Molecular and Biological Characterisation of Turnip mosaic virus Isolates Infecting Poppy (Papaver somniferum and P. rhoeas) in Slovakia
Abstract
1. Introduction
2. Material and Methods
2.1. Samples
2.2. Analysis of the Virome and Determination of the TuMV Full Length Genome Sequences by HTS
2.3. Inoculation of Oilseed Poppy and Experimental Plants
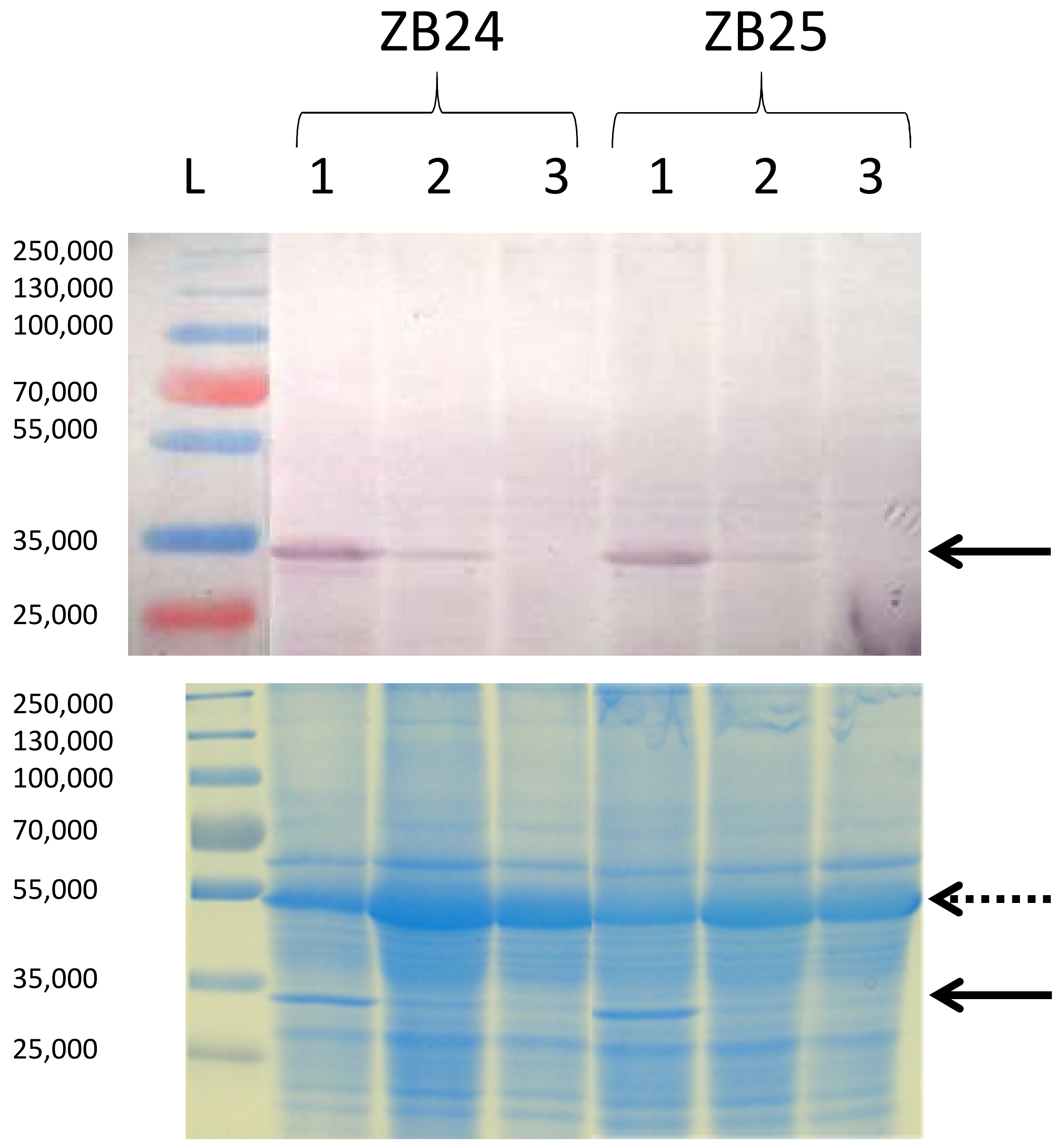
2.4. Estimation of Viral Accumulation in Plants
3. Results and Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Baroš, S.; Karšayová, B.; Jomová, K.; Gáspár, A.; Valko, M. Free radical scavenging capacity of Papaver somniferum L. and determination of pharmacologically active alkaloids using capillary electrophoresis. J. Microbiol. Biotechnol. Food Sci. 2012, 1, 725–732. [Google Scholar]
- Lančaričová, A.; Havrlentová, M.; Muchová, D.; Bednárová, A. Oil content and fatty acids composition of poppy seeds cultivated in two localities of Slovakia. Agriculture (Polnohospodárstvo) 2016, 62, 19–27. [Google Scholar] [CrossRef]
- Krošlák, E.; Maliar, T.; Nemeček, P.; Viskupičová, J.; Maliarová, M.; Havrlentová, M.; Kraic, J. Antioxidant and proteinase inhibitory activities of selected poppy (Papaver somniferum L.) genotypes. Chem. Biodivers. 2017, 14, e1700176. [Google Scholar] [CrossRef] [PubMed]
- Lutman, P.J.W.; Cussans, G.W.; Wright, K.J.; Wilson, B.J.; Lawson, H.M. The persistence of seeds of 16 weed species over six years in two arable fields. Weed Res. 2002, 42, 231–241. [Google Scholar] [CrossRef]
- Brunt, A.A.; Crabtree, K.; Dallwitz, M.J.; Gibbs, A.J.; Watson, L. Turnip Mosaic Potyvirus. Viruses of Plants; CABI: Wallingford, UK, 1996; pp. 1340–1343. [Google Scholar]
- Kubelková, D.; Špak, J. Virus diseases of poppy (Papaver somniferum L.) and some other species of the Papaveraceae family. Plant Prot. Sci. 1999, 35, 33–36. [Google Scholar] [CrossRef]
- Tang, J.; Lebas, B.; Liefting, L.; Veerakone, S.; Wei, T.; Ward, L. Opium poppy mosaic virus, a new umbravirus isolated from Papaver somniferum in New Zealand. Arch. Virol. 2016, 161, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Kumar, S.; Jaidi, M.; Raj, S.K.; Shukla, S.K. First Report of Tomato leaf curl New Delhi virus on Opium Poppy (Papaver somniferum) in India. Plant Dis. 2016, 100, 232. [Google Scholar] [CrossRef]
- Jenner, C.E.; Walsh, J.A. Pathotypic variation in Turnip mosaic virus with special reference to European isolates. Plant Pathol. 1996, 45, 848–856. [Google Scholar] [CrossRef]
- Walsh, J.A.; Jenner, C.E. Turnip mosaic virus and the quest for durable resistance. Mol. Plant Pathol. 2002, 3, 289–300. [Google Scholar] [CrossRef] [PubMed]
- Ohshima, K.; Tanaka, M.; Sako, N. The complete nucleotide sequence of Turnip mosaic virus RNA Japanese strain. Arch. Virol. 1996, 141, 1991–1997. [Google Scholar] [CrossRef] [PubMed]
- Chung, B.Y.; Miller, W.A.; Atkins, J.F.; Firth, A.E. An overlapping essential gene in the Potyviridae. Proc. Natl. Acad. Sci. USA 2008, 105, 5897–5902. [Google Scholar] [CrossRef] [PubMed]
- White, K.A. The polymerase slips and PIPO exists. EMBO Rep. 2015, 16, 885–886. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.D.; Tomitaka, Y.; Ho, S.Y.W.; Duchêne, S.; Vetten, H.-J.; Lesemann, D.; Walsh, J.A.; Gibbs, A.J.; Ohshima, K. Turnip mosaic potyvirus probably first spread to Eurasian brassica crops from wild orchids about 1000 years ago. PLoS ONE 2013, 8, e55336. [Google Scholar] [CrossRef] [PubMed]
- Yasaka, R.; Fukagawa, H.; Ikematsu, M.; Soda, H.; Korkmaz, S.; Golnaraghi, A.; Katis, N.; Ho, S.Y.W.; Gibbs, A.J.; Ohshima, K. The Timescale of Emergence and Spread of Turnip Mosaic Potyvirus. Sci. Rep. 2017, 7, 4240. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef] [PubMed]
- Glasa, M.; Labonne, G.; Quiot, J.-B. Effect of temperature on Plum pox virus infection. Acta Virol. 2003, 47, 49–52. [Google Scholar] [PubMed]
- Šubr, Z.; Glasa, M. Plum pox virus capsid protein mobility in SDS-polyacrylamide gel electrophoresis. Acta Virol. 1999, 43, 259–262. [Google Scholar] [PubMed]
- Adams, M.J.; Antoniw, J.F.; Fauquet, C.M. Molecular criteria for genus and species discrimination within the family Potyviridae. Arch. Virol. 2005, 150, 459–479. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Sun, Y.; Wang, Y.; Pan, H.; Wang, F.; Zhang, X.; Zhang, Y.; Liu, J. Molecular Characterization of the Complete Genome of Three Basal-BR Isolates of Turnip mosaic virus Infecting Raphanus sativus in China. Int. J. Mol. Sci. 2016, 17, 888. [Google Scholar] [CrossRef] [PubMed]
- Ohshima, K.; Yamaguchi, Y.; Hirota, R.; Hamamoto, T.; Tomimura, K.; Tan, Z.; Sano, T.; Azuhata, F.; Walsh, J.A.; Fletcher, J.; et al. Molecular evolution of Turnip mosaic virus: Evidence of host adaptation; genetic recombination and geographical spread. J. Gen. Virol. 2002, 83, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Ohshima, K.; Tomitaka, Y.; Wood, J.T.; Minematsu, Y.; Kajiyama, H.; Tomimura, K.; Gibbs, A.J. Patterns of recombination in Turnip mosaic virus genomic sequences indicate hotspots of recombination. J. Gen. Virol. 2007, 88, 298–315. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.D.; Tran, H.T.N.; Ohshima, K. Genetic variation of the Turnip mosaic virus population of Vietnam: A case study of founder; regional and local influences. Virus Res. 2013, 171, 138–149. [Google Scholar] [CrossRef] [PubMed]
- Tomimura, K.; Gibbs, A.J.; Jenner, C.E.; Walsh, J.A.; Ohshima, K. The phylogeny of Turnip mosaic virus; comparisons of 38 genomic sequences reveal a Eurasian origin and a recent ‘emergence’ in East Asia. Mol. Ecol. 2003, 12, 2099–2111. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Liu, J.L.; Gao, R.; Chen, J.; Shao, Y.H.; Li, X.D. Complete genomic sequence analyses of Turnip mosaic virus basal-BR isolates from China. Virus Genes 2009, 38, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Suehiro, N.; Natsuaki, T.; Watanabe, T.; Okuda, S. An important determinant of the ability of Turnip mosaic virus to infect Brassica spp. and/or Raphanus sativus is in its P3 protein. J. Gen. Virol. 2004, 85, 2087–2098. [Google Scholar] [CrossRef] [PubMed]
- Coutts, B.A.; Walsh, J.A.; Jones, R.A.C. Evaluation of resistance to Turnip mosaic virus in Australian Brassica napus genotypes. Aust. J. Agric. Res. 2007, 58, 67–74. [Google Scholar] [CrossRef]
- Kehoe, M.A.; Coutts, B.A.; Jones, R.A.C. Resistance phenotypes in diverse accessions; breeding lines; and cultivars of three mustard species inoculated with Turnip mosaic virus. Plant Dis. 2010, 94, 1290–1298. [Google Scholar] [CrossRef]
- Nyalugwe, E.P.; Barbetti, M.J.; Jones, R.A.C. Preliminary studies on resistance phenotypes to Turnip mosaic virus in Brassica napus and B. carinata from different continents and effects of temperature on their expression. Eur. J. Plant Pathol. 2014, 139, 687–706. [Google Scholar] [CrossRef]
- Guerret, M.G.L.; Nyalugwe, E.P.; Maina, S.; Barbetti, M.J.; van Leur, J.A.G.; Jones, R.A.C. Properties of a Turnip mosaic virus (TuMV) strain that breaks TuMV resistances in Brassica napus. Plant Dis. 2017, 101, 674–683. [Google Scholar] [CrossRef]
- Domingo, E.; Sheldon, J.; Perales, C. Viral quasispecies evolution. Microbiol. Mol. Biol. Rev. 2012, 76, 159–216. [Google Scholar] [CrossRef] [PubMed]
- Garcia, J.A.; Pallas, V. Viral factors involved in plant pathogenesis. Curr. Opin. Virol. 2015, 11, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Rodrigo, G.; Zwart, M.P.; Elena, S.F. Onset of virus systemic infection in plants is determined by speed of cell-to-cell movement and number of primary infection foci. J. R. Soc. Interface 2014, 11. [Google Scholar] [CrossRef] [PubMed]


| Papaver somniferum ZB24 | Papaver somniferum ZB25 | N. benthamianaa | R. sativus cv. Slovana a | R. sativus cv. Kulatá černá a | |||
|---|---|---|---|---|---|---|---|
| Isolate | Experiment 1 | Experiment 2 | Experiment 1 | Experiment 2 | |||
| HC9 | 12/7 b (2 m + 5 s) | 12/5 (2 m + 3 s) m = 0.552 ± 0.045 *s = 1.711 ± 0.129 | 10/4 (1 m + 3 s) | 12/6 (2 m + 4 s) m = 0.333 ± 0.049 s = 0.799 ± 0.184 | 10/10 | 20/0 | 20/0 |
| PK1 | 12/8 (4 m + 4 s) | 11/6 (4 m + 2 s) m = 0.652 ± 0.034 s = 1.589 ± 0.108 | 12/7 (3 m + 4 s) | 11/6 (3 m + 3 s) m = 0.417 ± 0.057 s = 0.983 ± 0.139 | 10/10 | 20/1 | 20/0 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Glasa, M.; Šoltys, K.; Predajňa, L.; Sihelská, N.; Nováková, S.; Šubr, Z.; Kraic, J.; Mihálik, D. Molecular and Biological Characterisation of Turnip mosaic virus Isolates Infecting Poppy (Papaver somniferum and P. rhoeas) in Slovakia. Viruses 2018, 10, 430. https://doi.org/10.3390/v10080430
Glasa M, Šoltys K, Predajňa L, Sihelská N, Nováková S, Šubr Z, Kraic J, Mihálik D. Molecular and Biological Characterisation of Turnip mosaic virus Isolates Infecting Poppy (Papaver somniferum and P. rhoeas) in Slovakia. Viruses. 2018; 10(8):430. https://doi.org/10.3390/v10080430
Chicago/Turabian StyleGlasa, Miroslav, Katarína Šoltys, Lukáš Predajňa, Nina Sihelská, Slavomíra Nováková, Zdeno Šubr, Ján Kraic, and Daniel Mihálik. 2018. "Molecular and Biological Characterisation of Turnip mosaic virus Isolates Infecting Poppy (Papaver somniferum and P. rhoeas) in Slovakia" Viruses 10, no. 8: 430. https://doi.org/10.3390/v10080430
APA StyleGlasa, M., Šoltys, K., Predajňa, L., Sihelská, N., Nováková, S., Šubr, Z., Kraic, J., & Mihálik, D. (2018). Molecular and Biological Characterisation of Turnip mosaic virus Isolates Infecting Poppy (Papaver somniferum and P. rhoeas) in Slovakia. Viruses, 10(8), 430. https://doi.org/10.3390/v10080430





