Contemporary Zika Virus Isolates Induce More dsRNA and Produce More Negative-Strand Intermediate in Human Astrocytoma Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Phylogenetic Analysis
2.2. Cells and Viruses
2.3. ZIKV Infections
2.4. Cell Viability
2.5. Flow Cytometry
2.6. Immunofluorescence Microscopy
2.7. Quantitative Reverse-Transcription PCR
2.8. Statistical Analysis
3. Results
3.1. Phylogenetic and Amino Acid Variance across ZIKV Isolates Selected for Comparative Analyses
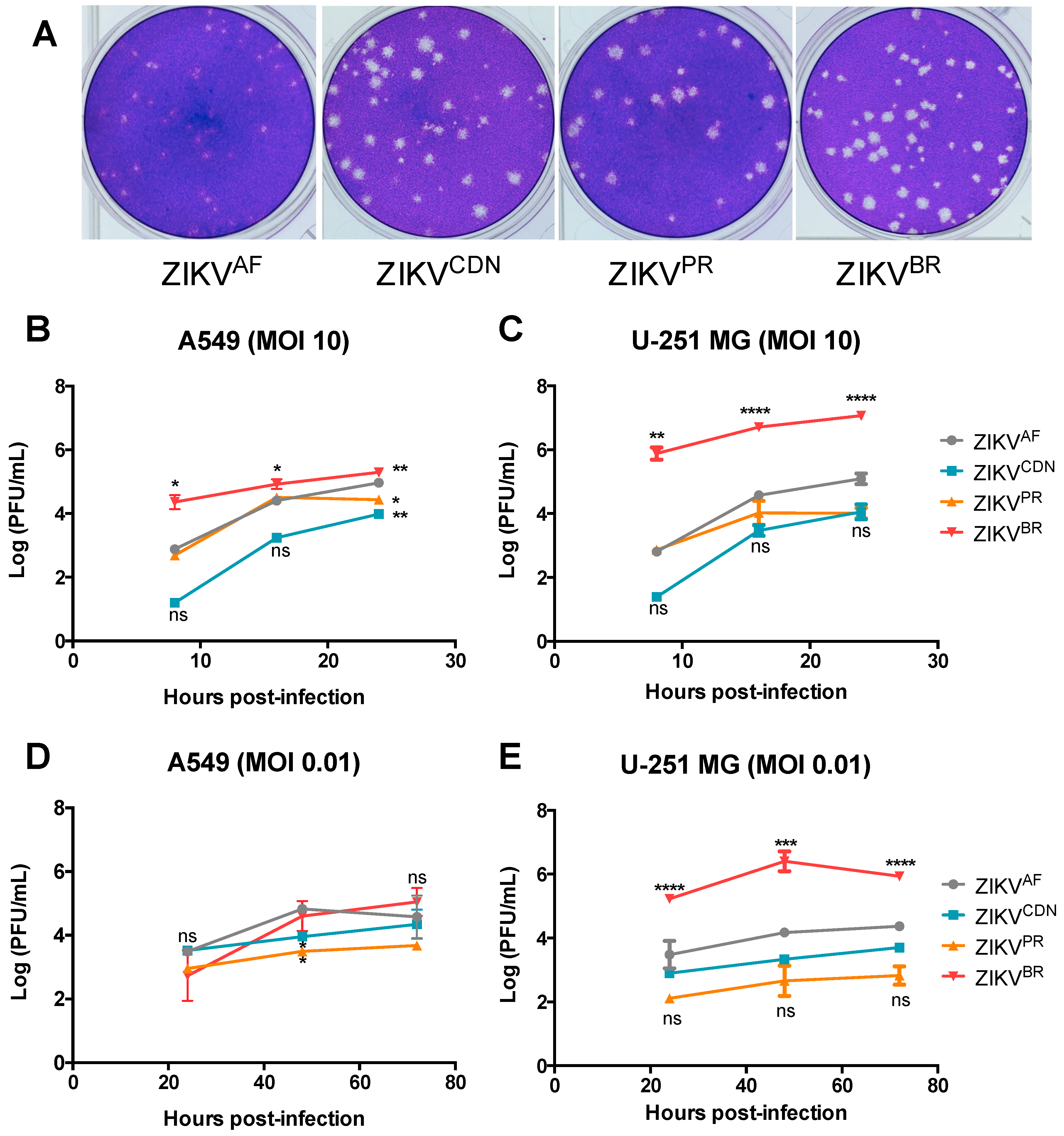
3.2. Zika Virus Isolates Display Unique Plaque Morphology and Different Growth Kinetics in Cell Culture
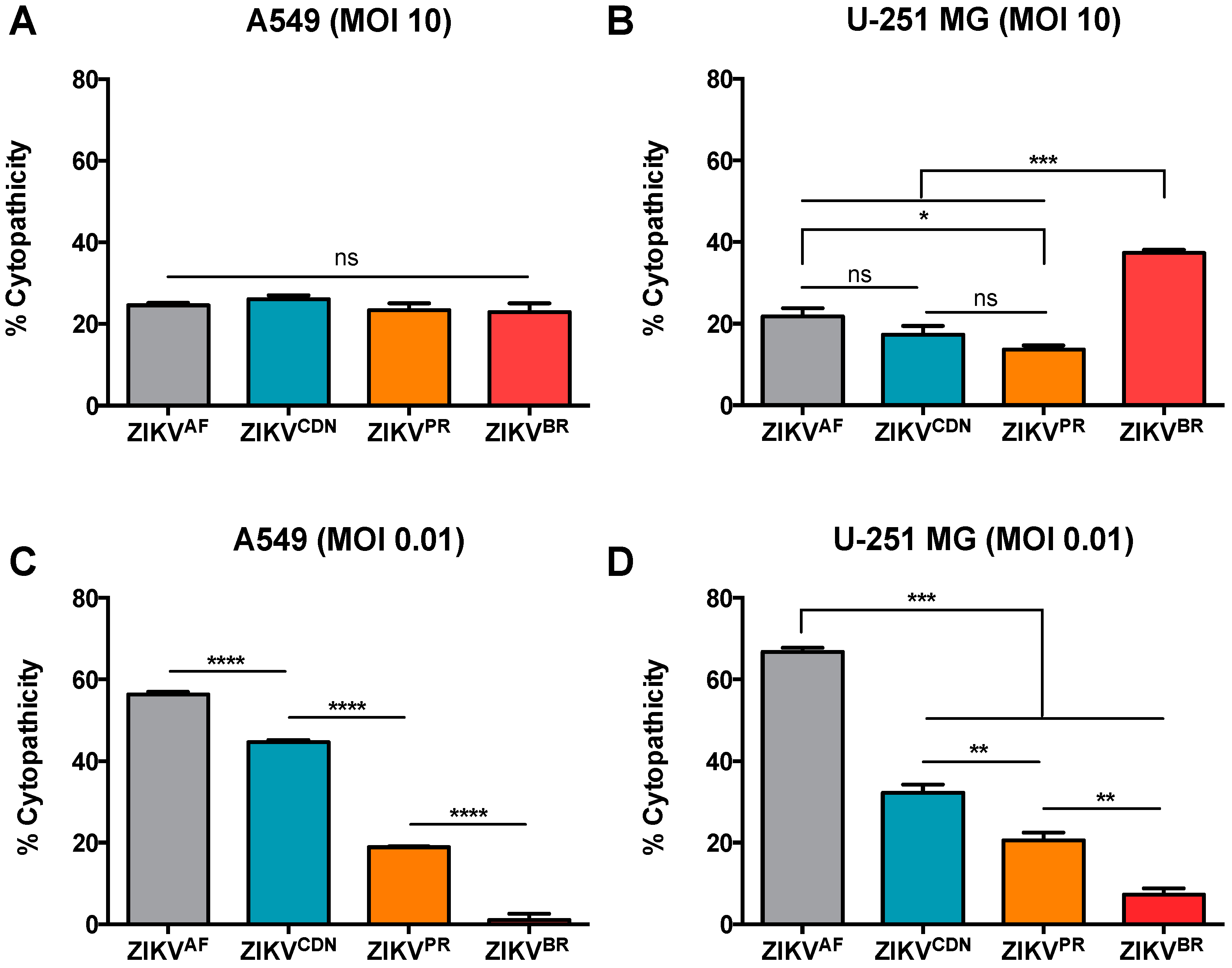
3.3. Cytopathic Effects Induced by ZIKV Isolates in Cell Culture Depends on MOI
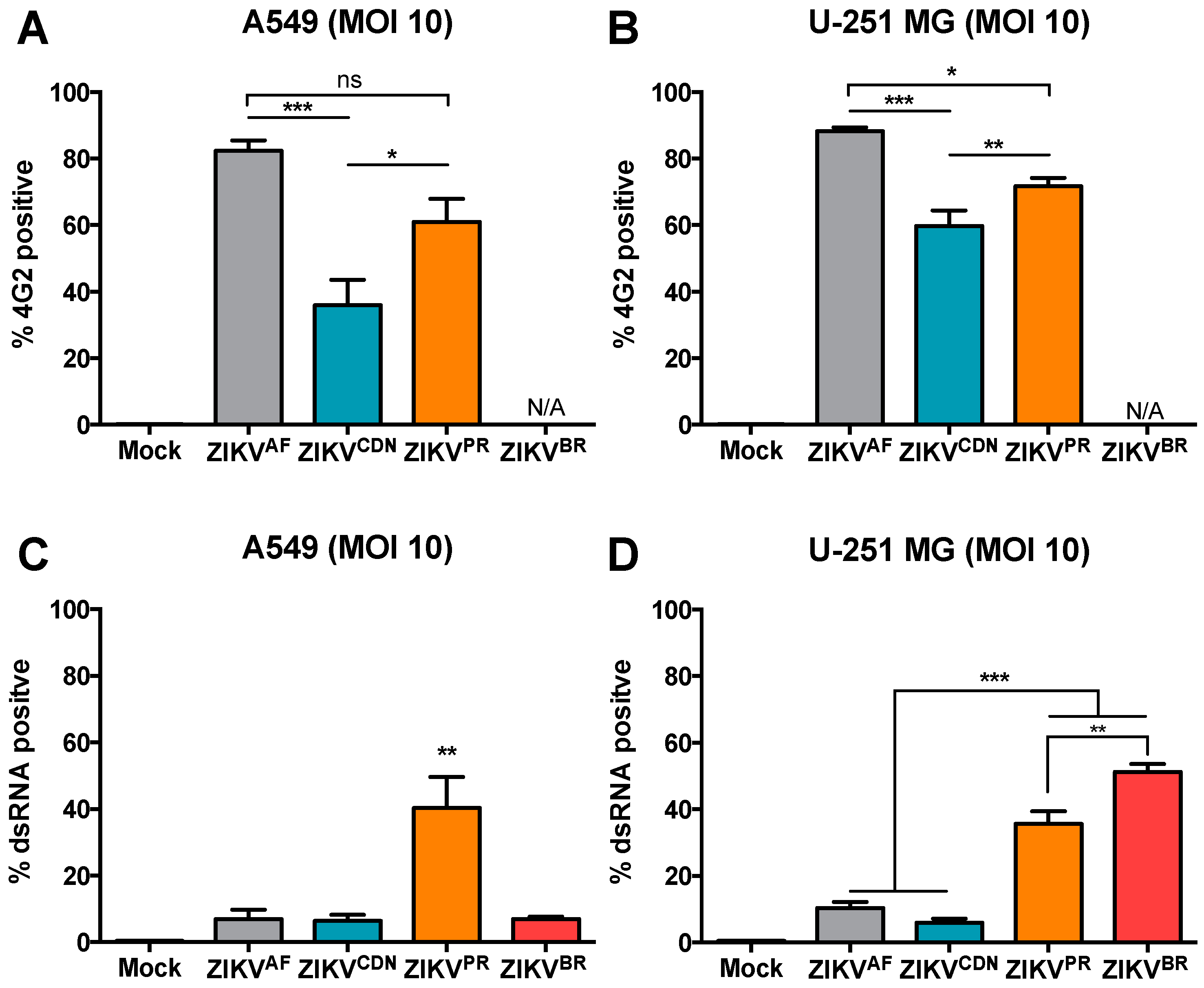
3.4. Flow Cytometry Analysis Reveals Strain-Specific Differences in Viral Infectivity and Cell Spread
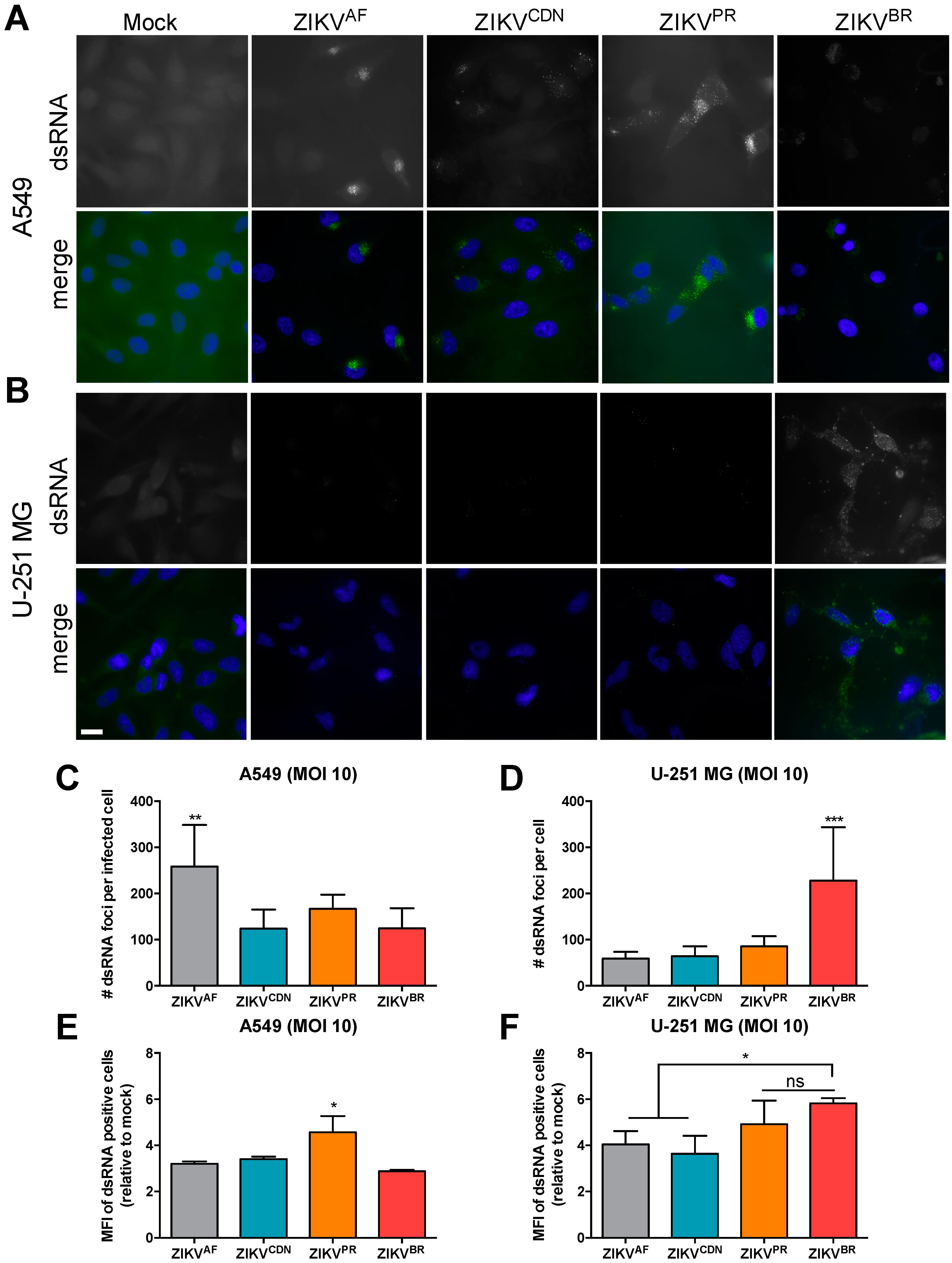
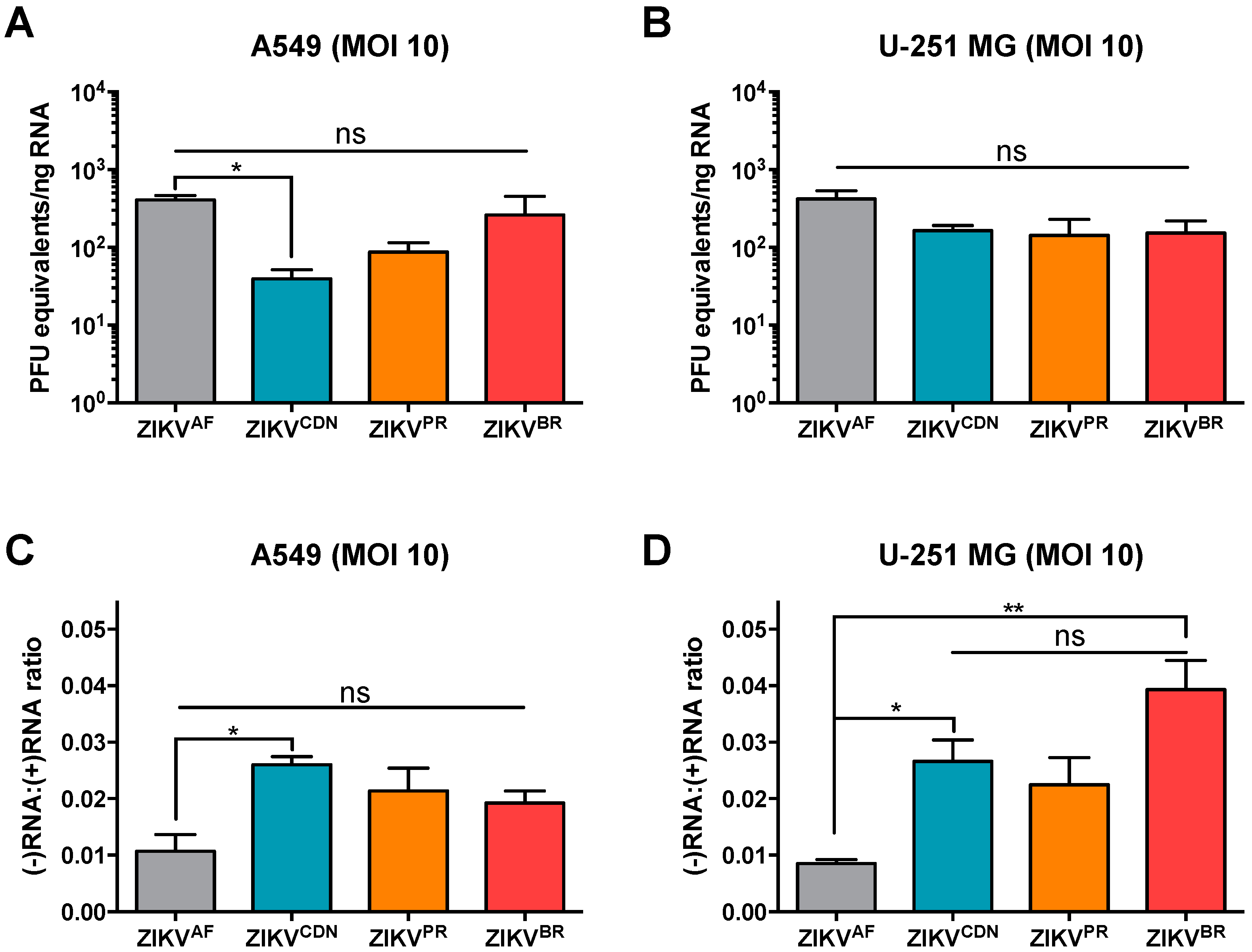
3.5. The Number of dsRNA Foci and Ratio of Negative- to Positive-Strand RNA Differ across African and Asian Lineage Isolates
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rajah, M.M.; Pardy, R.D.; Condotta, S.A.; Richer, M.J.; Sagan, S.M. Zika Virus: Emergence, Phylogenetics, Challenges and Opportunities. ACS Infect. Dis. 2016. [Google Scholar] [CrossRef]
- Dick, G.W.; Kitchen, S.F.; Haddow, A.J. Zika virus (I). Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 1952, 46, 509–520. [Google Scholar] [CrossRef]
- Hayes, E.B. Zika virus outside Africa. Emerg. Infect. Dis. 2009, 15, 1347–1350. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Ortiz, K.; Ansari, A.; Gershwin, M.E. The Zika outbreak of the 21st century. J. Autoimmun. 2016, 68, 1–13. [Google Scholar] [CrossRef]
- Duffy, M.R.; Chen, T.H.; Hancock, W.T.; Powers, A.M.; Kool, J.L.; Lanciotti, R.S.; Pretrick, M.; Marfel, M.; Holzbauer, S.; Dubray, C.; et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. N. Engl. J. Med. 2009, 360, 2536–2543. [Google Scholar] [CrossRef] [PubMed]
- Cao-Lormeau, V.M.; Blake, A.; Mons, S.; Lastere, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: A case-control study. Lancet 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- Fauci, A.S.; Morens, D.M. Zika Virus in the Americas—Yet Another Arbovirus Threat. N. Engl. J. Med. 2016, 374, 601–604. [Google Scholar] [CrossRef]
- Lazear, H.M.; Govero, J.; Smith, A.M.; Platt, D.J.; Fernandez, E.; Miner, J.J.; Diamond, M.S. A Mouse Model of Zika Virus Pathogenesis. Cell Host Microbe 2016. [Google Scholar] [CrossRef]
- Wakerley, B.R.; Yuki, N. Guillain-Barre syndrome. Expert Rev. Neurother. 2015, 15, 847–849. [Google Scholar] [CrossRef]
- Mlakar, J.; Korva, M.; Tul, N.; Popovic, M.; Poljsak-Prijatelj, M.; Mraz, J.; Kolenc, M.; Resman Rus, K.; Vesnaver Vipotnik, T.; Fabjan Vodusek, V.; et al. Zika Virus Associated with Microcephaly. N. Engl. J. Med. 2016, 374, 951–958. [Google Scholar] [CrossRef]
- Wise, J. Study links Zika virus to Guillain-Barre syndrome. BMJ 2016, 352, i1242. [Google Scholar] [CrossRef] [PubMed]
- Adams Waldorf, K.M.; Stencel-Baerenwald, J.E.; Kapur, R.P.; Studholme, C.; Boldenow, E.; Vornhagen, J.; Baldessari, A.; Dighe, M.K.; Thiel, J.; Merillat, S.; et al. Fetal brain lesions after subcutaneous inoculation of Zika virus in a pregnant nonhuman primate. Nat. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Weaver, S.C. Emergence of Epidemic Zika Virus Transmission and Congenital Zika Syndrome: Are Recently Evolved Traits to Blame? mBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Cugola, F.R.; Fernandes, I.R.; Russo, F.B.; Freitas, B.C.; Dias, J.L.; Guimaraes, K.P.; Benazzato, C.; Almeida, N.; Pignatari, G.C.; Romero, S.; et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 2016, 534, 267–271. [Google Scholar] [CrossRef]
- Hamel, R.; Ferraris, P.; Wichit, S.; Diop, F.; Talignani, L.; Pompon, J.; Garcia, D.; Liegeois, F.; Sall, A.A.; Yssel, H.; et al. African and Asian Zika virus strains differentially induce early antiviral responses in primary human astrocytes. Infect. Genet. Evol. 2017, 49, 134–137. [Google Scholar] [CrossRef] [PubMed]
- Limonta, D.; Jovel, J.; Kumar, A.; Airo, A.M.; Hou, S.; Saito, L.; Branton, W.; Ka-Shu Wong, G.; Mason, A.; Power, C.; et al. Human Fetal Astrocytes Infected with Zika Virus Exhibit Delayed Apoptosis and Resistance to Interferon: Implications for Persistence. Viruses 2018, 10, 646. [Google Scholar] [CrossRef]
- Tripathi, S.; Balasubramaniam, V.R.; Brown, J.A.; Mena, I.; Grant, A.; Bardina, S.V.; Maringer, K.; Schwarz, M.C.; Maestre, A.M.; Sourisseau, M.; et al. A novel Zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLoS Pathog. 2017, 13, e1006258. [Google Scholar] [CrossRef]
- Yuan, L.; Huang, X.Y.; Liu, Z.Y.; Zhang, F.; Zhu, X.L.; Yu, J.Y.; Ji, X.; Xu, Y.P.; Li, G.; Li, C.; et al. A single mutation in the prM protein of Zika virus contributes to fetal microcephaly. Science 2017. [Google Scholar] [CrossRef]
- Xia, H.; Luo, H.; Shan, C.; Muruato, A.E.; Nunes, B.T.D.; Medeiros, D.B.A.; Zou, J.; Xie, X.; Giraldo, M.I.; Vasconcelos, P.F.C.; et al. An evolutionary NS1 mutation enhances Zika virus evasion of host interferon induction. Nat. Commun. 2018, 9, 414. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, J.; Du, S.; Shan, C.; Nie, K.; Zhang, R.; Li, X.F.; Zhang, R.; Wang, T.; Qin, C.F.; et al. Evolutionary enhancement of Zika virus infectivity in Aedes aegypti mosquitoes. Nature 2017, 545, 482–486. [Google Scholar] [CrossRef]
- Frumence, E.; Roche, M.; Krejbich-Trotot, P.; El-Kalamouni, C.; Nativel, B.; Rondeau, P.; Misse, D.; Gadea, G.; Viranaicken, W.; Despres, P. The South Pacific epidemic strain of Zika virus replicates efficiently in human epithelial A549 cells leading to IFN-beta production and apoptosis induction. Virology 2016, 493, 217–226. [Google Scholar] [CrossRef]
- Airo, A.M.; Urbanowski, M.D.; Lopez-Orozco, J.; You, J.H.; Skene-Arnold, T.D.; Holmes, C.; Yamshchikov, V.; Malik-Soni, N.; Frappier, L.; Hobman, T.C. Expression of flavivirus capsids enhance the cellular environment for viral replication by activating Akt-signalling pathways. Virology 2018, 516, 147–157. [Google Scholar] [CrossRef]
- Kumar, A.; Hou, S.; Airo, A.M.; Limonta, D.; Mancinelli, V.; Branton, W.; Power, C.; Hobman, T.C. Zika virus inhibits type-I interferon production and downstream signaling. EMBO Rep. 2016, 17, 1766–1775. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, A.J.; Smith, J.L.; Haese, N.N.; Broeckel, R.M.; Parkins, C.J.; Kreklywich, C.; DeFilippis, V.R.; Denton, M.; Smith, P.P.; Messer, W.B.; et al. Zika Virus infection of rhesus macaques leads to viral persistence in multiple tissues. PLoS Pathog. 2017, 13, e1006219. [Google Scholar] [CrossRef] [PubMed]
- Retallack, H.; Di Lullo, E.; Arias, C.; Knopp, K.A.; Laurie, M.T.; Sandoval-Espinosa, C.; Mancia Leon, W.R.; Krencik, R.; Ullian, E.M.; Spatazza, J.; et al. Zika virus cell tropism in the developing human brain and inhibition by azithromycin. Proc. Natl. Acad. Sci. USA 2016, 113, 14408–14413. [Google Scholar] [CrossRef] [PubMed]
- Van den Pol, A.N.; Mao, G.; Yang, Y.; Ornaghi, S.; Davis, J.N. Zika Virus Targeting in the Developing Brain. J. Neurosci. 2017, 37, 2161–2175. [Google Scholar] [CrossRef]
- Hou, W.; Armstrong, N.; Obwolo, L.A.; Thomas, M.; Pang, X.; Jones, K.S.; Tang, Q. Determination of the Cell Permissiveness Spectrum, Mode of RNA Replication, and RNA-Protein Interaction of Zika Virus. BMC Infect. Dis. 2017, 17, 239. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Schwarz, M.C.; Sourisseau, M.; Espinosa, M.M.; Gray, E.S.; Chambers, M.T. Rescue of the 1947 Zika Virus Prototype Strain with a Cytomegalovirus Promoter-Driven cDNA Clone. mSphere 2016, 1, e00246-16. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, K.; Meatherall, B.; Zarra, D.; Drebot, M.; MacDonald, J.; Pabbaraju, K.; Wong, S.; Webster, P.; Lindsay, R.; Tellier, R. First case of Zika virus infection in a returning Canadian traveler. Am. J. Trop. Med. Hyg. 2014, 91, 1035–1038. [Google Scholar] [CrossRef] [PubMed]
- Heldt, C.L.; Hernandez, R.; Mudiganti, U.; Gurgel, P.V.; Brown, D.T.; Carbonell, R.G. A colorimetric assay for viral agents that produce cytopathic effects. J. Virol. Methods 2006, 135, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Lanciotti, R.S.; Kosoy, O.L.; Laven, J.J.; Velez, J.O.; Lambert, A.J.; Johnson, A.J.; Stanfield, S.M.; Duffy, M.R. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 2008, 14, 1232–1239. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Valderramos, S.G.; Wu, A.; Ouyang, S.; Li, C.; Brasil, P.; Bonaldo, M.; Coates, T.; Nielsen-Saines, K.; Jiang, T.; et al. From Mosquitos to Humans: Genetic Evolution of Zika Virus. Cell Host Microbe 2016. [Google Scholar] [CrossRef]
- Beaver, J.T.; Lelutiu, N.; Habib, R.; Skountzou, I. Evolution of Two Major Zika Virus Lineages: Implications for Pathology, Immune Response, and Vaccine Development. Front. Immunol. 2018, 9, 1640. [Google Scholar] [CrossRef] [PubMed]
- Lazear, H.M.; Diamond, M.S. Zika Virus: New Clinical Syndromes and Its Emergence in the Western Hemisphere. J. Virol. 2016, 90, 4864–4875. [Google Scholar] [CrossRef]
- Barnard, T.R.; Sagan, S.M. (McGill University, Montreal, QC, Canada). Personal Communication, 2018.
- Simonin, Y.; van Riel, D.; Van de Perre, P.; Rockx, B.; Salinas, S. Differential virulence between Asian and African lineages of Zika virus. PLoS Negl. Trop. Dis. 2017, 11, e0005821. [Google Scholar] [CrossRef]
- Alpuche-Lazcano, S.P.; McCullogh, C.R.; Del Corpo, O.; Rance, E.; Scarborough, R.J.; Mouland, A.J.; Sagan, S.M.; Teixeira, M.M.; Gatignol, A. Higher Cytopathic Effects of a Zika Virus Brazilian Isolate from Bahia Compared to a Canadian-Imported Thai Strain. Viruses 2018, 10, 53. [Google Scholar] [CrossRef]
- Kuivanen, S.; Korhonen, E.M.; Helisten, A.A.; Huhtamo, E.; Smura, T.; Vapalahti, O. Differences in the growth properties of Zika virus foetal brain isolate and related epidemic strains in vitro. J. Gen. Virol. 2017, 98, 1744–1748. [Google Scholar] [CrossRef]
- Bos, S.; Viranaicken, W.; Turpin, J.; El-Kalamouni, C.; Roche, M.; Krejbich-Trotot, P.; Despres, P.; Gadea, G. The structural proteins of epidemic and historical strains of Zika virus differ in their ability to initiate viral infection in human host cells. Virology 2018, 516, 265–273. [Google Scholar] [CrossRef]
- Sheridan, M.A.; Balaraman, V.; Schust, D.J.; Ezashi, T.; Roberts, R.M.; Franz, A.W.E. African and Asian strains of Zika virus differ in their ability to infect and lyse primitive human placental trophoblast. PLoS ONE 2018, 13, e0200086. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R.; Sprague, T.R.; Hollidge, B.S.; Valdez, S.M.; Padilla, S.L.; Bellanca, S.A.; Golden, J.W.; Coyne, S.R.; Kulesh, D.A.; Miller, L.J.; et al. African and Asian Zika Virus Isolates Display Phenotypic Differences Both In Vitro and In Vivo. Am. J. Trop. Med. Hyg. 2018, 98, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Simonin, Y.; Loustalot, F.; Desmetz, C.; Foulongne, V.; Constant, O.; Fournier-Wirth, C.; Leon, F.; Moles, J.P.; Goubaud, A.; Lemaitre, J.M.; et al. Zika Virus Strains Potentially Display Different Infectious Profiles in Human Neural Cells. EBioMedicine 2016, 12, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Quicke, K.M.; Bowen, J.R.; Johnson, E.L.; McDonald, C.E.; Ma, H.; O’Neal, J.T.; Rajakumar, A.; Wrammert, J.; Rimawi, B.H.; Pulendran, B.; et al. Zika Virus Infects Human Placental Macrophages. Cell Host Microbe 2016, 20, 83–90. [Google Scholar] [CrossRef]
- Nowakowski, T.J.; Pollen, A.A.; Di Lullo, E.; Sandoval-Espinosa, C.; Bershteyn, M.; Kriegstein, A.R. Expression Analysis Highlights AXL as a Candidate Zika Virus Entry Receptor in Neural Stem Cells. Cell Stem Cell 2016, 18, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Meertens, L.; Labeau, A.; Dejarnac, O.; Cipriani, S.; Sinigaglia, L.; Bonnet-Madin, L.; Le Charpentier, T.; Hafirassou, M.L.; Zamborlini, A.; Cao-Lormeau, V.M.; et al. Axl Mediates ZIKA Virus Entry in Human Glial Cells and Modulates Innate Immune Responses. Cell Rep. 2017, 18, 324–333. [Google Scholar] [CrossRef]
- Yun, S.I.; Song, B.H.; Frank, J.C.; Julander, J.G.; Olsen, A.L.; Polejaeva, I.A.; Davies, C.J.; White, K.L.; Lee, Y.M. Functional Genomics and Immunologic Tools: The Impact of Viral and Host Genetic Variations on the Outcome of Zika Virus Infection. Viruses 2018, 10, 422. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barnard, T.R.; Rajah, M.M.; Sagan, S.M. Contemporary Zika Virus Isolates Induce More dsRNA and Produce More Negative-Strand Intermediate in Human Astrocytoma Cells. Viruses 2018, 10, 728. https://doi.org/10.3390/v10120728
Barnard TR, Rajah MM, Sagan SM. Contemporary Zika Virus Isolates Induce More dsRNA and Produce More Negative-Strand Intermediate in Human Astrocytoma Cells. Viruses. 2018; 10(12):728. https://doi.org/10.3390/v10120728
Chicago/Turabian StyleBarnard, Trisha R., Maaran M. Rajah, and Selena M. Sagan. 2018. "Contemporary Zika Virus Isolates Induce More dsRNA and Produce More Negative-Strand Intermediate in Human Astrocytoma Cells" Viruses 10, no. 12: 728. https://doi.org/10.3390/v10120728
APA StyleBarnard, T. R., Rajah, M. M., & Sagan, S. M. (2018). Contemporary Zika Virus Isolates Induce More dsRNA and Produce More Negative-Strand Intermediate in Human Astrocytoma Cells. Viruses, 10(12), 728. https://doi.org/10.3390/v10120728




