The Effect of Permethrin Resistance on Aedes aegypti Transcriptome Following Ingestion of Zika Virus Infected Blood
Abstract
1. Introduction
2. Materials and Methods
2.1. Mosquito Strains
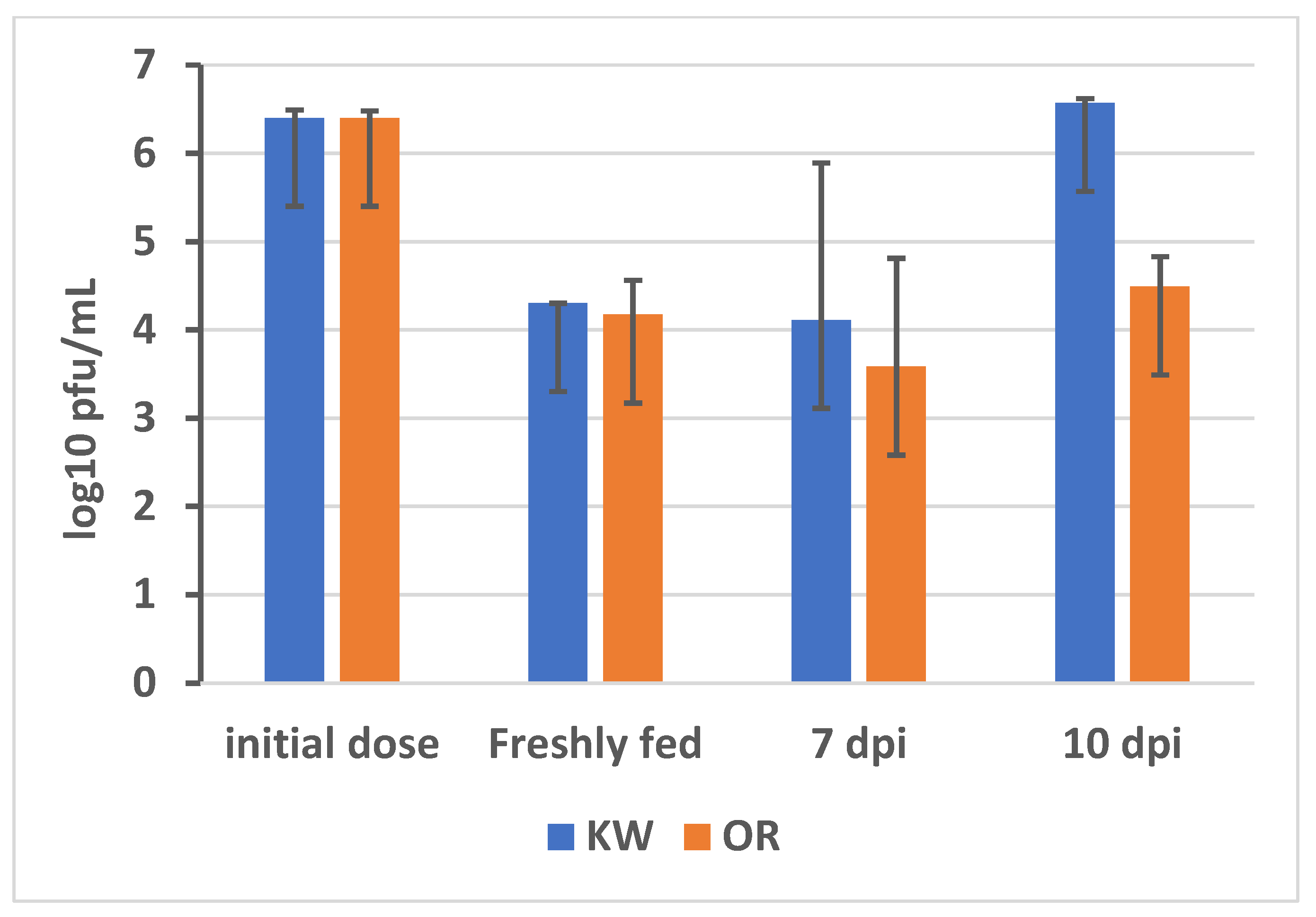
2.2. Zika Virus Infection
2.3. RNA Extraction
2.4. RNA-Seq Library Preparation and Sequencing
2.5. Data Mining and RNA-Seq Analysis
2.6. Assignments of Gene Ontology (GO) Terms and Pathway Analyses
2.7. C-DNA Synthesized and qPCR Amplification
3. Results
3.1. Global Changes in Transcriptome of the Aedes Aegypti Female Adult in Response to ZIKV Infection
3.1.1. Expression Profiles of Differentially Expressed (DE) Transcripts in Response to Blood Feeding (Control) between Two Aedes aegypti Strains, Resistant Versus Susceptible Strains
3.1.2. Expression Profiles of DE Transcripts in Aedes Aegypti, Resistant and Susceptible Strains, in Response to ZIKV
3.1.3. Expression Profiles of DE Transcripts in Response to ZIKV Infection between Two Strains Aedes Aegypti, Resistant Versus Susceptible Strains
3.1.4. DE Transcripts Related to Immunity in Response to ZIKV Infection
3.1.5. DE Transcripts Related to Detoxification in Response to ZIKV Infection
3.1.6. DE Transcripts Likely Related to Permethrin resistance in Response to ZIKV Infection
3.1.7. DE Transcripts Related to Cytoskeleton in Response to ZIKV Infection
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mousson, L.; Dauga, C.; Garrigues, T.; Schaffner, F.; Vazeille, M.; Failloux, A.B. Phylogeography of Aedes (Stegomyia) aegypti (L.) and Aedes (Stegomyia) albopictus (Skuse) (Diptera: Culicidae) based on mitochondrial DNA variations. Genet. Res. 2005, 86, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Goindin, D.; Delannay, C.; Ramdini, C.; Gustave, J.; Fouque, F. Parity and longevity of Aedes aegypti according to temperatures in controlled conditions and consequences on dengue transmission risks. PLoS ONE 2015, 10, e0135489. [Google Scholar] [CrossRef] [PubMed]
- Faye, O.; Freire, C.C.; Iamarino, A.; de Oliveira, J.V.; Diallo, M.; Zanotto, P.M.; Sall, A.A. Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl. Trop. Dis. 2014, 8, e2636. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Roche, C.; Teissier, A.; Robin, E.; Berry, A.L.; Mallet, H.P.; Sall, A.A.; Musso, D. Zika virus, French polynesia, South pacific, 2013. Emerg. Infect. Dis. 2014, 20, 1085–1086. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.S.; Bandeira, A.C.; Sardi, S.I. Zika Virus Outbreak, Bahia, Brazil. Emerg. Infect. Dis. 2015, 21, 1885–1886. [Google Scholar] [CrossRef] [PubMed]
- Sacramento, C.Q.; de Melo, G.R.; de Freitas, C.S.; Rocha, N.; Hoelz, L.V.; Miranda, M.; Fintelman-Rodrigues, N.; Marttorelli, A.; Ferreira, A.C.; Barbosa-Lima, G.; et al. The clinically approved antiviral drug sofosbuvir inhibits Zika virus replication. Sci. Rep. 2017, 7, 40920. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, E.L.; Tong, V.T.; Rozo, N.; Valencia, D.; Pacheco, O.; Gilboa, S.M.; Mercado, M.; Renquist, C.M.; González, M.; Ailes, E.C.; et al. Preliminary Report of Microcephaly Potentially Associated with Zika Virus Infection During Pregnancy—Colombia, January–November 2016. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 1409–1413. [Google Scholar] [CrossRef] [PubMed]
- Pinto-Díaz, C.A.; Rodríguez, Y.; Monsalve, D.M.; Acosta-Ampudia, Y.; Molano-González, N.; Anaya, J.M.; Ramírez-Santana, C. Autoimmunity in Guillain-Barré syndrome associated with Zika virus infection and beyond. Autoimmun. Rev. 2017, 16, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Estep, A.S.; Sanscrainte, N.D.; Waits, C.M.; Louton, J.E.; Becnel, J.J. Resistance Status and Resistance Mechanisms in a Strain of Aedes aegypti (Diptera: Culicidae) From Puerto Rico. J. Med. Entomol. 2017, 54, 1643–1648. [Google Scholar] [CrossRef] [PubMed]
- Antonio-Nkondjio, C.; Sonhafouo-Chiana, N.; Ngadjeu, C.S.; Doumbe-Belisse, P.; Talipouo, A.; Djamouko-Djonkam, L.; Kopya, E.; Bamou, R.; Awono-Ambene, P.; Wondji, C.S. Review of the evolution of insecticide resistance in main malaria vectors in Cameroon from 1990 to 2017. Parasit. Vectors 2017, 10, 472. [Google Scholar] [CrossRef] [PubMed]
- Ponce-García, G.; Del Río-Galvan, S.; Barrera, R.; Saavedra-Rodriguez, K.; Villanueva-Segura, K.; Felix, G.; Amador, M.; Flores, A.E. Knockdown Resistance Mutations in Aedes aegypti (Diptera: Culicidae) From Puerto Rico. J. Med. Entomol. 2016, 53, 1410–1414. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.M.; Chu, H.L.; Wang, G.; Zhu, X.J.; Guo, X.X.; Zhang, Y.M.; Xing, D.; Yan, T.; Zhao, M.H.; Dong, Y.D.; et al. Multiple-Insecticide Resistance and Classic Gene Mutations to Japanese Encephalitis Vector Culex tritaeniorhynchus from China. J. Am. Mosq. Control Assoc. 2016, 32, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Francis, S.; Saavedra-Rodriguez, K.; Perera, R.; Paine, M.; Black, W.C.; Delgoda, R. Insecticide resistance to permethrin and malathion and associated mechanisms in Aedes aegypti mosquitoes from St. Andrew Jamaica. PLoS ONE 2017, 12, e0179673. [Google Scholar]
- Seixas, G.; Grigoraki, L.; Weetman, D.; Vicente, J.L.; Silva, A.C.; Pinto, J.; Vontas, J.; Sousa, C.A. Insecticide resistance is mediated by multiple mechanisms in recently introduced Aedes aegypti from Madeira Island (Portugal). PLoS Negl. Trop. Dis. 2017, 11, e0005799. [Google Scholar] [CrossRef] [PubMed]
- Aguirre-Obando, O.A.; Martins, A.J.; Navarro-Silva, M.A. First report of the Phe1534Cys kdr mutation in natural populations of Aedes albopictus from Brazil. Parasit. Vectors 2017, 10, 160. [Google Scholar] [CrossRef] [PubMed]
- Liu, N. Insecticide resistance in mosquitoes: Impact, mechanisms, and research directions. Annu. Rev. Entomol. 2015, 60, 537–559. [Google Scholar] [CrossRef] [PubMed]
- Dang, K.; Doggett, S.L.; Veera Singham, G.; Lee, C.Y. Insecticide resistance and resistance mechanisms in bed bugs, Cimex spp. (Hemiptera: Cimicidae). Parasit. Vectors 2017, 10, 318. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Cupp, E.W.; Guo, A.; Liu, N. Insecticide resistance in Alabama and Florida mosquito strains of Aedes albopictus. J. Med. Entomol. 2004, 41, 946–952. [Google Scholar] [CrossRef] [PubMed]
- Hemingway, J.; Hawkes, N.J.; McCarroll, L.; Ranson, H. The molecular basis of insecticide resistance in mosquitoes. Insect. Biochem. Mol. Biol. 2004, 34, 653–665. [Google Scholar] [CrossRef] [PubMed]
- Alemayehu, E.; Asale, A.; Eba, K.; Getahun, K.; Tushune, K.; Bryon, A.; Morou, E.; Vontas, J.; van Leeuwen, T.; Duchateau, L.; et al. Mapping insecticide resistance and characterization of resistance mechanisms in Anopheles arabiensis (Diptera: Culicidae) in Ethiopia. Parasit. Vectors 2017, 10, 407. [Google Scholar] [CrossRef] [PubMed]
- Pignatelli, P.; Ingham, V.A.; Balabanidou, V.; Vontas, J.; Lycett, G.; Ranson, H. The Anopheles gambiae ATP-binding cassette transporter family: Phylogenetic analysis and tissue localization provide clues on function and role in insecticide resistance. Insect. Mol. Biol. 2017, 27, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Zhong, D.; Chang, X.; Zhou, G.; He, Z.; Fu, F.; Yan, Z.; Zhu, G.; Xu, T.; Bonizzoni, M.; Wang, M.H.; et al. Relationship between knockdown resistance, metabolic detoxification and organismal resistance to pyrethroids in Anopheles sinensis. PLoS ONE 2013, 8, e55475. [Google Scholar] [CrossRef] [PubMed]
- Mastrantonio, V.; Ferrari, M.; Epis, S.; Negri, A.; Scuccimarra, G.; Montagna, M.; Favia, G.; Porretta, D.; Urbanelli, S.; Bandi, C. Gene expression modulation of ABC transporter genes in response to permethrin in adults of the mosquito malaria vector Anopheles stephensi. Acta Trop. 2017, 171, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Grigoraki, L.; Pipini, D.; Labbé, P.; Chaskopoulou, A.; Weill, M.; Vontas, J. Carboxylesterase gene amplifications associated with insecticide resistance in Aedes albopictus: Geographical distribution and evolutionary origin. PLoS Negl. Trop. Dis. 2017, 11, e0005533. [Google Scholar] [CrossRef] [PubMed]
- Grigoraki, L.; Lagnel, J.; Kioulos, I.; Kampouraki, A.; Morou, E.; Labbé, P.; Weill, M.; Vontas, J. Transcriptome Profiling and Genetic Study Reveal Amplified Carboxylesterase Genes Implicated in Temephos Resistance, in the Asian Tiger Mosquito Aedes albopictus. PLoS Negl. Trop. Dis. 2015, 9, e0003771. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Liu, N. Regulation of P450-mediated permethrin resistance in Culex quinquefasciatus by the GPCR/Gαs/AC/cAMP/PKA signaling cascade. Biochem. Biophys. Rep. 2017, 12, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Huang, Y.; Zou, F.; Liu, B.; Tian, M.; Ye, W.; Guo, J.; Sun, X.; Zhou, D.; Sun, Y.; et al. The role of miR-2∼13∼71 cluster in resistance to deltamethrin in Culex pipiens pallens. Insect. Biochem. Mol. Biol. 2017, 84, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Al Nazawi, A.M.; Aqili, J.; Alzahrani, M.; McCall, P.J.; Weetman, D. Combined target site (kdr) mutations play a primary role in highly pyrethroid resistant phenotypes of Aedes aegypti from Saudi Arabia. Parasit. Vectors 2017, 10, 161. [Google Scholar] [CrossRef] [PubMed]
- Gong, Y.; Li, T.; Feng, Y.; Liu, N. The function of two P450s, CYP9M10 and CYP6AA7, in the permethrin resistance of Culex quinquefasciatus. Sci. Rep. 2017, 7, 587. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.S.; Kim, H.C.; Klein, T.A.; Ju, Y.R. Insecticide resistance and cytochrome-P450 activation in unfed and blood-fed laboratory and field populations of Culex pipiens pallens. J. Pest. Sci. 2017, 90, 759–771. [Google Scholar] [CrossRef] [PubMed]
- Faucon, F.; Gaude, T.; Dusfour, I.; Navratil, V.; Corbel, V.; Juntarajumnong, W.; Girod, R.; Poupardin, R.; Boyer, F.; Reynaud, S.; et al. In the hunt for genomic markers of metabolic resistance to pyrethroids in the mosquito Aedes aegypti: An integrated next-generation sequencing approach. PLoS Negl. Trop. Dis. 2017, 11, e0005526. [Google Scholar] [CrossRef] [PubMed]
- Faucon, F.; Dusfour, I.; Gaude, T.; Navratil, V.; Boyer, F.; Chandre, F.; Sirisopa, P.; Thanispong, K.; Juntarajumnong, W.; Poupardin, R.; et al. Identifying genomic changes associated with insecticide resistance in the dengue mosquito Aedes aegypti by deep targeted sequencing. Genome. Res. 2015, 25, 1347–1359. [Google Scholar] [CrossRef] [PubMed]
- Reid, W.R.; Zhang, L.; Gong, Y.; Li, T.; Liu, N. Gene expression profiles of the Southern house mosquito Culex quinquefasciatus during exposure to permethrin. Insect Sci. 2018, 25, 439–453. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Guo, J.; Ye, W.; Guo, Q.; Huang, Y.; Ma, L.; Zhou, D.; Shen, B.; Sun, Y.; Zhu, C. Cuticle genes CpCPR63 and CpCPR47 may confer resistance to deltamethrin in Culex pipiens pallens. Parasitol. Res. 2017, 116, 2175–2179. [Google Scholar] [CrossRef] [PubMed]
- Ishak, I.H.; Riveron, J.M.; Ibrahim, S.S.; Stott, R.; Longbottom, J.; Irving, H.; Wondji, C.S. The Cytochrome P450 gene CYP6P12 confers pyrethroid resistance in kdr-free Malaysian populations of the dengue vector Aedes albopictus. Sci. Rep. 2016, 6, 24707. [Google Scholar] [CrossRef] [PubMed]
- Smith, L.B.; Kasai, S.; Scott, J.G. Voltage-sensitive sodium channel mutations S989P + V1016G in Aedes aegypti confer variable resistance to pyrethroids, DDT and oxadiazines. Pest. Manag. Sci. 2017, 74, 737–745. [Google Scholar] [CrossRef] [PubMed]
- Hamid, P.H.; Prastowo, J.; Widyasari, A.; Taubert, A.; Hermosilla, C. Knockdown resistance (kdr) of the voltage-gated sodium channel gene of Aedes aegypti population in Denpasar, Bali, Indonesia. Parasit. Vectors 2017, 10, 283. [Google Scholar] [CrossRef] [PubMed]
- Ingham, V.A.; Pignatelli, P.; Moore, J.D.; Wagstaff, S.; Ranson, H. The transcription factor Maf-S regulates metabolic resistance to insecticides in the malaria vector Anopheles gambiae. BMC Genom. 2017, 18, 669. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Behura, S.K.; Franz, A.W.E. The midgut transcriptome of Aedes aegypti fed with saline or protein meals containing chikungunya virus reveals genes potentially involved in viral midgut escape. BMC Genom. 2017, 18, 382. [Google Scholar] [CrossRef] [PubMed]
- Shin, D.; Civana, A.; Acevedo, C.; Smartt, C.T. Transcriptomics of differential vector competence: West Nile virus infection in two populations of Culex pipiens quinquefasciatus linked to ovary development. BMC Genom. 2014, 15, 513. [Google Scholar] [CrossRef] [PubMed]
- Shrinet, J.; Srivastava, P.; Sunil, S. Transcriptome analysis of Aedes aegypti in response to mono-infections and co-infections of dengue virus-2 and chikungunya virus. Biochem. Biophys. Res. Commun. 2017, 492, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Shrinet, J.; Jain, S.; Jain, J.; Bhatnagar, R.K.; Sunil, S. Next generation sequencing reveals regulation of distinct Aedes microRNAs during chikungunya virus development. PLoS Negl. Trop. Dis. 2014, 8, e2616. [Google Scholar] [CrossRef] [PubMed]
- Etebari, K.; Hegde, S.; Saldaña, M.A.; Widen, S.G.; Wood, T.G.; Asgari, S.; Hughes, G.L. Global Transcriptome Analysis of Aedes aegypti Mosquitoes in Response to Zika Virus Infection. mSphere 2017, 2, e00456-17. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Smagghe, G.; Meeus, I. The role of a single gene encoding the Single von Willebrand factor C-domain protein (SVC) in bumblebee immunity extends beyond antiviral defense. Insect Biochem. Mol. Biol. 2017, 91, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Angleró-Rodríguez, Y.I.; MacLeod, H.J.; Kang, S.; Carlson, J.S.; Jupatanakul, N.; Dimopoulos, G. Molecular Responses to Zika Virus: Modulation of Infection by the Toll and Jak/Stat Immune Pathways and Virus Host Factors. Front. Microbiol. 2017, 8, 2050. [Google Scholar] [CrossRef] [PubMed]
- Tongluan, N.; Ramphan, S.; Wintachai, P.; Jaresitthikunchai, J.; Khongwichit, S.; Wikan, N.; Rajakam, S.; Yoksan, S.; Wongsiriroj, N.; Roytrakul, S.; et al. Involvement of fatty acid synthase in dengue virus infection. Virol. J. 2017, 14, 28. [Google Scholar] [CrossRef] [PubMed]
- Saldaña, M.A.; Etebari, K.; Hart, C.E.; Widen, S.G.; Wood, T.G.; Thangamani, S.; Asgari, S.; Hughes, G.L. Zika virus alters the microRNA expression profile and elicits an RNAi response in Aedes aegypti mosquitoes. PLoS Negl. Trop. Dis. 2017, 11, e0005760. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Alto, B.W.; Smartt, C.T.; Shin, D. Transcription Profiling for Defensins of Aedes aegypti (Diptera: Culicidae) During Development and in Response to Infection With Chikungunya and Zika Viruses. J. Med. Entomol. 2017, 55, 78–89. [Google Scholar]
- Dusfour, I.; Zorrilla, P.; Guidez, A.; Issaly, J.; Girod, R.; Guillaumot, L.; Robello, C.; Strode, C. Deltamethrin Resistance Mechanisms in Aedes aegypti Populations from Three French Overseas Territories Worldwide. PLoS Negl. Trop. Dis. 2015, 9, e0004226. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.Q.; Yu, F. DEB: A web interface for RNA-seq digital gene expression analysis. Bioinformation 2011, 7, 44–45. [Google Scholar] [CrossRef] [PubMed]
- Cerenius, L.; Söderhäll, K. The prophenoloxidase-activating system in invertebrates. Immunol. Rev. 2004, 198, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Fischetti, V.A. Streptococcal M protein: Molecular design and biological behavior. Clin. Microbiol. Rev. 1989, 2, 285–314. [Google Scholar] [CrossRef] [PubMed]
- Glinton, K.; Beck, J.; Liang, Z.; Qiu, C.; Lee, S.W.; Ploplis, V.A.; Castellino, F.J. Variable region in streptococcal M-proteins provides stable binding with host fibrinogen for plasminogen-mediated bacterial invasion. J. Biol. Chem. 2017, 292, 6775–6785. [Google Scholar] [CrossRef] [PubMed]
- Llimargas, M.; Strigini, M.; Katidou, M.; Karagogeos, D.; Casanova, J. Lachesin is a component of a septate junction-based mechanism that controls tube size and epithelial integrity in the Drosophila tracheal system. Development 2004, 131, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.; Lee, S.; Dziarski, R.; Chakravarti, S. A novel antimicrobial peptidoglycan recognition protein in the cornea. Invest. Ophthalmol. Vis. Sci. 2009, 50, 4185–4191. [Google Scholar] [CrossRef] [PubMed]
- Moyes, C.L.; Vontas, J.; Martins, A.J.; Ng, L.C.; Koou, S.Y.; Dusfour, I.; Raghavendra, K.; Pinto, J.; Corbel, V.; David, J.P.; et al. Contemporary status of insecticide resistance in the major Aedes vectors of arboviruses infecting humans. PLoS Negl. Trop. Dis. 2017, 11, e0005625. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, B.J.; Pignatelli, P.; Nikou, D.; Paine, M.J. Pinpointing P450s associated with pyrethroid metabolism in the dengue vector, Aedes aegypti: Developing new tools to combat insecticide resistance. PLoS Negl. Trop. Dis. 2012, 6, e1595. [Google Scholar] [CrossRef] [PubMed]
- Bariami, V.; Jones, C.M.; Poupardin, R.; Vontas, J.; Ranson, H. Gene amplification, ABC transporters and cytochrome P450s: Unraveling the molecular basis of pyrethroid resistance in the dengue vector, Aedes aegypti. PLoS Negl. Trop. Dis. 2012, 6, e1692. [Google Scholar] [CrossRef] [PubMed]
- Etebari, K.; Osei-Amo, S.; Blomberg, S.P.; Asgari, S. Dengue virus infection alters post-transcriptional modification of microRNAs in the mosquito vector Aedes aegypti. Sci. Rep. 2015, 5, 15968. [Google Scholar] [CrossRef] [PubMed]
- Acharya, D.; Paul, A.M.; Anderson, J.F.; Huang, F.; Bai, F. Loss of Glycosaminoglycan Receptor Binding after Mosquito Cell Passage Reduces Chikungunya Virus Infectivity. PLoS Negl. Trop. Dis. 2015, 9, e0004139. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.W.; He, Z.B.; Yan, Z.T.; Si, F.L.; Zhou, Y.; Chen, B. Genome-wide and expression-profiling analyses suggest the main cytochrome P450 genes related to pyrethroid resistance in the malaria vector, Anopheles sinensis (Diptera Culicidae). Pest. Manag. Sci. 2018, 74, 1810–1820. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Ochomo, E.; Dunn, W.A.; Britton, M.; Afrane, Y.; Zhou, G.; Hartsel, J.; Lee, M.C.; Xu, J.; Githeko, A.; et al. RNA-seq analyses of changes in the Anopheles gambiae transcriptome associated with resistance to pyrethroids in Kenya: Identification of candidate-resistance genes and candidate-resistance SNPs. Parasit. Vectors 2015, 8, 474. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Wang, W.; Hong, S.; Lei, Z.; Fang, F.; Guo, Q.; Hu, S.; Tian, M.; Liu, B.; Zhang, D.; et al. Comparative transcriptome analyses of deltamethrin-susceptible and -resistant Culex pipiens pallens by RNA-seq. Mol. Genet. Genom. 2016, 291, 309–321. [Google Scholar] [CrossRef] [PubMed]
- Bonizzoni, M.; Afrane, Y.; Dunn, W.A.; Atieli, F.K.; Zhou, G.; Zhong, D.; Li, J.; Githeko, A.; Yan, G. Comparative transcriptome analyses of deltamethrin-resistant and -susceptible Anopheles gambiae mosquitoes from Kenya by RNA-Seq. PLoS ONE 2012, 7, e44607. [Google Scholar] [CrossRef] [PubMed]
- Rivero, A.; Vézilier, J.; Weill, M.; Read, A.F.; Gandon, S. Insecticide control of vector-borne diseases: When is insecticide resistance a problem? PLoS Pathog. 2010, 6, e1001000. [Google Scholar] [CrossRef] [PubMed]
- Alout, H.; Ndam, N.T.; Sandeu, M.M.; Djégbe, I.; Chandre, F.; Dabiré, R.K.; Djogbénou, L.S.; Corbel, V.; Cohuet, A. Insecticide resistance alleles affect vector competence of Anopheles gambiae s.s. for Plasmodium falciparum field isolates. PLoS ONE 2013, 8, e63849. [Google Scholar] [CrossRef] [PubMed]
- McCarroll, L.; Hemingway, J. Can insecticide resistance status affect parasite transmission in mosquitoes? Insect. Biochem. Mol. Biol. 2002, 32, 1345–1351. [Google Scholar] [CrossRef]
- McCarroll, L.; Paton, M.G.; Karunaratne, S.H.; Jayasuryia, H.T.; Kalpage, K.S.; Hemingway, J. Insecticides and mosquito-borne disease. Nature. 2000, 407, 961–962. [Google Scholar] [CrossRef] [PubMed]
- Mondotte, J.A.; Saleh, M.C. Antiviral Immune Response and the Route of Infection in Drosophila melanogaster. Adv. Virus Res. 2018, 100, 247–278. [Google Scholar] [PubMed]
- Yoneyama, M.; Fujita, T. Recognition of viral nucleic acids in innate immunity. Rev. Med. Virol. 2010, 20, 4–22. [Google Scholar] [CrossRef] [PubMed]
- Onomoto, K.; Yoneyama, M.; Fujita, T. Recognition of viral nucleic acids and regulation of type I. IFN expression. Nihon Rinsho 2006, 64, 1236–1243. [Google Scholar] [PubMed]
- Dziarski, R. Peptidoglycan recognition proteins (PGRPs). Mol. Immunol. 2004, 40, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Kanost, M.R.; Jiang, H. Clip-domain serine proteases as immune factors in insect hemolymph. Curr. Opin. Insect Sci. 2015, 11, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Conway, M.J.; Watson, A.M.; Colpitts, T.M.; Dragovic, S.M.; Li, Z.; Wang, P.; Feitosa, F.; Shepherd, D.T.; Ryman, K.D.; Klimstra, W.B.; et al. Mosquito saliva serine protease enhances dissemination of dengue virus into the mammalian host. J. Virol. 2014, 88, 164–175. [Google Scholar] [CrossRef] [PubMed]
- Shin, S.W.; Bian, G.; Raikhel, A.S. A toll receptor and a cytokine, Toll5A and Spz1C, are involved in toll antifungal immune signaling in the mosquito Aedes aegypti. J. Biol. Chem. 2006, 281, 39388–39395. [Google Scholar] [CrossRef] [PubMed]
- Zou, Z.; Shin, S.W.; Alvarez, K.S.; Kokoza, V.; Raikhel, A.S. Distinct melanization pathways in the mosquito Aedes aegypti. Immunity 2010, 32, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, Y.; Morita, T. Structure and function of snake venom cysteine-rich secretory proteins. Toxicon 2004, 44, 227–231. [Google Scholar] [CrossRef] [PubMed]
- Londono-Renteria, B.; Troupin, A.; Conway, M.J.; Vesely, D.; Ledizet, M.; Roundy, C.M.; Cloherty, E.; Jameson, S.; Vanlandingham, D.; Higgs, S.; et al. Dengue Virus Infection of Aedes aegypti Requires a Putative Cysteine Rich Venom Protein. PLoS Pathog. 2015, 11, e1005202. [Google Scholar] [CrossRef] [PubMed]
- Itokawa, K.; Komagata, O.; Kasai, S.; Kawada, H.; Mwatele, C.; Dida, G.O.; Njenga, S.M.; Mwandawiro, C.; Tomita, T. Global spread and genetic variants of the two CYP9M10 haplotype forms associated with insecticide resistance in Culex quinquefasciatus Say. Heredity (Edinb) 2013, 111, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Itokawa, K.; Komagata, O.; Kasai, S.; Ogawa, K.; Tomita, T. Testing the causality between CYP9M10 and pyrethroid resistance using the TALEN and CRISPR/Cas9 technologies. Sci. Rep. 2016, 6, 24652. [Google Scholar] [CrossRef] [PubMed]
- David, J.P.; Faucon, F.; Chandor-Proust, A.; Poupardin, R.; Riaz, M.A.; Bonin, A.; Navratil, V.; Reynaud, S. Comparative analysis of response to selection with three insecticides in the dengue mosquito Aedes aegypti using mRNA sequencing. BMC Genom. 2014, 15, 174. [Google Scholar] [CrossRef] [PubMed]
- Riaz, M.A.; Chandor-Proust, A.; Dauphin-Villemant, C.; Poupardin, R.; Jones, C.M.; Strode, C.; Régent-Kloeckner, M.; David, J.P.; Reynaud, S. Molecular mechanisms associated with increased tolerance to the neonicotinoid insecticide imidacloprid in the dengue vector Aedes aegypti. Aquat. Toxicol. 2013, 126, 326–337. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Liu, N. Genome analysis of cytochrome P450s and their expression profiles in insecticide resistant mosquitoes, Culex quinquefasciatus. PLoS ONE 2011, 6, e29418. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.; Zhong, D.; Cao, J.; Zhou, H.; Li, J.; Liu, Y.; Bai, L.; Xu, S.; Wang, M.H.; Zhou, G.; et al. Transcriptome profiling of pyrethroid resistant and susceptible mosquitoes in the malaria vector, Anopheles sinensis. BMC Genom. 2014, 15, 448. [Google Scholar] [CrossRef] [PubMed]
- Cudmore, S.; Reckmann, I.; Way, M. Viral manipulations of the actin cytoskeleton. Trends Microbiol. 1997, 5, 142–148. [Google Scholar] [CrossRef]
- Sanders, M.C.; Theriot, J.A. Tails from the hall of infection: Actin-based motility of pathogens. Trends Microbiol. 1996, 4, 211–213. [Google Scholar] [CrossRef]
- Luftig, R.B.; Lupo, L.D. Viral interactions with the host-cell cytoskeleton: The role of retroviral proteases. Trends Microbiol. 1994, 2, 178–182. [Google Scholar] [CrossRef]
- Zhang, J.; Wu, N.; Gao, N.; Yan, W.; Sheng, Z.; Fan, D.; An, J. Small G Rac1 is involved in replication cycle of dengue serotype 2 virus in EAhy926 cells via the regulation of actin cytoskeleton. Sci. China Life Sci. 2016, 59, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Brice, A.; Whelan, D.R.; Ito, N.; Shimizu, K.; Wiltzer-Bach, L.; Lo, C.Y.; Blondel, D.; Jans, D.A.; Bell, T.D.; Moseley, G.W. Quantitative Analysis of the Microtubule Interaction of Rabies Virus P3 Protein: Roles in Immune Evasion and Pathogenesis. Sci. Rep. 2016, 6, 33493. [Google Scholar] [CrossRef] [PubMed]
- Oberhofer, A.; Spieler, P.; Rosenfeld, Y.; Stepp, W.L.; Cleetus, A.; Hume, A.N.; Mueller-Planitz, F.; Ökten, Z. Myosin Va’s adaptor protein melanophilin enforces track selection on the microtubule and actin networks in vitro. Proc. Natl. Acad. Sci. USA 2017, 114, E4714–E4723. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Xiao, X.; Yin, D.; Zhang, X. The interaction between viral protein and host actin facilitates the virus infection to host. Gene 2012, 507, 139–145. [Google Scholar] [CrossRef] [PubMed]
| Gene ID | logFC | p-adj | Gene Description | Gene Name | Publications |
|---|---|---|---|---|---|
| A | |||||
| AAEL012457 | 2.2934 | 3.09 × 10−10 | alcohol dehydrogenase | Faucon et al., 2015 [32] | |
| AAEL009044 | 2.5174 | 4.04 × 10−11 | amine oxidase | Faucon et al., 2015 [32] | |
| AAEL002385 | 2.0902 | 2.33 × 10−3 | Carboxy/choline esterase | CCEAE3B | Dusfour, 2015 [49] |
| AAEL001960 | 2.1283 | 7.7 × 10−7 | cytochrome P450 | CYP6M9 | Faucon et al., 2015 [32] |
| AAEL002031 | 1.837 | 1.37 × 10−19 | cytochrome P450 | CYP12F7 | Faucon et al., 2015 [32] |
| AAEL006798 | 2.8656 | 8.05 × 10−3 | cytochrome P450 | CYPJ10 | Faucon et al., 2017 [31] |
| AAEL006805 | 2.6907 | 6.0 × 10−18 | cytochrome P450 | CYP9J2 | Faucon et al., 2015 [32] |
| AAEL006815 | 2.4201 | 6.0 × 10−17 | cytochrome P450 | CYP9J16 | Faucon et al., [31,32] |
| AAEL007473 | 2.1909 | 5.5 × 10−15 | cytochrome P450 | CYP6AH1 | Faucon et al., 2015 [32] |
| AAEL009018 | 2.5349 | 4.5 × 10−8 | cytochrome P450 | CYP6CB1 | Faucon et al., 2015 [32] |
| AAEL009123 | 2.4747 | 7.8 × 10−9 | cytochrome P450 | CYP6Z6 | Faucon et al., 2017 [31] |
| AAEL009125 | 2.6551 | 7.0 × 10−13 | cytochrome P450 | CYP6M10 | Faucon et al., 2017 [31] |
| AAEL009129 | 2.4377 | 2.6 × 10−34 | cytochrome P450 | CYP6Z9 | Faucon et al., 2017 [31] |
| AAEL014603 | 2.4789 | 2.24 × 10−3 | cytochrome P450 | CYP9J30 | Faucon et al., [31,32] |
| AAEL014607 | 2.1961 | 4.6 × 10−4 | cytochrome P450 | CYP9J? | Faucon et al., 2015 [32] |
| AAEL014608 | 1.9517 | 8.2 × 10−3 | cytochrome P450 | CYP9J? | Faucon et al., 2017 [31] |
| AAEL014609 | 2.4159 | 1.1 × 10−12 | cytochrome P450 | CYP9J26 | Faucon et al., [31,32] |
| AAEL014614 | 3.9363 | 4.1 × 10−8 | cytochrome P450 | CYP9J? | Faucon et al., 2015 [32] |
| AAEL014617 | 2.3975 | 5.7 × 10−4 | cytochrome P450 | CYP9J28 | Faucon et al., 2015 [32] |
| AAEL014893 | 2.2053 | 2.1 × 10−8 | cytochrome P450 | CYP6BB2 | Faucon et al., 2015 [32] |
| AAEL015663 | 4.0198 | 9.0 × 10−6 | cytochrome P450 | CYP25? | Faucon et al., 2015 [32] |
| AAEL017297 | 3.2113 | 1.7 × 10−6 | cytochrome P450 | CYP6M9 | Faucon et al., [31,32] |
| AAEL003099 | 3.0651 | 1.4 × 10−4 | glucosyl/glucuronosyl transferases | Faucon et al., 2015 [32] | |
| B | |||||
| AAEL001312 | 2.0651 | 2.5 × 10−4 | cytochrome P450 | CYP9M6 | Faucon et al., 2017 [31] |
| AAEL006798 | 3.6029 | 5.1 × 10−4 | cytochrome P450 | CYP9J10 | Faucon et al., 2015 [32] |
| AAEL006811 | 2.5971 | 2.37 × 10−3 | cytochrome P450 | CYP9J8 | Faucon et al., [31,32] |
| AAEL014606 | 1.9372 | 1.49 × 10−6 | cytochrome P450 | CYPJ7 | Faucon et al., 2015 [32] |
| AAEL014617 | 2.2615 | 3.35 × 10−3 | cytochrome P450 | CYPJ28 | Faucon et al., 2015 [32] |
| AAEL014891 | −2.8122 | 3.8 × 10−4 | cytochrome P450 | CYP6P12 | Faucon et al., 2017 [31] |
| AAEL007947 | 2.5390 | 8.31 × 10−32 | glutathione transferase | GSTE | Faucon et al., 2017 [31] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, L.; Alto, B.W.; Shin, D.; Yu, F. The Effect of Permethrin Resistance on Aedes aegypti Transcriptome Following Ingestion of Zika Virus Infected Blood. Viruses 2018, 10, 470. https://doi.org/10.3390/v10090470
Zhao L, Alto BW, Shin D, Yu F. The Effect of Permethrin Resistance on Aedes aegypti Transcriptome Following Ingestion of Zika Virus Infected Blood. Viruses. 2018; 10(9):470. https://doi.org/10.3390/v10090470
Chicago/Turabian StyleZhao, Liming, Barry W. Alto, Dongyoung Shin, and Fahong Yu. 2018. "The Effect of Permethrin Resistance on Aedes aegypti Transcriptome Following Ingestion of Zika Virus Infected Blood" Viruses 10, no. 9: 470. https://doi.org/10.3390/v10090470
APA StyleZhao, L., Alto, B. W., Shin, D., & Yu, F. (2018). The Effect of Permethrin Resistance on Aedes aegypti Transcriptome Following Ingestion of Zika Virus Infected Blood. Viruses, 10(9), 470. https://doi.org/10.3390/v10090470





