A Single-Nucleotide Polymorphism of αVβ3 Integrin Is Associated with the Andes Virus Infection Susceptibility
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
Three Sets of Subjects Were Evaluated
2.2. Ethical Statement
2.3. DNA Extraction and Genotyping
2.4. Statistical Analysis
3. Results
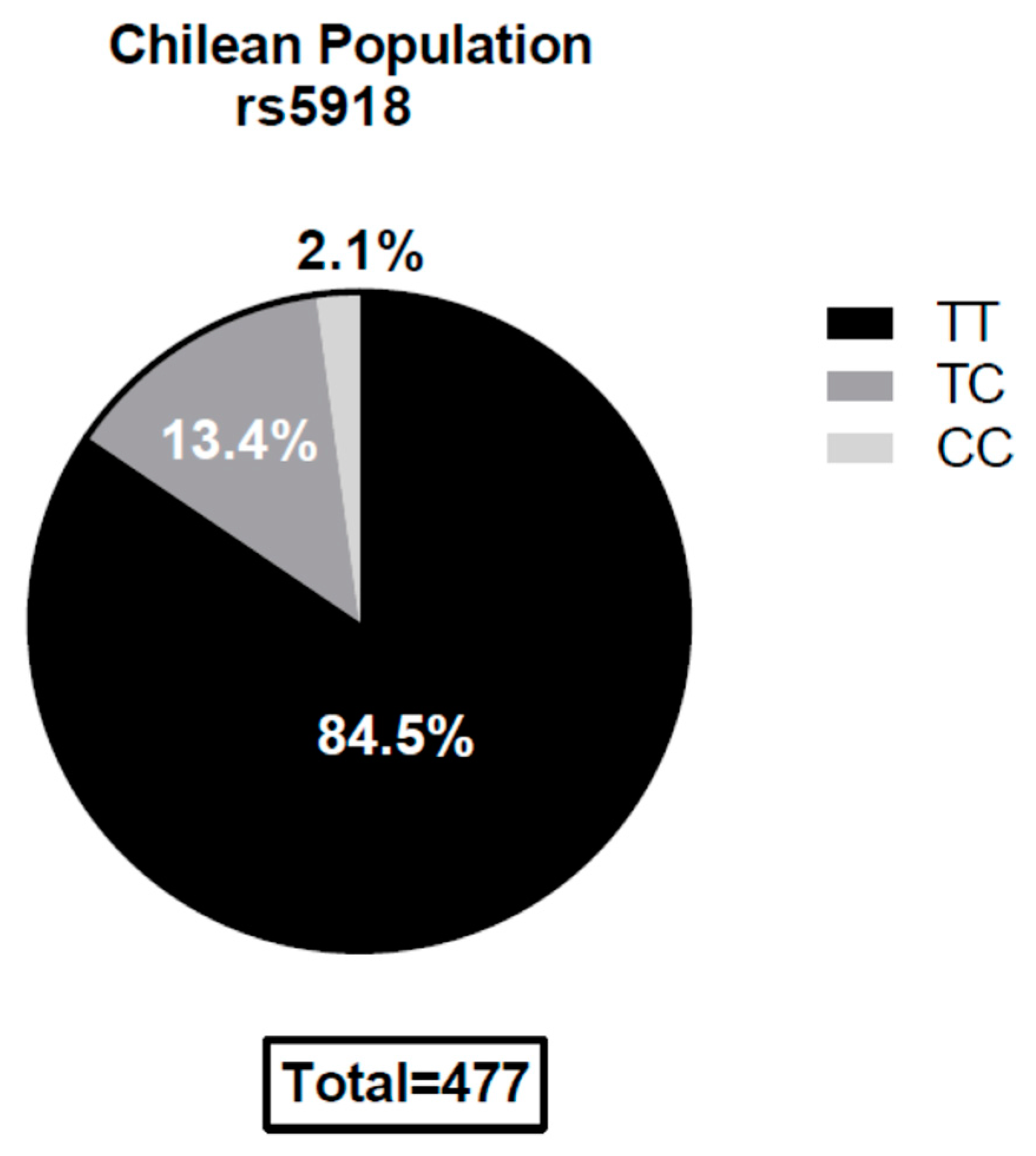
3.1. Genotype Distribution in the General Population
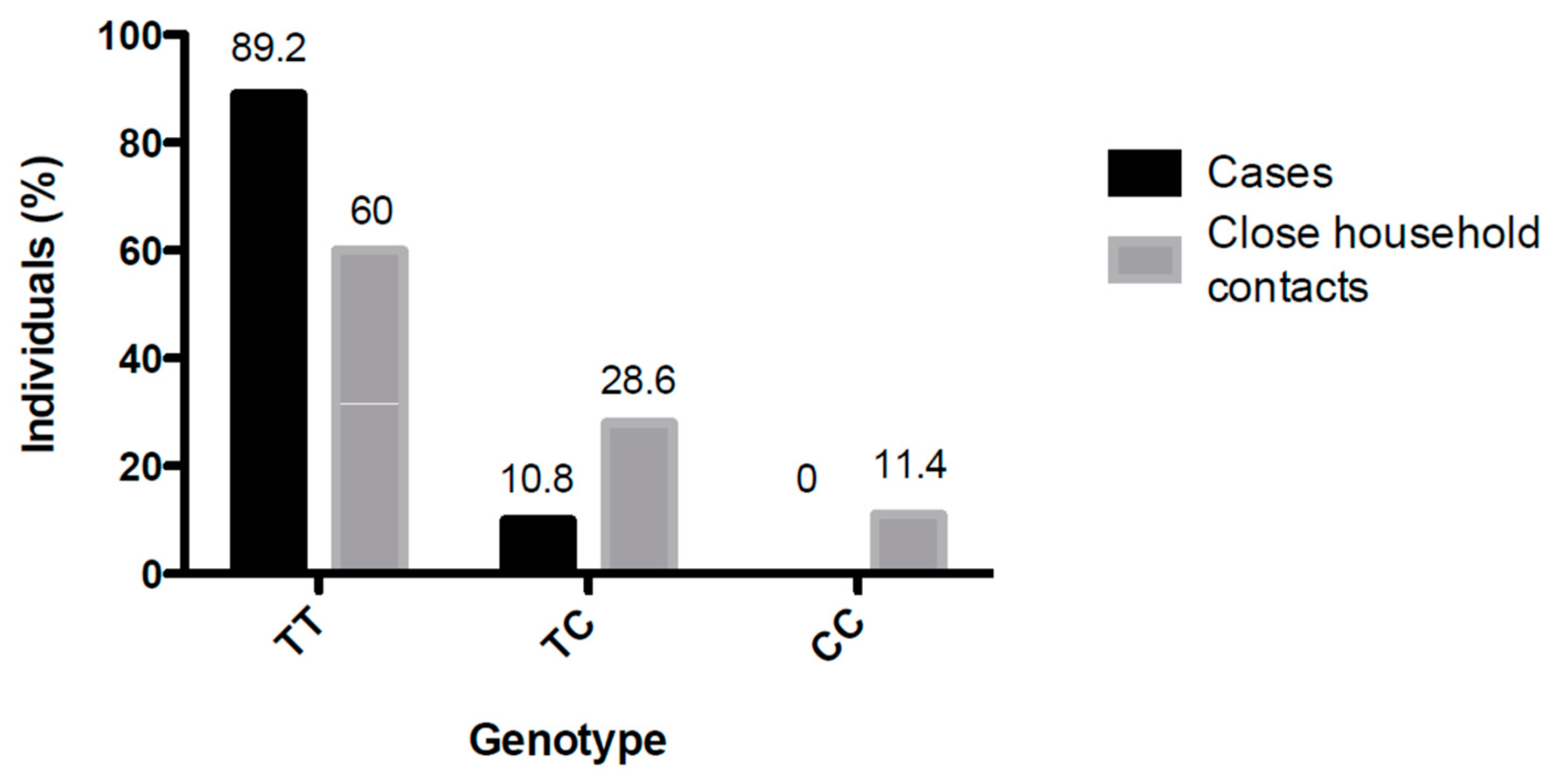
3.2. Analysis of SNP rs5918 Distribution Among Study Group 2 (Cases and Close-Household Contacts) and Study Group 3 (11 Infected Close-Household Contacts)
3.3. ANDV Infection Risk Assessment and Risk Models for SNP rs5918 Genotype and Infection Among Cases and Close-Household Contacts
3.4. ANDV Infection Risk Assessment Among Cases and Uninfected Close-Household Contacts Exposed to the Same Risk Activity
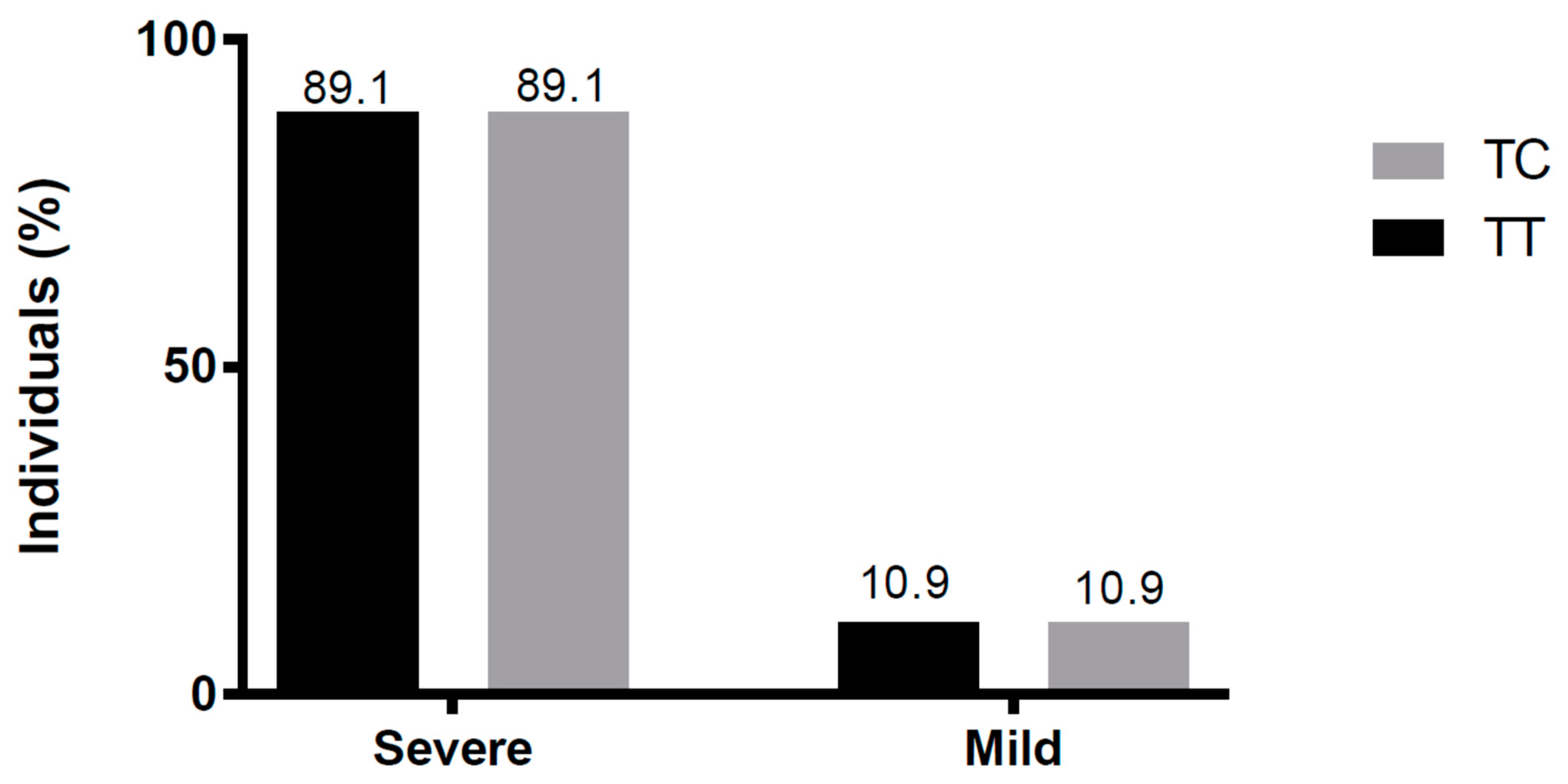
3.5. Severity of ANDV-Induced Disease and SNP rs5918 Genotype Distribution
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schmaljohn, C. Hantaviruses: A Global Disease Problem. Emerg. Infect. Dis. 1997, 3, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Knipe, D.M.; Howley, P. Fields Virology (Knipe, Fields Virology)-2 Volume Set; LWW: Philadelphia, PA, USA, 2013; ISBN 978-1-4511-0563-6. [Google Scholar]
- Medina, R.A.; Torres-Perez, F.; Galeno, H.; Navarrete, M.; Vial, P.A.; Palma, R.E.; Ferres, M.; Cook, J.A.; Hjelle, B. Ecology, Genetic Diversity, and Phylogeographic Structure of Andes Virus in Humans and Rodents in Chile. J. Virol. 2009, 83, 2446–2459. [Google Scholar] [CrossRef] [PubMed]
- Toro, J.; Vega, J.D.; Khan, A.S.; Mills, J.N.; Padula, P.; Terry, W.; Yadón, Z.; Valderrama, R.; Ellis, B.A.; Pavletic, C.; et al. An Outbreak of Hantavirus Pulmonary Syndrome, Chile, 1997. Emerg. Infect. Dis. 1998, 4, 687–694. [Google Scholar] [CrossRef] [PubMed]
- Síndrome Cardiopulmonar por Hantavirus. Available online: http://epi.minsal.cl/hantavirus-materiales-relacionados/ (accessed on 10 August 2018).
- Ferrés, M.; Vial, P.; Marco, C.; Yañez, L.; Godoy, P.; Castillo, C.; Hjelle, B.; Delgado, I.; Lee, S.; Mertz, G.J.; et al. Prospective Evaluation of Household Contacts of Persons with Hantavirus Cardiopulmonary Syndrome in Chile. J. Infect. Dis. 2007, 195, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Valdebenito, C.; Calvo, M.; Vial, C.; Mansilla, R.; Marco, C.; Palma, R.E.; Vial, P.A.; Valdivieso, F.; Mertz, G.; Ferrés, M. Person-to-Person Household and Nosocomial Transmission of Andes Hantavirus, Southern Chile, 2011. Emerg. Infect. Dis. 2014, 20, 1637–1644. [Google Scholar] [CrossRef] [PubMed]
- Vial, P.A.; Valdivieso, F.; Mertz, G.; Castillo, C.; Belmar, E.; Delgado, I.; Tapia, M.; Ferrés, M. Incubation Period of Hantavirus Cardiopulmonary Syndrome. Emerg. Infect. Dis. 2006, 12, 3. [Google Scholar] [CrossRef] [PubMed]
- Núñez, J.J.; Fritz, C.L.; Knust, B.; Buttke, D.; Enge, B.; Novak, M.G.; Kramer, V.; Osadebe, L.; Messenger, S.; Albariño, C.G.; et al. Hantavirus Infections among Overnight Visitors to Yosemite National Park, California, USA, 2012. Emerg. Infect. Dis. 2014, 20, 386–393. [Google Scholar] [CrossRef]
- Ferres, M.; Valdivieso, F.; Vial, P.A. Manejo del Paciente Crítico con Sindróme Cardiopulmonar por Hantavirus, Primera ed.; Editorial Salesiani: Ate District, Peru, 2004. [Google Scholar]
- Ferrés, M.; Vial, P. Hantavirus infection in children. Emerg. Infect. Dis. 1998, 4, 85–87. [Google Scholar]
- Vaheri, A.; Strandin, T.; Hepojoki, J.; Sironen, T.; Henttonen, H.; Mäkelä, S.; Mustonen, J. Uncovering the mysteries of hantavirus infections. Nat. Rev. Microbiol. 2013, 11, 539–550. [Google Scholar] [CrossRef]
- Borges, E.; Jan, Y.; Ruoslahti, E. Platelet-derived Growth Factor Receptor β and Vascular Endothelial Growth Factor Receptor 2 Bind to the β 3 Integrin through Its Extracellular Domain. J. Biol. Chem. 2000, 275, 39867–39873. [Google Scholar] [CrossRef]
- Matthys, V.S.; Gorbunova, E.E.; Gavrilovskaya, I.N.; Mackow, E.R. Andes Virus Recognition of Human and Syrian Hamster 3 Integrins Is Determined by an L33P Substitution in the PSI Domain. J. Virol. 2010, 84, 352–360. [Google Scholar] [CrossRef] [PubMed]
- Raftery, M.J.; Lalwani, P.; Krautkrӓmer, E.; Peters, T.; Scharffetter-Kochanek, K.; Krüger, R.; Hofmann, J.; Seeger, K.; Krüger, D.H.; Schönrich, G. β2 integrin mediates hantavirus-induced release of neutrophil extracellular traps. J. Exp. Med. 2014, 211, 1485–1497. [Google Scholar] [CrossRef] [PubMed]
- Krautkrämer, E.; Zeier, M. Hantavirus causing hemorrhagic fever with renal syndrome enters from the apical surface and requires decay-accelerating factor (DAF/CD55). J. Virol. 2008, 82, 4257–4264. [Google Scholar] [CrossRef] [PubMed]
- Jangra, R.K.; Herbert, A.S.; Li, R.; Jae, L.T.; Kleinfelter, L.M.; Slough, M.M.; Barker, S.L.; Guardado-Calvo, P.; Román-Sosa, G.; Dieterle, M.E.; et al. Protocadherin-1 is essential for cell entry by New World hantaviruses. Nature 2018, 563, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Gavrilovskaya, I.N.; Brown, E.J.; Ginsberg, M.H.; Mackow, E.R. Cellular Entry of Hantaviruses Which Cause Hemorrhagic Fever with Renal Syndrome Is Mediated by β3 Integrins. J. Virol. 1999, 73, 9. [Google Scholar]
- Gavrilovskaya, I.N.; Gorbunova, E.E.; Mackow, N.A.; Mackow, E.R. Hantaviruses Direct Endothelial Cell Permeability by Sensitizing Cells to the Vascular Permeability Factor VEGF, while Angiopoietin 1 and Sphingosine 1-Phosphate Inhibit Hantavirus-Directed Permeability. J. Virol. 2008, 82, 5797–5806. [Google Scholar] [CrossRef] [PubMed]
- Raymond, T.; Gorbunova, E.; Gavrilovskaya, I.N.; Mackow, E.R. Pathogenic hantaviruses bind plexin-semaphorin-integrin domains present at the apex of inactive, bent v 3 integrin conformers. Proc. Natl. Acad. Sci. USA 2005, 102, 1163–1168. [Google Scholar] [CrossRef]
- Miquel, J.F.; Covarrubias, C.; Villaroel, L.; Mingrone, G.; Greco, A.V.; Puglielli, L.; Carvallo, P.; Marshall, G.; Pino, G.D.; Nervi, F. Genetic epidemiology of cholesterol cholelithiasis among Chilean Hispanics, Amerindians, and Maoris. Gastroenterology 1998, 115, 937–946. [Google Scholar] [CrossRef]
- Valle, M.O.D.; Edelstein, A.; Segura, E.L.; Rossi, C.M.; Colavecchia, S.B.; Rabinovich, R.D.; MartíNez, P.V.; Miguel, S.D.L.; Padula, P.J. Development and evaluation of a solid-phase enzyme immunoassay based on Andes hantavirus recombinant nucleoprotein. J. Med. Microbiol. 2000, 49, 149–155. [Google Scholar]
- Vial, C.; Martinez-Valdebenito, C.; Rios, S.; Martinez, J.; Vial, P.A.; Ferres, M.; Rivera, J.C.; Perez, R.; Valdivieso, F. Molecular method for the detection of Andes hantavirus infection: validation for clinical diagnostics. Diagn. Microbiol. Infect. Dis. 2016, 84, 36–39. [Google Scholar] [CrossRef]
- Angulo, J.; Pino, K.; Pavez, C.; Biel, F.; Labbé, P.; Miquel, J.F.; Soza, A.; López-Lastra, M. Genetic variations in host IL28B links to the detection of peripheral blood mononuclear cells-associated hepatitis C virus RNA in chronically infected patients. J. Viral Hepat. 2013, 20, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Reference SNP (refSNP) Cluster Report: rs5918** with Pathogenic Allele**. Available online: https://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=rs5918 (accessed on 19 November 2018).
- Gavrilovskaya, I.N.; Peresleni, T.; Geimonen, E.; Mackow, E.R. Pathogenic hantaviruses selectively inhibit β3 integrin directed endothelial cell migration. Arch. Virol. 2002, 147, 1913–1931. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Gao, M.; Han, Q.; Fang, J.; Zhao, Q.; Zhang, N. Intensity of Platelet β3 Integrin in Patients with Hemorrhagic Fever with Renal Syndrome and Its Correlation with Disease Severity. Viral Immunol. 2008, 21, 255–262. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Gao, M.; Han, Q.; Lou, S.; Fang, J. Platelet glycoprotein IIb/IIIa (HPA-1 and HPA-3) polymorphisms in patients with hemorrhagic fever with renal syndrome. Hum. Immunol. 2009, 70, 452–456. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.-P.; Xiong, H.-R.; Zhu, N.; Chen, Q.-Z.; Wang, H.; Zhong, C.-J.; Wang, M.-R.; Lu, S.; Luo, F.; Hou, W. Lack of association between integrin αVβ3 gene polymorphisms and hemorrhagic fever with renal syndrome in Han Chinese from Hubei, China. Virol. Sin. 2017, 32, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Eyheramendy, S.; Martinez, F.I.; Manevy, F.; Vial, C.; Repetto, G.M. Genetic structure characterization of Chileans reflects historical immigration patterns. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Chapman, S.J.; Hill, A.V.S. Human genetic susceptibility to infectious disease. Nat. Rev. Genet. 2012, 13, 175–188. [Google Scholar] [CrossRef]
| ANDV Patients n/Total (%) | Close-Household Individuals n/Total (%) | |||
|---|---|---|---|---|
| SNP | TT | 66/74 (89.2) | 63/105 (60) | p < 0.05 |
| TC | 8/74 (10.8) | 30/105 (28.6) | ||
| CC | 0/74 (0.0) | 12/105 (11.4) | ||
| Age (mean) | Years old | 36.1 (CI:32–40) | 32.1 (CI:28.6–35.6) | |
| Sex | M | 51/74 (68.9) | 37/105 (35.2) | p < 0.05 |
| Ethnicity | Hispanic | 61/74 (82.4) | 83/105 (79) | |
| Native | 6/74 (8.1) | 7/105 (6.7) | ||
| Other | 7/74 (9.5) | 15/105 (14.3) | ||
| Type of residence | Rural | 38/74 (51.4) | 40/105 (38.1) | |
| Work activities | High risk (forestry and agriculture) | 26/74 (35.1) | 8/105 (7.6) | p < 0.05 |
| Risk activities | Visit rural areas | 65/74 (87.8) | 71/96 (74) | |
| See rodents | 31/73 (42.5) | 17/97 (17.5) | p < 0.05 | |
| See or touch rodent’s excrement | 15/73 (20.5) | 10/97 (10.3) | ||
| Handle gnawed food | 12/73 (16.4) | 3/97 (3.1) | p < 0.05 | |
| Eat gnawed food | 3/73 (4.1) | 1/97 (1) | ||
| Rat extermination activities | 10/73 (13.7) | 3/97 (3.1) | ||
| Enter into abandoned shelter | 43/73 (58.9) | 21/97 (21.6) | p < 0.05 | |
| Clean abandoned shelter | 17/73 (23.3) | 6/97 (6.2) | p < 0.05 | |
| Clean up rodent’s feces | 20/72 (27.8) | 7/97 (7.2) | p < 0.05 | |
| Forestry activities | 20/73 (27.4) | 12/97 (12.4) | ||
| Agricultural Activities | 38/73 (52.1) | 16/97 (16.5) | p < 0.05 | |
| Handle wood | 45/73 (61.6) | 32/97 (33) | p < 0.05 | |
| Walk into heavy forest | 40/73 (54.8) | 24/94 (25.5) | p < 0.05 | |
| Collect wild fruits | 17/73 (23.3) | 15/97 (15.5) | p < 0.05 | |
| Demolition activities | 3/73 (4.1) | 3/97 (3.1) | ||
| Camping in unauthorized areas | 7/73 (9.6) | 8/97 (8.2) | ||
| Sleep outdoors | 6/74 (8.1) | 3/97 (3.1) | ||
| OR Crude (CI) | OR1 | OR2 | OR3 | |
|---|---|---|---|---|
| SNP (TT) | 6.2 (2.7–14.1) * | 19.7 (3–131) * | 12.6 (2.9–55.3) * | 12.6 (2.9–54.5) * |
| Sex (M) | 4.1 (2.2–7.7) * | 1.3 (0.3–5.1) | 1.5 (0.4–4.7) | 1.3 (0.4–4.4) |
| Type of residence (R) | 1.7 (0.9–3.1) | 0.1 (0.02–0.8) | ND | ND |
| Work activities | 8.8 (3.4–22.5) * | 21 (2.5–178) * | 7.9 (1.6–39.4) | 8.1 (1.6–40.6) |
| Visit rural areas | 2.5 (1.1–5.9) * | 0.4 (0.05–3.6) | 1.4 (0.4–5.1) | 1 (0.2–5.6) |
| See rodents | 3.5 (1.7–7) * | 1 (0.2–5.4) | ND | ND |
| See or touch rodent’s excrement | 2.2 (0.9–5.4) | 0 | ND | ND |
| Handle gnawed food | 6.2 (1.7–22.7) * | 8.3 (0.3–248) | 1 (0–10.5) | ND |
| Eat gnawed food | 4.1 (0.4–40.4) | 1.9 (0.04–87) | ND | ND |
| Rat extermination activities | 5 (1.3–18.8) * | 4.4 (0.2–88) | 1.2 (0–19.2) | |
| Get into abandoned shelter | 5.2 (2.6–10.1) * | 11 (2–60) * | 3.3 (1–10.7) | 4 (1.2–13) |
| Clean abandoned shelter | 4.6 (1.7–12.4) * | 0.6 (0.1–5.4) | 0.8 (0.1–5.4) | 0.8 (0.1–4.6) |
| Clean up rodent’s feces | 5 (2–12.5) * | 4.5 (0.3–68) | 1.7 (0.2–17.1) | 2.4 (0.4–14.7) |
| Forestry activities | 2.7 (1.2–5.9) * | 0.1 (0–0.6) | 0.2 (0–1) | 0.2 (0–1.2) |
| Agricultural activities | 3.1 (1.5–6.4) * | 0.2 (0–1.5) | 0.9 (0.2–3.3) | 0.8 (0.2–3.1) |
| Handle wood | 3.2 (1.7–6.2) * | 3.4 (0.7–16.3) | 2.8 (0.8–9.8) | 2.5 (0.7–9.4) |
| Walk into heavy forest | 3.5 (1.8–6.8) * | 38 (4–349) * | 6.6 (1.8–23.4) | 7.2 (2–25.3) |
| Collect wild fruits | 1.7 (0.8–3.6) | 3.7 (0.6–22) | ND | 1.8 (0.5–6.8) |
| Demolition activities | 1.3 (0.3–6.8) | 0.3 (0–5) | ND | 0.5 (0–8.5) |
| Camping in unauthorized areas | 1.1 (0.4–3.4) | 3.2 (0.3–33) | ND | 1.2 (0.2–8.7) |
| Sleep outdoors | 2.8 (0.7–11.4) | 0.5 (0–6) | ND | ND |
| Genotype | Access to Abandoned Places 1 | Handle Wood 2 | ||
|---|---|---|---|---|
| Cases n (%) | Close-household contacts n (%) | Cases n (%) | Close-household contacts n (%) | |
| Susceptible (TT) | 39 (90.7) | 12 (57.1) | 38 (84.4) | 19 (59.4) |
| Protective (TC/CC) | 4 (9.3) | 9 (42.9) | 7 (15.6) | 13 (40.6) |
| Total | 43 | 21 | 45 | 32 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martínez-Valdebenito, C.; Angulo, J.; Le Corre, N.; Marco, C.; Vial, C.; Miquel, J.F.; Cerda, J.; Mertz, G.; Vial, P.; Lopez-Lastra, M.; et al. A Single-Nucleotide Polymorphism of αVβ3 Integrin Is Associated with the Andes Virus Infection Susceptibility. Viruses 2019, 11, 169. https://doi.org/10.3390/v11020169
Martínez-Valdebenito C, Angulo J, Le Corre N, Marco C, Vial C, Miquel JF, Cerda J, Mertz G, Vial P, Lopez-Lastra M, et al. A Single-Nucleotide Polymorphism of αVβ3 Integrin Is Associated with the Andes Virus Infection Susceptibility. Viruses. 2019; 11(2):169. https://doi.org/10.3390/v11020169
Chicago/Turabian StyleMartínez-Valdebenito, Constanza, Jenniffer Angulo, Nicole Le Corre, Claudia Marco, Cecilia Vial, Juan Francisco Miquel, Jaime Cerda, Gregory Mertz, Pablo Vial, Marcelo Lopez-Lastra, and et al. 2019. "A Single-Nucleotide Polymorphism of αVβ3 Integrin Is Associated with the Andes Virus Infection Susceptibility" Viruses 11, no. 2: 169. https://doi.org/10.3390/v11020169
APA StyleMartínez-Valdebenito, C., Angulo, J., Le Corre, N., Marco, C., Vial, C., Miquel, J. F., Cerda, J., Mertz, G., Vial, P., Lopez-Lastra, M., & Ferrés, M. (2019). A Single-Nucleotide Polymorphism of αVβ3 Integrin Is Associated with the Andes Virus Infection Susceptibility. Viruses, 11(2), 169. https://doi.org/10.3390/v11020169





