New Adenovirus Groups in Western Palaearctic Bats
Abstract
1. Introduction
2. Materials and Methods
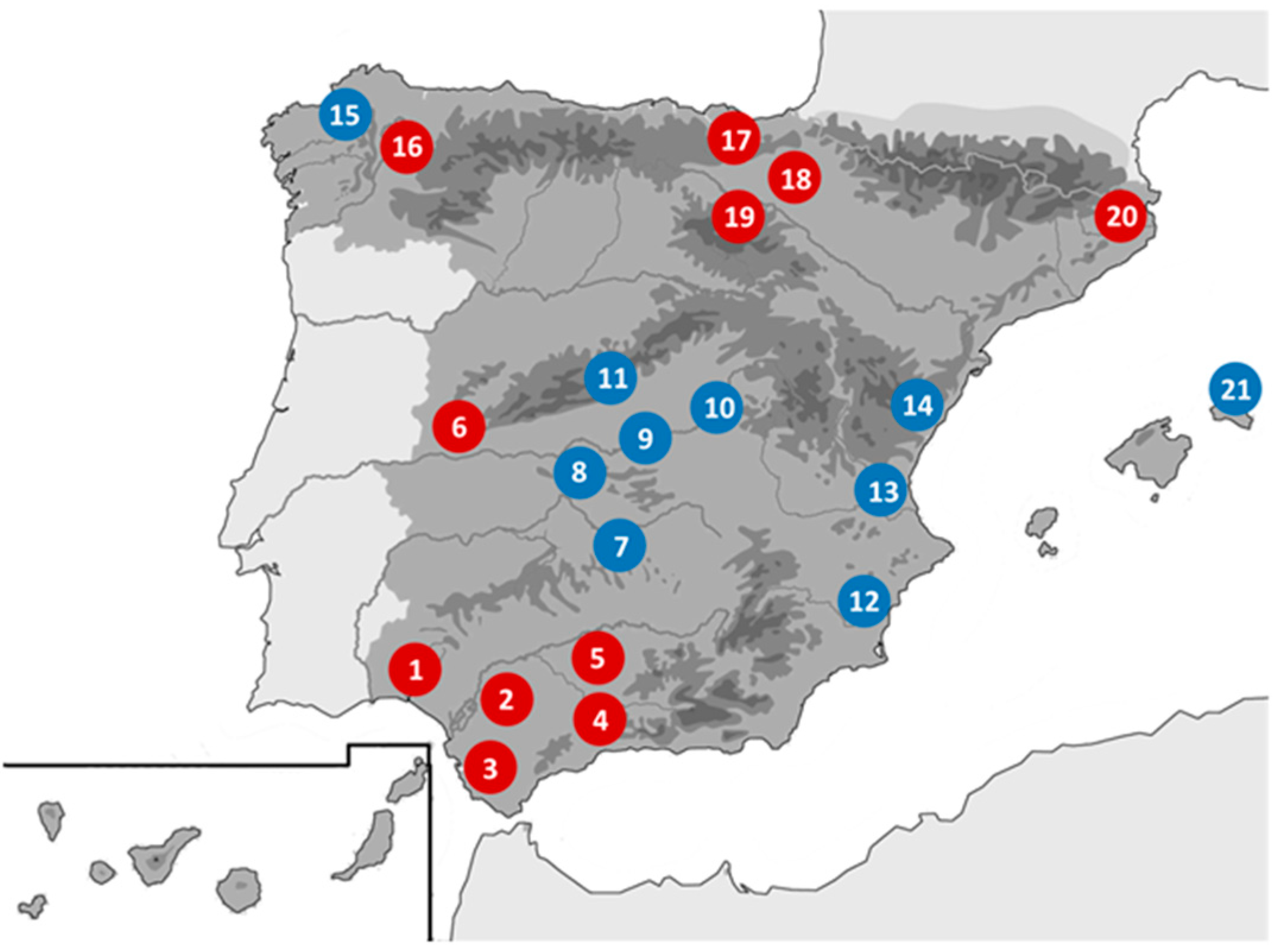
2.1. Origin of Samples and Preparation
2.2. Adenovirus Detection by Generic PCR Methods
2.3. Sequence and Phylogenetic Analysis
3. Results
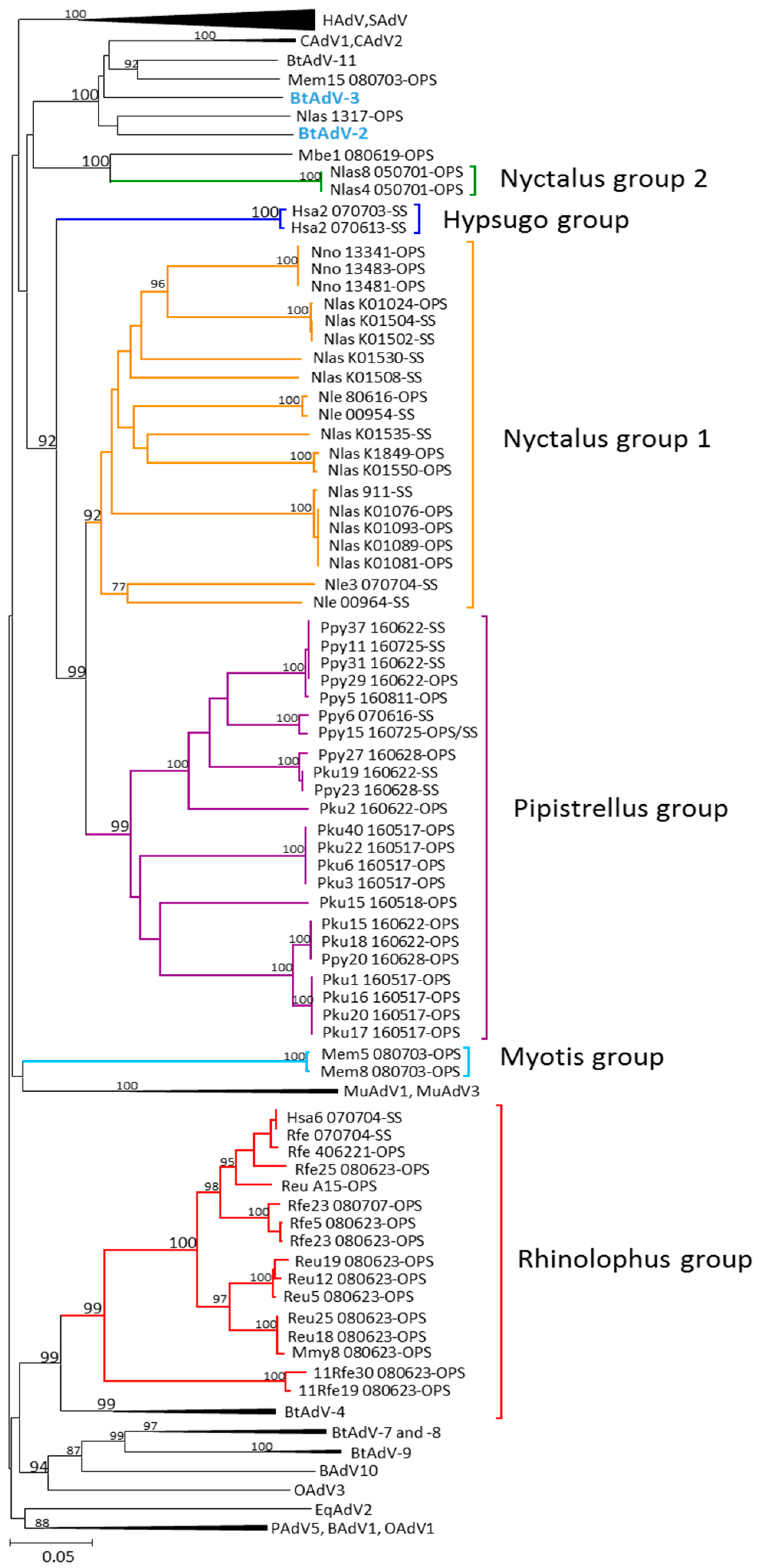
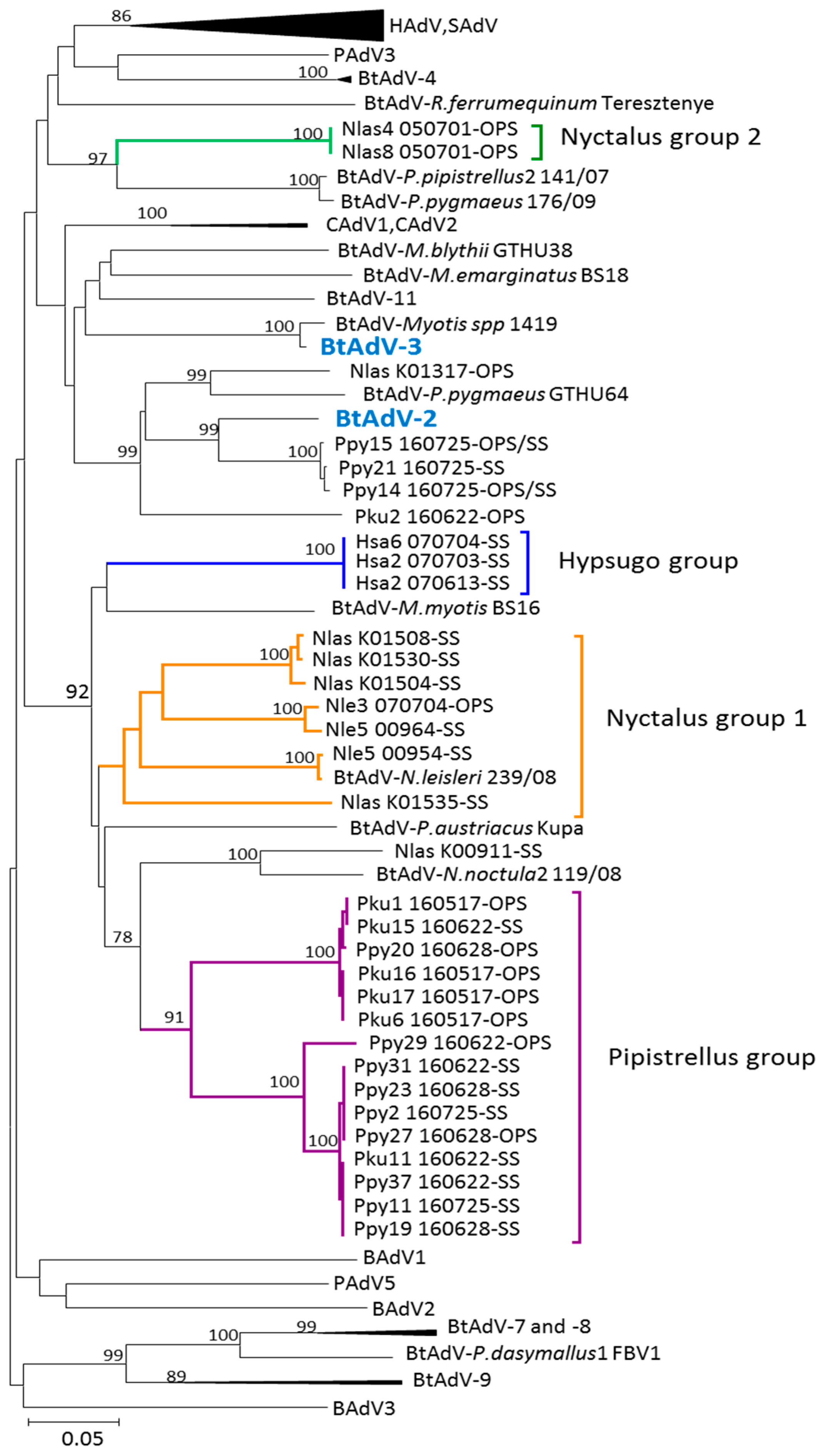
3.1. Phylogenetic Analysis of Bat AdV Sequences
3.2. Partial AdV Hexon Gene Sequence Analysis
3.3. Partial DNA-Dependent DNA Polymerase Gene Sequences
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- IUCN SSC Bat Specialist Group. Available online: http://www.iucnbsg.org/ (accessed on 14 May 2018).
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.; Lau, S.; Woo, P.; Yuen, K.Y. Bats as a continuing source of emerging infections in humans. Rev. Med. Virol. 2007. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Corman, V.M.; Wegner, T.; Tateno, A.F.; Zerbinati, R.M.; Gloza-Rausch, F.; Seebens, A.; Müller, M.A.; Drosten, C. Amplification of emerging viruses in a bat colony. Emerg. Infect. Dis. 2011, 17, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Jánoska, M.; Vidovszky, M.; Molnár, V.; Liptovszky, M.; Harrach, B.; Benkö, M. Novel adenoviruses and herpesviruses detected in bats. Vet. J. 2011, 189, 118–121. [Google Scholar] [CrossRef] [PubMed]
- Tan, B.; Yang, X.-L.; Ge, X.-Y.; Peng, C.; Zhang, Y.-Z.; Zhang, L.-B.; Shi, Z.-L. Novel bat adenoviruses with an extremely large E3 gene. J. Gen. Virol. 2016, 97, 1625–1635. [Google Scholar] [CrossRef] [PubMed]
- Waruhiu, C.; Ommeh, S.; Obanda, V.; Agwanda, B.; Gakuya, F.; Ge, X.Y.; Yang, X.L.; Wu, L.J.; Zohaib, A.; Hu, B.; et al. Molecular detection of viruses in Kenyan bats and discovery of novel astroviruses, caliciviruses and rotaviruses. Virol. Sin. 2017, 32, 101–114. [Google Scholar] [CrossRef] [PubMed]
- Harrach, B.; Benkö, M.; Both, G.; Brown, M.; Davison, A.J.; Echavarría, M.; Hess, M.; Jones, M.S.; Kajon, A.; Lehmkuhl, A.D.; et al. Family Adenoviridae. In Virus Taxonomy: Classification and Nomenclature of Viruses. Ninth report of the International Committee of Taxonomy of Viruses; Elsevier: San Diego, CA, USA, 2011; pp. 125–141. [Google Scholar]
- Maeda, K.; Hondo, E.; Terakawa, J.; Kiso, Y.; Nakaichi, N.; Endoh, D.; Sakai, K.; Morikawa, S.; Mizutani, T. Isolation of novel adenovirus from fruit bat (Pteropus dasymallus yayeyamae). Emerg. Infect. Dis. 2008, 14, 347–349. [Google Scholar] [CrossRef] [PubMed]
- Sonntag, M.; Mühldorfer, K.; Speck, S.; Wibbelt, G.; Kurth, A. New adenovirus in bats, Germany. Emerg. Infect. Dis. 2009, 15, 2052–2055. [Google Scholar] [CrossRef] [PubMed]
- Kohl, C.; Vidovszky, M.; Mühldorfer, K.; Dabrowski, P.W.; Radonic, A.; Nitsche, A.; Wibbelt, G.; Kurth, A.; Harrach, B. Genome analysis of bat adenovirus 2: Indications of interspecies transmission. J. Virol. 2012, 86, 1888–1892. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Ge, X.; Zhang, H.; Zhou, P.; Zhu, Y.; Zhang, Y.; Yuan, J.; Wang, L.F.; Shi, Z. Host range, prevalence, and genetic diversity of adenoviruses in bats. J. Virol. 2010, 84, 3889–3897. [Google Scholar] [CrossRef] [PubMed]
- Lima, F.E.; Cibulski, S.P.; Elesbao, F.; Carnieli Junior, P.; Batista, H.B.; Roehe, P.M.; Franco, A.C. First detection of adenovirus in the vampire bat (Desmodus rotundus) in Brazil. Virus Genes 2013, 47, 378–381. [Google Scholar] [CrossRef] [PubMed]
- Vidovszky, M.; Kohl, C.; Boldogh, S.; Görföl, T.; Wibbelt, G.; Kurth, A.; Harrach, B. Random sampling of the Central European bat fauna reveals the existence of numerous hitherto unknown adenoviruses. Acta Vet. Hung. 2015, 63, 508–525. [Google Scholar] [CrossRef] [PubMed]
- Tan, B.; Yang, X.L.; Ge, X.Y.; Peng, C.; Liu, H.Z.; Zhang, Y.Z.; Zhang, L.B.; Shi, Z.L. Novel bat adenoviruses with low G + C content shed new light on the evolution of adenoviruses. J. Gen. Virol. 2017, 98, 739–748. [Google Scholar] [CrossRef] [PubMed]
- Baker, K.S.; Leggett, R.M.; Bexfield, N.H.; Alston, M.; Daly, G.; Todd, S.; Tachedjian, M.; Holmes, C.E.G.; Crameri, S.; Wang, L.-F.; et al. Metagenomic study of the viruses of African straw-coloured fruit bats: Detection of a chiropteran poxvirus and isolation of a novel adenovirus. Virology 2013, 441, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, H.; Kajihara, M.; Nao, N.; Shigeno, A.; Fujikura, D.; Hang’ombe, B.M.; Mweene, A.S.; Mutemwa, A.; Squarre, D.; Yamada, M.; et al. Characterization of a Novel Bat Adenovirus Isolated from Straw-Colored Fruit Bat (Eidolon helvum). Viruses 2017, 9, 371. [Google Scholar] [CrossRef] [PubMed]
- Van Vuren, P.J.; Allam, M.; Wiley, M.R.; Ismail, A.; Storm, N.; Birkhead, M.; Markotter, W.; Palacios, G.; Paweska, J.T. A novel adenovirus isolated from the Egyptian fruit bat in South Africa is closely related to recent isolates from China. Sci. Rep. 2018, 8, 9584. [Google Scholar] [CrossRef] [PubMed]
- Hackenbrack, N.; Rogers, M.B.; Ashley, R.E.; Keel, M.K.; Kubiski, S.V.; Bryan, J.A.; Ghedin, E.; Holmes, E.C.; Hafenstein, S.L.; Allison, A.B. Evolution and Cryo-electron Microscopy Capsid Structure of a North American Bat Adenovirus and Its Relationship to Other Mastadenoviruses. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Mingo-Casas, P.; Sandonís, V.; Vázquez-Morón, S.; Berciano, J.M.; Juste, J.; Echevarría, J.E. Rabies in Spain. A Peculiarity in Eurasia. Ann. Virol. Res. 2017, 3, 1030. [Google Scholar]
- Juste, J.; Bilgin, R.; Muñoz, J.; Ibáñez, C. Mitochondrial DNA signatures at different spatial scales: From the effects of the Straits of Gibraltar to population structure in the meridional serotine bat (Eptesicus isabellinus). Heredity 2009, 103, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Morón, S.; Juste, J.; Ibáñez, C.; Berciano, J.M.; Echevarría, J.E. Phylogeny of European Bat Lyssavirus 1 in Eptesicus isabellinus Bats, Spain. Emerg. Infect. Dis. 2011, 17, 520–523. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Morón, S.; Juste, J.; Ibáñez, C.; Ruiz-Villamor, E.; Avellón, A.; Vera, M.; Echevarría, J.E. Endemic Circulation of European Bat Lyssavirus Type 1 in Serotine Bats, Spain. Emerg. Infect. Dis. 2008, 14, 1263–1266. [Google Scholar] [CrossRef] [PubMed]
- Ceballos, N.A.; Morón, S.V.; Berciano, J.M.; Nicolás, O.; López, C.A.; Juste, J.; Nevado, C.R.; Setién, Á.A.; Echevarría, J.E. Novel Lyssavirus in Bat, Spain. Emerg. Infect. Dis. 2013, 19, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Negredo, A.; Palacios, G.; Vázquez-Morón, S.; González, F.; Dopazo, H.; Molero, F.; Juste, J.; Quetglas, J.; Savji, N.; de la Cruz Martínez, M.; et al. Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog. 2011, 7, e1002304. [Google Scholar] [CrossRef] [PubMed]
- Falcón, A.; Vázquez-Morón, S.; Casas, I.; Aznar, C.; Ruiz, G.; Pozo, F.; Perez-Breña, P.; Juste, J.; Ibáñez, C.; Garin, I.; et al. Detection of alpha and betacoronaviruses in multiple Iberian bat species. Arch. Virol. 2011, 156, 1883–1890. [Google Scholar] [CrossRef] [PubMed]
- Pozo, F.; Juste, J.; Vázquez-Morón, S.; Aznar-López, C.; Ibáñez, C.; Garin, I.; Aihartza, J.; Casas, I.; Tenorio, A.; Echevarría, J.E. Identification of Novel Betaherpesviruses in Iberian Bats Reveals Parallel Evolution. PLoS ONE 2016, 11, e0169153. [Google Scholar] [CrossRef] [PubMed]
- Aznar-Lopez, C.; Vazquez-Moron, S.; Marston, D.A.; Juste, J.; Ibanez, C.; Berciano, J.M.; Salsamendi, E.; Aihartza, J.; Banyard, A.C.; McElhinney, L.; et al. Detection of rhabdovirus viral RNA in oropharyngeal swabs and ectoparasites of Spanish bats. J. Gen. Virol. 2013, 94, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Casas, I.; Powell, L.; Klapper, P.E.; Cleator, G.M. New method for the extraction of viral RNA and DNA from cerebrospinal fluid for use in the polymerase chain reaction assay. J. Virol. Methods 1995, 53, 25–36. [Google Scholar] [CrossRef]
- Calvo, C.; García-García, M.L.; Sanchez-Dehesa, R.; Román, C.; Tabares, A.; Pozo, F.; Casas, I. Eight Year Prospective Study of Adenoviruses Infections in Hospitalized Children. Comparison with Other Respiratory Viruses. PLoS ONE 2015, 10, e0132162. [Google Scholar] [CrossRef] [PubMed]
- Casas, I.; Avellon, A.; Mosquera, M.; Jabado, O.; Echevarria, J.E.; Campos, R.H.; Rewers, M.; Perez-Breña, P.; Lipkin, W.I.; Palacios, G. Molecular Identification of Adenoviruses in Clinical Samples by Analyzing a Partial Hexon Genomic Region. J. Clin. Microbiol. 2005, 43, 6176–6182. [Google Scholar] [CrossRef] [PubMed]
- Posada, D.; Crandall, K.A. MODELTEST: Testing the model of DNA substitution. Bioinformatics 1998, 14, 817–818. [Google Scholar] [CrossRef] [PubMed]
- García-Mudarra, J.L.; Ibáñez, C.; Juste, J. The Straits of Gibraltar: Barrier or bridge to Ibero-Moroccan bat diversity? Biol. J. Linn. Soc. 2009, 96, 434–450. [Google Scholar] [CrossRef]
- Zhang, Y.; Bergelson, J.M. Adenovirus Receptors. J. Virol. 2005, 79, 12125–12131. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H. Viral Polymerases. Adv. Exp. Med. Biol. 2012, 726, 267–304. [Google Scholar] [CrossRef] [PubMed]
- Roberts, D.M.; Nanda, A.; Havenga, M.J.; Abbink, P.; Lynch, D.M.; Ewald, B.A.; Liu, J.; Thorner, A.R.; Swanson, P.E.; Gorgone, D.A.; et al. Hexon-chimaeric adenovirus serotype 5 vectors circumvent pre-existing anti-vector immunity. Nature 2006, 441, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Rux, J.J.; Kuser, P.R.; Burnett, R.M. Structural and phylogenetic analysis of adenovirus hexons by use of high-resolution X-ray crystallographic, molecular modeling, and sequence-based methods. J. Virol. 2003, 77, 9553–9566. [Google Scholar] [CrossRef] [PubMed]
- Kajon, A.E.; Dickson, L.M.; Murtagh, P.; Viale, D.; Carballal, G.; Echavarria, M. Molecular Characterization of an Adenovirus 3–16 Intertypic Recombinant Isolated in Argentina from an Infant Hospitalized with Acute Respiratory Infection. J. Clin. Microbiol. 2010, 48, 1494–1496. [Google Scholar] [CrossRef] [PubMed]
- Arroyo-Cabrales, J.; Álvarez-Castañeda, S.T. Corynorhinus rafinesquii. The IUCN Red List of Threatened Species 2017; International Union for Conservation of Nature: Gland, Switzerland, 2017. [Google Scholar]
- Stadelmann, B.; Lin, L.-K.; Kunz, T.H.; Ruedi, M. Molecular phylogeny of New World Myotis (Chiroptera, Vespertilionidae) inferred from mitochondrial and nuclear DNA genes. Mol. Phylogenet. Evol. 2007, 43, 32–48. [Google Scholar] [CrossRef] [PubMed]
| Iberian Bat Species | Abb 1 | OPS 2 | SS 3 | Year 4 | Hexon Sequences 5 | DNA-Pol Sequences 6 | |
|---|---|---|---|---|---|---|---|
| Family | Name | ||||||
| Vespertilionidae | Barbastella barbastellus | Bba | 0/38 | 0/4 | 07,08 | N/A | N/A |
| Eptesicus isabellinus | Eis | 0 | 0/8 | 04,07 | N/A | N/A | |
| Eptesicus serotinus | Ese | 0 | 0/14 | 03,07 | N/A | N/A | |
| Hypsugo savii | Hsa | 0/31 | 3/26 | 07 | HM856338,41,42 | JX065121, 22, MG208122 | |
| Myotis alcathoe | Mal | 0 | 0/1 | 07 | N/A | N/A | |
| Myotis bechsteinii | Mbe | 1/18 | 0/2 | 07 | MF540611 | N/A | |
| Myotis blythii | Mbl | 0/29 | 0 | 04 | N/A | N/A | |
| Myotis capaccinii | Mca | 0/15 | 0 | 04,07 | N/A | N/A | |
| Myotis daubentonii | Mda | 0/63 | 0/41 | 04,07 | N/A | N/A | |
| Myotis emarginatus | Mem | 3/56 | 0 | 08 | MF540608-10 | N/A | |
| Myotis escalerai | Mes | 0/13 | 0 | 04,07 | N/A | N/A | |
| Myotis myotis | Mmy | 1/79 | 0/1 | 04,07 | HM856353 | N/A | |
| Myotis mystacinus | Mmt | 0/2 | 0/8 | 07 | N/A | N/A | |
| Myotis nattereri | Mna | 0/36 | 0/3 | 07 | N/A | N/A | |
| Nyctalus noctula | Nno | 3/122 | 0 | 07 | MF540597-99 | N/A | |
| Nyctalus lasiopterus | Nlas | 10/139 | 6/40 | 07 | HM856327-34,39-40,43,45-47, 50, MG132211 | JX065117-20, 23,25-26,28 | |
| Nyctalus leisleri | Nle | 1/19 | 3/26 | 07 | HM856344,48, 51-52 | JX065124,27,29 | |
| Pipistrellus kuhlii | Pku | 12/350 | 2/4 | 07,16 | MF540577-85,87,89 | MF404970-73,75,86 | |
| Pipistrellus pipistrellus | Ppi | 0/29 | 0/4 | 07,16 | HM856349 | N/A | |
| Pipistrellus pygmaeus | Ppy | 6/36 | 11/120 | 07,16 | MF540575-76, 86,88,90-96 | MF404968-69,74, 76-79, 80-85,87-89 | |
| Plecotus auritus | Pau | 0/11 | 0/8 | 04,07 | N/A | N/A | |
| Plecotus austriacus | Pas | 0/10 | 0/6 | 04,07 | N/A | N/A | |
| Miniopteridae | Miniopterus schreibersii | Msc | 0/152 | 0/2 | 04,07, 16 | N/A | N/A |
| Rhinolophidae | Rhinolophus euryale | Reu | 6/49 | 0 | 04,07, 08 | MF540600-02,12-13 HM856335 | N/A |
| Rhinolophus ferrumequinum | Rfe | 7/90 | 1/3 | 04,07 | MF540603-07,14 HM856336-37 | N/A | |
| Rhinolophus hipposideros | Rhi | 0/4 | 0/4 | 07,08 | N/A | N/A | |
| Rhinolophus mehelyi | Rme | 0/1 | 0 | 07,08 | N/A | N/A | |
| Total | 27/32 | 50/1392 | 26/325 | 69 49OPS + 20SS | 35 14OPS + 21SS | ||
| Group of Bat AdV | Tentative Virus Name | Abb Name | % aa Pairwise Distances 1 | |
|---|---|---|---|---|
| BtAdV-3 | BtAdV-2 | |||
| bat AdVs associated with BtAdV-2 | Bat mastadenovirus P. pygmaeus 14 160725 | Ppy14 160725 | 31.4 | 11.8 |
| Bat mastadenovirus P. pygmaeus 15 160725 | Ppy15 160725 | 31.4 | 12.7 | |
| Bat mastadenovirus P. pygmaeus 21 160725 | Ppy21 160725 | 32 | 12.2 | |
| Potentially novel bat AdVs | Bat mastadenovirus N. lasiopterus K01317 | Nlas K01317 | 33.8 | 25.5 |
| Bat mastadenovirus N. lasiopterus K01508 | Nlas K01508 | 35 | 41.8 | |
| Bat mastadenovirus N. lasiopterus K01530 | Nlas K01530 | 35.7 | 42.5 | |
| Bat mastadenovirus N. lasiopterus K01504 | Nlas K01504 | 35.7 | 42.5 | |
| Bat mastadenovirus N. leisleri 3 070704 | Nle3 070704 | 32.5 | 42.5 | |
| Bat mastadenovirus N. leisleri 00964 | Nle 00964 | 34.5 | 44 | |
| Bat mastadenovirus N. leisleri 5 00954 | Nle5 00954 | 39.7 | 41.8 | |
| Bat mastadenovirus N. lasiopterus K01535 | Nlas K01535 | 41.8 | 52.1 | |
| Bat mastadenovirus N. lasiopterus K00911 | Nlas K00911 | 43.5 | 46.1 | |
| Bat mastadenovirus N. lasiopterus 4 050701 | Nlas4 050701 | 42.6 | 38.8 | |
| Bat mastadenovirus N. lasiopterus 8 050701 | Nlas8 050701 | 42.6 | 38.8 | |
| Bat mastadenovirus P. kuhlii 2 160622 | Pku2 160622 | 37.5 | 23.8 | |
| Bat mastadenovirus P. kuhlii 1 160517 | Pku1 160517 | 39.4 | 39 | |
| Bat mastadenovirus P. kuhlii 15 160622 | Pku15 160622 | 39.4 | 39 | |
| Bat mastadenovirus P. pygmaeus 20 160628 | Ppy20 160628 | 38.7 | 39 | |
| Bat mastadenovirus P. kuhlii 16 160517 | Pku16 160517 | 38.7 | 38.3 | |
| Bat mastadenovirus P. kuhlii 17 160517 | Pku17 160517 | 38.7 | 38.3 | |
| Bat mastadenovirus P. kuhlii 6 160517 | Pku6 160517 | 38.7 | 38.3 | |
| Bat mastadenovirus P. pygmaeus 29 160622 | Ppy29 160622 | 44.8 | 36.5 | |
| Bat mastadenovirus P. pygmaeus 31 160622 | Ppy31 160622 | 41.2 | 41.6 | |
| Bat mastadenovirus P. pygmaeus 23 160628 | Ppy23 160628 | 41.2 | 41.6 | |
| Bat mastadenovirus P. pygmaeus 2 160725 | Ppy2 160725 | 41.2 | 41.6 | |
| Bat mastadenovirus P. pygmaeus 27 160628 | Ppy27 160628 | 41.2 | 41.6 | |
| Bat mastadenovirus P. kuhlii 11 160622 | Pku11 160622 | 42 | 40.9 | |
| Bat mastadenovirus P. pygmaeus 37 160622 | Ppy37 160622 | 42 | 40.9 | |
| Bat mastadenovirus P. pygmaeus 11 160725 | Ppy11 160725 | 42 | 40.9 | |
| Bat mastadenovirus P. pygmaeus 19 160628 | Ppy19 160628 | 42 | 40.9 | |
| Bat mastadenovirus H. savii 6 070704 | Hsa6 070704 | 41 | 41.7 | |
| Bat mastadenovirus H. savii 2 070613 | Hsa2 070613 | 41 | 41.7 | |
| Bat mastadenovirus H. savii 2 070703 | Hsa2 070703 | 41 | 41.7 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iglesias-Caballero, M.; Juste, J.; Vázquez-Morón, S.; Falcon, A.; Aznar-Lopez, C.; Ibáñez, C.; Pozo, F.; Ruiz, G.; Berciano, J.M.; Garin, I.; et al. New Adenovirus Groups in Western Palaearctic Bats. Viruses 2018, 10, 443. https://doi.org/10.3390/v10080443
Iglesias-Caballero M, Juste J, Vázquez-Morón S, Falcon A, Aznar-Lopez C, Ibáñez C, Pozo F, Ruiz G, Berciano JM, Garin I, et al. New Adenovirus Groups in Western Palaearctic Bats. Viruses. 2018; 10(8):443. https://doi.org/10.3390/v10080443
Chicago/Turabian StyleIglesias-Caballero, Maria, Javier Juste, Sonia Vázquez-Morón, Ana Falcon, Carolina Aznar-Lopez, Carlos Ibáñez, Francisco Pozo, Guillermo Ruiz, Jose M. Berciano, Inazio Garin, and et al. 2018. "New Adenovirus Groups in Western Palaearctic Bats" Viruses 10, no. 8: 443. https://doi.org/10.3390/v10080443
APA StyleIglesias-Caballero, M., Juste, J., Vázquez-Morón, S., Falcon, A., Aznar-Lopez, C., Ibáñez, C., Pozo, F., Ruiz, G., Berciano, J. M., Garin, I., Aihartza, J., Echevarría, J. E., & Casas, I. (2018). New Adenovirus Groups in Western Palaearctic Bats. Viruses, 10(8), 443. https://doi.org/10.3390/v10080443






