From SARS to MERS, Thrusting Coronaviruses into the Spotlight
Abstract
1. Introduction
2. Epidemiology of SARS-CoV and MERS-CoV
3. Pathogenesis of SARS-CoV and MERS-CoV
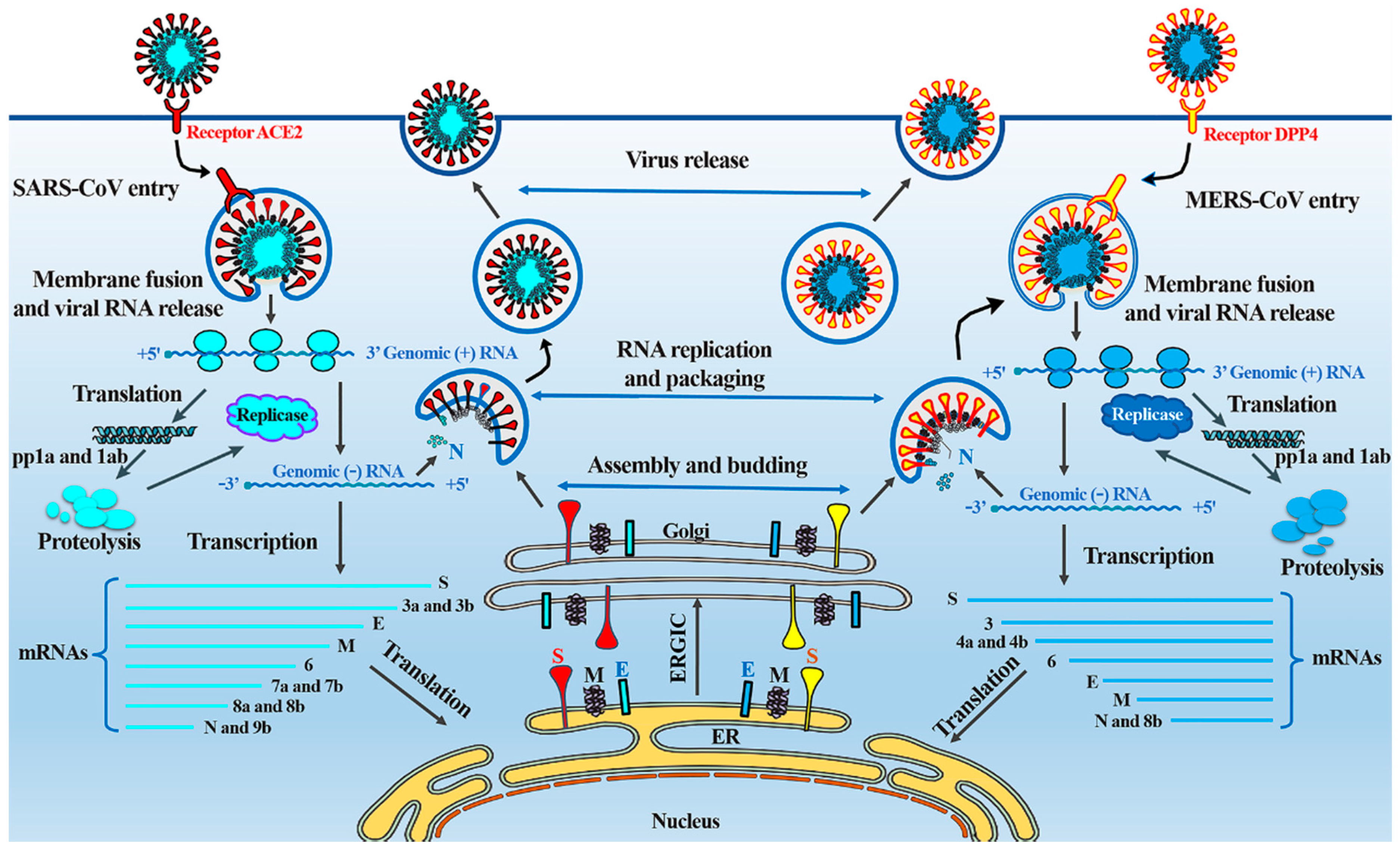
4. Comparative Pathology and Life Cycles of SARS-CoV and MERS-CoV
5. SARS and MERS-CoV Spike Protein: A Key Target for Antivirals
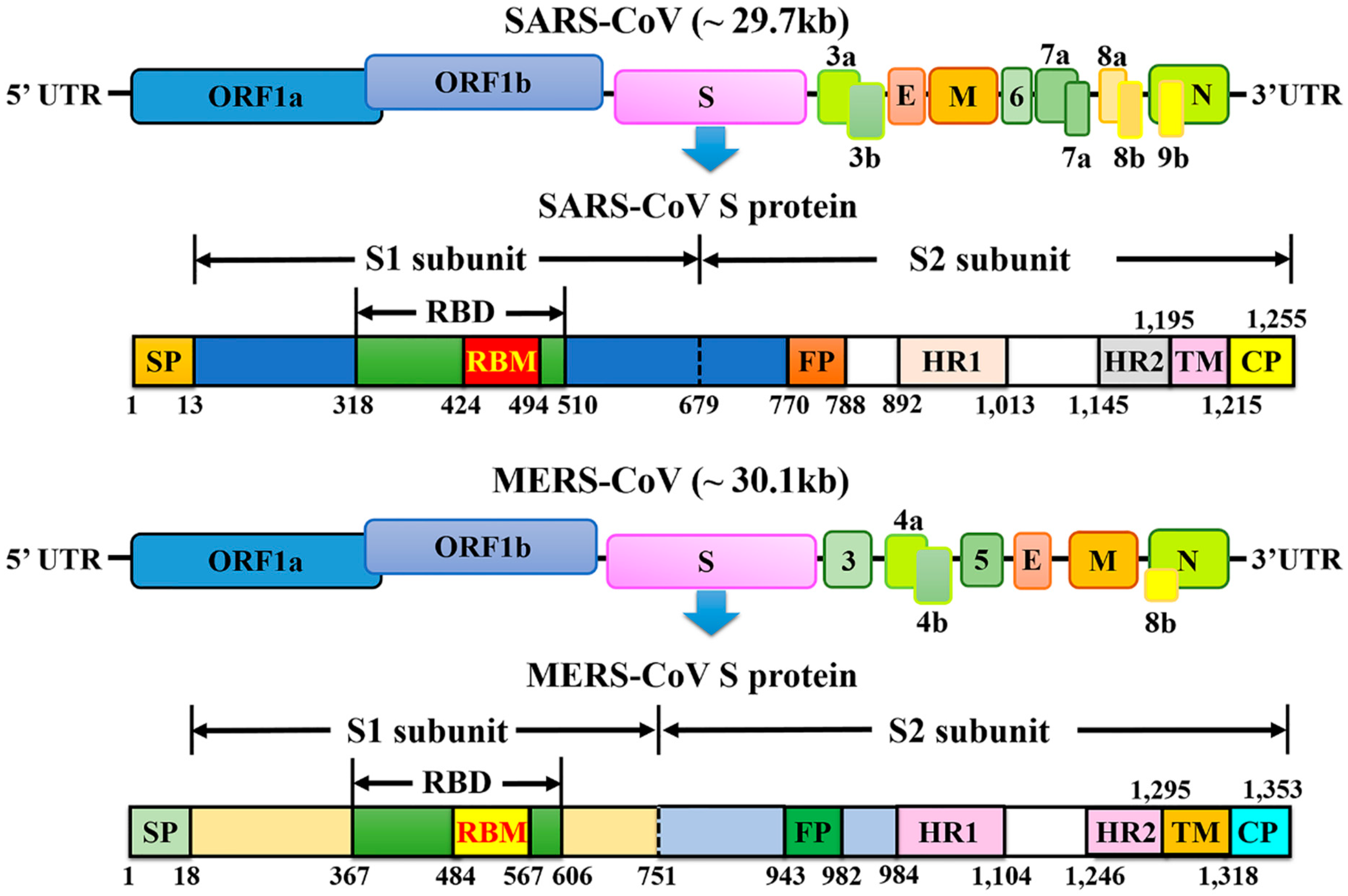
5.1. Structure of the SARS-CoV and MERS-CoV Spike Protein
5.2. Functions of the SARS-CoV and MERS-CoV S Protein
5.3. Vaccines Based on the SARS-CoV and MERS-CoV S Protein
5.4. S Protein-Based Therapeutics for SARS-CoV and MERS-CoV
6. The Animal Models of SARS and MERS-CoV
6.1. Animal Models of SARS-CoV
6.1.1. Mouse Models
6.1.2. Hamster Model
6.1.3. Ferret Model
6.1.4. Non-Human Primate Models
6.2. Animal Models of MERS-CoV
6.2.1. Mouse Model for MERS Infection
6.2.2. Camelidae
6.2.3. Non-Human Primates
7. Role of Host Receptors in Animal Models of SARS-CoV and MERS-CoV
8. Outlook and Summary
Funding
Acknowledgments
Conflicts of Interest
References
- Yin, Y.; Wunderink, R.G. MERS, SARS and other coronaviruses as causes of pneumonia. Respirology 2018, 23, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Drosten, C.; Gunther, S.; Preiser, W.; van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef] [PubMed]
- Ksiazek, T.G.; Erdman, D.; Goldsmith, C.S.; Zaki, S.R.; Peret, T.; Emery, S.; Tong, S.; Urbani, C.; Comer, J.A.; Lim, W.; et al. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1953–1966. [Google Scholar] [CrossRef] [PubMed]
- Rota, P.A. Characterization of a Novel Coronavirus Associated with Severe Acute Respiratory Syndrome. Science 2003, 300, 1394–1399. [Google Scholar] [CrossRef] [PubMed]
- Zhong, N.S.; Zheng, B.J.; Li, Y.M.; Poon, L.L.; Xie, Z.H.; Chan, K.H.; Li, P.H.; Tan, S.Y.; Chang, Q.; Xie, J.P.; et al. Epidemiology and cause of severe acute respiratory syndrome (SARS) in Guangdong, People’s Republic of China, in February, 2003. Lancet 2003, 362, 1353–1358. [Google Scholar] [CrossRef]
- Lee, N.; Hui, D.; Wu, A.; Chan, P.; Cameron, P.; Joynt, G.M.; Ahuja, A.; Yung, M.Y.; Leung, C.B.; To, K.F.; et al. A major outbreak of severe acute respiratory syndrome in Hong Kong. N. Engl. J. Med. 2003, 348, 1986–1994. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.Y.; Li, J.L.; Yang, X.L.; Chmura, A.A.; Zhu, G.; Epstein, J.H.; Mazet, J.K.; Hu, B.; Zhang, W.; Peng, C.; et al. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 2013, 503, 535–538. [Google Scholar] [CrossRef]
- Menachery, V.D.; Yount, B.J.; Debbink, K.; Agnihothram, S.; Gralinski, L.E.; Plante, J.A.; Graham, R.L.; Scobey, T.; Ge, X.Y.; Donaldson, E.F.; et al. A SARS-like cluster of circulating bat coronaviruses shows potential for human emergence. Nat. Med. 2015, 21, 1508–1513. [Google Scholar] [CrossRef]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- de Wit, E.; van Doremalen, N.; Falzarano, D.; Munster, V.J. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef]
- World Health Organization. WHO Guidelines for the Global Surveillance of Severe Acute Respiratory Syndrome (SARS). Updated Recommendations. October 2004. Available online: http://www.who.int/mediacentre/factsheets/mers-cov/en/ (accessed on 8 October 2018).
- Guan, Y.; Zheng, B.J.; He, Y.Q.; Liu, X.L.; Zhuang, Z.X.; Cheung, C.L.; Luo, S.W.; Li, P.H.; Zhang, L.J.; Guan, Y.J.; et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 2003, 302, 276–278. [Google Scholar] [CrossRef]
- Azhar, E.I.; El-Kafrawy, S.A.; Farraj, S.A.; Hassan, A.M.; Al-Saeed, M.S.; Hashem, A.M.; Madani, T.A. Evidence for camel-to-human transmission of MERS coronavirus. N. Engl. J. Med. 2014, 370, 2499–2505. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Alshukairi, A.N.; Zheng, J.; Zhao, J.; Nehdi, A.; Baharoon, S.A.; Layqah, L.; Bokhari, A.; Al, J.S.; Samman, N.; Boudjelal, M.; et al. High Prevalence of MERS-CoV Infection in Camel Workers in Saudi Arabia. mBio 2018, 9, e01985-18. [Google Scholar] [CrossRef] [PubMed]
- Assiri, A.; Al-Tawfiq, J.A.; Al-Rabeeah, A.A.; Al-Rabiah, F.A.; Al-Hajjar, S.; Al-Barrak, A.; Flemban, H.; Al-Nassir, W.N.; Balkhy, H.H.; Al-Hakeem, R.F.; et al. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. Lancet Infect. Dis. 2013, 13, 752–761. [Google Scholar] [CrossRef]
- Arabi, Y.M.; Arifi, A.A.; Balkhy, H.H.; Najm, H.; Aldawood, A.S.; Ghabashi, A.; Hawa, H.; Alothman, A.; Khaldi, A.; Al, R.B. Clinical course and outcomes of critically ill patients with Middle East respiratory syndrome coronavirus infection. Ann. Intern. Med. 2014, 160, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Al-Abdallat, M.M.; Payne, D.C.; Alqasrawi, S.; Rha, B.; Tohme, R.A.; Abedi, G.R.; Al, N.M.; Iblan, I.; Jarour, N.; Farag, N.H.; et al. Hospital-associated outbreak of Middle East respiratory syndrome coronavirus: A serologic, epidemiologic, and clinical description. Clin. Infect. Dis. 2014, 59, 1225–1233. [Google Scholar] [CrossRef] [PubMed]
- Al, H.F.; Pringle, K.; Al, M.M.; Kim, L.; Pham, H.; Alami, N.N.; Khudhair, A.; Hall, A.J.; Aden, B.; El, S.F.; et al. Response to Emergence of Middle East Respiratory Syndrome Coronavirus, Abu Dhabi, United Arab Emirates, 2013–2014. Emerg. Infect. Dis. 2016, 22, 1162–1168. [Google Scholar]
- Assiri, A.; McGeer, A.; Perl, T.M.; Price, C.S.; Al, R.A.; Cummings, D.A.; Alabdullatif, Z.N.; Assad, M.; Almulhim, A.; Makhdoom, H.; et al. Hospital outbreak of Middle East respiratory syndrome coronavirus. N. Engl. J. Med. 2013, 369, 407–416. [Google Scholar] [CrossRef]
- Drosten, C.; Muth, D.; Corman, V.M.; Hussain, R.; Al, M.M.; HajOmar, W.; Landt, O.; Assiri, A.; Eckerle, I.; Al, S.A.; et al. An observational, laboratory-based study of outbreaks of middle East respiratory syndrome coronavirus in Jeddah and Riyadh, kingdom of Saudi Arabia, 2014. Clin. Infect. Dis. 2015, 60, 369–377. [Google Scholar] [CrossRef]
- Park, H.Y.; Lee, E.J.; Ryu, Y.W.; Kim, Y.; Kim, H.; Lee, H.; Yi, S.J. Epidemiological investigation of MERS-CoV spread in a single hospital in South Korea, May to June 2015. Eurosurveillance 2015, 20, 21169. [Google Scholar] [CrossRef] [PubMed]
- Skowronski, D.M.; Astell, C.; Brunham, R.C.; Low, D.E.; Petric, M.; Roper, R.L.; Talbot, P.J.; Tam, T.; Babiuk, L. Severe acute respiratory syndrome (SARS): A year in review. Annu. Rev. Med. 2005, 56, 357–381. [Google Scholar] [CrossRef] [PubMed]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar]
- Wang, Y.; Sun, J.; Zhu, A.; Zhao, J.; Zhao, J. Current understanding of middle east respiratory syndrome coronavirus infection in human and animal models. J. Thorac. Dis. 2018, 10, S2260–S2271. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Yang, Y.; Zhou, Y.; Lu, L.; Li, F.; Jiang, S. MERS-CoV spike protein: A key target for antivirals. Expert Opin. Ther. Targets 2017, 21, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; He, Y.; Zhou, Y.; Liu, S.; Zheng, B.; Jiang, S. The spike protein of SARS-CoV—A target for vaccine and therapeutic development. Nat. Rev. Microbiol. 2009, 7, 226–236. [Google Scholar] [CrossRef]
- Zumla, A.; Hui, D.S.; Perlman, S. Middle East respiratory syndrome. Lancet 2015, 386, 995–1007. [Google Scholar] [CrossRef]
- Jeffers, S.A.; Tusell, S.M.; Gillim-Ross, L.; Hemmila, E.M.; Achenbach, J.E.; Babcock, G.J.; Thomas, W.J.; Thackray, L.B.; Young, M.D.; Mason, R.J.; et al. CD209L (L-SIGN) is a receptor for severe acute respiratory syndrome coronavirus. Proc. Natl. Acad. Sci. USA 2004, 101, 15748–15753. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wei, Q.; Alvarez, X.; Wang, H.; Du, Y.; Zhu, H.; Jiang, H.; Zhou, J.; Lam, P.; Zhang, L.; et al. Epithelial Cells Lining Salivary Gland Ducts Are Early Target Cells of Severe Acute Respiratory Syndrome Coronavirus Infection in the Upper Respiratory Tracts of Rhesus Macaques. J. Virol. 2011, 85, 4025–4030. [Google Scholar] [CrossRef] [PubMed]
- Kuba, K.; Imai, Y.; Rao, S.; Gao, H.; Guo, F.; Guan, B.; Huan, Y.; Yang, P.; Zhang, Y.; Deng, W.; et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 2005, 11, 875–879. [Google Scholar] [CrossRef] [PubMed]
- Qinfen, Z.; Jinming, C.; Xiaojun, H.; Huanying, Z.; Jicheng, H.; Ling, F.; Kunpeng, L.; Jingqiang, Z. The life cycle of SARS coronavirus in Vero E6 cells. J. Med. Virol. 2004, 73, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Korteweg, C.; McNutt, M.A.; Gu, J. Pathogenetic mechanisms of severe acute respiratory syndrome. Virus Res. 2008, 133, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Korteweg, C. Pathology and pathogenesis of severe acute respiratory syndrome. Am. J. Pathol. 2007, 170, 1136–1147. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; He, L.; Zhang, Q.; Huang, Z.; Che, X.; Hou, J.; Wang, H.; Shen, H.; Qiu, L.; Li, Z.; et al. Organ distribution of severe acute respiratory syndrome (SARS) associated coronavirus (SARS-CoV) in SARS patients: Implications for pathogenesis and virus transmission pathways. J. Pathol. 2004, 203, 622–630. [Google Scholar] [CrossRef] [PubMed]
- Meyerholz, D.K.; Lambertz, A.M.; McCray, P.J. Dipeptidyl Peptidase 4 Distribution in the Human Respiratory Tract: Implications for the Middle East Respiratory Syndrome. Am. J. Pathol. 2016, 186, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Widagdo, W.; Raj, V.S.; Schipper, D.; Kolijn, K.; van Leenders, G.; Bosch, B.J.; Bensaid, A.; Segales, J.; Baumgartner, W.; Osterhaus, A.; et al. Differential Expression of the Middle East Respiratory Syndrome Coronavirus Receptor in the Upper Respiratory Tracts of Humans and Dromedary Camels. J. Virol. 2016, 90, 4838–4842. [Google Scholar] [CrossRef]
- Oboho, I.K.; Tomczyk, S.M.; Al-Asmari, A.M.; Banjar, A.A.; Al-Mugti, H.; Aloraini, M.S.; Alkhaldi, K.Z.; Almohammadi, E.L.; Alraddadi, B.M.; Gerber, S.I.; et al. 2014 MERS-CoV outbreak in Jeddah—A link to health care facilities. N. Engl. J. Med. 2015, 372, 846–854. [Google Scholar] [CrossRef]
- Chu, K.H.; Tsang, W.K.; Tang, C.S.; Lam, M.F.; Lai, F.M.; To, K.F.; Fung, K.S.; Tang, H.L.; Yan, W.W.; Chan, H.W.; et al. Acute renal impairment in coronavirus-associated severe acute respiratory syndrome. Kidney Int. 2005, 67, 698–705. [Google Scholar] [CrossRef]
- Saad, M.; Omrani, A.S.; Baig, K.; Bahloul, A.; Elzein, F.; Matin, M.A.; Selim, M.A.; Al, M.M.; Al, N.D.; Al, A.A.; et al. Clinical aspects and outcomes of 70 patients with Middle East respiratory syndrome coronavirus infection: A single-center experience in Saudi Arabia. Int. J. Infect. Dis. 2014, 29, 301–306. [Google Scholar] [CrossRef]
- Alagaili, A.N.; Briese, T.; Mishra, N.; Kapoor, V.; Sameroff, S.C.; Burbelo, P.D.; de Wit, E.; Munster, V.J.; Hensley, L.E.; Zalmout, I.S.; et al. Middle East respiratory syndrome coronavirus infection in dromedary camels in Saudi Arabia. mBio 2014, 5, e814–e884. [Google Scholar] [CrossRef]
- Lambeir, A.M.; Durinx, C.; Scharpe, S.; De Meester, I. Dipeptidyl-peptidase IV from bench to bedside: An update on structural properties, functions, and clinical aspects of the enzyme DPP IV. Crit. Rev. Clin. Lab. Sci. 2003, 40, 209–294. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Zhou, J.; Wong, B.H.; Li, C.; Cheng, Z.S.; Lin, X.; Poon, V.K.; Sun, T.; Lau, C.C.; Chan, J.F.; et al. Productive replication of Middle East respiratory syndrome coronavirus in monocyte-derived dendritic cells modulates innate immune response. Virology 2014, 454–455, 197–205. [Google Scholar] [CrossRef]
- Zhou, J.; Chu, H.; Li, C.; Wong, B.H.; Cheng, Z.S.; Poon, V.K.; Sun, T.; Lau, C.C.; Wong, K.K.; Chan, J.Y.; et al. Active replication of Middle East respiratory syndrome coronavirus and aberrant induction of inflammatory cytokines and chemokines in human macrophages: Implications for pathogenesis. J. Infect. Dis. 2014, 209, 1331–1342. [Google Scholar] [CrossRef]
- Chu, H.; Zhou, J.; Wong, B.H.; Li, C.; Chan, J.F.; Cheng, Z.S.; Yang, D.; Wang, D.; Lee, A.C.; Li, C.; et al. Middle East Respiratory Syndrome Coronavirus Efficiently Infects Human Primary T Lymphocytes and Activates the Extrinsic and Intrinsic Apoptosis Pathways. J. Infect. Dis. 2016, 213, 904–914. [Google Scholar] [CrossRef]
- Yeung, M.; Yao, Y.; Jia, L.; Chan, J.F.W.; Chan, K.; Cheung, K.; Chen, H.; Poon, V.K.M.; Tsang, A.K.L.; To, K.K.W.; et al. MERS coronavirus induces apoptosis in kidney and lung by upregulating Smad7 and FGF2. Nat. Microbiol. 2016, 1, 16004. [Google Scholar] [CrossRef]
- Liu, W.J.; Lan, J.; Liu, K.; Deng, Y.; Yao, Y.; Wu, S.; Chen, H.; Bao, L.; Zhang, H.; Zhao, M.; et al. Protective T Cell Res.ponses Featured by Concordant Recognition of Middle East Respiratory Syndrome Coronavirus-Derived CD8+ T Cell Epitopes and Host MHC. J. Immunol. 2017, 198, 873–882. [Google Scholar] [CrossRef] [PubMed]
- Chan, R.W.; Chan, M.C.; Agnihothram, S.; Chan, L.L.; Kuok, D.I.; Fong, J.H.; Guan, Y.; Poon, L.L.; Baric, R.S.; Nicholls, J.M.; et al. Tropism of and innate immune responses to the novel human betacoronavirus lineage C virus in human ex vivo respiratory organ cultures. J. Virol. 2013, 87, 6604–6614. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Lau, C.C.; Chan, K.H.; Li, C.P.; Chen, H.; Jin, D.Y.; Chan, J.F.; Woo, P.C.; Yuen, K.Y. Delayed induction of proinflammatory cytokines and suppression of innate antiviral response by the novel Middle East respiratory syndrome coronavirus: Implications for pathogenesis and treatment. J. Gen. Virol. 2013, 94, 2679–2690. [Google Scholar] [CrossRef] [PubMed]
- Marra, M.A.; Jones, S.J.; Astell, C.R.; Holt, R.A.; Brooks-Wilson, A.; Butterfield, Y.S.; Khattra, J.; Asano, J.K.; Barber, S.A.; Chan, S.Y.; et al. The Genome sequence of the SARS-associated coronavirus. Science 2003, 300, 1399–1404. [Google Scholar] [CrossRef] [PubMed]
- Qu, X.X.; Hao, P.; Song, X.J.; Jiang, S.M.; Liu, Y.X.; Wang, P.G.; Rao, X.; Song, H.D.; Wang, S.Y.; Zuo, Y.; et al. Identification of two critical amino acid residues of the severe acute respiratory syndrome coronavirus spike protein for its variation in zoonotic tropism transition via a double substitution strategy. J. Biol. Chem. 2005, 280, 29588–29595. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhang, C.; Sui, J.; Kuhn, J.H.; Moore, M.J.; Luo, S.; Wong, S.K.; Huang, I.C.; Xu, K.; Vasilieva, N.; et al. Receptor and viral determinants of SARS-coronavirus adaptation to human ACE2. EMBO J. 2005, 24, 1634–1643. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Li, W.; Farzan, M.; Harrison, S.C. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science 2005, 309, 1864–1868. [Google Scholar] [CrossRef] [PubMed]
- Li, F. Receptor recognition mechanisms of coronaviruses: A decade of structural studies. J. Virol. 2015, 89, 1954–1964. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Hu, Y.; Wang, Q.; Qi, J.; Gao, F.; Li, Y.; Zhang, Y.; Zhang, W.; Yuan, Y.; Bao, J.; et al. Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 2013, 500, 227–231. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Lu, G.; Qi, J.; Li, Y.; Wu, Y.; Deng, Y.; Geng, H.; Li, H.; Wang, Q.; Xiao, H.; et al. Structure of the fusion core and inhibition of fusion by a heptad repeat peptide derived from the S protein of Middle East respiratory syndrome coronavirus. J. Virol. 2013, 87, 13134–13140. [Google Scholar] [CrossRef]
- Wang, N.; Shi, X.; Jiang, L.; Zhang, S.; Wang, D.; Tong, P.; Guo, D.; Fu, L.; Cui, Y.; Liu, X.; et al. Structure of MERS-CoV spike receptor-binding domain complexed with human receptor DPP4. Cell Res. 2013, 23, 986–993. [Google Scholar] [CrossRef]
- Zhang, N.; Jiang, S.; Du, L. Current advancements and potential strategies in the development of MERS-CoV vaccines. Expert Rev. Vaccines 2014, 13, 761–774. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Zhao, G.; Kou, Z.; Ma, C.; Sun, S.; Poon, V.K.; Lu, L.; Wang, L.; Debnath, A.K.; Zheng, B.J.; et al. Identification of a receptor-binding domain in the S protein of the novel human coronavirus Middle East respiratory syndrome coronavirus as an essential target for vaccine development. J. Virol. 2013, 87, 9939–9942. [Google Scholar] [CrossRef] [PubMed]
- Raj, V.S.; Mou, H.; Smits, S.L.; Dekkers, D.H.; Muller, M.A.; Dijkman, R.; Muth, D.; Demmers, J.A.; Zaki, A.; Fouchier, R.A.; et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 2013, 495, 251–254. [Google Scholar] [CrossRef]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef]
- Chen, Y.; Rajashankar, K.R.; Yang, Y.; Agnihothram, S.S.; Liu, C.; Lin, Y.L.; Baric, R.S.; Li, F. Crystal structure of the receptor-binding domain from newly emerged Middle East respiratory syndrome coronavirus. J. Virol. 2013, 87, 10777–10783. [Google Scholar] [CrossRef]
- Lu, L.; Liu, Q.; Zhu, Y.; Chan, K.H.; Qin, L.; Li, Y.; Wang, Q.; Chan, J.F.; Du, L.; Yu, F.; et al. Structure-based discovery of Middle East respiratory syndrome coronavirus fusion inhibitor. Nat. Commun. 2014, 5, 3067. [Google Scholar] [CrossRef]
- Zheng, Q.; Deng, Y.; Liu, J.; van der Hoek, L.; Berkhout, B.; Lu, M. Core structure of S2 from the human coronavirus NL63 spike glycoprotein. Biochemistry 2006, 45, 15205–15215. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Lou, Z.; Liu, Y.; Pang, H.; Tien, P.; Gao, G.F.; Rao, Z. Crystal structure of severe acute respiratory syndrome coronavirus spike protein fusion core. J. Biol. Chem. 2004, 279, 49414–49419. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Liu, Y.; Lou, Z.; Qin, L.; Li, X.; Bai, Z.; Pang, H.; Tien, P.; Gao, G.F.; Rao, Z. Structural basis for coronavirus-mediated membrane fusion. Crystal structure of mouse hepatitis virus spike protein fusion core. J. Biol. Chem. 2004, 279, 30514–30522. [Google Scholar] [CrossRef]
- Hofmann, H.; Hattermann, K.; Marzi, A.; Gramberg, T.; Geier, M.; Krumbiegel, M.; Kuate, S.; Uberla, K.; Niedrig, M.; Pohlmann, S. S protein of severe acute respiratory syndrome-associated coronavirus mediates entry into hepatoma cell lines and is targeted by neutralizing antibodies in infected patients. J. Virol. 2004, 78, 6134–6142. [Google Scholar] [CrossRef]
- Holmes, K.V. SARS-associated coronavirus. N. Engl. J. Med. 2003, 348, 1948–1951. [Google Scholar] [CrossRef]
- Li, F.; Berardi, M.; Li, W.; Farzan, M.; Dormitzer, P.R.; Harrison, S.C. Conformational states of the severe acute respiratory syndrome coronavirus spike protein ectodomain. J. Virol. 2006, 80, 6794–6800. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.K.; Li, W.; Moore, M.J.; Choe, H.; Farzan, M. A 193-amino acid fragment of the SARS coronavirus S protein efficiently binds angiotensin-converting enzyme 2. J. Biol. Chem. 2004, 279, 3197–3201. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Chakraborti, S.; Dimitrov, A.S.; Gramatikoff, K.; Dimitrov, D.S. The SARS-CoV S glycoprotein: Expression and functional characterization. Biochem. Biophys. Res. Commun. 2003, 312, 1159–1164. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zheng, N.; Hao, P.; Cao, Y.; Zhong, Y. A molecular docking model of SARS-CoV S1 protein in complex with its receptor, human ACE2. Comput. Biol. Chem. 2005, 29, 254–257. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Li, J.; Jiang, S. A single amino acid substitution (R441A) in the receptor-binding domain of SARS coronavirus spike protein disrupts the antigenic structure and binding activity. Biochem. Biophys. Res. Commun. 2006, 344, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Sainz, B.J.; Rausch, J.M.; Gallaher, W.R.; Garry, R.F.; Wimley, W.C. Identification and characterization of the putative fusion peptide of the severe acute respiratory syndrome-associated coronavirus spike protein. J. Virol. 2005, 79, 7195–7206. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.Y.; Huang, Y.; Ganesh, L.; Leung, K.; Kong, W.P.; Schwartz, O.; Subbarao, K.; Nabel, G.J. pH-dependent entry of severe acute respiratory syndrome coronavirus is mediated by the spike glycoprotein and enhanced by dendritic cell transfer through DC-SIGN. J. Virol. 2004, 78, 5642–5650. [Google Scholar] [CrossRef] [PubMed]
- Han, D.P.; Lohani, M.; Cho, M.W. Specific asparagine-linked glycosylation sites are critical for DC-SIGN- and L-SIGN-mediated severe acute respiratory syndrome coronavirus entry. J. Virol. 2007, 81, 12029–12039. [Google Scholar] [CrossRef]
- Graham, R.L.; Donaldson, E.F.; Baric, R.S. A decade after SARS: Strategies for controlling emerging coronaviruses. Nat. Rev. Microbiol. 2013, 11, 836–848. [Google Scholar] [CrossRef]
- Keng, C.T.; Zhang, A.; Shen, S.; Lip, K.M.; Fielding, B.C.; Tan, T.H.; Chou, C.F.; Loh, C.B.; Wang, S.; Fu, J.; et al. Amino acids 1055 to 1192 in the S2 region of severe acute respiratory syndrome coronavirus S protein induce neutralizing antibodies: Implications for the development of vaccines and antiviral agents. J. Virol. 2005, 79, 3289–3296. [Google Scholar] [CrossRef]
- Zhou, T.; Wang, H.; Luo, D.; Rowe, T.; Wang, Z.; Hogan, R.J.; Qiu, S.; Bunzel, R.J.; Huang, G.; Mishra, V.; et al. An exposed domain in the severe acute respiratory syndrome coronavirus spike protein induces neutralizing antibodies. J. Virol. 2004, 78, 7217–7226. [Google Scholar] [CrossRef]
- Bukreyev, A.; Lamirande, E.W.; Buchholz, U.J.; Vogel, L.N.; Elkins, W.R.; St, C.M.; Murphy, B.R.; Subbarao, K.; Collins, P.L. Mucosal immunisation of African green monkeys (Cercopithecus aethiops) with an attenuated parainfluenza virus expressing the SARS coronavirus spike protein for the prevention of SARS. Lancet 2004, 363, 2122–2127. [Google Scholar] [CrossRef]
- Yang, Z.Y.; Kong, W.P.; Huang, Y.; Roberts, A.; Murphy, B.R.; Subbarao, K.; Nabel, G.J. A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 2004, 428, 561–564. [Google Scholar] [CrossRef]
- Kam, Y.W.; Kien, F.; Roberts, A.; Cheung, Y.C.; Lamirande, E.W.; Vogel, L.; Chu, S.L.; Tse, J.; Guarner, J.; Zaki, S.R.; et al. Antibodies against trimeric S glycoprotein protect hamsters against SARS-CoV challenge despite their capacity to mediate FcgammaRII-dependent entry into B cells in vitro. Vaccine 2007, 25, 729–740. [Google Scholar] [CrossRef] [PubMed]
- Czub, M.; Weingartl, H.; Czub, S.; He, R.; Cao, J. Evaluation of modified vaccinia virus Ankara based recombinant SARS vaccine in ferrets. Vaccine 2005, 23, 2273–2279. [Google Scholar] [CrossRef]
- Weingartl, H.; Czub, M.; Czub, S.; Neufeld, J.; Marszal, P.; Gren, J.; Smith, G.; Jones, S.; Proulx, R.; Deschambault, Y.; et al. Immunization with modified vaccinia virus Ankara-based recombinant vaccine against severe acute respiratory syndrome is associated with enhanced hepatitis in ferrets. J. Virol. 2004, 78, 12672–12676. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.; Wang, Y.; Fett, C.A.; Zhao, G.; Li, F.; Perlman, S.; Jiang, S.; Zhou, Y.; Du, L. Recombinant Receptor-Binding Domains of Multiple Middle East Respiratory Syndrome Coronaviruses (MERS-CoVs) Induce Cross-Neutralizing Antibodies against Divergent Human and Camel MERS-CoVs and Antibody Escape Mutants. J. Virol. 2017, 91, e01651-16. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shi, W.; Joyce, M.G.; Modjarrad, K.; Zhang, Y.; Leung, K.; Lees, C.R.; Zhou, T.; Yassine, H.M.; Kanekiyo, M.; et al. Evaluation of candidate vaccine approaches for MERS-CoV. Nat. Commun. 2015, 6, 7712. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, S.C.; Warimwe, G.M. Rapid development of vaccines against emerging pathogens: The replication-deficient simian adenovirus platform technology. Vaccine 2017, 35, 4461–4464. [Google Scholar] [CrossRef]
- Kim, E.; Okada, K.; Kenniston, T.; Raj, V.S.; AlHajri, M.M.; Farag, E.A.; AlHajri, F.; Osterhaus, A.D.; Haagmans, B.L.; Gambotto, A. Immunogenicity of an adenoviral-based Middle East Respiratory Syndrome coronavirus vaccine in BALB/c mice. Vaccine 2014, 32, 5975–5982. [Google Scholar] [CrossRef]
- Chi, H.; Zheng, X.; Wang, X.; Wang, C.; Wang, H.; Gai, W.; Perlman, S.; Yang, S.; Zhao, J.; Xia, X. DNA vaccine encoding Middle East respiratory syndrome coronavirus S1 protein induces protective immune responses in mice. Vaccine 2017, 35, 2069–2075. [Google Scholar] [CrossRef]
- Al-Amri, S.S.; Abbas, A.T.; Siddiq, L.A.; Alghamdi, A.; Sanki, M.A.; Al-Muhanna, M.K.; Alhabbab, R.Y.; Azhar, E.I.; Li, X.; Hashem, A.M. Immunogenicity of Candidate MERS-CoV DNA Vaccines Based on the Spike Protein. Sci. Rep. 2017, 7, 44875. [Google Scholar] [CrossRef]
- Modjarrad, K. MERS-CoV vaccine candidates in development: The current landscape. Vaccine 2016, 34, 2982–2987. [Google Scholar] [CrossRef]
- Zhao, J.; Zhao, J.; Mangalam, A.K.; Channappanavar, R.; Fett, C.; Meyerholz, D.K.; Agnihothram, S.; Baric, R.S.; David, C.S.; Perlman, S. Airway Memory CD4+ T Cells Mediate Protective Immunity against Emerging Respiratory Coronaviruses. Immunity 2016, 44, 1379–1391. [Google Scholar] [CrossRef]
- Roper, R.L.; Rehm, K.E. SARS vaccines: Where are we? Expert Rev. Vaccines 2009, 8, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Zhang, J.; Su, N.; Xu, J.; Wang, N.; Chen, J.; Chen, X.; Liu, Y.; Gao, H.; Jia, Y.; et al. Safety and immunogenicity from a phase I trial of inactivated severe acute respiratory syndrome coronavirus vaccine. Antivir. Ther. 2007, 12, 1107–1113. [Google Scholar] [PubMed]
- Deng, Y.; Lan, J.; Bao, L.; Huang, B.; Ye, F.; Chen, Y.; Yao, Y.; Wang, W.; Qin, C.; Tan, W. Enhanced protection in mice induced by immunization with inactivated whole viruses compare to spike protein of middle east respiratory syndrome coronavirus. Emerg. Microbes Infect. 2018, 7, 60. [Google Scholar] [CrossRef] [PubMed]
- See, R.H.; Zakhartchouk, A.N.; Petric, M.; Lawrence, D.J.; Mok, C.P.; Hogan, R.J.; Rowe, T.; Zitzow, L.A.; Karunakaran, K.P.; Hitt, M.M.; et al. Comparative evaluation of two severe acute respiratory syndrome (SARS) vaccine candidates in mice challenged with SARS coronavirus. J. Gen. Virol. 2006, 87, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Enjuanes, L.; Dediego, M.L.; Alvarez, E.; Deming, D.; Sheahan, T.; Baric, R. Vaccines to prevent severe acute respiratory syndrome coronavirus-induced disease. Virus Res. 2008, 133, 45–62. [Google Scholar] [CrossRef] [PubMed]
- Bolles, M.; Deming, D.; Long, K.; Agnihothram, S.; Whitmore, A.; Ferris, M.; Funkhouser, W.; Gralinski, L.; Totura, A.; Heise, M.; et al. A double-inactivated severe acute respiratory syndrome coronavirus vaccine provides incomplete protection in mice and induces increased eosinophilic proinflammatory pulmonary response upon challenge. J. Virol. 2011, 85, 12201–12215. [Google Scholar] [CrossRef] [PubMed]
- Ishioka, T.; Kimura, H.; Kita, H.; Obuchi, M.; Hoshino, H.; Noda, M.; Nishina, A.; Kozawa, K.; Kato, M. Effects of respiratory syncytial virus infection and major basic protein derived from eosinophils in pulmonary alveolar epithelial cells (A549). Cell Biol. Int. 2011, 35, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Fett, C.; DeDiego, M.L.; Regla-Nava, J.A.; Enjuanes, L.; Perlman, S. Complete protection against severe acute respiratory syndrome coronavirus-mediated lethal respiratory disease in aged mice by immunization with a mouse-adapted virus lacking E protein. J. Virol. 2013, 87, 6551–6559. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.L.; Becker, M.M.; Eckerle, L.D.; Bolles, M.; Denison, M.R.; Baric, R.S. A live, impaired-fidelity coronavirus vaccine protects in an aged, immunocompromised mouse model of lethal disease. Nat. Med. 2012, 18, 1820–1826. [Google Scholar] [CrossRef]
- Almazan, F.; DeDiego, M.L.; Sola, I.; Zuniga, S.; Nieto-Torres, J.L.; Marquez-Jurado, S.; Andres, G.; Enjuanes, L. Engineering a replication-competent, propagation-defective Middle East respiratory syndrome coronavirus as a vaccine candidate. mBio 2013, 4, e613–e650. [Google Scholar] [CrossRef] [PubMed]
- Vignuzzi, M.; Wendt, E.; Andino, R. Engineering attenuated virus vaccines by controlling replication fidelity. Nat. Med. 2008, 14, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.; Vogel, L.; Guarner, J.; Hayes, N.; Murphy, B.; Zaki, S.; Subbarao, K. Severe acute respiratory syndrome coronavirus infection of golden Syrian hamsters. J. Virol. 2005, 79, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Deming, D.; Sheahan, T.; Heise, M.; Yount, B.; Davis, N.; Sims, A.; Suthar, M.; Harkema, J.; Whitmore, A.; Pickles, R.; et al. Vaccine efficacy in senescent mice challenged with recombinant SARS-CoV bearing epidemic and zoonotic spike variants. PLoS Med. 2006, 3, e525. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhang, L.; Qin, C.; Ba, L.; Yi, C.E.; Zhang, F.; Wei, Q.; He, T.; Yu, W.; Yu, J.; et al. Recombinant modified vaccinia virus Ankara expressing the spike glycoprotein of severe acute respiratory syndrome coronavirus induces protective neutralizing antibodies primarily targeting the receptor binding region. J. Virol. 2005, 79, 2678–2688. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Yao, Y.; Deng, Y.; Chen, H.; Lu, G.; Wang, W.; Bao, L.; Deng, W.; Wei, Q.; Gao, G.F.; et al. Recombinant Receptor Binding Domain Protein Induces Partial Protective Immunity in Rhesus Macaques Against Middle East Respiratory Syndrome Coronavirus Challenge. EBioMedicine 2015, 2, 1438–1446. [Google Scholar] [CrossRef]
- Jiaming, L.; Yanfeng, Y.; Yao, D.; Yawei, H.; Linlin, B.; Baoying, H.; Jinghua, Y.; Gao, G.F.; Chuan, Q.; Wenjie, T. The recombinant N-terminal domain of spike proteins is a potential vaccine against Middle East respiratory syndrome coronavirus (MERS-CoV) infection. Vaccine 2017, 35, 10–18. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Subbarao, K.; Moss, B. Neutralizing antibody and protective immunity to SARS coronavirus infection of mice induced by a soluble recombinant polypeptide containing an N-terminal segment of the spike glycoprotein. Virology 2005, 334, 160–165. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Tsoi, H.W.; Chen, Z.W.; Wong, B.H.; Zhang, L.; Chan, J.K.; Wong, L.P.; He, W.; Ma, C.; et al. SARS coronavirus spike polypeptide DNA vaccine priming with recombinant spike polypeptide from Escherichia coli as booster induces high titer of neutralizing antibody against SARS coronavirus. Vaccine 2005, 23, 4959–4968. [Google Scholar] [CrossRef]
- Shi, S.; Peng, J.; Li, Y.; Qin, C.; Liang, G.; Xu, L.; Yang, Y.; Wang, J.; Sun, Q. The expression of membrane protein augments the specific responses induced by SARS-CoV nucleocapsid DNA immunization. Mol. Immunol. 2006, 43, 1791–1798. [Google Scholar] [CrossRef]
- Gupta, V.; Tabiin, T.M.; Sun, K.; Chandrasekaran, A.; Anwar, A.; Yang, K.; Chikhlikar, P.; Salmon, J.; Brusic, V.; Marques, E.T.; et al. SARS coronavirus nucleocapsid immunodominant T-cell epitope cluster is common to both exogenous recombinant and endogenous DNA-encoded immunogens. Virology 2006, 347, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, M.; Jiang, H.; Suzuki, Y.; Li, X.; Xiao, P.; Tanaka, T.; Ling, H.; Yang, B.; Saitoh, H.; Zhang, L.; et al. Procyanidins and butanol extract of Cinnamomi Cortex inhibit SARS-CoV infection. Antivir. Res. 2009, 82, 73–81. [Google Scholar] [CrossRef]
- Hu, H.; Li, L.; Kao, R.Y.; Kou, B.; Wang, Z.; Zhang, L.; Zhang, H.; Hao, Z.; Tsui, W.H.; Ni, A.; et al. Screening and identification of linear B-cell epitopes and entry-blocking peptide of severe acute respiratory syndrome (SARS)-associated coronavirus using synthetic overlapping peptide library. J. Comb. Chem. 2005, 7, 648–656. [Google Scholar] [CrossRef]
- Zheng, B.J.; Guan, Y.; Hez, M.L.; Sun, H.; Du, L.; Zheng, Y.; Wong, K.L.; Chen, H.; Chen, Y.; Lu, L.; et al. Synthetic peptides outside the spike protein heptad repeat regions as potent inhibitors of SARS-associated coronavirus. Antivir. Ther. 2005, 10, 393–403. [Google Scholar] [PubMed]
- Bosch, B.J.; Martina, B.E.; Van Der Zee, R.; Lepault, J.; Haijema, B.J.; Versluis, C.; Heck, A.J.; De Groot, R.; Osterhaus, A.D.; Rottier, P.J. Severe acute respiratory syndrome coronavirus (SARS-CoV) infection inhibition using spike protein heptad repeat-derived peptides. Proc. Natl. Acad. Sci. USA 2004, 101, 8455–8460. [Google Scholar] [CrossRef]
- Yuan, K.; Yi, L.; Chen, J.; Qu, X.; Qing, T.; Rao, X.; Jiang, P.; Hu, J.; Xiong, Z.; Nie, Y.; et al. Suppression of SARS-CoV entry by peptides corresponding to heptad regions on spike glycoprotein. Biochem. Biophys. Res. Commun. 2004, 319, 746–752. [Google Scholar] [CrossRef]
- McRoy, W.C.; Baric, R.S. Amino acid substitutions in the S2 subunit of mouse hepatitis virus variant V51 encode determinants of host range expansion. J. Virol. 2008, 82, 1414–1424. [Google Scholar] [CrossRef]
- He, Y.; Li, J.; Heck, S.; Lustigman, S.; Jiang, S. Antigenic and immunogenic characterization of recombinant baculovirus-expressed severe acute respiratory syndrome coronavirus spike protein: Implication for vaccine design. J. Virol. 2006, 80, 5757–5767. [Google Scholar] [CrossRef]
- He, Y.; Li, J.; Li, W.; Lustigman, S.; Farzan, M.; Jiang, S. Cross-neutralization of human and palm civet severe acute respiratory syndrome coronaviruses by antibodies targeting the receptor-binding domain of spike protein. J. Immunol. 2006, 176, 6085–6092. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Zhu, Q.; Liu, S.; Zhou, Y.; Yang, B.; Li, J.; Jiang, S. Identification of a critical neutralization determinant of severe acute respiratory syndrome (SARS)-associated coronavirus: Importance for designing SARS vaccines. Virology 2005, 334, 74–82. [Google Scholar] [CrossRef]
- Lai, S.C.; Chong, P.C.; Yeh, C.T.; Liu, L.S.; Jan, J.T.; Chi, H.Y.; Liu, H.W.; Chen, A.; Wang, Y.C. Characterization of neutralizing monoclonal antibodies recognizing a 15-residues epitope on the spike protein HR2 region of severe acute respiratory syndrome coronavirus (SARS-CoV). J. Biomed. Sci. 2005, 12, 711–727. [Google Scholar] [CrossRef] [PubMed]
- Nie, Y.; Wang, G.; Shi, X.; Zhang, H.; Qiu, Y.; He, Z.; Wang, W.; Lian, G.; Yin, X.; Du, L.; et al. Neutralizing antibodies in patients with severe acute respiratory syndrome-associated coronavirus infection. J. Infect. Dis. 2004, 190, 1119–1126. [Google Scholar] [CrossRef]
- Traggiai, E.; Becker, S.; Subbarao, K.; Kolesnikova, L.; Uematsu, Y.; Gismondo, M.R.; Murphy, B.R.; Rappuoli, R.; Lanzavecchia, A. An efficient method to make human monoclonal antibodies from memory B cells: Potent neutralization of SARS coronavirus. Nat. Med. 2004, 10, 871–875. [Google Scholar] [CrossRef]
- Rockx, B.; Corti, D.; Donaldson, E.; Sheahan, T.; Stadler, K.; Lanzavecchia, A.; Baric, R. Structural basis for potent cross-neutralizing human monoclonal antibody protection against lethal human and zoonotic severe acute respiratory syndrome coronavirus challenge. J. Virol. 2008, 82, 3220–3235. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Chakraborti, S.; He, Y.; Roberts, A.; Sheahan, T.; Xiao, X.; Hensley, L.E.; Prabakaran, P.; Rockx, B.; Sidorov, I.A.; et al. Potent cross-reactive neutralization of SARS coronavirus isolates by human monoclonal antibodies. Proc. Natl. Acad. Sci. USA 2007, 104, 12123–12128. [Google Scholar] [CrossRef] [PubMed]
- Coughlin, M.; Lou, G.; Martinez, O.; Masterman, S.K.; Olsen, O.A.; Moksa, A.A.; Farzan, M.; Babcook, J.S.; Prabhakar, B.S. Generation and characterization of human monoclonal neutralizing antibodies with distinct binding and sequence features against SARS coronavirus using XenoMouse. Virology 2007, 361, 93–102. [Google Scholar] [CrossRef]
- van den Brink, E.N.; Ter Meulen, J.; Cox, F.; Jongeneelen, M.A.; Thijsse, A.; Throsby, M.; Marissen, W.E.; Rood, P.M.; Bakker, A.B.; Gelderblom, H.R.; et al. Molecular and biological characterization of human monoclonal antibodies binding to the spike and nucleocapsid proteins of severe acute respiratory syndrome coronavirus. J. Virol. 2005, 79, 1635–1644. [Google Scholar] [CrossRef] [PubMed]
- Sui, J.; Li, W.; Murakami, A.; Tamin, A.; Matthews, L.J.; Wong, S.K.; Moore, M.J.; Tallarico, A.S.; Olurinde, M.; Choe, H.; et al. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc. Natl. Acad. Sci. USA 2004, 101, 2536–2541. [Google Scholar] [CrossRef]
- Liu, S.; Xiao, G.; Chen, Y.; He, Y.; Niu, J.; Escalante, C.R.; Xiong, H.; Farmar, J.; Debnath, A.K.; Tien, P.; et al. Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: Implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet 2004, 363, 938–947. [Google Scholar] [CrossRef]
- Corti, D.; Zhao, J.; Pedotti, M.; Simonelli, L.; Agnihothram, S.; Fett, C.; Fernandez-Rodriguez, B.; Foglierini, M.; Agatic, G.; Vanzetta, F.; et al. Prophylactic and postexposure efficacy of a potent human monoclonal antibody against MERS coronavirus. Proc. Natl. Acad. Sci. USA 2015, 112, 10473–10478. [Google Scholar] [CrossRef]
- Li, Y.; Wan, Y.; Liu, P.; Zhao, J.; Lu, G.; Qi, J.; Wang, Q.; Lu, X.; Wu, Y.; Liu, W.; et al. A humanized neutralizing antibody against MERS-CoV targeting the receptor-binding domain of the spike protein. Cell Res. 2015, 25, 1237–1249. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Zhao, G.; Yang, Y.; Qiu, H.; Wang, L.; Kou, Z.; Tao, X.; Yu, H.; Sun, S.; Tseng, C.T.; et al. A conformation-dependent neutralizing monoclonal antibody specifically targeting receptor-binding domain in Middle East respiratory syndrome coronavirus spike protein. J. Virol. 2014, 88, 7045–7053. [Google Scholar] [CrossRef] [PubMed]
- Raj, V.S.; Smits, S.L.; Provacia, L.B.; van den Brand, J.M.; Wiersma, L.; Ouwendijk, W.J.; Bestebroer, T.M.; Spronken, M.I.; van Amerongen, G.; Rottier, P.J.; et al. Adenosine deaminase acts as a natural antagonist for dipeptidyl peptidase 4-mediated entry of the Middle East respiratory syndrome coronavirus. J. Virol. 2014, 88, 1834–1838. [Google Scholar] [CrossRef] [PubMed]
- Zhou, N.; Pan, T.; Zhang, J.; Li, Q.; Zhang, X.; Bai, C.; Huang, F.; Peng, T.; Zhang, J.; Liu, C.; et al. Glycopeptide Antibiotics Potently Inhibit Cathepsin L in the Late Endosome/Lysosome and Block the Entry of Ebola Virus, Middle East Respiratory Syndrome Coronavirus (MERS-CoV), and Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV). J. Biol. Chem. 2016, 291, 9218–9232. [Google Scholar] [CrossRef]
- Gierer, S.; Bertram, S.; Kaup, F.; Wrensch, F.; Heurich, A.; Kramer-Kuhl, A.; Welsch, K.; Winkler, M.; Meyer, B.; Drosten, C.; et al. The spike protein of the emerging betacoronavirus EMC uses a novel coronavirus receptor for entry, can be activated by TMPRSS2, and is targeted by neutralizing antibodies. J. Virol. 2013, 87, 5502–5511. [Google Scholar] [CrossRef] [PubMed]
- Shirato, K.; Kawase, M.; Matsuyama, S. Middle East respiratory syndrome coronavirus infection mediated by the transmembrane serine protease TMPRSS2. J. Virol. 2013, 87, 12552–12561. [Google Scholar] [CrossRef]
- Zhou, Y.; Vedantham, P.; Lu, K.; Agudelo, J.; Carrion, R.; Nunneley, J.W.; Barnard, D.; Pöhlmann, S.; McKerrow, J.H.; Renslo, A.R.; Simmons, G. Protease inhibitors targeting coronavirus and filovirus entry. Antivir. Res. 2015, 116, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Momattin, H.; Mohammed, K.; Zumla, A.; Memish, Z.A.; Al-Tawfiq, J.A. Therapeutic Options for Middle East Respiratory Syndrome Coronavirus (MERS-CoV)–possible lessons from a systematic review of SARS-CoV therapy. Int. J. Infect. Dis. 2013, 17, e792–e798. [Google Scholar] [CrossRef]
- Hart, B.J.; Dyall, J.; Postnikova, E.; Zhou, H.; Kindrachuk, J.; Johnson, R.F.; Olinger, G.J.; Frieman, M.B.; Holbrook, M.R.; Jahrling, P.B.; et al. Interferon-beta and mycophenolic acid are potent inhibitors of Middle East respiratory syndrome coronavirus in cell-based assays. J. Gen. Virol. 2014, 95, 571–577. [Google Scholar] [CrossRef]
- ter Meulen, J.; van den Brink, E.N.; Poon, L.L.; Marissen, W.E.; Leung, C.S.; Cox, F.; Cheung, C.Y.; Bakker, A.Q.; Bogaards, J.A.; van Deventer, E.; et al. Human monoclonal antibody combination against SARS coronavirus: Synergy and coverage of escape mutants. PLoS Med. 2006, 3, e237. [Google Scholar] [CrossRef]
- Zhang, J.S.; Chen, J.T.; Liu, Y.X.; Zhang, Z.S.; Gao, H.; Liu, Y.; Wang, X.; Ning, Y.; Liu, Y.F.; Gao, Q.; et al. A serological survey on neutralizing antibody titer of SARS convalescent sera. J. Med. Virol. 2005, 77, 147–150. [Google Scholar] [CrossRef]
- Miyoshi-Akiyama, T.; Ishida, I.; Fukushi, M.; Yamaguchi, K.; Matsuoka, Y.; Ishihara, T.; Tsukahara, M.; Hatakeyama, S.; Itoh, N.; Morisawa, A.; et al. Fully Human Monoclonal Antibody Directed to Proteolytic Cleavage Site in Severe Acute Respiratory Syndrome (SARS) Coronavirus S Protein Neutralizes the Virus in a Rhesus Macaque SARS Model. J. Infect. Dis. 2011, 203, 1574–1581. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, L.; Kuwahara, K.; Li, L.; Liu, Z.; Li, T.; Zhu, H.; Liu, J.; Xu, Y.; Xie, J.; et al. Immunodominant SARS Coronavirus Epitopes in Humans Elicited both Enhancing and Neutralizing Effects on Infection in Non-human Primates. ACS Infect. Dis. 2016, 2, 361–376. [Google Scholar] [CrossRef]
- Chen, Z.; Bao, L.; Chen, C.; Zou, T.; Xue, Y.; Li, F.; Lv, Q.; Gu, S.; Gao, X.; Cui, S.; et al. Human Neutralizing Monoclonal Antibody Inhibition of Middle East Respiratory Syndrome Coronavirus Replication in the Common Marmoset. J. Infect. Dis. 2017, 215, 1807–1815. [Google Scholar] [CrossRef]
- Stockman, L.J.; Bellamy, R.; Garner, P.; Low, D. SARS: Systematic Review of Treatment Effects. PLoS Med. 2006, 3, e343. [Google Scholar] [CrossRef]
- Chan, P.K.S.; Tang, J.W.; Hui, D.S.C. SARS: Clinical presentation, transmission, pathogenesis and treatment options. Clin. Sci. 2006, 110, 193–204. [Google Scholar] [CrossRef]
- Chan, J.F.; Yao, Y.; Yeung, M.; Deng, W.; Bao, L.; Jia, L.; Li, F.; Xiao, C.; Gao, H.; Yu, P.; et al. Treatment with Lopinavir/Ritonavir or Interferon-β1b Improves Outcome of MERS-CoV Infection in a Nonhuman Primate Model of Common Marmoset. J. Infect. Dis. 2015, 212, 1904–1913. [Google Scholar] [CrossRef]
- Subbarao, K.; McAuliffe, J.; Vogel, L.; Fahle, G.; Fischer, S.; Tatti, K.; Packard, M.; Shieh, W.J.; Zaki, S.; Murphy, B. Prior infection and passive transfer of neutralizing antibody prevent replication of severe acute respiratory syndrome coronavirus in the respiratory tract of mice. J. Virol. 2004, 78, 3572–3577. [Google Scholar] [CrossRef]
- Wentworth, D.E.; Gillim-Ross, L.; Espina, N.; Bernard, K.A. Mice susceptible to SARS coronavirus. Emerg. Infect. Dis. 2004, 10, 1293–1296. [Google Scholar] [CrossRef]
- Glass, W.G.; Subbarao, K.; Murphy, B.; Murphy, P.M. Mechanisms of host defense following severe acute respiratory syndrome-coronavirus (SARS-CoV) pulmonary infection of mice. J. Immunol. 2004, 173, 4030–4039. [Google Scholar] [CrossRef]
- Yang, X.H.; Deng, W.; Tong, Z.; Liu, Y.X.; Zhang, L.F.; Zhu, H.; Gao, H.; Huang, L.; Liu, Y.L.; Ma, C.M.; et al. Mice transgenic for human angiotensin-converting enzyme 2 provide a model for SARS coronavirus infection. Comp. Med. 2007, 57, 450–459. [Google Scholar] [PubMed]
- Roberts, A.; Paddock, C.; Vogel, L.; Butler, E.; Zaki, S.; Subbarao, K. Aged BALB/c mice as a model for increased severity of severe acute respiratory syndrome in elderly humans. J. Virol. 2005, 79, 5833–5838. [Google Scholar] [CrossRef] [PubMed]
- Hogan, R.J.; Gao, G.; Rowe, T.; Bell, P.; Flieder, D.; Paragas, J.; Kobinger, G.P.; Wivel, N.A.; Crystal, R.G.; Boyer, J.; et al. Resolution of primary severe acute respiratory syndrome-associated coronavirus infection requires Stat1. J. Virol. 2004, 78, 11416–11421. [Google Scholar] [CrossRef]
- Frieman, M.; Yount, B.; Agnihothram, S.; Page, C.; Donaldson, E.; Roberts, A.; Vogel, L.; Woodruff, B.; Scorpio, D.; Subbarao, K.; et al. Molecular determinants of severe acute respiratory syndrome coronavirus pathogenesis and virulence in young and aged mouse models of human disease. J. Virol. 2012, 86, 884–897. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, J.; Legge, K.; Perlman, S. Age-related increases in PGD(2) expression impair respiratory DC migration, resulting in diminished T cell responses upon respiratory virus infection in mice. J. Clin. Investig. 2011, 121, 4921–4930. [Google Scholar] [CrossRef]
- Zhao, J.; Zhao, J.; Perlman, S. T cell responses are required for protection from clinical disease and for virus clearance in severe acute respiratory syndrome coronavirus-infected mice. J. Virol. 2010, 84, 9318–9325. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, J.; Van Rooijen, N.; Perlman, S. Evasion by stealth: Inefficient immune activation underlies poor T cell response and severe disease in SARS-CoV-infected mice. PLoS Pathog. 2009, 5, e1000636. [Google Scholar] [CrossRef] [PubMed]
- Sheahan, T.; Morrison, T.E.; Funkhouser, W.; Uematsu, S.; Akira, S.; Baric, R.S.; Heise, M.T. MyD88 is required for protection from lethal infection with a mouse-adapted SARS-CoV. PLoS Pathog. 2008, 4, e1000240. [Google Scholar] [CrossRef]
- Li, C.K.; Wu, H.; Yan, H.; Ma, S.; Wang, L.; Zhang, M.; Tang, X.; Temperton, N.J.; Weiss, R.A.; Brenchley, J.M.; et al. T cell responses to whole SARS coronavirus in humans. J. Immunol. 2008, 181, 5490–5500. [Google Scholar] [CrossRef]
- Chen, W.; Xu, Z.; Mu, J.; Yang, L.; Gan, H.; Mu, F.; Fan, B.; He, B.; Huang, S.; You, B.; et al. Antibody response and viraemia during the course of severe acute respiratory syndrome (SARS)-associated coronavirus infection. J. Med. Microbiol. 2004, 53, 435–438. [Google Scholar] [CrossRef]
- Spruth, M.; Kistner, O.; Savidis-Dacho, H.; Hitter, E.; Crowe, B.; Gerencer, M.; Bruhl, P.; Grillberger, L.; Reiter, M.; Tauer, C.; et al. A double-inactivated whole virus candidate SARS coronavirus vaccine stimulates neutralising and protective antibody responses. Vaccine 2006, 24, 652–661. [Google Scholar] [CrossRef]
- Greenough, T.C.; Babcock, G.J.; Roberts, A.; Hernandez, H.J.; Thomas, W.J.; Coccia, J.A.; Graziano, R.F.; Srinivasan, M.; Lowy, I.; Finberg, R.W.; et al. Development and characterization of a severe acute respiratory syndrome-associated coronavirus-neutralizing human monoclonal antibody that provides effective immunoprophylaxis in mice. J. Infect. Dis. 2005, 191, 507–514. [Google Scholar] [CrossRef]
- Kapadia, S.U.; Rose, J.K.; Lamirande, E.; Vogel, L.; Subbarao, K.; Roberts, A. Long-term protection from SARS coronavirus infection conferred by a single immunization with an attenuated VSV-based vaccine. Virology 2005, 340, 174–182. [Google Scholar] [CrossRef]
- Stadler, K.; Roberts, A.; Becker, S.; Vogel, L.; Eickmann, M.; Kolesnikova, L.; Klenk, H.D.; Murphy, B.; Rappuoli, R.; Abrignani, S.; et al. SARS vaccine protective in mice. Emerg. Infect. Dis. 2005, 11, 1312–1314. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef]
- Schaecher, S.R.; Stabenow, J.; Oberle, C.; Schriewer, J.; Buller, R.M.; Sagartz, J.E.; Pekosz, A. An immunosuppressed Syrian golden hamster model for SARS-CoV infection. Virology 2008, 380, 312–321. [Google Scholar] [CrossRef]
- Luo, D.; Ni, B.; Zhao, G.; Jia, Z.; Zhou, L.; Pacal, M.; Zhang, L.; Zhang, S.; Xing, L.; Lin, Z.; et al. Protection from infection with severe acute respiratory syndrome coronavirus in a Chinese hamster model by equine neutralizing F(ab’)2. Viral Immunol. 2007, 20, 495–502. [Google Scholar] [CrossRef]
- Subbarao, K.; Roberts, A. Is there an ideal animal model for SARS? Trends Microbiol. 2006, 14, 299–303. [Google Scholar] [CrossRef]
- Roberts, A.; Thomas, W.D.; Guarner, J.; Lamirande, E.W.; Babcock, G.J.; Greenough, T.C.; Vogel, L.; Hayes, N.; Sullivan, J.L.; Zaki, S.; et al. Therapy with a severe acute respiratory syndrome-associated coronavirus-neutralizing human monoclonal antibody reduces disease severity and viral burden in golden Syrian hamsters. J. Infect. Dis. 2006, 193, 685–692. [Google Scholar] [CrossRef]
- Martina, B.E.; Haagmans, B.L.; Kuiken, T.; Fouchier, R.A.; Rimmelzwaan, G.F.; Van Amerongen, G.; Peiris, J.S.; Lim, W.; Osterhaus, A.D. Virology: SARS virus infection of cats and ferrets. Nature 2003, 425, 915. [Google Scholar] [CrossRef]
- van den Brand, J.M.; Haagmans, B.L.; Leijten, L.; van Riel, D.; Martina, B.E.; Osterhaus, A.D.; Kuiken, T. Pathology of experimental SARS coronavirus infection in cats and ferrets. Vet. Pathol. 2008, 45, 551–562. [Google Scholar] [CrossRef]
- Darnell, M.E.; Plant, E.P.; Watanabe, H.; Byrum, R.; St, C.M.; Ward, J.M.; Taylor, D.R. Severe acute respiratory syndrome coronavirus infection in vaccinated ferrets. J. Infect. Dis. 2007, 196, 1329–1338. [Google Scholar] [CrossRef]
- ter Meulen, J.; Bakker, A.B.; van den Brink, E.N.; Weverling, G.J.; Martina, B.E.; Haagmans, B.L.; Kuiken, T.; de Kruif, J.; Preiser, W.; Spaan, W.; et al. Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet 2004, 363, 2139–2141. [Google Scholar] [CrossRef]
- See, R.H.; Petric, M.; Lawrence, D.J.; Mok, C.P.; Rowe, T.; Zitzow, L.A.; Karunakaran, K.P.; Voss, T.G.; Brunham, R.C.; Gauldie, J.; et al. Severe acute respiratory syndrome vaccine efficacy in ferrets: Whole killed virus and adenovirus-vectored vaccines. J. Gen. Virol. 2008, 89, 2136–2146. [Google Scholar] [CrossRef]
- Qin, C.; Wang, J.; Wei, Q.; She, M.; Marasco, W.A.; Jiang, H.; Tu, X.; Zhu, H.; Ren, L.; Gao, H.; et al. An animal model of SARS produced by infection of Macaca mulatta with SARS coronavirus. J. Pathol. 2005, 206, 251–259. [Google Scholar] [CrossRef]
- McAuliffe, J.; Vogel, L.; Roberts, A.; Fahle, G.; Fischer, S.; Shieh, W.J.; Butler, E.; Zaki, S.; St, C.M.; Murphy, B.; et al. Replication of SARS coronavirus administered into the respiratory tract of African Green, rhesus and cynomolgus monkeys. Virology 2004, 330, 8–15. [Google Scholar] [CrossRef]
- Rowe, T.; Gao, G.; Hogan, R.J.; Crystal, R.G.; Voss, T.G.; Grant, R.L.; Bell, P.; Kobinger, G.P.; Wivel, N.A.; Wilson, J.M. Macaque model for severe acute respiratory syndrome. J. Virol. 2004, 78, 11401–11404. [Google Scholar] [CrossRef]
- Li, B.; Tang, Q.; Cheng, D.; Qin, C.; Xie, F.Y.; Wei, Q.; Xu, J.; Liu, Y.; Zheng, B.; Woodle, M.C.; et al. Using siRNA in prophylactic and therapeutic regimens against SARS coronavirus in Rhesus macaque. Nat. Med. 2005, 11, 944–951. [Google Scholar] [CrossRef]
- Liu, L.; Wei, Q.; Nishiura, K.; Peng, J.; Wang, H.; Midkiff, C.; Alvarez, X.; Qin, C.; Lackner, A.; Chen, Z. Spatiotemporal interplay of severe acute respiratory syndrome coronavirus and respiratory mucosal cells drives viral dissemination in rhesus macaques. Mucosal Immunol. 2016, 9, 1089–1101. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Kuiken, T.; Schutten, M.; van Amerongen, G.; van Doornum, G.J.; van den Hoogen, B.G.; Peiris, M.; Lim, W.; Stohr, K.; Osterhaus, A.D. Aetiology: Koch’s postulates fulfilled for SARS virus. Nature 2003, 423, 240. [Google Scholar] [CrossRef]
- Kuiken, T.; Fouchier, R.A.; Schutten, M.; Rimmelzwaan, G.F.; van Amerongen, G.; van Riel, D.; Laman, J.D.; de Jong, T.; van Doornum, G.; Lim, W.; et al. Newly discovered coronavirus as the primary cause of severe acute respiratory syndrome. Lancet 2003, 362, 263–270. [Google Scholar] [CrossRef]
- Greenough, T.C.; Carville, A.; Coderre, J.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Mansfield, K. Pneumonitis and multi-organ system disease in common marmosets (Callithrix jacchus) infected with the severe acute respiratory syndrome-associated coronavirus. Am. J. Pathol. 2005, 167, 455–463. [Google Scholar] [CrossRef]
- Lawler, J.V.; Endy, T.P.; Hensley, L.E.; Garrison, A.; Fritz, E.A.; Lesar, M.; Baric, R.S.; Kulesh, D.A.; Norwood, D.A.; Wasieloski, L.P.; et al. Cynomolgus macaque as an animal model for severe acute respiratory syndrome. PLoS Med. 2006, 3, e149. [Google Scholar] [CrossRef]
- Roberts, A.; Subbarao, K. Animal models for SARS. Adv. Exp. Med. Biol. 2006, 581, 463–471. [Google Scholar]
- Chen, Y.; Liu, L.; Wei, Q.; Zhu, H.; Jiang, H.; Tu, X.; Qin, C.; Chen, Z. Rhesus angiotensin converting enzyme 2 supports entry of severe acute respiratory syndrome coronavirus in Chinese macaques. Virology 2008, 381, 89–97. [Google Scholar] [CrossRef]
- Cockrell, A.S.; Peck, K.M.; Yount, B.L.; Agnihothram, S.S.; Scobey, T.; Curnes, N.R.; Baric, R.S.; Heise, M.T. Mouse dipeptidyl peptidase 4 is not a functional receptor for Middle East respiratory syndrome coronavirus infection. J. Virol. 2014, 88, 5195–5199. [Google Scholar] [CrossRef]
- Coleman, C.M.; Matthews, K.L.; Goicochea, L.; Frieman, M.B. Wild-type and innate immune-deficient mice are not susceptible to the Middle East respiratory syndrome coronavirus. J. Gen. Virol. 2014, 95, 408–412. [Google Scholar] [CrossRef]
- de Wit, E.; Feldmann, F.; Horne, E.; Martellaro, C.; Haddock, E.; Bushmaker, T.; Rosenke, K.; Okumura, A.; Rosenke, R.; Saturday, G.; et al. Domestic Pig Unlikely Reservoir for MERS-CoV. Emerg. Infect. Dis. 2017, 23, 985–988. [Google Scholar] [CrossRef]
- Peck, K.M.; Scobey, T.; Swanstrom, J.; Jensen, K.L.; Burch, C.L.; Baric, R.S.; Heise, M.T. Permissivity of Dipeptidyl Peptidase 4 Orthologs to Middle East Respiratory Syndrome Coronavirus Is Governed by Glycosylation and Other Complex Determinants. J. Virol. 2017, 91, e00534-17. [Google Scholar] [CrossRef]
- Haagmans, B.L.; van den Brand, J.M.; Provacia, L.B.; Raj, V.S.; Stittelaar, K.J.; Getu, S.; de Waal, L.; Bestebroer, T.M.; van Amerongen, G.; Verjans, G.M.; et al. Asymptomatic Middle East respiratory syndrome coronavirus infection in rabbits. J. Virol. 2015, 89, 6131–6135. [Google Scholar] [CrossRef]
- de Wit, E.; Prescott, J.; Baseler, L.; Bushmaker, T.; Thomas, T.; Lackemeyer, M.G.; Martellaro, C.; Milne-Price, S.; Haddock, E.; Haagmans, B.L.; et al. The Middle East respiratory syndrome coronavirus (MERS-CoV) does not replicate in Syrian hamsters. PLoS ONE 2013, 8, e69127. [Google Scholar] [CrossRef]
- Li, K.; Wohlford-Lenane, C.L.; Channappanavar, R.; Park, J.E.; Earnest, J.T.; Bair, T.B.; Bates, A.M.; Brogden, K.A.; Flaherty, H.A.; Gallagher, T.; et al. Mouse-adapted MERS coronavirus causes lethal lung disease in human DPP4 knockin mice. Proc. Natl. Acad. Sci. USA 2017, 114, E3119–E3128. [Google Scholar] [CrossRef]
- Zhao, G.; Jiang, Y.; Qiu, H.; Gao, T.; Zeng, Y.; Guo, Y.; Yu, H.; Li, J.; Kou, Z.; Du, L.; et al. Multi-Organ Damage in Human Dipeptidyl Peptidase 4 Transgenic Mice Infected with Middle East Respiratory Syndrome-Coronavirus. PLoS ONE 2015, 10, e145561. [Google Scholar] [CrossRef]
- Zhao, J.; Li, K.; Wohlford-Lenane, C.; Agnihothram, S.S.; Fett, C.; Zhao, J.; Gale, M.J.; Baric, R.S.; Enjuanes, L.; Gallagher, T.; et al. Rapid generation of a mouse model for Middle East respiratory syndrome. Proc. Natl. Acad. Sci. USA 2014, 111, 4970–4975. [Google Scholar] [CrossRef]
- Tseng, C.T.; Huang, C.; Newman, P.; Wang, N.; Narayanan, K.; Watts, D.M.; Makino, S.; Packard, M.M.; Zaki, S.R.; Chan, T.S.; et al. Severe acute respiratory syndrome coronavirus infection of mice transgenic for the human Angiotensin-converting enzyme 2 virus receptor. J. Virol. 2007, 81, 1162–1173. [Google Scholar] [CrossRef]
- Cockrell, A.S.; Yount, B.L.; Scobey, T.; Jensen, K.; Douglas, M.; Beall, A.; Tang, X.C.; Marasco, W.A.; Heise, M.T.; Baric, R.S. A mouse model for MERS coronavirus-induced acute respiratory distress syndrome. Nat. Microbiol. 2016, 2, 16226. [Google Scholar] [CrossRef]
- Haagmans, B.L.; van den Brand, J.M.; Raj, V.S.; Volz, A.; Wohlsein, P.; Smits, S.L.; Schipper, D.; Bestebroer, T.M.; Okba, N.; Fux, R.; et al. An orthopoxvirus-based vaccine reduces virus excretion after MERS-CoV infection in dromedary camels. Science 2016, 351, 77–81. [Google Scholar] [CrossRef]
- Adney, D.R.; Bielefeldt-Ohmann, H.; Hartwig, A.E.; Bowen, R.A. Infection, Replication, and Transmission of Middle East Respiratory Syndrome Coronavirus in Alpacas. Emerg. Infect. Dis. 2016, 22, 1031–1037. [Google Scholar] [CrossRef]
- Crameri, G.; Durr, P.A.; Klein, R.; Foord, A.; Yu, M.; Riddell, S.; Haining, J.; Johnson, D.; Hemida, M.G.; Barr, J.; et al. Experimental Infection and Response to Rechallenge of Alpacas with Middle East Respiratory Syndrome Coronavirus. Emerg. Infect. Dis. 2016, 22, 1071–1074. [Google Scholar] [CrossRef]
- Yao, Y.; Bao, L.; Deng, W.; Xu, L.; Li, F.; Lv, Q.; Yu, P.; Chen, T.; Xu, Y.; Zhu, H.; et al. An Animal Model of MERS Produced by Infection of Rhesus Macaques with MERS Coronavirus. J. Infect. Dis. 2013, 209, 236–242. [Google Scholar] [CrossRef]
- Yu, P.; Xu, Y.; Deng, W.; Bao, L.; Huang, L.; Xu, Y.; Yao, Y.; Qin, C. Comparative pathology of rhesus macaque and common marmoset animal models with Middle East respiratory syndrome coronavirus. PLoS ONE 2017, 12, e172093. [Google Scholar] [CrossRef] [PubMed]
- Falzarano, D.; de Wit, E.; Feldmann, F.; Rasmussen, A.L.; Okumura, A.; Peng, X.; Thomas, M.J.; van Doremalen, N.; Haddock, E.; Nagy, L.; et al. Infection with MERS-CoV causes lethal pneumonia in the common marmoset. PLoS Pathog. 2014, 10, e1004250. [Google Scholar] [CrossRef] [PubMed]
- Munster, V.J.; de Wit, E.; Feldmann, H. Pneumonia from human coronavirus in a macaque model. N. Engl. J. Med. 2013, 368, 1560–1562. [Google Scholar] [CrossRef] [PubMed]
- Falzarano, D.; de Wit, E.; Rasmussen, A.L.; Feldmann, F.; Okumura, A.; Scott, D.P.; Brining, D.; Bushmaker, T.; Martellaro, C.; Baseler, L.; et al. Treatment with interferon-alpha2b and ribavirin improves outcome in MERS-CoV-infected rhesus macaques. Nat. Med. 2013, 19, 1313–1317. [Google Scholar] [CrossRef]
- Johnson, R.F.; Via, L.E.; Kumar, M.R.; Cornish, J.P.; Yellayi, S.; Huzella, L.; Postnikova, E.; Oberlander, N.; Bartos, C.; Ork, B.L.; et al. Intratracheal exposure of common marmosets to MERS-CoV Jordan-n3/2012 or MERS-CoV EMC/2012 isolates does not result in lethal disease. Virology 2015, 485, 422–430. [Google Scholar] [CrossRef]
- Li, W.; Greenough, T.C.; Moore, M.J.; Vasilieva, N.; Somasundaran, M.; Sullivan, J.L.; Farzan, M.; Choe, H. Efficient replication of severe acute respiratory syndrome coronavirus in mouse cells is limited by murine angiotensin-converting enzyme 2. J. Virol. 2004, 78, 11429–11433. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Tu, C.; Xin, C.; Xuan, H.; Meng, Q.; Liu, Y.; Yu, Y.; Guan, Y.; Jiang, Y.; Yin, X.; et al. Civets are equally susceptible to experimental infection by two different severe acute respiratory syndrome coronavirus isolates. J. Virol. 2005, 79, 2620–2625. [Google Scholar] [CrossRef]
- Barlan, A.; Zhao, J.; Sarkar, M.K.; Li, K.; McCray, P.J.; Perlman, S.; Gallagher, T. Receptor variation and susceptibility to Middle East respiratory syndrome coronavirus infection. J. Virol. 2014, 88, 4953–4961. [Google Scholar] [CrossRef]
- van Doremalen, N.; Miazgowicz, K.L.; Milne-Price, S.; Bushmaker, T.; Robertson, S.; Scott, D.; Kinne, J.; McLellan, J.S.; Zhu, J.; Munster, V.J. Host species restriction of Middle East respiratory syndrome coronavirus through its receptor, dipeptidyl peptidase 4. J. Virol. 2014, 88, 9220–9232. [Google Scholar] [CrossRef]
| SARS-CoV | MERS-CoV | ||
|---|---|---|---|
| Genus | Beta-CoVs, lineage B | Beta-CoVs, lineage C | |
| Possible Natural Reservoir | Bat | Bat | |
| Possible Intermediary Host | Palm civet | Dromedary camel | |
| Origin | Guangdong province, China | Arabian Peninsula | |
| Clinical Epidemiology | Total global number reported to WHO | More than 8098 people | 2254 (from 2012 through 16 September 2018) |
| Affected countries | 29 | 27 | |
| Number of deaths | 916 | 800 | |
| Mortality | More than 10% | More than 35% | |
| Transmission region | Globally | Regionally | |
| Transmission patterns | From animal to human; from human to human | ||
| The predominant receptor | Human angiotensin-converting enzyme 2 (ACE2) | Human dipeptidyl peptidase 4 (DPP4 or CD26) | |
| Receptor distribution | Arterial and venous endothelium; arterial smooth muscle; small intestine; respiratory tract epithelium; alveolar monocytes and macrophages | Respiratory tract epithelium; kidney; small intestine; liver and prostate; activated leukocytes | |
| Cell line susceptibility | Respiratory tract; kidney; liver | Respiratory tract; intestinal tract; genitourinary tract; liver, kidney, neurons; monocyte; Tlymphocyte; and histiocytic cell lines | |
| Viral replication efficiency | High | Higher | |
| Ability to inhibit IFN production | Delayed recognition and proinflammatory response | Delayed recognition and proinflammatory response | |
| SARS-CoV | MERS-CoV | ||
|---|---|---|---|
| Length of nucleotides | 29,727 | 30,119 | |
| Open reading frames (ORFs) | 11 | 11 | |
| Structural protein | 4 | 4 | |
| Spike protein (length of amino acids) | 1255 | 1353 | |
| S1 subunit | Receptor-binding domain (RBD) | 318–510 | 367–588 |
| Receptor-binding motif (RBM) | 424–494 | 484–567 | |
| S2 subunit | Heptad repeat 1 (HR1) domains | 892–1013 | 984–1104 |
| Heptad repeat 2 (HR2) domains | 1145–1195 | 1246–1295 | |
| Non-structural proteins (NSPs) | At least 5 | 16 | |
| Accessory proteins | 8 | 5 | |
| A characteristic gene order | 5′-replicase ORF1ab, spike (S), envelope (E), membrane (M), and nucleocapsid (N)-3′ | ||
| Vaccine Strategy | Process of Production | References | Advantages | Disadvantages | |
|---|---|---|---|---|---|
| SARS | MERS | ||||
| Inactivated virus vaccines | Virus particles are inactivated by heat, chemicals, or radiation | Whole virus, with or without adjuvant (promote an effective immune response against the inactivated pathogen) [93,94] | Whole virus, with or without adjuvant (promote an effective immune response against the inactivated pathogen) [91,95] | Maintained virus particles structure; rapidly develop; easy to prepare; safety; high-titer neutralizing antibodies [93]; protection with adjuvant [96,97]. | Potential inappropriate for highly immunosuppressed individuals; possible TH2 cell-distortive immune response [98,99]. |
| Live-attenuated virus vaccines | Attenuated the virulence, but still keeping it viable by mutagenesis or targeted deletions | Envelope protein deletion [100]; non-structural protein 14 (nsp14) and exonuclease (ExoN) inactivation [101] | Full-length infectious cDNA clone or mutant viruses [102] | Inexpensive; quick immunity; less adverse effect; activates all phases of the immune system [103]; more durable immunity; more targeted [77]. | Phenotypic or genotypic reversion possible; need sufficient viral replication [77]. |
| Viral vector vaccines | Genetically engineered unrelated viral genome with deficient packaging elements for encoding targeted gene | Spike and nucleocapsid proteins [100,104] | Spike and nucleocapsid proteins [87,88] | Safety; stronger and specific cellular and humoral immune responses [77]. | Varies inoculation routes may produce different immune responses [96]; possibly incomplete protection; may fail in aged vaccinees; possible TH2 cell-distortive immune response [105]. |
| Subunit vaccines | Antigenic components inducing the immune system without introducing viral particles, whole or otherwise. | Spike and nucleocapsid proteins [53,59,106] | Spike and nucleocapsid proteins [85,86,107,108] | High safety; consistent production; can induce cellular and humoral immune responses; high-titer neutralizing antibodies [109]. | Uncertain cost-effectiveness; relatively lower immunogenicity; need appropriate adjuvants [77]. |
| DNA vaccines | Genetically engineered DNA for directly producing an antigen | Spike and nucleocapsid proteins [110,111] | Spike and nucleocapsid proteins [89,90] | Easier to design; high safety; high-titer neutralizing antibodies [110]. | Lower immune responses; potential TH2 cell-distortive immune response results; potential ineffective; possibly delayed-type hypersensitivity [112]. |
| Treatment | Stage of Development | |
|---|---|---|
| SARS (Notes) | MERS (Notes) | |
| Host protease inhibitors | Effective in mouse models [138] | In vitro inhibition [138] |
| Viral protease inhibitors | In vitro inhibition [139] | In vitro inhibition [140] |
| Monoclonal and polyclonal antibodies | Effective in mouse, ferrets, golden Syrian hamster [124,141,142] and non-human primate models [143,144] | Effective in mouse, rabbit, and non-human primate models [10,145] |
| Convalescent plasma | Off-label use in patients [146,147] | Effective in a mouse model; clinical trial approved [10] |
| Interferons | Off-label use in patients (often in combination with immunoglobulins or thymosins) [146,147] | Effective in non-human primate models; off-label use in patients (often in combination with a broad-spectrum antibiotic and oxygen) [10] |
| Ribavirin | Off-label use in patients (often in combination with corticosteroids) [146,147] | Effective in a non-human primate model; off-label use in patients (often in combination with a broad-spectrum antibiotic and oxygen) [10] |
| Lopinavir and ritonavir | Off-label use in patients (improved the outcome in combination with ribavirin) [146,147] | Effective in a non-human primate model; off-label use in patients [10,148] |
| Common Feature | None of these therapeutic agents are approved for commercial use in humans | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, Z.; Xu, Y.; Bao, L.; Zhang, L.; Yu, P.; Qu, Y.; Zhu, H.; Zhao, W.; Han, Y.; Qin, C. From SARS to MERS, Thrusting Coronaviruses into the Spotlight. Viruses 2019, 11, 59. https://doi.org/10.3390/v11010059
Song Z, Xu Y, Bao L, Zhang L, Yu P, Qu Y, Zhu H, Zhao W, Han Y, Qin C. From SARS to MERS, Thrusting Coronaviruses into the Spotlight. Viruses. 2019; 11(1):59. https://doi.org/10.3390/v11010059
Chicago/Turabian StyleSong, Zhiqi, Yanfeng Xu, Linlin Bao, Ling Zhang, Pin Yu, Yajin Qu, Hua Zhu, Wenjie Zhao, Yunlin Han, and Chuan Qin. 2019. "From SARS to MERS, Thrusting Coronaviruses into the Spotlight" Viruses 11, no. 1: 59. https://doi.org/10.3390/v11010059
APA StyleSong, Z., Xu, Y., Bao, L., Zhang, L., Yu, P., Qu, Y., Zhu, H., Zhao, W., Han, Y., & Qin, C. (2019). From SARS to MERS, Thrusting Coronaviruses into the Spotlight. Viruses, 11(1), 59. https://doi.org/10.3390/v11010059




