Genes for Membrane Transport Proteins: Not So Rare in Viruses
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Channels
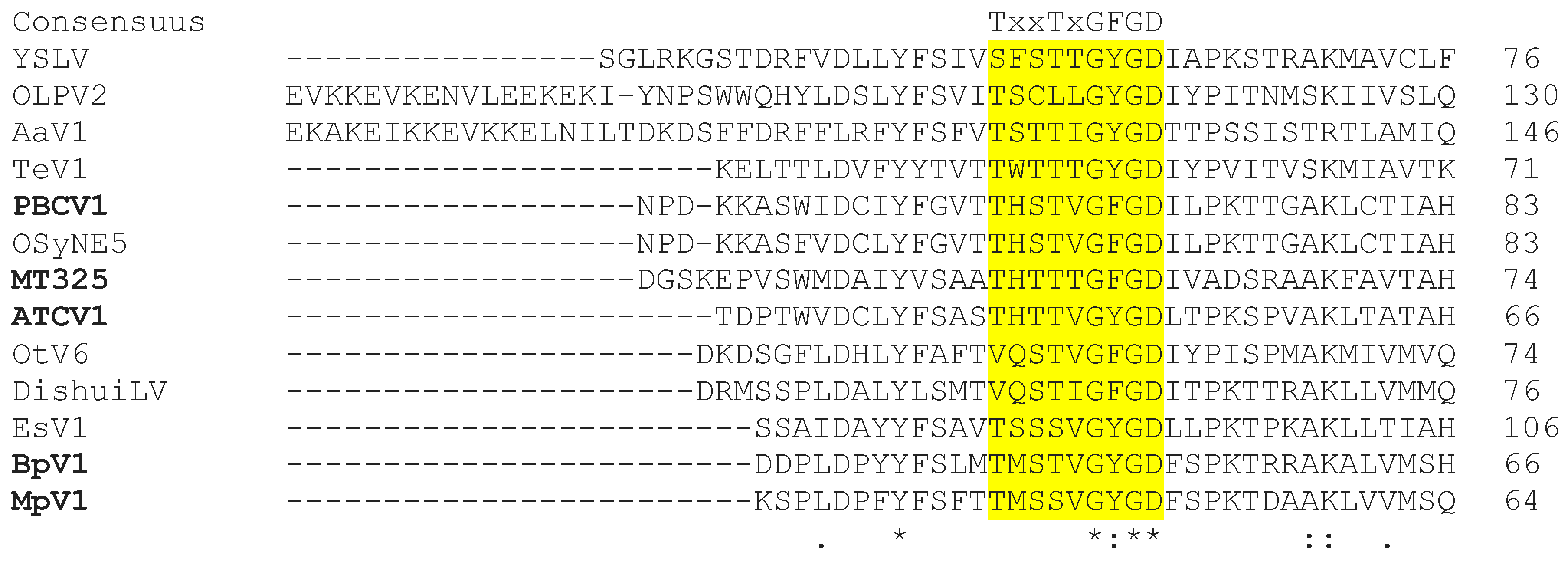
3.1.1. Potassium Channels
3.1.2. Chloride Channels
3.1.3. Ligand-Gated Ion Channels
3.1.4. Aquaglyceroporins
3.1.5. Mechanosensitive Channels
3.1.6. Other Ion Channels
3.2. ATPases and ABC Transporters
3.2.1. ATPases
3.2.2. ABC Transporters
3.3. Transporters
3.3.1. Potassium Transporters
3.3.2. Magnesium Transporters
3.3.3. Ammonium Transporters
3.3.4. Sodium/Phosphate Symporters
3.3.5. Sodium/Glucose Transporters
3.3.6. Sodium/Calcium Symporter
3.3.7. Nicotinamide Mononucleotide Transporters
3.3.8. Nucleoside Transporters
3.3.9. Amino Acid Transporters
3.4. Other Proteins
Putative Transport Proteins
- (i)
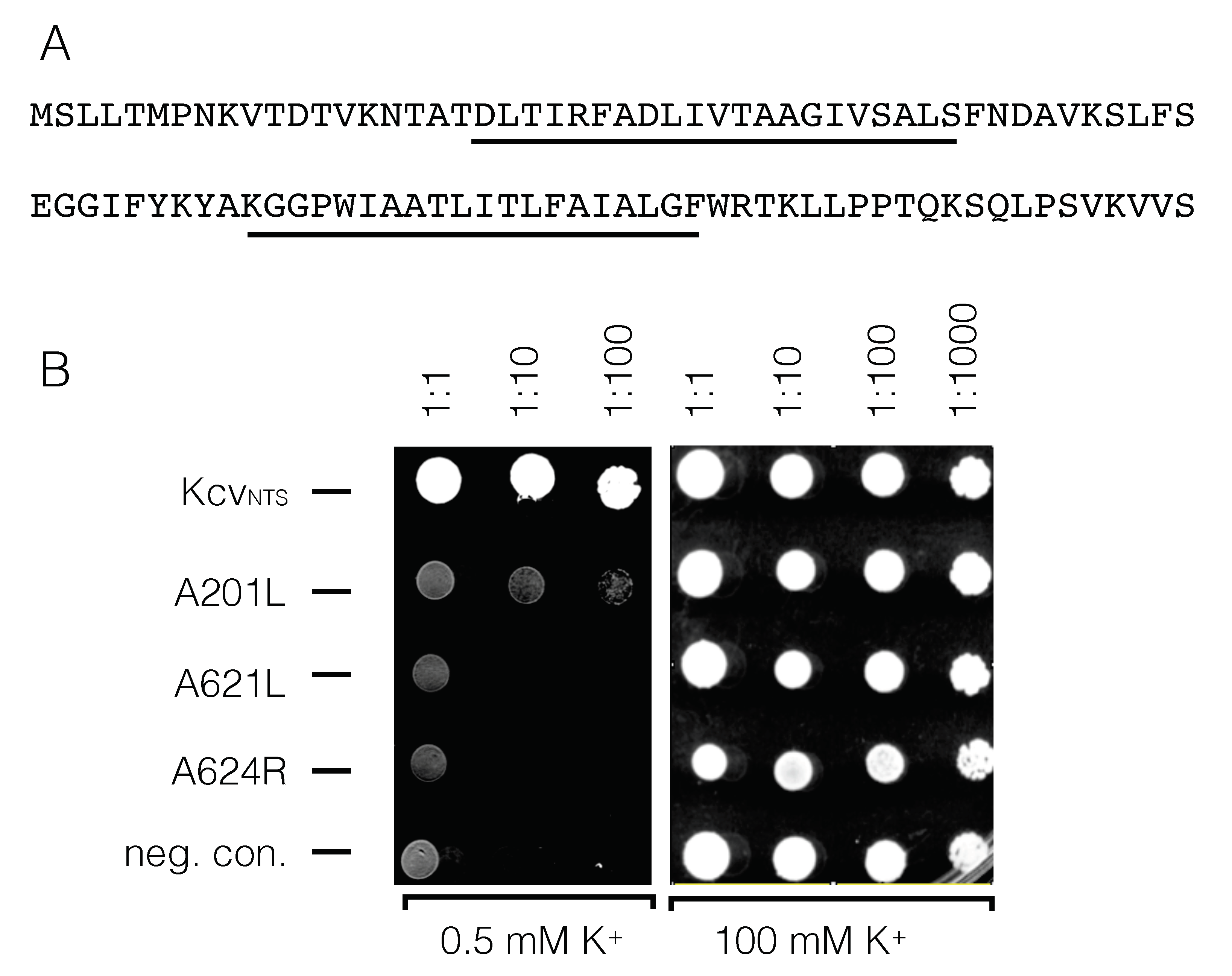
- ORF A201L (accession #AAC96569.1) codes for a 94 aa protein with two predicted TMs. It has weak similarity to proteins from Archeae.
- (ii)
- ORF A621L (accession #AAC96952.2) codes for a 117 aa protein with two TMs and weak similarity to the potassium channel Kcv. There are no non-viral homologs so far in the databases.
- (iii)
- ORF A624R (accession #AAC96955.1) codes for a 121 aa protein with two or three predicted TMs. It also contains the DUF2177 conserved domain that is so far not characterized.
4. Discussion
4.1. Abundance of Viral Transport Proteins and Function
4.2. The Evolutionary Origin of Viral Transport Proteins
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pinto, L.H.; Holsinger, L.J.; Lamb, R.A. Influenza virus M2 protein has ion channel activity. Cell 1992, 69, 517–528. [Google Scholar] [CrossRef]
- Fischer, W.B.; Sansom, M.S. Viral ion channels: Structure and function. Biochim. Biophys. Acta 2002, 1561, 27–45. [Google Scholar] [CrossRef]
- Gonzalez, M.E.; Carrasco, L. Viroporins. FEBS Lett. 2003, 552, 28–34. [Google Scholar] [CrossRef]
- Wang, K.; Xie, S.Q.; Sun, B. Viral proteins function as ion channels. Biochim. Biophys. Acta 2011, 1808, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Nieva, J.L.; Madan, V.; Carrasco, L. Viroporins: Structure and biological functions. Nat. Rev. Microbiol. 2012, 10, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Sze, C.W.; Tan, Y.J. Viral membrane channels: Roles and function in the virus life cycle. Viruses 2015, 7, 3261–3284. [Google Scholar] [CrossRef] [PubMed]
- Takeda, M.; Pekosz, A.; Shuck, K.; Pinto, L.H.; Lamb, R.A. Influenza A virus M2 ion channel activity is essential for efficient replication in tissue culture. J. Virol. 2002, 76, 1391–1399. [Google Scholar] [CrossRef] [PubMed]
- Thomaston, J.L.; DeGrado, W.F. Crystal structure of the drug-resistant S31N influenza M2 proton channel. Protein Sci. 2016, 25, 1551–1554. [Google Scholar] [CrossRef] [PubMed]
- Schubert, U.; Bour, S.; Ferrer-Montiel, A.V.; Montal, M.; Maldarell, F.; Strebel, K. The two biological activities of human immunodeficiency virus type Vpu protein involve two separable structural domains. J. Virol. 1996, 70, 809–819. [Google Scholar] [PubMed]
- Greiner, T.; Bolduan, S.; Gross, C.; Hamacher, K.; Schubert, U.; Moroni, A.; Thiel, G. Ion channel activity of Vpu proteins is conserved throughout evolution of HIV-1 and SIV. Viruses 2016, 8, 325. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lin, E.C.; Das, B.B.; Tian, Y.; Opella, S.J. Structural determination of virus protein U from HIV-1 by NMR in membrane environments. Biochim. Biophys. Acta 2015, 1848, 3007–3018. [Google Scholar] [CrossRef] [PubMed]
- Plugge, B.; Gazzarini, S.; Cerana, R.; Van Etten, J.; Nelson, M.; DiFrancesco, D.; Moroni, A.; Thiel, G. A potassium ion channel protein encoded by chlorella virus PBCV-1. Science 2000, 287, 1641–1644. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G.; Baumeister, D.; Schroeder, I.; Kast, S.M.; Van Etten, J.L.; Moroni, A. Minimal art: Or why small viral K+ channels are good tools for understanding basic structure and function relations. Biochim. Biophys. Acta 2011, 1808, 580–588. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G.; Greiner, T.; Dunigan, D.D.; Moroni, A.; Van Etten, J.L. Large dsDNA chloroviruses encode several membrane transport proteins. Virology 2015, 479–480, 38–45. [Google Scholar] [CrossRef] [PubMed]
- Manzoor, R.; Igarashi, M.; Takada, A. Influenza A virus M2 protein: Roles from ingress to egress. Int. J. Mol. Sci. 2017, 18, E2649. [Google Scholar] [CrossRef] [PubMed]
- Greiner, T.; Frohns, F.; Kang, M.; Van Etten, J.L.; Käsmann, A.; Moroni, A.; Hertel, B.; Thiel, G. Chlorella viruses prevent multiple infections by membrane depolarization of the host. J. Gen. Virol. 2009, 90, 2033–2039. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Lee, H.H.; Thiel, G.; Van Etten, J.L.; Saraf, R.F. Noninvasive measurement of a single viral infection in a cell. ACS Nano 2016, 10, 5123–5130. [Google Scholar] [CrossRef] [PubMed]
- Gazzarrini, S.; Kang, M.; Epimashko, S.; Van Etten, J.L.; Dainty, J.; Thiel, G.; Moroni, A. Chlorella virus MT325 encodes water and potassium channels that interact synergistically. Proc. Natl. Acad. Sci. USA 2006, 103, 5355–5360. [Google Scholar] [CrossRef] [PubMed]
- Bonza, M.C.; Martin, H.; Kang, M.; Lewis, G.; Greiner, T.; Giacometti, S.; Van Etten, J.L.; De Michelis, M.I.; Thiel, G.; Moroni, A. A functional calcium-transporting ATPase encoded by chlorella viruses. J. Gen. Virol. 2010, 91, 2620–2629. [Google Scholar] [CrossRef] [PubMed]
- Greiner, T.; Ramos, J.; Alvarez, M.C.; Gurnon, J.R.; Kang, M.; Van Etten, J.L.; Moroni, A.; Thiel, G. Functional HAK/KUP/KT-like potassium transporters encoded by chlorella viruses. Plant J. 2011, 68, 977–986. [Google Scholar] [CrossRef] [PubMed]
- Forterre, P. Giant viruses: Conflicts in revisiting the virus concept. Intervirology 2010, 53, 362–378. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Wolf, Y.I.; Koonin, E.V. Origin of giant viruses from smaller DNA viruses not from a fourth domain of cellular life. Virology 2014, 466–467, 38–52. [Google Scholar] [CrossRef] [PubMed]
- Claverie, J.-M.; Abergel, C. Giant viruses: The difficult breaking of multiple epistemological barriers. Stud. Hist. Philos. Biol. Biomed. Sci. 2016, 59, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Hamacher, K.; Greiner, T.; Van Etten, J.L.; Gebhardt, M.; Cosentino, C.; Moroni, A.; Thiel, G. Phycodnavirus potassium ion channel proteins question the virus molecular piracy hypothesis. PLoS ONE 2012, 7, e38826. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G.; Moroni, A.; Blanc, G.; Van Etten, J.L. Potassium ion channels: Could they have evolved from viruses? Plant Physiol. 2013, 162, 1215–1224. [Google Scholar] [CrossRef] [PubMed]
- Lindell, D.; Sullivan, M.B.; Johnson, Z.I.; Tolonen, A.C.; Rohwer, F.; Chisholm, S.W. Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc. Natl. Acad. Sci. USA 2004, 101, 11013–11018. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 19 July 2018).
- Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins (accessed on 19 July 2018).
- Available online: http://www.cbs.dtu.dk/services/TMHMM/ (accessed on 19 July 2018).
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7. Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Gebhardt, M.; Hoffgaard, F.; Hamacher, K.; Kast, S.M.; Moroni, A.; Thiel, G. Membrane anchoring and interaction between transmembrane domains is crucial for K+ channel function. J. Biol. Chem. 2011, 286, 11299–11306. [Google Scholar] [CrossRef] [PubMed]
- Hille, B. Ion Channels of Excitable Membranes, 3rd ed.; Sinauer Associates Inc.: Sunderland, MA, USA, 2001. [Google Scholar]
- Quispe, C.F.; Esmael, A.; Sonderman, O.; McQuinn, M.; Agarkova, I.; Battah, M.; Duncan, G.A.; Dunigan, D.D.; Smith, T.P.L.; De Castro, C.; et al. Characterization of a new chlorovirus type with permissive and non-permissive features on phylogenetically related algal strains. Virology 2017, 500, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Schvarcz, C.R.; Steward, G.F. A giant virus infecting green algae encodes key fermentation genes. Virology 2018, 518, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Shealy, R.T.; Murphy, A.D.; Ramaratham, R.; Jakobsson, E.; Subramaniam, S. Sequence-function analysis of the K+ selective family of ion channels using a comprehensive alignment and the KcsA channel structure. Biophys. J. 2003, 84, 2929–2942. [Google Scholar] [CrossRef]
- Doyle, D.A.; Morais Cabral, J.; Pfuetzner, R.A.; Kuo, A.; Gulbis, J.M.; Cohen, S.L.; Chait, B.T.; MacKinnon, R. The structure of the potassium channel: Molecular basis of K+ conduction and selectivity. Science 1998, 280, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Available online: http://phagesdb.org/hosts/genera/1/?sequenced=True (accessed on 19 July 2018).
- Helaers, R.; Milinkovitch, M.C. MetaPIGA v2.0: Maximum likelihood large phylogeny estimation using the metapopulation genetic algorithm and other stochastic heuristics. BMC Bioinform. 2010, 11, 379. [Google Scholar] [CrossRef] [PubMed]
- Siotto, F.; Martin, C.; Rauh, O.; Van Etten, J.L.; Schroeder, I.; Moroni, A.; Thiel, G. Viruses infecting marine picoplankton encode functional potassium ion channels. Virology 2014, 466–467, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Heginbotham, L.; Lu, Z.; Abramson, T.; MacKinnon, R. Mutations in the K+ channel signature sequence. Biophys. J. 1994, 66, 1061–1067. [Google Scholar] [CrossRef]
- Duran, C.; Thompson, C.H.; Xiao, Q.; Hartzell, C. Chloride channels: Often enigmatic, rarely predictable. Annu. Rev. Physiol. 2010, 72, 95–121. [Google Scholar] [CrossRef] [PubMed]
- Tovar, K.R.; Westbrook, G.L. Ligand-gated ion channels. In Cell Physiology Source Book, 4th ed.; Elsevier: Amsterdam, The Netherlands, 2012; pp. 549–562. [Google Scholar]
- Tapken, D.; Anschütz, U.; Liu, L.-H.; Hülsken, T.; Seebohm, G.; Becker, D.; Hollmann, M. A plant homolog of animal glutamate receptors is an ion channel gated by multiple hydrophobic amino acids. Sci. Signal. 2013, 6, ra47. [Google Scholar] [CrossRef] [PubMed]
- Verkman, A.S.; Anderson, M.O.; Papadopulos, M.C. Aquaporins: Important but elusive drug targets. Nat. Rev. Drug Discov. 2014, 13, 259–277. [Google Scholar] [CrossRef] [PubMed]
- Martinac, B. Mechanosensitive ion channels: Molecules of mechanotransduction. J. Cell Sci. 2004, 117, 2449–2460. [Google Scholar] [CrossRef] [PubMed]
- Carrara, G.; Saraiva, N.; Parson, M.; Byrne, B.; Prole, D.; Taylor, C.W.; Smith, G.L. Golgi anti-apoptotic proteins are highly conserved ion channels that affect apoptosis and cell migration. J. Biol. Chem. 2015, 290, 11785–11801. [Google Scholar] [CrossRef] [PubMed]
- Saraiva, N.; Prole, D.L.; Carrara, G.; Maluquer de Motes, C.; Johnson, B.F.; Byrne, B.; Taylor, C.W.; Smith, G.L. Human and viral Golgi anti-apoptotic proteins (GAAPs) oligomerize via different mechanisms and monomeric GAAP inhibits apoptosis and modulates calcium. J. Biol. Chem. 2013, 288, 13057–13067. [Google Scholar] [CrossRef] [PubMed]
- Wilkens, S. Structure and mechanism of ABC transporters. F1000Prime Rep. 2015, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Schulz, F.; Yutin, N.; Ivanova, N.N.; Ortega, D.R.; Lee, T.K.; Vierheilig, J.; Daims, H.; Horn, M.; Wagner, M.; Jensen, G.J.; et al. Giant viruses with expanded complement of translation system components. Science 2017, 356, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Mitsuhashi, W.; Asano, S.; Miyamoto, K.; Wada, S. Further research on the biological function of inclusion bodies of Anomala cuprea entomopoxvirus, with special reference to the effect on the insecticidal activity of Bacillus thurgiensis formulation. Pest Manag. Sci. 2014, 70, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Hediger, M.A.; Romero, M.F.; Peng, J.-B.; Rolfs, A.; Takanaga, H.; Bruford, E.A. The ABC of solute carriers: Physiological, pathological and therapeutic implications of human membrane transport proteins. Pflügers Arch. Eur. J. Physiol. 2004, 447, 465–468. [Google Scholar] [CrossRef] [PubMed]
- Fan, W.; Guo, W.; Van Etten, J.L.; Mower, J.P. Multiple origins of endosymbionts in Chlorellaceae with no reductive effects on the plastide or mitochondrial genomes. Sci. Rep. 2017, 7, 10101. [Google Scholar] [CrossRef] [PubMed]
- Maguire, M.E. Magnesium transporters: Properties, regulation and structure. Front. Biosci. 2006, 11, 3149–3163. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.L.; Banks, J.L.; Snavely, M.D.; Maguire, M.E. Sequence and topology of the CorA magnesium transport systems of Salmonella typhimurium and Escherichia coli. Identification of a new class of transport protein. J. Biol. Chem. 1993, 268, 14071–14080. [Google Scholar] [PubMed]
- Monier, A.; Chambouvet, A.; Milner, D.S.; Attah, V.; Terrado, R.; Lovejoy, C.; Moreau, H.; Santoro, A.E.; Derelle, E.; Richards, T.A. Host-derived viral transporter protein for nitrogen uptake in infected marine phytoplankton. Proc. Natl. Acad. Sci. USA 2017, 114, E7489–E7498. [Google Scholar] [CrossRef] [PubMed]
- Jauniaux, J.-C.; Grenson, M. GAP1, the general amino acid permease gene of Saccharomyces cerevisiae. Eur. J. Biochem. 1990, 190, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Chatelain, F.C.; Gazzarrini, S.; Fujiwara, Y.; Arrigoni, C.; Domigan, C.; Ferrara, G.; Pantoja, C.; Thiel, G.; Moroni, A.; Minor, D.L. Selection of inhibitor-resistant viral potassium channels identifies a selectivity filter site that affects barium and amantadine block. PLoS ONE 2009, 4, e7496. [Google Scholar] [CrossRef] [PubMed]
- Madan, V.; Bartenschläger, R. Structural and functional properties of the hepatitis C virus p7 viroporin. Viruses 2015, 7, 4461–4481. [Google Scholar] [CrossRef] [PubMed]
- Bolduan, S.; Votteler, J.; Lodermeyer, V.; Greiner, T.; Koppensteiner, H.; Schindler, M.; Thiel, G.; Schubert, U. Ion channel activity of HIV-1 Vpu is dispensable for counteraction of CD317. Virology 2011, 416, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Hsu, K.; Seharaseyon, J.; Dong, P.; Bour, S.; Marbán, E. Mutual functional destruction of HIV-1 and host TASK-1 channel. Mol. Cell 2004, 23, 259–267. [Google Scholar] [CrossRef]
- Strebel, K. HIV-1 Vpu—An ion channel in search of a job. Biochim. Biophys. Acta 2014, 1838, 1074–1081. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G.; Moroni, A.; Dunigan, D.; Van Etten, J.L. Initial events associated with virus PBCV-1 infection of Chlorella NC64A. Prog. Bot. 2010, 71, 169–183. [Google Scholar] [CrossRef] [PubMed]
- Balss, J.; Mehmel, M.; Baumeister, D.; Hertel, B.; Delaroque, N.; Chatelain, F.C.; Minor, D.J.; Van Etten, J.L.; Moroni, A.; Thiel, G. Transmembrane domain length of viral K+ channels is a signal for mitochondria targeting. Proc. Natl. Acad. Sci. USA 2008, 105, 12313–12318. [Google Scholar] [CrossRef] [PubMed]
- Delaroque, N.; Wolf, S.; Müller, D.G.; Knippers, R. The brown alga virus EsV-1 particle contains a putative hybrid histidine kinase. Virology 2000, 273, 383–390. [Google Scholar] [CrossRef] [PubMed]
- Boulanger, P.; Lettelier, L. Characterisation of ion channels involved in the penetration of phage T4 DNA into Escherichia coli cells. J. Biol. Chem. 1988, 263, 9767–9775. [Google Scholar] [PubMed]
- Boulanger, P.; Lettelier, L. Ion channels are likely to be involved in the two steps of phage T5 DNA penetration into Escherichia coli cells. J. Biol. Chem. 1992, 267, 3168–3172. [Google Scholar] [PubMed]
- Van Etten, J.L.; Van Etten, C.H.; Johnson, J.K.; Burbank, D.E. A survey for viruses from fresh water that infect a eukaryotic Chlorella-like green alga. Appl. Environ. Microbiol. 1985, 49, 1326–1328. [Google Scholar] [PubMed]
- Zhang, Y.; Burbank, D.E.; Van Etten, J.L. Chlorella viruses isolated in China. Appl. Environ. Microbiol. 1988, 54, 2170–2173. [Google Scholar] [PubMed]





© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Greiner, T.; Moroni, A.; Van Etten, J.L.; Thiel, G. Genes for Membrane Transport Proteins: Not So Rare in Viruses. Viruses 2018, 10, 456. https://doi.org/10.3390/v10090456
Greiner T, Moroni A, Van Etten JL, Thiel G. Genes for Membrane Transport Proteins: Not So Rare in Viruses. Viruses. 2018; 10(9):456. https://doi.org/10.3390/v10090456
Chicago/Turabian StyleGreiner, Timo, Anna Moroni, James L Van Etten, and Gerhard Thiel. 2018. "Genes for Membrane Transport Proteins: Not So Rare in Viruses" Viruses 10, no. 9: 456. https://doi.org/10.3390/v10090456
APA StyleGreiner, T., Moroni, A., Van Etten, J. L., & Thiel, G. (2018). Genes for Membrane Transport Proteins: Not So Rare in Viruses. Viruses, 10(9), 456. https://doi.org/10.3390/v10090456





