Comparative Study of the Temperature Sensitive, Cold Adapted and Attenuated Mutations Present in the Master Donor Viruses of the Two Commercial Human Live Attenuated Influenza Vaccines
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Plasmids
2.3. Minigenome Assays
2.4. Virus Rescue
2.5. Virus Growth Kinetics
2.6. Plaque Assays
2.7. Animal Experiments
2.8. Enzyme-Linked Immunosorbent Assays (ELISAs)
2.9. Hemagglutination Inhibition (HAI) Assays
2.10. Evaluation of T Cell Responses
3. Results
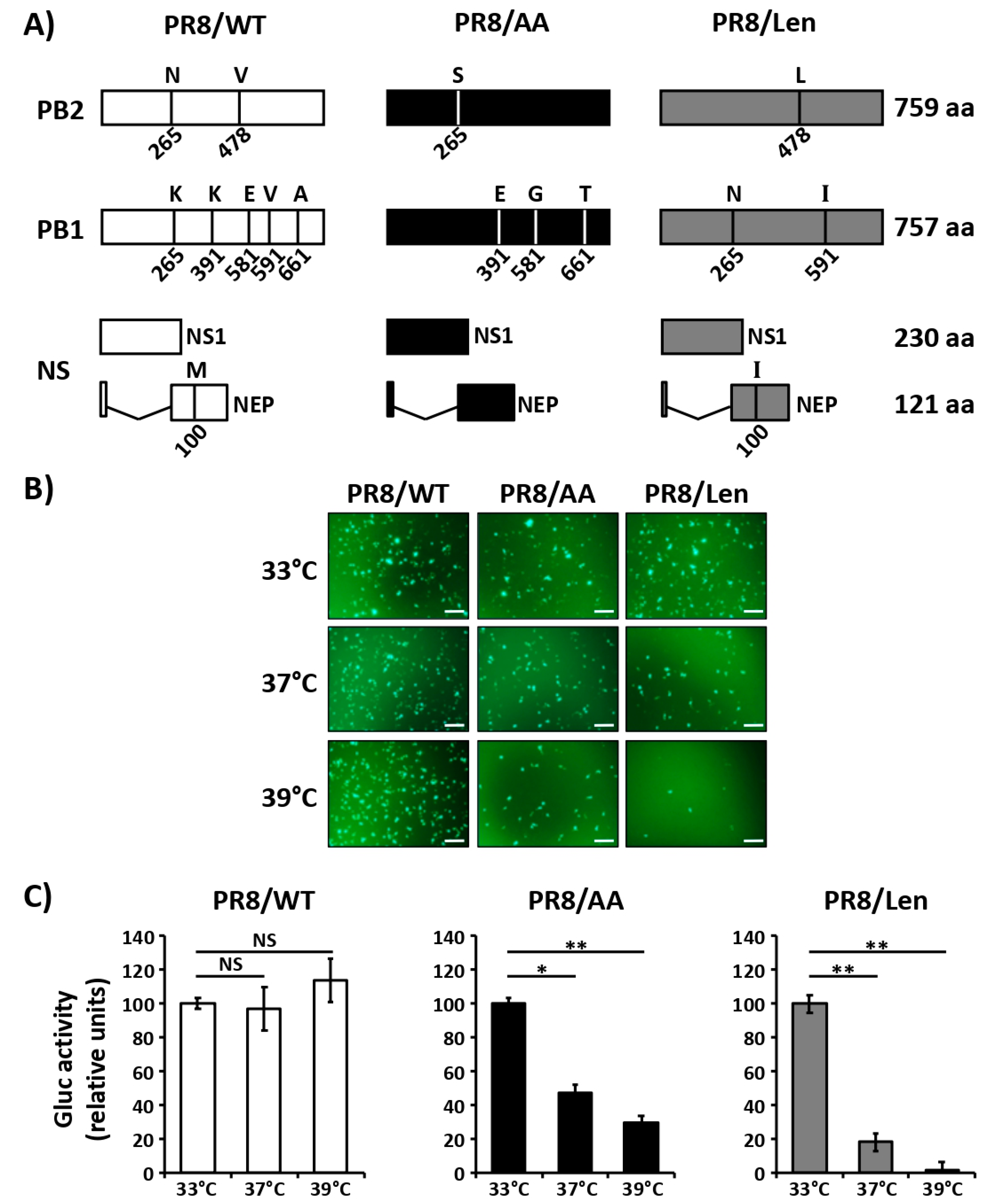
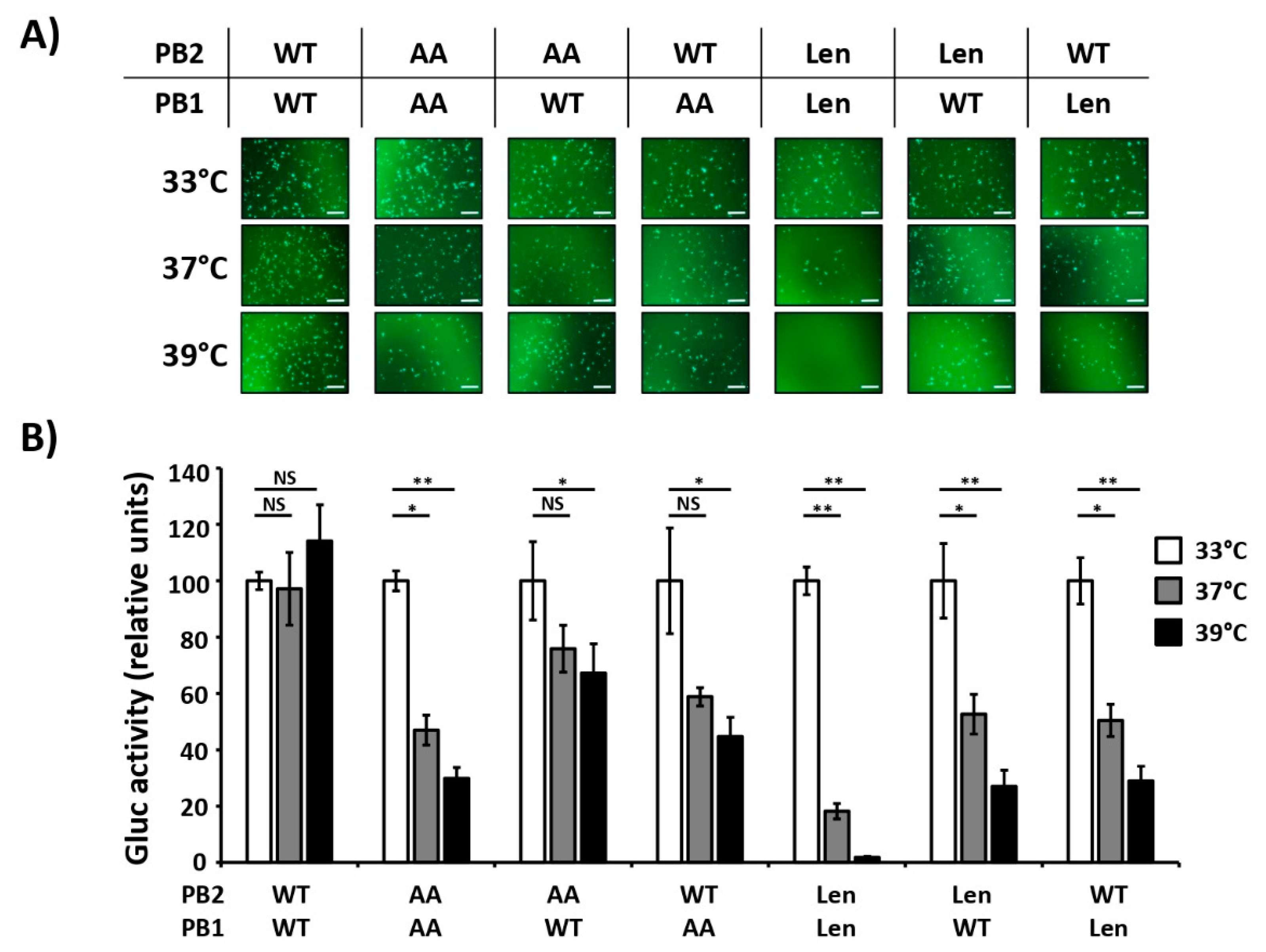
3.1. Mutations of the US AA MDV and Russian Len MDV Confer Different Levels of Temperature Sensitivity to the Polymerase of PR8
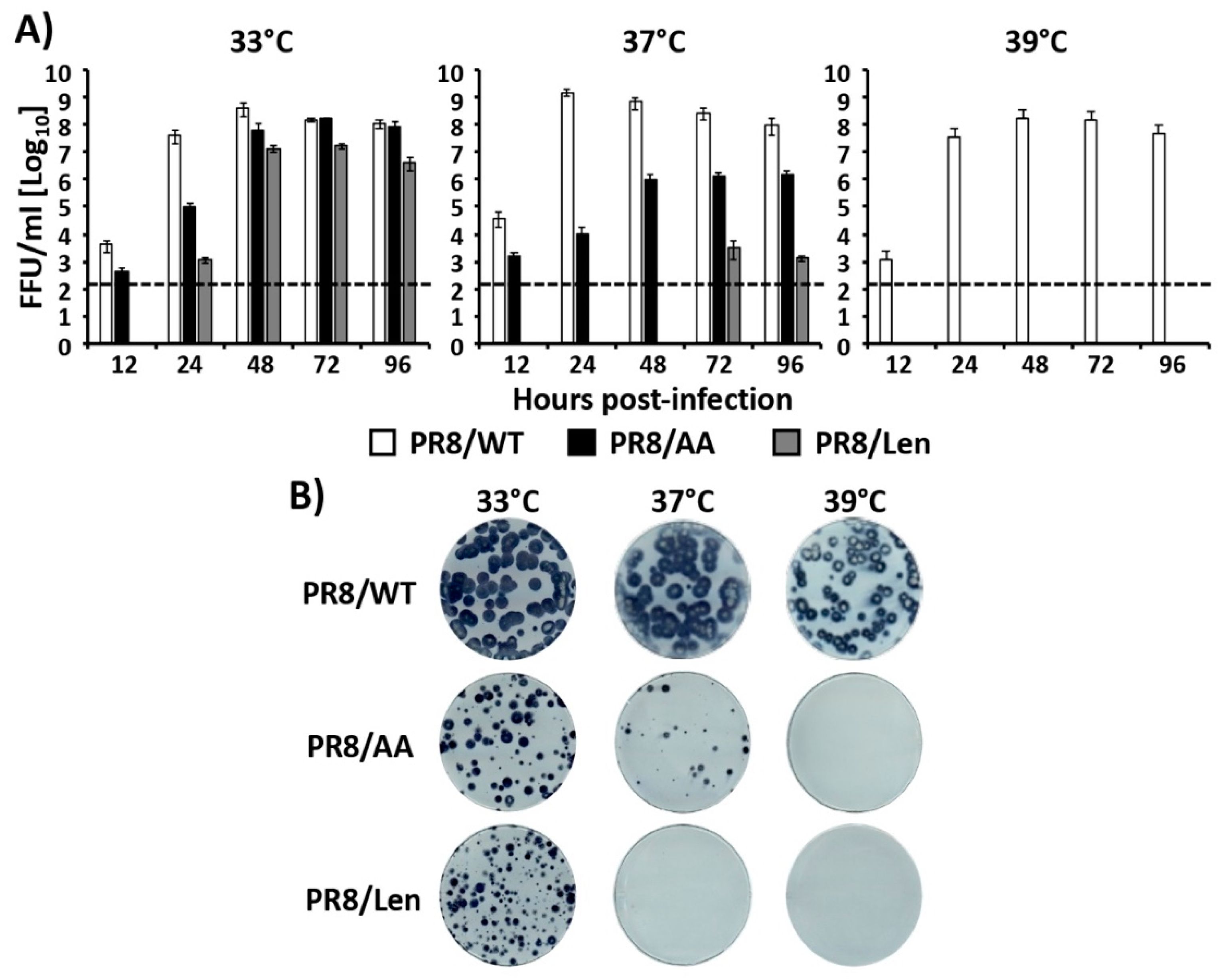
3.2. Recombinant PR8/Len is More Temperature Sensitive than PR8/AA
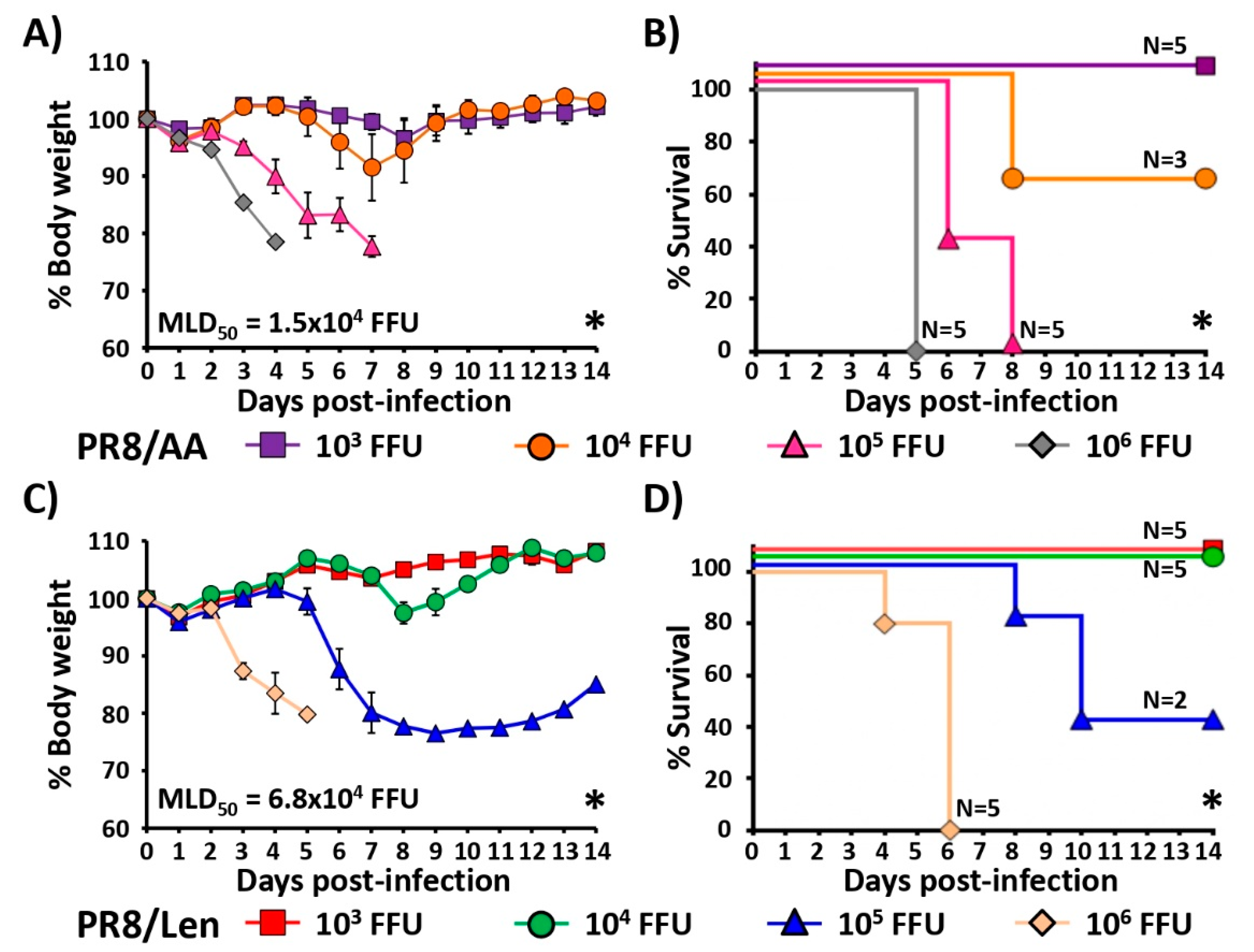
3.3. Attenuation of PR8/Len and PR8/AA in a Mouse Model of Influenza Infection
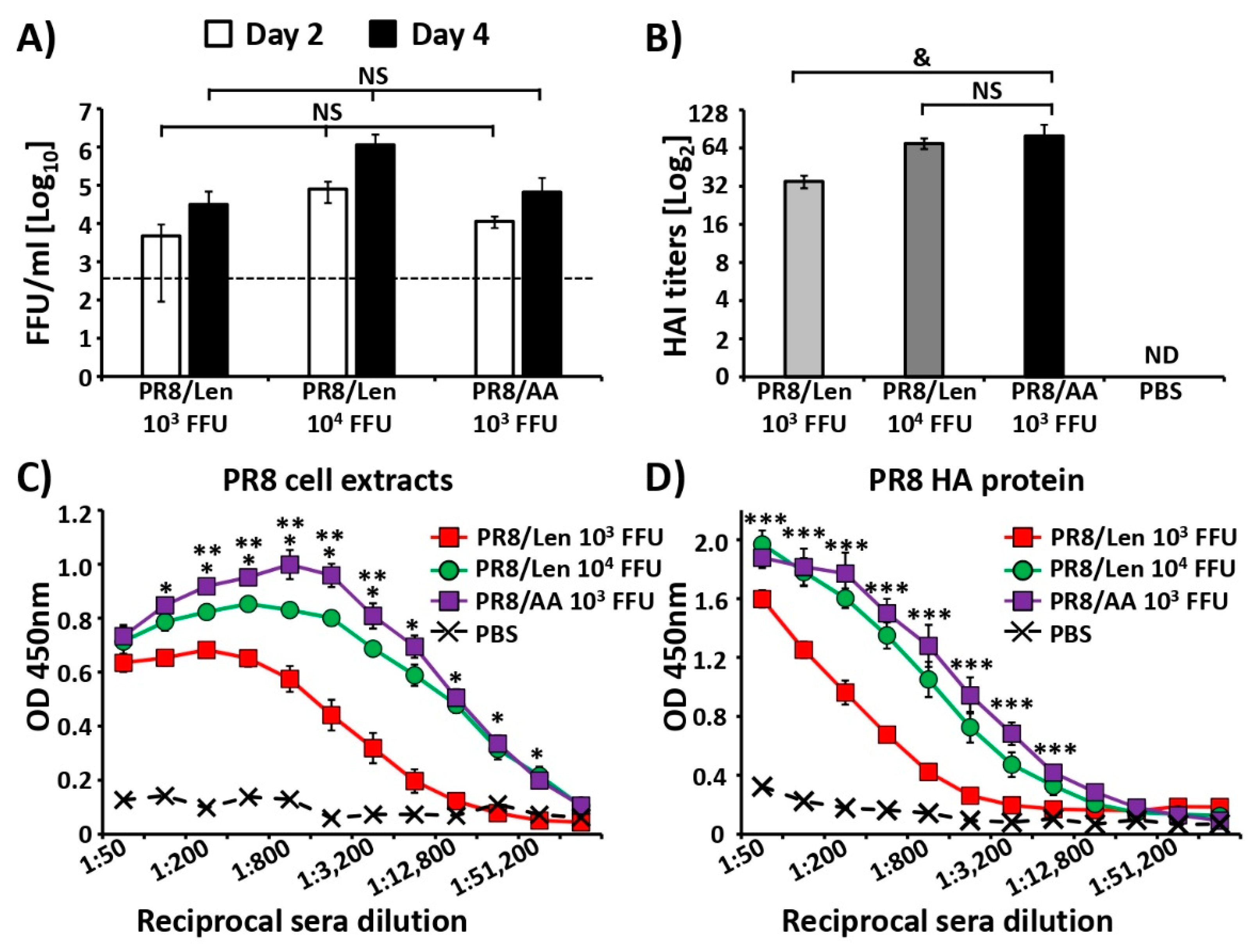
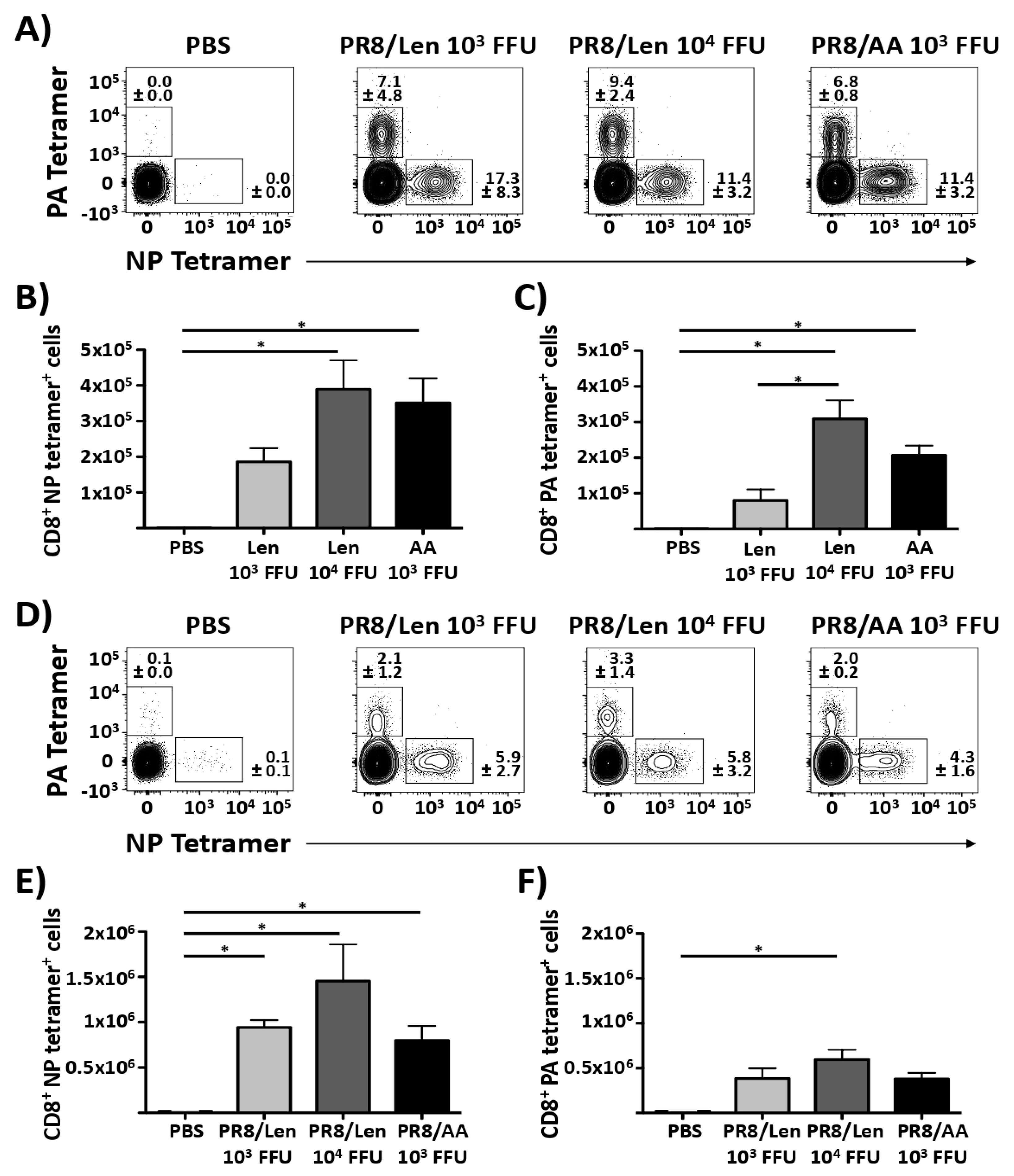
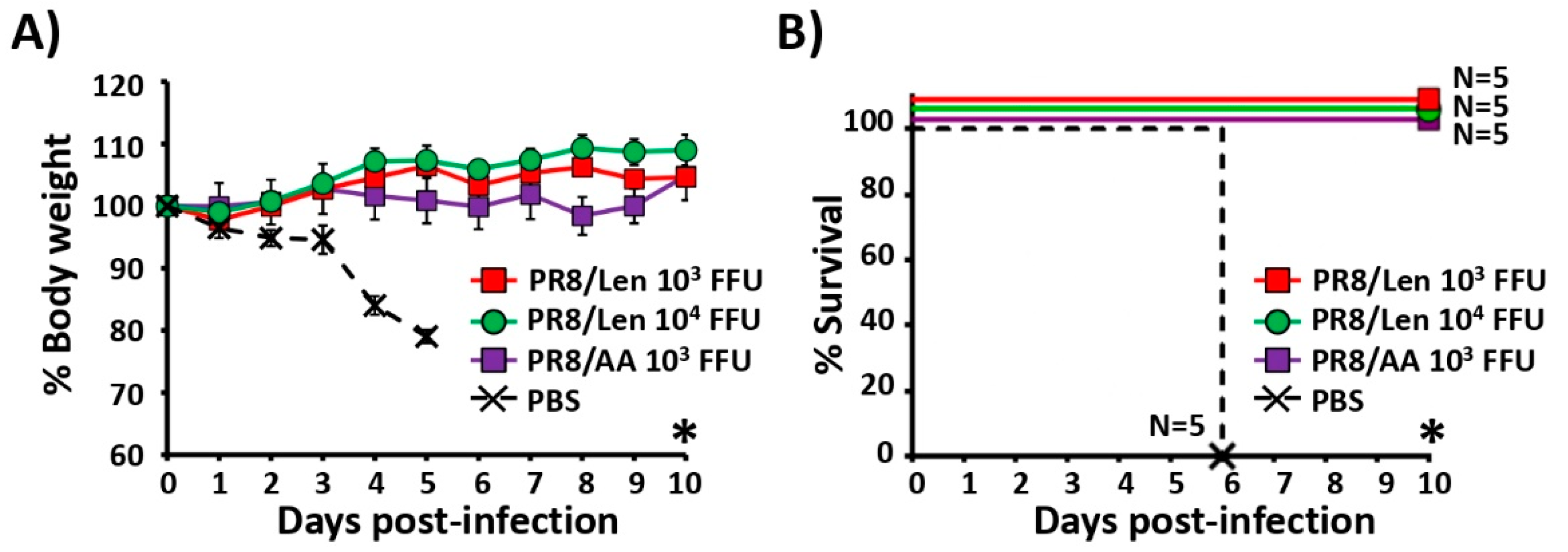
3.4. Immune Responses and Protection Efficacy of PR8/Len and PR8/AA in Mice against a Homologous Viral Challenge
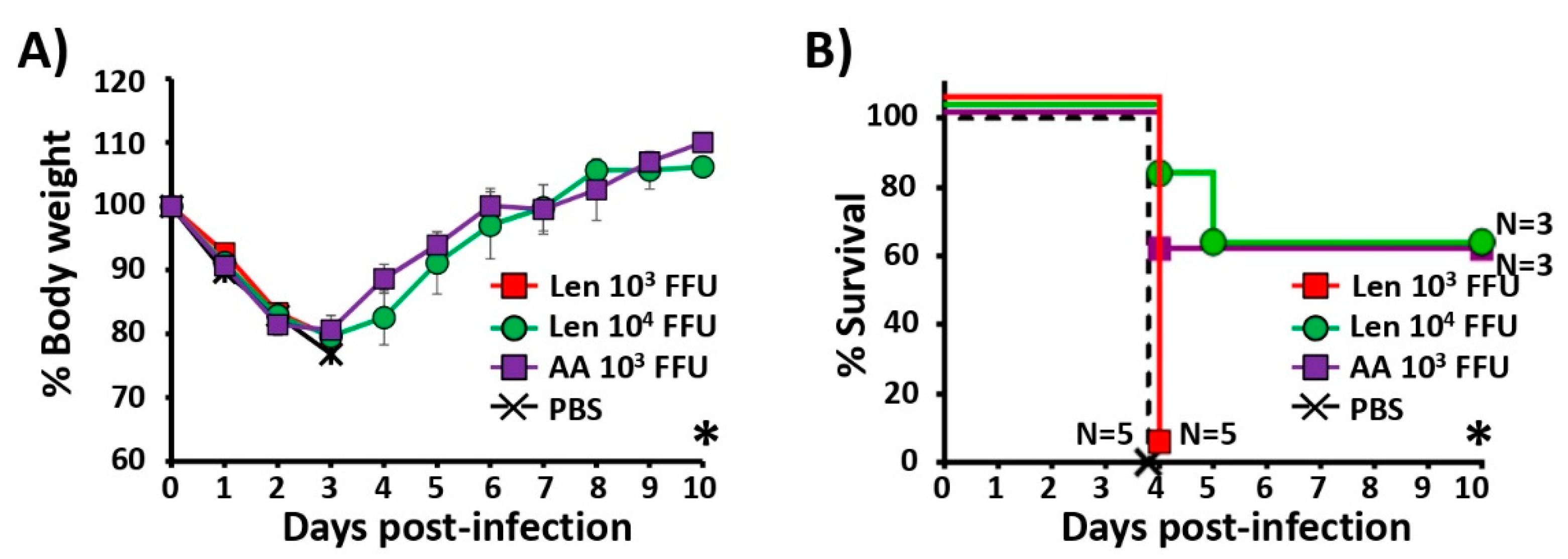
3.5. Immune Responses and Protection Efficacy of PR8/Len and PR8/AA Against a Heterologous Viral Challenge
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- CDC. Disease Burden of Influenza. 2018. Available online: https://www.cdc.gov/flu/about/disease/burden.htm#economic (accessed on 9 October 2019).
- Lynch, J.P., III; Walsh, E.E. Influenza: Evolving strategies in treatment and prevention. Semin. Respir. Crit. Care Med. 2007, 28, 144–158. [Google Scholar] [CrossRef] [PubMed]
- Barr, I.G.; McCauley, J.; Cox, N.; Daniels, R.; Engelhardt, O.G.; Fukuda, K.; Grohmann, G.; Hay, A.; Kelso, A.; Klimov, A.; et al. Epidemiological, antigenic and genetic characteristics of seasonal influenza A(H1N1), A(H3N2) and B influenza viruses: Basis for the WHO recommendation on the composition of influenza vaccines for use in the 2009–2010 Northern Hemisphere season. Vaccine 2010, 28, 1156–1167. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.J.; Vijaykrishna, D.; Bahl, J.; Lycett, S.J.; Worobey, M.; Pybus, O.G.; Ma, S.K.; Cheung, C.L.; Raghwani, J.; Bhatt, S.; et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 2009, 459, 1122–1125. [Google Scholar] [CrossRef] [PubMed]
- Pica, N.; Palese, P. Toward a universal influenza virus vaccine: Prospects and challenges. Annu. Rev. Med. 2013, 64, 189–202. [Google Scholar] [CrossRef] [PubMed]
- Belongia, E.A.; Kieke, B.A.; Donahue, J.G.; Greenlee, R.T.; Balish, A.; Foust, A.; Lindstrom, S.; Shay, D.K. Effectiveness of inactivated influenza vaccines varied substantially with antigenic match from the 2004–2005 season to the 2006–2007 season. J. Infect. Dis. 2009, 199, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Osterholm, M.T.; Kelley, N.S.; Sommer, A.; Belongia, E.A. Efficacy and effectiveness of influenza vaccines: A systematic review and meta-analysis. Lancet Infect. Dis. 2012, 12, 36–44. [Google Scholar] [CrossRef]
- Cox, M.M.; Patriarca, P.A.; Treanor, J. FluBlok, a recombinant hemagglutinin influenza vaccine. Influenza Other Respir Viruses 2008, 2, 211–219. [Google Scholar] [CrossRef]
- Palese, P.S.; Megan, L. Fields Virology, 5th ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; Volume 2. [Google Scholar]
- Hoft, D.F.; Babusis, E.; Worku, S.; Spencer, C.T.; Lottenbach, K.; Truscott, S.M.; Abate, G.; Sakala, I.G.; Edwards, K.M.; Creech, C.B.; et al. Live and inactivated influenza vaccines induce similar humoral responses, but only live vaccines induce diverse T-cell responses in young children. J. Infect. Dis. 2011, 204, 845–853. [Google Scholar] [CrossRef]
- Nogales, A.; Rodriguez, L.; Chauche, C.; Huang, K.; Reilly, E.C.; Topham, D.J.; Murcia, P.R.; Parrish, C.R.; Martinez-Sobrido, L. A temperature sensitive live-attenuated canine influenza virus H3N8 vaccine. J. Virol. 2016, 91. [Google Scholar] [CrossRef]
- Rodriguez, L.; Nogales, A.; Reilly, E.C.; Topham, D.J.; Murcia, P.R.; Parrish, C.R.; Martinez Sobrido, L. A live-attenuated influenza vaccine for H3N2 canine influenza virus. Virology 2017, 504, 96–106. [Google Scholar] [CrossRef]
- Rodriguez, L.; Reedy, S.; Nogales, A.; Murcia, P.R.; Chambers, T.M.; Martinez-Sobrido, L. Development of a novel equine influenza virus live-attenuated vaccine. Virology 2018, 516, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Maassab, H.F. Adaptation and growth characteristics of influenza virus at 25 degrees c. Nature 1967, 213, 612–614. [Google Scholar] [CrossRef] [PubMed]
- Aleksandrova, G.I.; Mikutskaia, B.A.; Pleshanova, R.A.; Panova, N.G.; Smorodintsev, A.A. The reactive and immunogenic properties and epidemiological effectiveness of further attenuated influenza virus vaccinal strains (observed in preschool children). Vopr. Virusol. 1965, 10, 67–73. [Google Scholar] [PubMed]
- Wilson, W.D.; Robinson, D. Field safety of a modified-live, cold-adapted intranasal equine influenza vaccine (HESKAFlu Avert IN vaccine) in horses. J. Equine Vet. Sci. 2000, 20, 8–10. [Google Scholar] [CrossRef]
- Youngner, J.S.; Whitaker-Dowling, P.; Chambers, T.M.; Rushlow, K.E.; Sebring, R. Derivation and characterization of a live attenuated equine influenza vaccine virus. Am. J. Vet. Res. 2001, 62, 1290–1294. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.; Zhou, H.; Kemble, G.; Jin, H. The cold adapted and temperature sensitive influenza A/Ann Arbor/6/60 virus, the master donor virus for live attenuated influenza vaccines, has multiple defects in replication at the restrictive temperature. Virology 2008, 380, 304–311. [Google Scholar] [CrossRef]
- Murphy, B.R.; Coelingh, K. Principles underlying the development and use of live attenuated cold-adapted influenza A and B virus vaccines. Viral Immunol. 2002, 15, 295–323. [Google Scholar] [CrossRef] [PubMed]
- Kohlmeier, J.E.; Woodland, D.L. Immunity to respiratory viruses. Annu. Rev. Immunol. 2009, 27, 61–82. [Google Scholar] [CrossRef]
- Rodriguez, L.; Nogales, A.; Murcia, P.R.; Parrish, C.R.; Martinez-Sobrido, L. A bivalent live-attenuated influenza vaccine for the control and prevention of H3N8 and H3N2 canine influenza viruses. Vaccine 2017, 35, 4374–4381. [Google Scholar] [CrossRef]
- Cheng, X.; Zengel, J.R.; Suguitan, A.L., Jr.; Xu, Q.; Wang, W.; Lin, J.; Jin, H. Evaluation of the humoral and cellular immune responses elicited by the live attenuated and inactivated influenza vaccines and their roles in heterologous protection in ferrets. J. Infect. Dis. 2013, 208, 594–602. [Google Scholar] [CrossRef] [PubMed]
- Victor, S.T.; Watanabe, S.; Katsura, H.; Ozawa, M.; Kawaoka, Y. A replication-incompetent PB2-knockout influenza A virus vaccine vector. J. Virol. 2012. [Google Scholar] [CrossRef]
- De Villiers, P.J.; Steele, A.D.; Hiemstra, L.A.; Rappaport, R.; Dunning, A.J.; Gruber, W.C.; Forrest, B.D. Efficacy and safety of a live attenuated influenza vaccine in adults 60 years of age and older. Vaccine 2009, 28, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Baker, S.F.; Guo, H.; Albrecht, R.A.; Garcia-Sastre, A.; Topham, D.J.; Martinez-Sobrido, L. Protection against Lethal Influenza with a Viral Mimic. J. Virol. 2013, 87, 8591–8605. [Google Scholar] [CrossRef] [PubMed]
- Powell, T.J.; Silk, J.D.; Sharps, J.; Fodor, E.; Townsend, A.R. Pseudotyped influenza A virus as a vaccine for the induction of heterotypic immunity. J. Virol. 2012, 86, 13397–13406. [Google Scholar] [CrossRef] [PubMed]
- Uraki, R.; Kiso, M.; Iwatsuki-Horimoto, K.; Fukuyama, S.; Takashita, E.; Ozawa, M.; Kawaoka, Y. A novel bivalent vaccine based on a PB2-knockout influenza virus protects mice from pandemic H1N1 and highly pathogenic H5N1 virus challenges. J. Virol. 2013, 87, 7874–7881. [Google Scholar] [CrossRef]
- Katsura, H.; Iwatsuki-Horimoto, K.; Fukuyama, S.; Watanabe, S.; Sakabe, S.; Hatta, Y.; Murakami, S.; Shimojima, M.; Horimoto, T.; Kawaoka, Y. A replication-incompetent virus possessing an uncleavable hemagglutinin as an influenza vaccine. Vaccine 2012, 30, 6027–6033. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Recommendations to assure the quality, safety and efficacy of influenza vaccines (human, live attenuated) for intranasal administration. WHO Tech. Rep. Ser. 2013, 977, 153–227. [Google Scholar]
- World Health Organization (WHO). WHO Meeting on Live Attenuated Influenza Vaccine Effectiveness; WHO: Geneva, Switzerland, 2016. [Google Scholar]
- Rudenko, L.; Yeolekar, L.; Kiseleva, I.; Isakova-Sivak, I. Development and approval of live attenuated influenza vaccines based on Russian master donor viruses: Process challenges and success stories. Vaccine 2016, 34, 5436–5441. [Google Scholar] [CrossRef]
- Cox, N.J.; Kitame, F.; Kendal, A.P.; Maassab, H.F.; Naeve, C. Identification of sequence changes in the cold-adapted, live attenuated influenza vaccine strain, A/Ann Arbor/6/60 (H2N2). Virology 1988, 167, 554–567. [Google Scholar] [CrossRef]
- Snyder, M.H.; Betts, R.F.; DeBorde, D.; Tierney, E.L.; Clements, M.L.; Herrington, D.; Sears, S.D.; Dolin, R.; Maassab, H.F.; Murphy, B.R. Four viral genes independently contribute to attenuation of live influenza A/Ann Arbor/6/60 (H2N2) cold-adapted reassortant virus vaccines. J. Virol. 1988, 62, 488–495. [Google Scholar]
- Isakova-Sivak, I.; Chen, L.M.; Matsuoka, Y.; Voeten, J.T.; Kiseleva, I.; Heldens, J.G.; den Bosch, H.; Klimov, A.; Rudenko, L.; Cox, N.J.; et al. Genetic bases of the temperature-sensitive phenotype of a master donor virus used in live attenuated influenza vaccines: A/Leningrad/134/17/57 (H2N2). Virology 2011, 412, 297–305. [Google Scholar] [CrossRef] [PubMed]
- Cox, A.; Baker, S.F.; Nogales, A.; Martinez-Sobrido, L.; Dewhurst, S. Development of a mouse-adapted live attenuated influenza virus that permits in vivo analysis of enhancements to the safety of live attenuated influenza virus vaccine. J. Virol. 2015, 89, 3421–3426. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Zhou, H.; Lu, B.; Kemble, G. Imparting temperature sensitivity and attenuation in ferrets to A/Puerto Rico/8/34 influenza virus by transferring the genetic signature for temperature sensitivity from cold-adapted A/Ann Arbor/6/60. J. Virol. 2004, 78, 995–998. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Li, Y.; Speer, S.D.; Subba, A.; Lin, X.; Wentworth, D.E. Engineering temperature sensitive live attenuated influenza vaccines from emerging viruses. Vaccine 2012, 30, 3691–3702. [Google Scholar] [CrossRef] [PubMed]
- Nogales, A.; Rodriguez, L.; DeDiego, M.L.; Topham, D.J.; Martinez-Sobrido, L. Interplay of PA-X and NS1 Proteins in Replication and Pathogenesis of a Temperature-Sensitive 2009 Pandemic H1N1 Influenza A Virus. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Broadbent, A.J.; Santos, C.P.; Godbout, R.A.; Subbarao, K. The temperature-sensitive and attenuation phenotypes conferred by mutations in the influenza virus PB2, PB1, and NP genes are influenced by the species of origin of the PB2 gene in reassortant viruses derived from influenza A/California/07/2009 and A/WSN/33 viruses. J. Virol. 2014, 88, 12339–12347. [Google Scholar] [CrossRef] [PubMed]
- Czako, R.; Vogel, L.; Sutton, T.; Matsuoka, Y.; Krammer, F.; Chen, Z.; Jin, H.; Subbarao, K. H5N2 vaccine viruses on Russian and US live attenuated influenza virus backbones demonstrate similar infectivity, immunogenicity and protection in ferrets. Vaccine 2018, 36, 1871–1879. [Google Scholar] [CrossRef]
- Chen, Z.; Baz, M.; Lu, J.; Paskel, M.; Santos, C.; Subbarao, K.; Jin, H.; Matsuoka, Y. Development of a high-yield live attenuated H7N9 influenza virus vaccine that provides protection against homologous and heterologous H7 wild-type viruses in ferrets. J. Virol. 2014, 88, 7016–7023. [Google Scholar] [CrossRef]
- Peng, Y.; Wang, B.; Talaat, K.; Karron, R.; Powell, T.J.; Zeng, H.; Dong, D.; Luke, C.J.; McMichael, A.; Subbarao, K.; et al. Boosted Influenza-Specific T Cell Responses after H5N1 Pandemic Live Attenuated Influenza Virus Vaccination. Front. Immunol. 2015, 6, 287. [Google Scholar] [CrossRef]
- Martinez-Sobrido, L.; Garcia-Sastre, A. Generation of recombinant influenza virus from plasmid DNA. J. Vis. Exp. 2010. [Google Scholar] [CrossRef]
- Grimm, D.; Staeheli, P.; Hufbauer, M.; Koerner, I.; Martinez-Sobrido, L.; Solorzano, A.; Garcia-Sastre, A.; Haller, O.; Kochs, G. Replication fitness determines high virulence of influenza A virus in mice carrying functional Mx1 resistance gene. Proc. Natl. Acad. Sci. USA 2007, 104, 6806–6811. [Google Scholar] [CrossRef] [PubMed]
- Kilbourne, E.D. Future influenza vaccines and the use of genetic recombinants. Bull. World Health Organ. 1969, 41, 643–645. [Google Scholar] [PubMed]
- Nogales, A.; Baker, S.F.; Ortiz-Riano, E.; Dewhurst, S.; Topham, D.J.; Martinez-Sobrido, L. Influenza A Virus Attenuation by Codon Deoptimization of the NS Gene for Vaccine Development. J. Virol. 2014, 88, 10525–10540. [Google Scholar] [CrossRef] [PubMed]
- Schickli, J.H.; Flandorfer, A.; Nakaya, T.; Martinez-Sobrido, L.; Garcia-Sastre, A.; Palese, P. Plasmid-only rescue of influenza A virus vaccine candidates. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2001, 356, 1965–1973. [Google Scholar] [CrossRef] [PubMed]
- National Research Council (U.S.); Committee for the Update of the Guide for the Care and Use of Laboratory Animals; Institute for Laboratory Animal Research (U.S.); National Academies Press (U.S.). Guide for the Care and Use of Laboratory Animals, 8th ed.; National Academies Press: Washington, DC, USA, 2011; p. 220.
- Rodriguez, L.; Nogales, A.; Martinez-Sobrido, L. Influenza A Virus Studies in a Mouse Model of Infection. J. Vis. Exp. 2017. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J.; Muench, H. A simple method of estimating fifty percent endpoints. Am. J. Hyg. 1938, 27, 493–497. [Google Scholar]
- Nogales, A.; DeDiego, M.L.; Topham, D.J.; Martinez-Sobrido, L. Rearrangement of Influenza Virus Spliced Segments for the Development of Live-Attenuated Vaccines. J. Virol. 2016, 90, 6291–6302. [Google Scholar] [CrossRef]
- Reilly, E.C.; Lambert-Emo, K.; Topham, D.J. The Effects of Acute Neutrophil Depletion on Resolution of Acute Influenza Infection, Establishment of Tissue Resident Memory (TRM), and Heterosubtypic Immunity. PLoS ONE 2016, 11, e0164274. [Google Scholar] [CrossRef]
- Belshe, R.; Lee, M.S.; Walker, R.E.; Stoddard, J.; Mendelman, P.M. Safety, immunogenicity and efficacy of intranasal, live attenuated influenza vaccine. Expert Rev. Vaccines 2004, 3, 643–654. [Google Scholar] [CrossRef]
- Belshe, R.B.; Edwards, K.M.; Vesikari, T.; Black, S.V.; Walker, R.E.; Hultquist, M.; Kemble, G.; Connor, E.M.; Group, C.T.C.E.S. Live attenuated versus inactivated influenza vaccine in infants and young children. N. Engl. J. Med. 2007, 356, 685–696. [Google Scholar] [CrossRef]
- Gorse, G.J.; Belshe, R.B.; Munn, N.J. Superiority of live attenuated compared with inactivated influenza A virus vaccines in older, chronically ill adults. Chest 1991, 100, 977–984. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.G.; Slepushkin, A.N.; Monto, A.S.; Kendal, A.P.; Grigorieva, E.P.; Burtseva, E.P.; Rekstin, A.R.; Beljaev, A.L.; Bragina, V.E.; Cox, N.; et al. Efficacy of live attenuated and inactivated influenza vaccines in schoolchildren and their unvaccinated contacts in Novgorod, Russia. J. Infect. Dis. 1993, 168, 881–887. [Google Scholar] [CrossRef] [PubMed]
- Slepushkin, A.N.; Obrosova-Serova, N.P.; Burtseva, E.I.; Rudenko, L.G.; Govorkova, E.A.; Vartanyan, R.V.; Verestsinsky, A.I.; Lonskaya, N.I.; Harmon, M.W.; Torok, T.; et al. Comparison of live attenuated and inactivated influenza vaccines in schoolchildren in Russia: I. Safety and efficacy in two Moscow schools, 1987/88. Vaccine 1993, 11, 323–328. [Google Scholar] [CrossRef]
- Lukarska, M.; Fournier, G.; Pflug, A.; Resa-Infante, P.; Reich, S.; Naffakh, N.; Cusack, S. Structural basis of an essential interaction between influenza polymerase and Pol II CTD. Nature 2017, 541, 117–121. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. The PyMOL Molecular Graphics System; Schrodinger LLC: New York, NY, USA, 2002. [Google Scholar]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodriguez, L.; Blanco-Lobo, P.; Reilly, E.C.; Maehigashi, T.; Nogales, A.; Smith, A.; Topham, D.J.; Dewhurst, S.; Kim, B.; Martínez-Sobrido, L. Comparative Study of the Temperature Sensitive, Cold Adapted and Attenuated Mutations Present in the Master Donor Viruses of the Two Commercial Human Live Attenuated Influenza Vaccines. Viruses 2019, 11, 928. https://doi.org/10.3390/v11100928
Rodriguez L, Blanco-Lobo P, Reilly EC, Maehigashi T, Nogales A, Smith A, Topham DJ, Dewhurst S, Kim B, Martínez-Sobrido L. Comparative Study of the Temperature Sensitive, Cold Adapted and Attenuated Mutations Present in the Master Donor Viruses of the Two Commercial Human Live Attenuated Influenza Vaccines. Viruses. 2019; 11(10):928. https://doi.org/10.3390/v11100928
Chicago/Turabian StyleRodriguez, Laura, Pilar Blanco-Lobo, Emma C. Reilly, Tatsuya Maehigashi, Aitor Nogales, Andrew Smith, David J. Topham, Stephen Dewhurst, Baek Kim, and Luis Martínez-Sobrido. 2019. "Comparative Study of the Temperature Sensitive, Cold Adapted and Attenuated Mutations Present in the Master Donor Viruses of the Two Commercial Human Live Attenuated Influenza Vaccines" Viruses 11, no. 10: 928. https://doi.org/10.3390/v11100928
APA StyleRodriguez, L., Blanco-Lobo, P., Reilly, E. C., Maehigashi, T., Nogales, A., Smith, A., Topham, D. J., Dewhurst, S., Kim, B., & Martínez-Sobrido, L. (2019). Comparative Study of the Temperature Sensitive, Cold Adapted and Attenuated Mutations Present in the Master Donor Viruses of the Two Commercial Human Live Attenuated Influenza Vaccines. Viruses, 11(10), 928. https://doi.org/10.3390/v11100928





