of IV Epidemics in 2010–2017 in Northern Europe
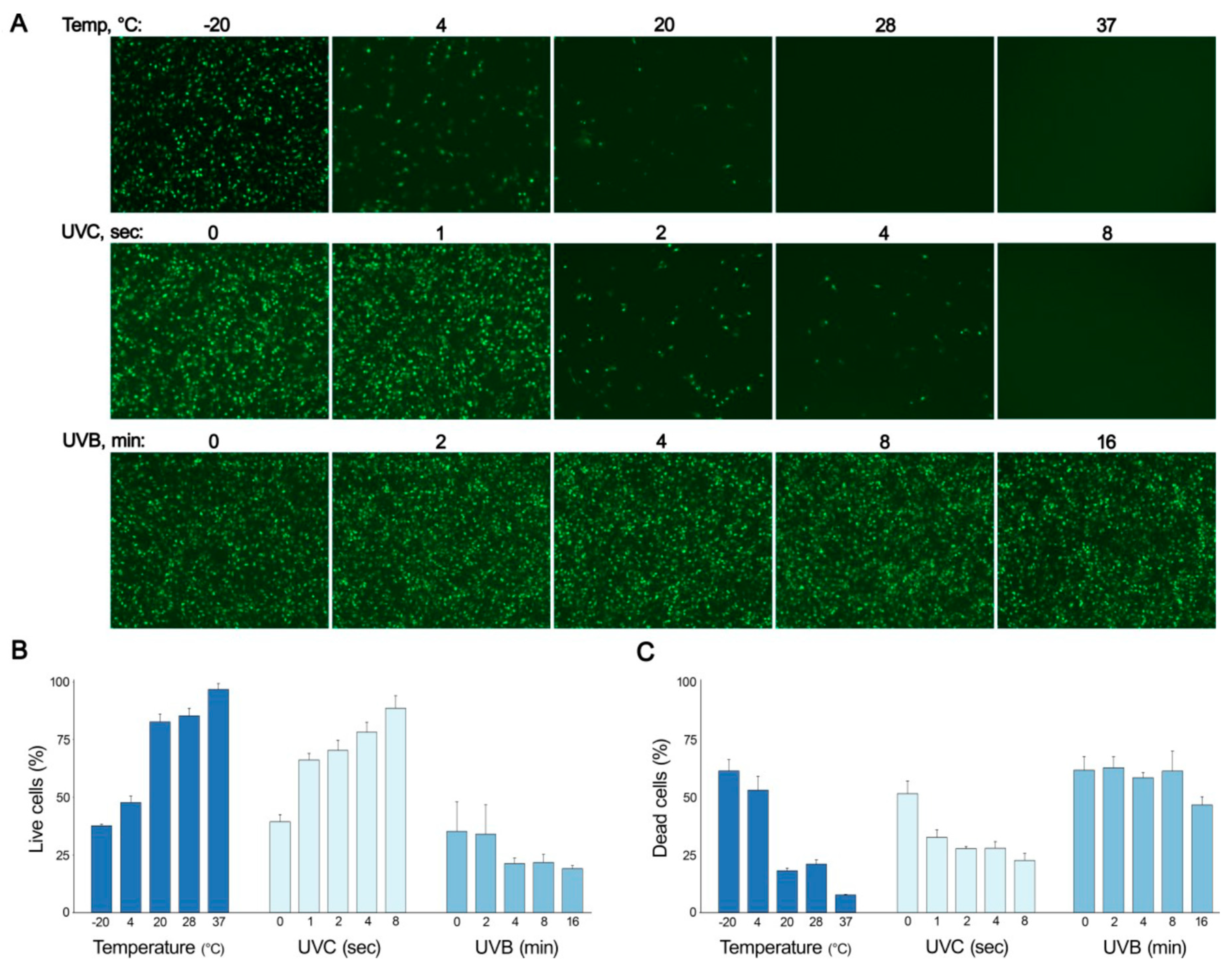
3.4. Low Temperature and Low UV Radiation Preserve IV Infectivity In Vitro
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| IV | influenza virus |
| IAV | influenza A virus |
| UV | ultraviolet |
| GFP | green fluorescent protein |
| RPE | human telomerase reverse transcriptase-immortalized retinal pigment cells |
| moi | multiplicity of infection |
| hpi | hours post infection |
| WHO | World Health Organization |
| NIC | National Influenza Center |
| GISRS | Global Influenza Surveillance and Response System |
| NWS | National Weather Service |
| PWS | personal weather stations |
| TEMIS | Tropospheric Emission Monitoring Internet Service |
| RF | Random Forest |
| RMSE | root-mean-square deviation |
| DMEM | Dulbecco’s Modified Eagle’s medium |
| TPCK-trypsin | l-1-tosylamido-2-phenylethyl chloromethyl ketone-trypsin |
References
- Influenza. Available online: http://www.who.int/influenza/en/ (accessed on 10 October 2018).
- Types of Influenza Viruses. Available online: https://www.cdc.gov/flu/about/viruses/types.htm (accessed on 10 October 2018).
- Iuliano, A.D.; Roguski, K.M.; Chang, H.H.; Muscatello, D.J.; Palekar, R.; Tempia, S.; Cohen, C.; Gran, J.M.; Schanzer, D.; Cowling, B.J.; et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet 2018, 391, 1285–1300. [Google Scholar] [CrossRef]
- Kash, J.C.; Taubenberger, J.K. The role of viral, host, and secondary bacterial factors in influenza pathogenesis. Am. J. Pathol. 2015, 185, 1528–1536. [Google Scholar] [CrossRef] [PubMed]
- Sooryanarain, H.; Elankumaran, S. Environmental role in influenza virus outbreaks. Annu. Rev. Anim. Biosci. 2015, 3, 347–373. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Barroso, D.; Leon-Gomez, I.; Delgado-Sanz, C.; Larrauri, A. Climatic Factors and Influenza Transmission, Spain, 2010–2015. Int. J. Environ. Res. Public Health 2017, 14, 1469. [Google Scholar] [CrossRef] [PubMed]
- Deyle, E.R.; Maher, M.C.; Hernandez, R.D.; Basu, S.; Sugihara, G. Global environmental drivers of influenza. Proc. Natl. Acad. Sci. USA 2016, 113, 13081–13086. [Google Scholar] [CrossRef] [PubMed]
- Roussel, M.; Pontier, D.; Cohen, J.M.; Lina, B.; Fouchet, D. Quantifying the role of weather on seasonal influenza. BMC Public Health 2016, 16, 441. [Google Scholar] [CrossRef] [PubMed]
- Soebiyanto, R.P.; Gross, D.; Jorgensen, P.; Buda, S.; Bromberg, M.; Kaufman, Z.; Prosenc, K.; Socan, M.; Vega Alonso, T.; Widdowson, M.A.; et al. Associations between Meteorological Parameters and Influenza Activity in Berlin (Germany), Ljubljana (Slovenia), Castile and Leon (Spain) and Israeli Districts. PLoS ONE 2015, 10, e0134701. [Google Scholar] [CrossRef] [PubMed]
- Lowen, A.C.; Steel, J. Roles of humidity and temperature in shaping influenza seasonality. J. Virol. 2014, 88, 7692–7695. [Google Scholar] [CrossRef] [PubMed]
- Peci, A.; Winter, A.L.; Li, L.; Gnaneshan, S.; Liu, J.; Mubareka, S.; Gubbay, J.B. Effect of absolute and relative humidity, temperature and wind speed on influenza activity in Toronto, Canada. Appl. Environ. Microbiol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Charland, K.M.; Buckeridge, D.L.; Sturtevant, J.L.; Melton, F.; Reis, B.Y.; Mandl, K.D.; Brownstein, J.S. Effect of environmental factors on the spatio-temporal patterns of influenza spread. Epidemiol. Infect. 2009, 137, 1377–1387. [Google Scholar] [CrossRef] [PubMed]
- Sagripanti, J.L.; Lytle, C.D. Inactivation of influenza virus by solar radiation. Photochem. Photobiol. 2007, 83, 1278–1282. [Google Scholar] [CrossRef] [PubMed]
- FluNet. Available online: http://www.who.int/influenza/gisrs_laboratory/flunet/en/ (accessed on 10 October 2018).
- Denisova, O.V.; Kakkola, L.; Feng, L.; Stenman, J.; Nagaraj, A.; Lampe, J.; Yadav, B.; Aittokallio, T.; Kaukinen, P.; Ahola, T.; et al. Obatoclax, saliphenylhalamide, and gemcitabine inhibit influenza a virus infection. J. Biol. Chem. 2012, 287, 35324–35332. [Google Scholar] [CrossRef] [PubMed]
- Pirhonen, J.; Sareneva, T.; Kurimoto, M.; Julkunen, I.; Matikainen, S. Virus infection activates IL-1 beta and IL-18 production in human macrophages by a caspase-1-dependent pathway. J. Immunol. 1999, 162, 7322–7329. [Google Scholar] [PubMed]
- Reimer, C.B.; Baker, R.S.; van Frank, R.M.; Newlin, T.E.; Cline, G.B.; Anderson, N.G. Purification of large quantities of influenza virus by density gradient centrifugation. J. Virol. 1967, 1, 1207–1216. [Google Scholar] [PubMed]
- Kittel, C.; Sereinig, S.; Ferko, B.; Stasakova, J.; Romanova, J.; Wolkerstorfer, A.; Katinger, H.; Egorov, A. Rescue of influenza virus expressing GFP from the NS1 reading frame. Virology 2004, 324, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Lakspere, T.; Tynell, J.; Kaloinen, M.; Vanlede, M.; Parsons, A.; Ikonen, N.; Kallio-Kokko, H.; Kantele, A.; Mattila, P.; Almusa, H.; et al. Full-Genome Sequences of Influenza A(H1N1)pdm09 Viruses Isolated from Finnish Patients from 2009 to 2013. Genome Announc. 2014, 2, e01004-13. [Google Scholar] [CrossRef] [PubMed]
- Tamerius, J.D.; Shaman, J.; Alonso, W.J.; Bloom-Feshbach, K.; Uejio, C.K.; Comrie, A.; Viboud, C. Environmental predictors of seasonal influenza epidemics across temperate and tropical climates. PLoS Pathog. 2013, 9, e1003194. [Google Scholar] [CrossRef]
- Ballester, J.; Rodó, X.; Robine, J.M.; Herrmann, F.R. European seasonal mortality and influenza incidence due to winter temperature variability. Nat. Clim. Chang. 2016, 6, 927–930. [Google Scholar] [CrossRef]
- Towers, S.; Chowell, G.; Hameed, R.; Jastrebski, M.; Khan, M.; Meeks, J.; Mubayi, A.; Harris, G. Climate change and influenza: The likelihood of early and severe influenza seasons following warmer than average winters. PLoS Curr. 2013, 5. [Google Scholar] [CrossRef] [PubMed]
- Geier, D.A.; Kern, J.K.; Geier, M.R. A longitudinal ecological study of seasonal influenza deaths in relation to climate conditions in the United States from 1999 through 2011. Infect. Ecol. Epidemiol. 2018, 8, 1474708. [Google Scholar] [CrossRef] [PubMed]
- European Monitoring of Excess Mortality for Public Health Action. Available online: www.euromomo.eu (accessed on 10 October 2018).
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ianevski, A.; Zusinaite, E.; Shtaida, N.; Kallio-Kokko, H.; Valkonen, M.; Kantele, A.; Telling, K.; Lutsar, I.; Letjuka, P.; Metelitsa, N.; et al. Low Temperature and Low UV Indexes Correlated with Peaks of Influenza Virus Activity in Northern Europe during 2010–2018. Viruses 2019, 11, 207. https://doi.org/10.3390/v11030207
Ianevski A, Zusinaite E, Shtaida N, Kallio-Kokko H, Valkonen M, Kantele A, Telling K, Lutsar I, Letjuka P, Metelitsa N, et al. Low Temperature and Low UV Indexes Correlated with Peaks of Influenza Virus Activity in Northern Europe during 2010–2018. Viruses. 2019; 11(3):207. https://doi.org/10.3390/v11030207
Chicago/Turabian StyleIanevski, Aleksandr, Eva Zusinaite, Nastassia Shtaida, Hannimari Kallio-Kokko, Miia Valkonen, Anu Kantele, Kaidi Telling, Irja Lutsar, Pille Letjuka, Natalja Metelitsa, and et al. 2019. "Low Temperature and Low UV Indexes Correlated with Peaks of Influenza Virus Activity in Northern Europe during 2010–2018" Viruses 11, no. 3: 207. https://doi.org/10.3390/v11030207
APA StyleIanevski, A., Zusinaite, E., Shtaida, N., Kallio-Kokko, H., Valkonen, M., Kantele, A., Telling, K., Lutsar, I., Letjuka, P., Metelitsa, N., Oksenych, V., Dumpis, U., Vitkauskiene, A., Stašaitis, K., Öhrmalm, C., Bondeson, K., Bergqvist, A., Cox, R. J., Tenson, T., ... Kainov, D. E. (2019). Low Temperature and Low UV Indexes Correlated with Peaks of Influenza Virus Activity in Northern Europe during 2010–2018. Viruses, 11(3), 207. https://doi.org/10.3390/v11030207






