Strawberry Vein Banding Virus P6 Protein Is a Translation Trans-Activator and Its Activity Can be Suppressed by FveIF3g
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Vector Construction
2.3. Agro-Infiltration and GFP Fluorescence Assay
2.4. Confocal Microscopy
2.5. Protein Extraction and Western Blot Assays
2.6. RNA Extraction and Northern Blot Assays
2.7. Yeast Two-Hybrid and Bimolecular Fluorescence Complementation (BIFC) Assays
3. Results
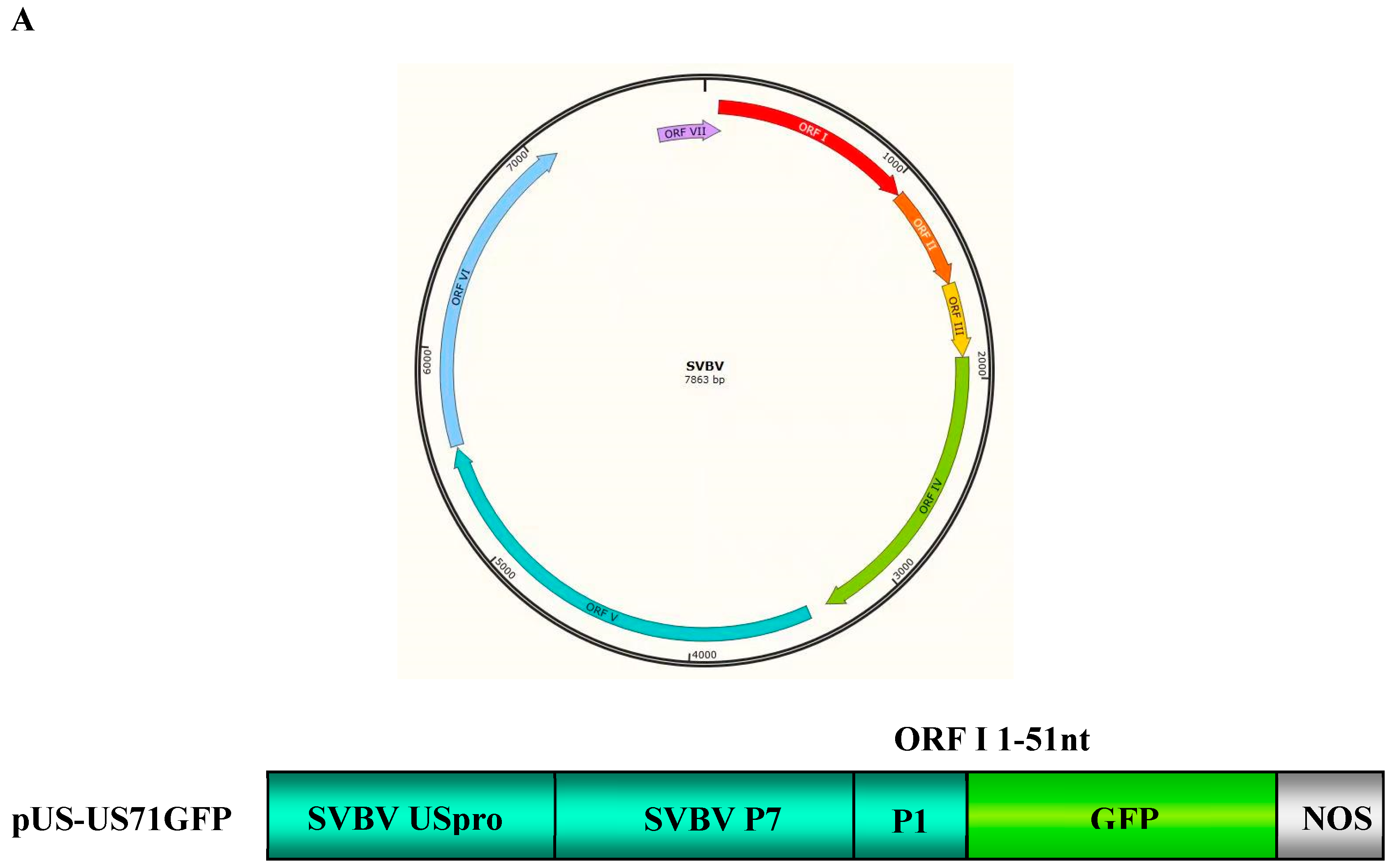
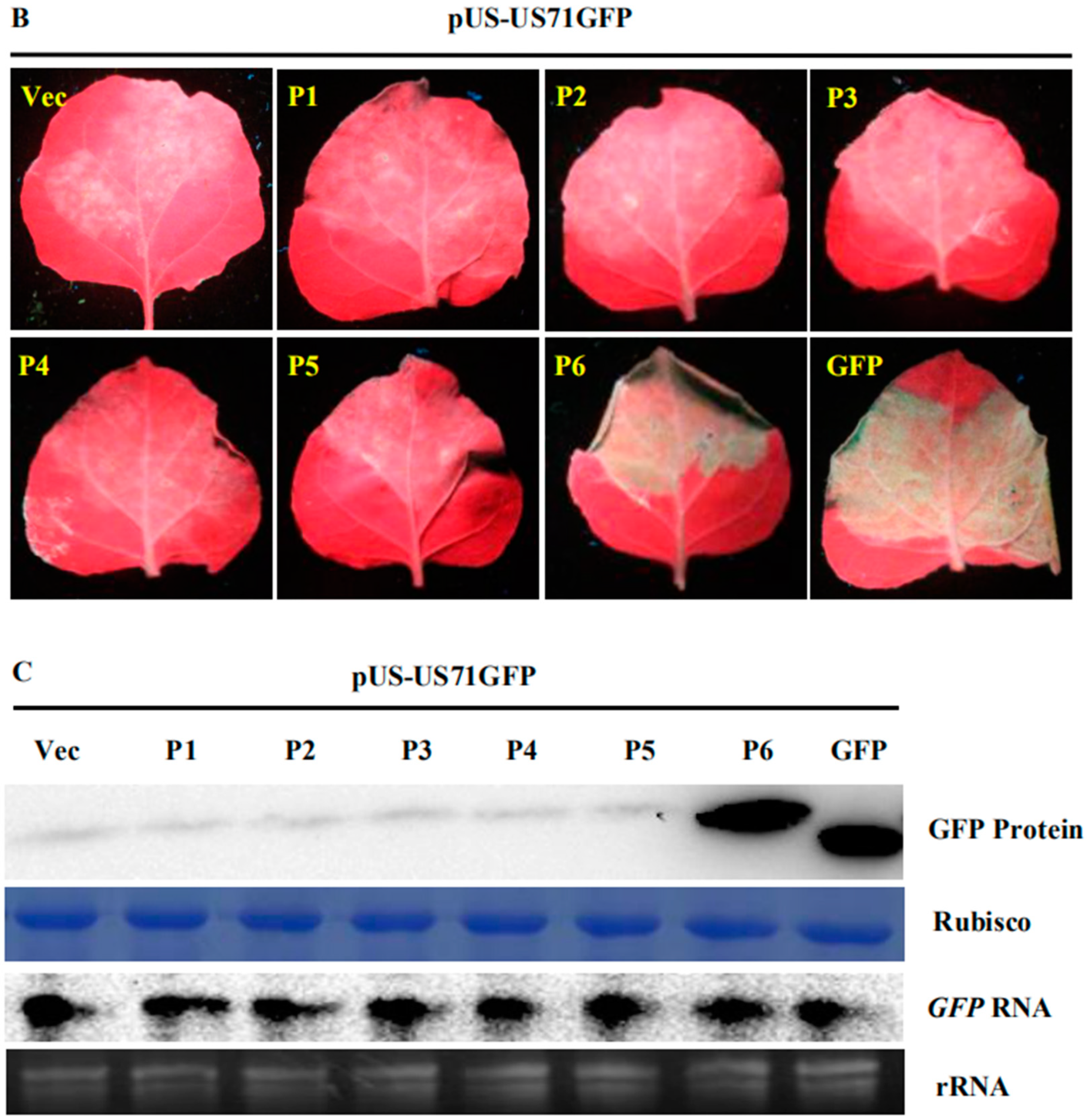
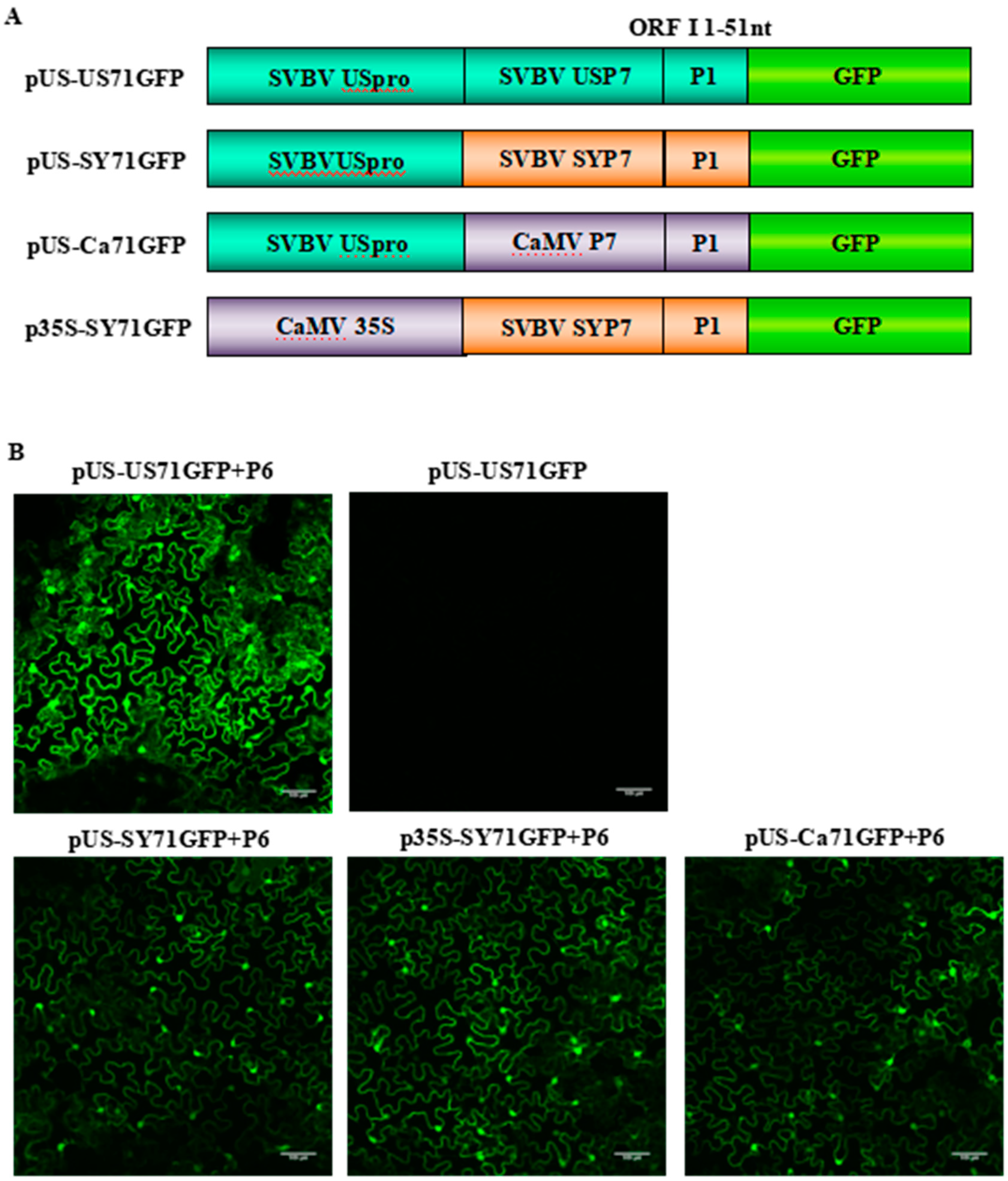
3.1. SVBV P6 is a Trans-Activator of RNA Translation
3.2. P6 Can Trans-Activate the Translation of the gfp RNA from Different Bicistrons
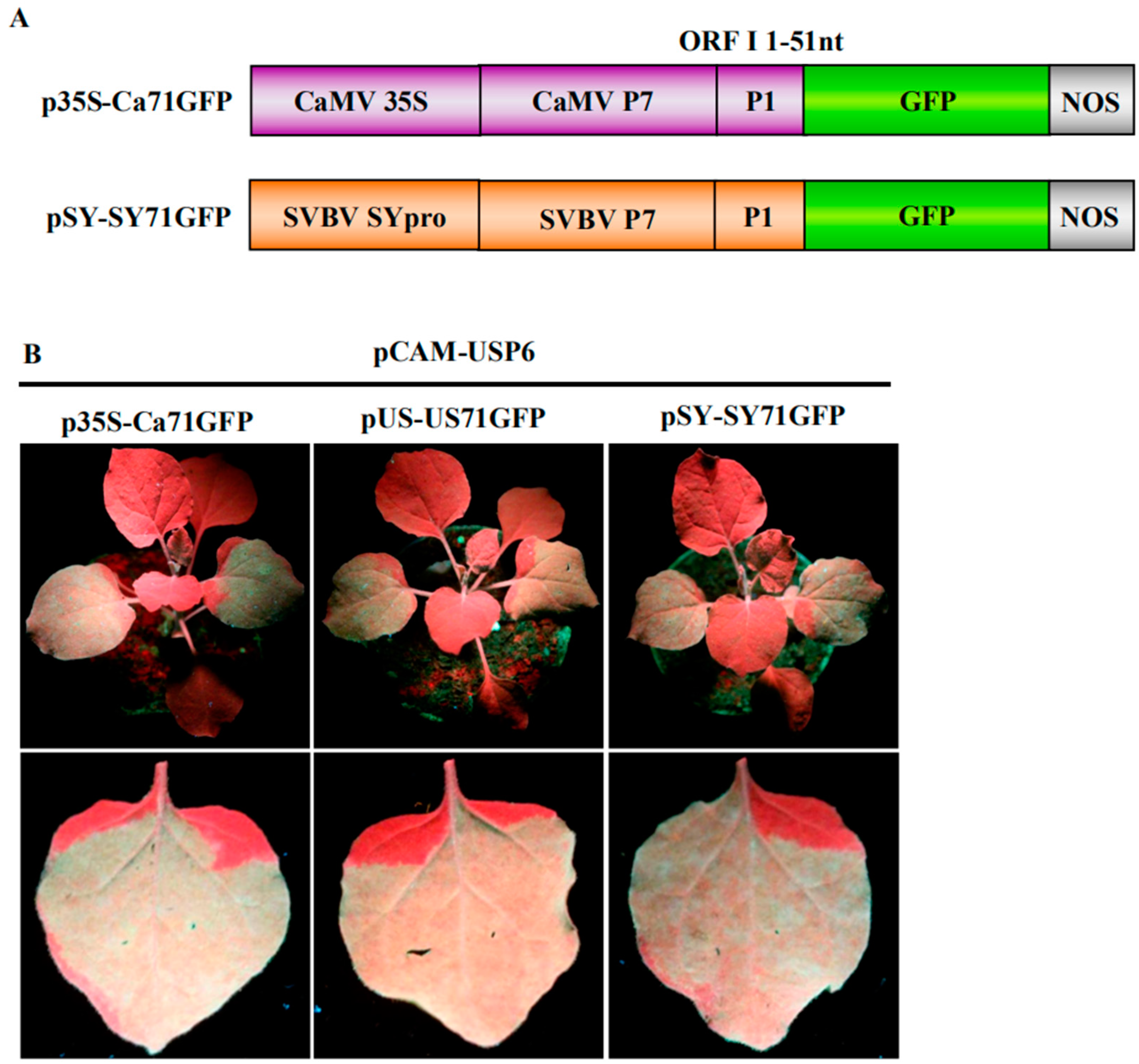
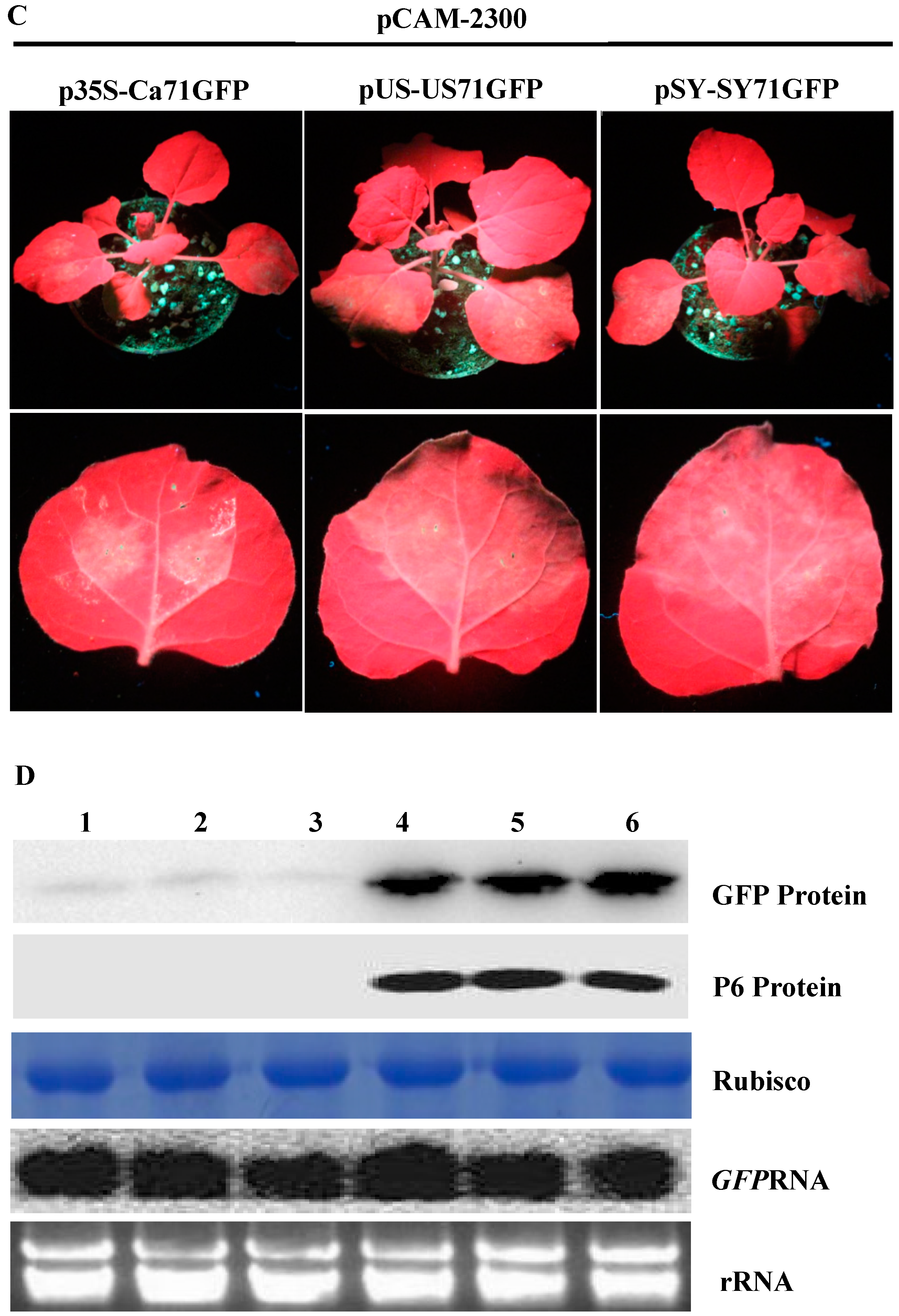
3.3. P6 Can Trans-Activate the Translation of gfp mRNA in Bicistrons Expressed Through a Heterozygous Promoter
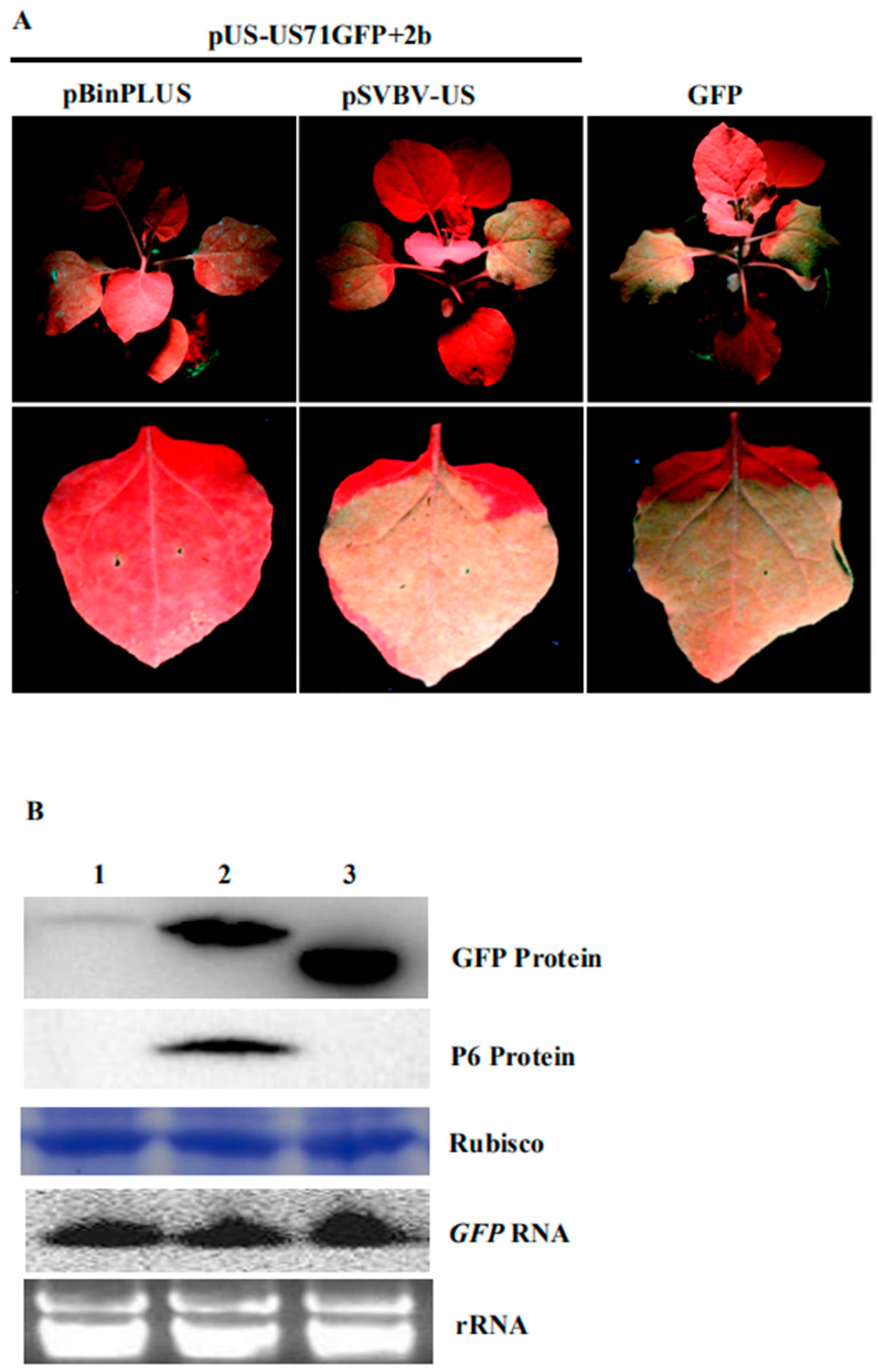
3.4. P6 Protein Encoded by SVBV Can Also Trans-Activate the Translation of gfp mRNA
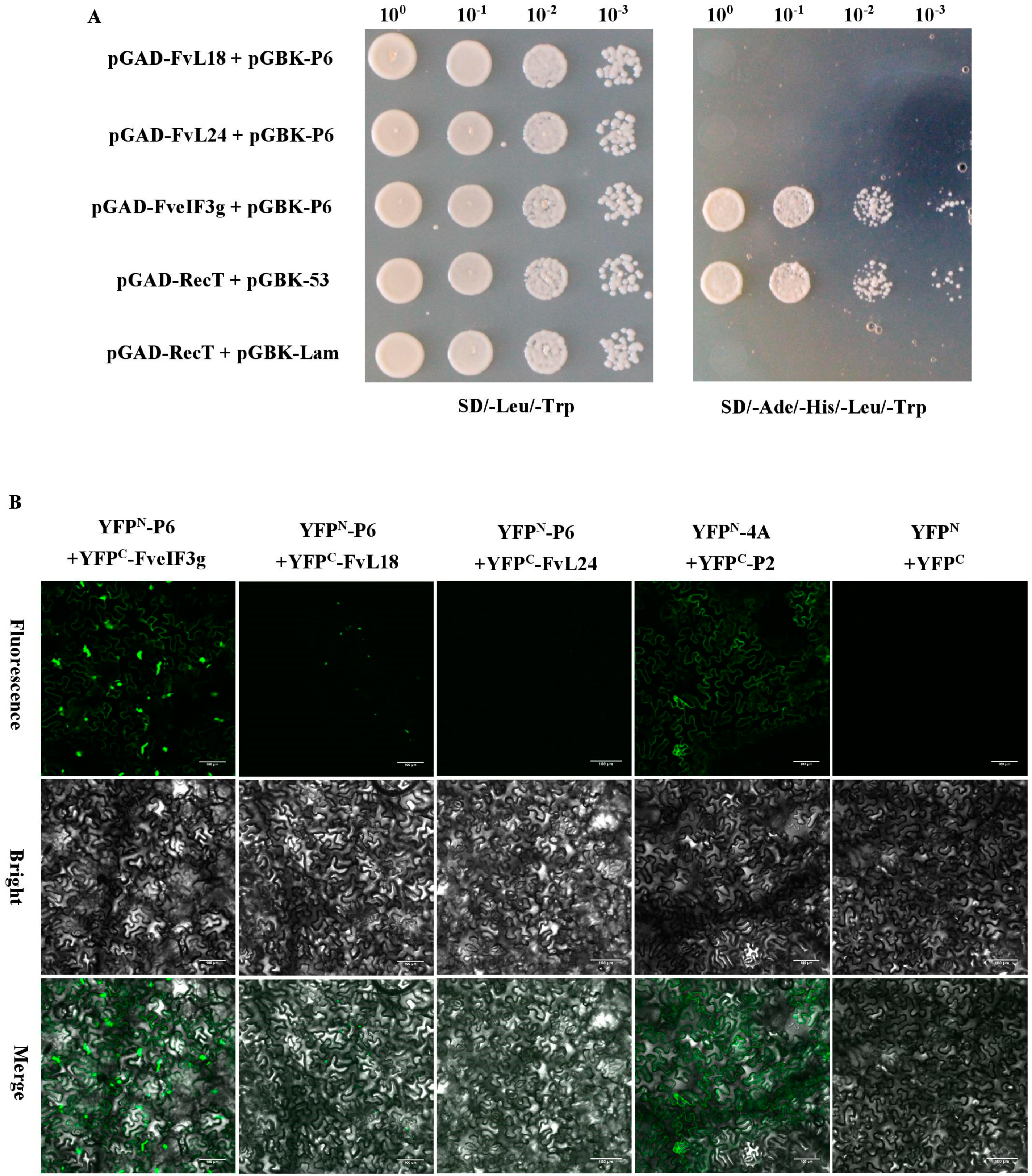
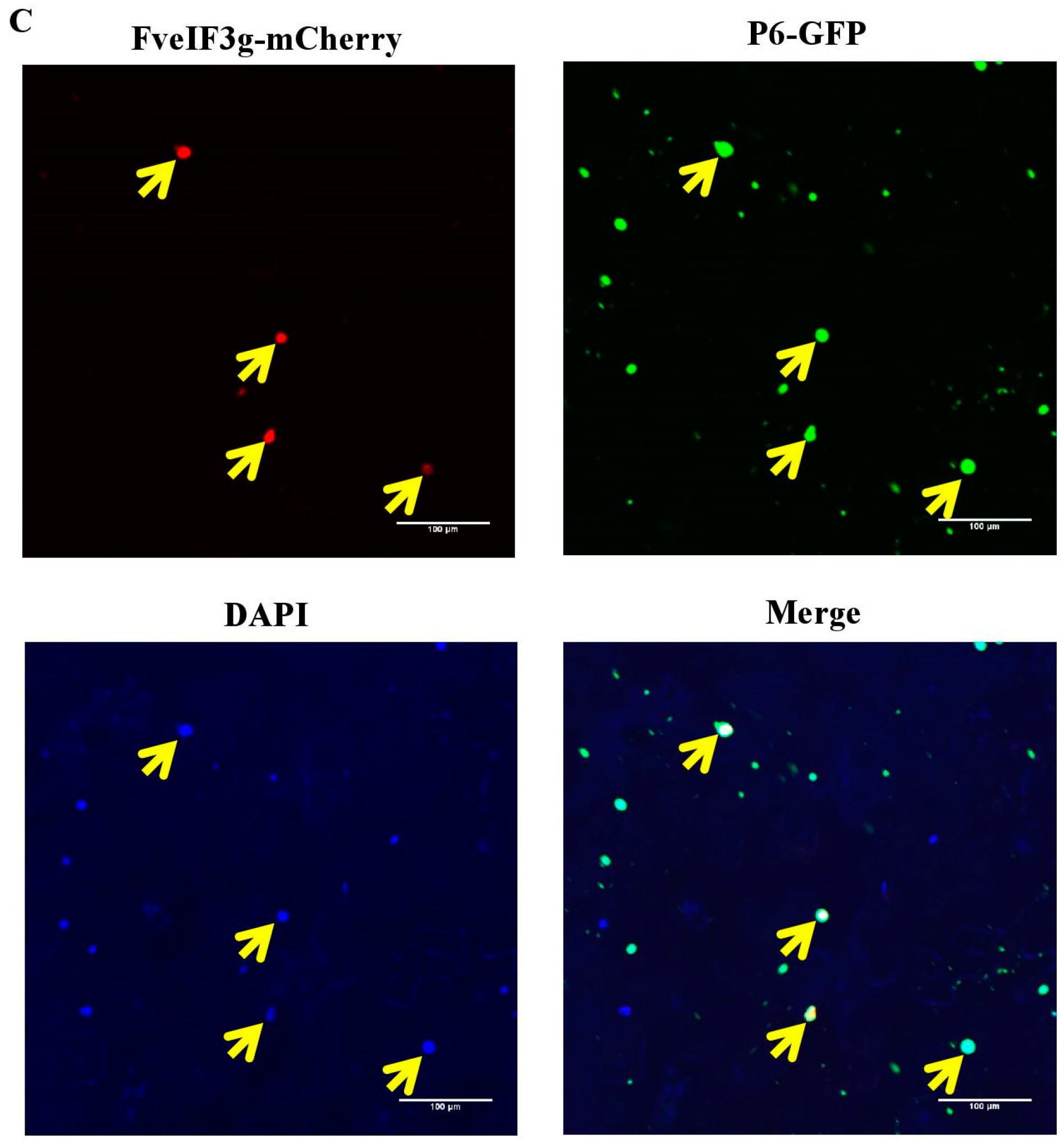
3.5. P6 Protein Can Interact with FveIF3g of F. vesca Both In Vitro and In Vivo
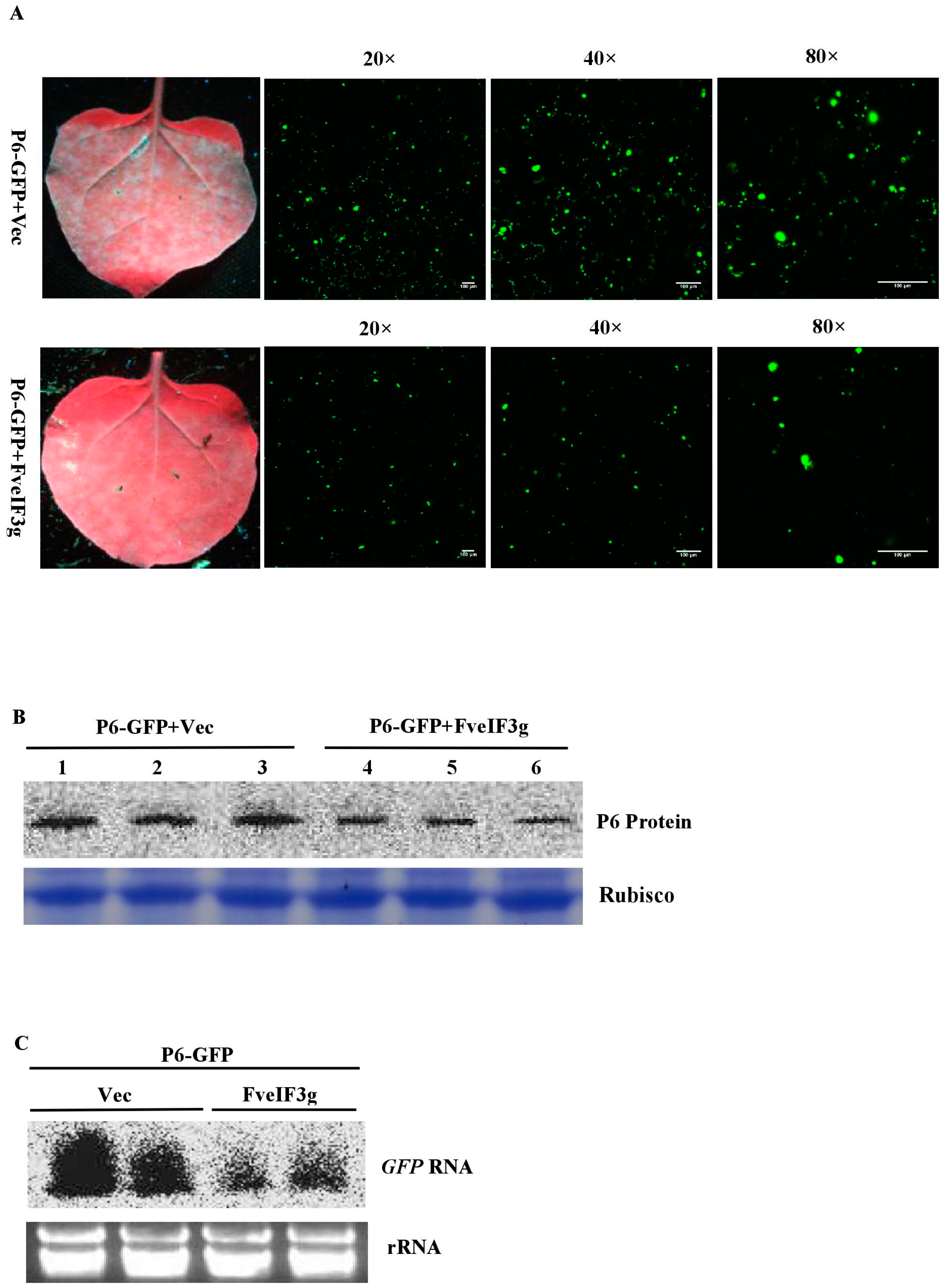
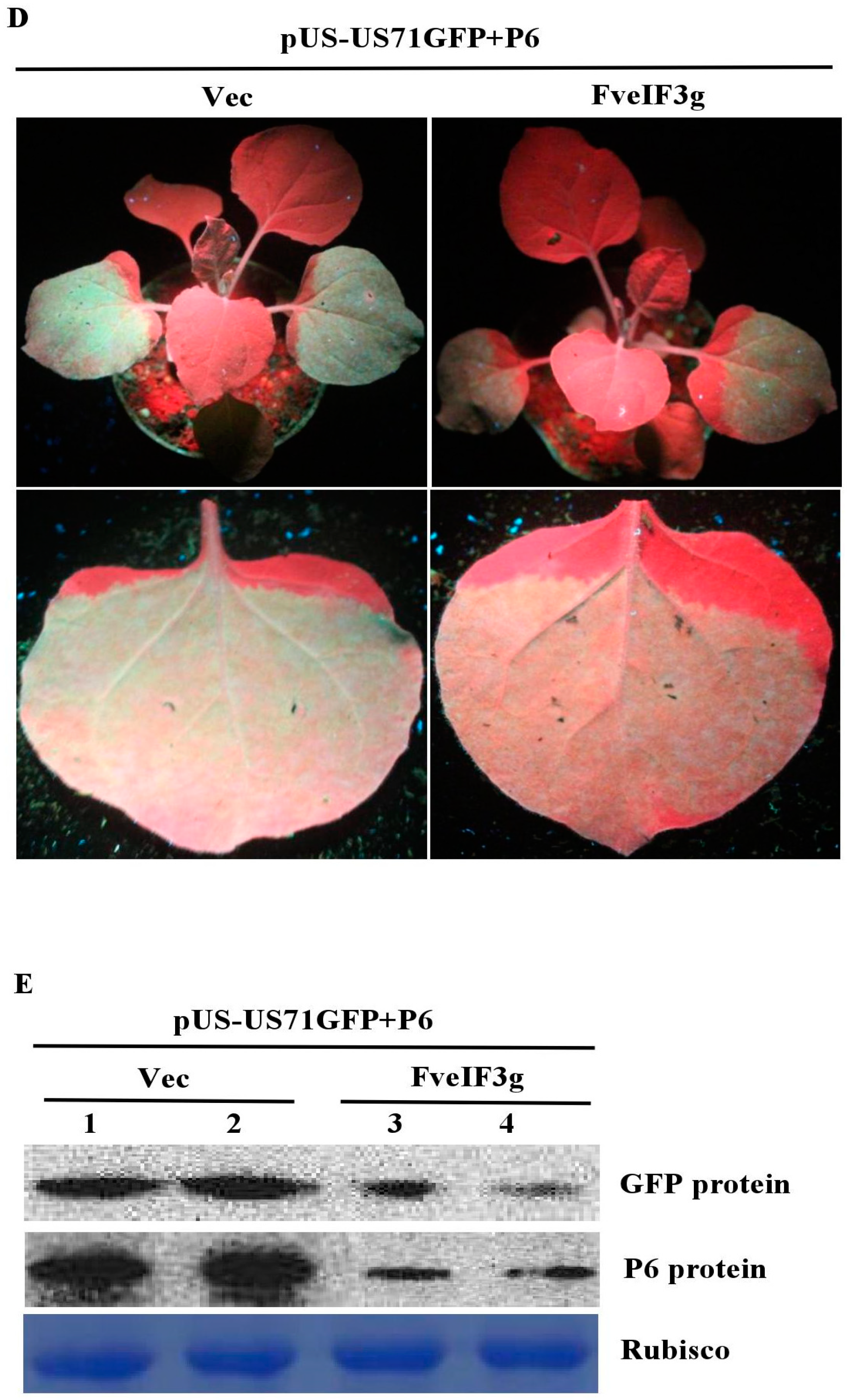
3.6. FveIF3g Affects P6 Protein Accumulation and Inhibits the Trans-Activation Activity of P6 Protein
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Converse, R.H. Modern approaches to strawberry virus research. Acta Hortic. 1992, 308, 19–30. [Google Scholar] [CrossRef]
- Honetšlegrová, J.; Mráz, I.; Špak, J. Detection and isolation of Strawberry vein banding virus in the Czech Republic. Acta Hortic. 1995, 385, 29–32. [Google Scholar] [CrossRef]
- Feng, M.F.; Zhang, H.P.; Pan, Y.; Hu, Y.H.; Chen, J.; Zuo, D.P.; Jiang, T. Complete nucleotide sequence of Strawberry vein banding virus China isolate and infectivity of its full-length DNA clone. Virol. J. 2016, 13, 164. [Google Scholar] [CrossRef] [PubMed]
- Vašková, D.; Špak, J.; Klerks, M.M.; Schoen, C.D.; Thompson, J.P.; Jelkmann, W. Real-time NASBA for detection of Strawberry vein banding virus. Eur. J. Plant Pathol. 2004, 110, 213–221. [Google Scholar] [CrossRef]
- Petrzik, K.; Benes, V.; Mraz, I.; Honetslegrova-Franova, J.; Ansorge, W.; Spak, J. Strawberry vein banding virus-definitive member of the genus Caulimovirus. Virus Genes 1998, 16, 303–305. [Google Scholar] [CrossRef] [PubMed]
- Špak, J.; Petrzik, K. Variability in sequence of Strawberry vein banding virus. Biol. Plant. 2006, 50, 660–666. [Google Scholar]
- Bolton, A.T. Effects of three virus diseases and their combinations on fruit yield of strawberries. Can. J. Plant Sci. 1974, 54, 271–275. [Google Scholar] [CrossRef]
- Thompson, J.R.; Wetzel, S.; Klerks, M.M.; Vaskova, D.; Schoen, C.D.; Spak, J.; Jelkmann, W. Multiplex RT-PCR detection of four aphid-borne strawberry viruses in Fragaria spp. in combination with a plant mRNA specific internal control. J. Virol. Methods 2003, 111, 85–93. [Google Scholar] [CrossRef]
- Pattanaik, S.; Dey, N.; Bhattacharyya, S.; Maiti, I.B. Isolation of full-length transcript promoter from the Strawberry vein banding virus (SVBV) and expression analysis by protoplasts transient assays and in transgenic plants. Plant Sci. 2004, 167, 427–438. [Google Scholar] [CrossRef]
- Schoelz, J.; Shepherd, R.J.; Daubert, S. Region VI of cauliflower mosaic virus encodes a host range determinant. Mol. Cell. Biol. 1986, 6, 2632–2637. [Google Scholar] [CrossRef]
- Feng, M.F.; Zuo, D.P.; Jiang, X.Z.; Li, S.; Chen, J.; Jiang, L.; Zhou, X.P.; Jiang, T. Identification of Strawberry vein banding virus encoded P6 as an RNA silencing suppressor. Virology 2018, 520, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Zhou, X.H.; Li, S.; Feng, M.F.; Shi, M.L.; Zuo, D.P.; Jiang, X.Z.; Ceng, J.; Hu, Y.H.; Zhang, X.X.; et al. Strawberry vein banding virus P6 protein intracellular transport and an important domain identifcation. J. Integr. Agric. 2018, 17, 2031–2041. [Google Scholar] [CrossRef]
- Baughman, G.A.; Jacobs, J.D.; Howell, S.H. Cauliflower mosaic virus gene VI produces a symptomatic phenotype in transgenic tobacco plants. Proc. Natl. Acad. Sci. USA 1988, 85, 733–737. [Google Scholar] [CrossRef] [PubMed]
- Leh, V.; Yot, P.; Keller, M. The Cauliflower mosaic virus translational transactivator interacts with the 60S ribosomal subunit protein L18 of Arabidopsis thaliana. Virology 2000, 266, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Angel, C.A.; Lutz, L.; Yang, X.H.; Rodriguez, A.; Adair, A.; Zhang, Y.; Leisner, S.M.; Nelson, R.S.; Schoelz, J.E. The P6 protein of Cauliflower mosaic virus interacts with CHUP1, a plant protein which moves chloroplasts on actin microfilaments. Virology 2013, 443, 363–374. [Google Scholar] [CrossRef]
- Asano, K.; Clayton, J.; Shalev, A.; Hinnebusch, A.G. A multifactor complex of eukaryotic initiation factors, eIF1, eIF2, eIF3, eIF5, and initiator tRNA Met is an important translation initiation intermediate in vivo. Genes Dev. 2000, 14, 2534–2546. [Google Scholar] [CrossRef]
- Hinnebusch, A.G. Translational regulation of yeast GCN4: A window on factors that control initiator-tRNA binding to the ribosome. J. Biol. Chem. 1997, 270, 21661–21664. [Google Scholar] [CrossRef]
- Pestova, T.V.; Borukhov, S.I.; Hellen, C.U.T. Eukaryotic ribosomes require initiation factors 1 and 1A to locate initiation codons. Nature 1998, 394, 854–859. [Google Scholar] [CrossRef]
- Pestova, T.V.; Shatsky, I.N.; Fletcher, S.P.; Jackson, R.J.; Hellen, C.U.T. A prokaryotic-like mode of cytoplasmic eukaryotic ribosome binding to the initiation codon during internal translation initiation of hepatitis C and classical swine fever virus RNAs. Genes Dev. 1998, 12, 67–83. [Google Scholar] [CrossRef]
- Scholthof, H.B.; Gowda, S.; Wu, F.C.; Shepherd, R.J. The full-length transcript of a caulimovirus is a polycistronic mRNA whose genes are transactivated by the product of gene VI. J. Virol. 1992, 66, 3131–3139. [Google Scholar]
- Zijlstra, C.; Hohn, T. Cauliflower mosaic virus gene VI controls translation from dicistronic expression units in transgenic Arabidopsis plants. Plant Cell 1992, 4, 1471–1484. [Google Scholar] [CrossRef] [PubMed]
- Hohn, T.; Park, H.S.; Peraza, O.G.; Stavolones, L.; Pooggin, M.M.; Kobayashi, K.; Ryabova, L.A. Shunting and controlled reinitiation: The encounter of cauliflower mosaic virus with the translational machinery. Cold Spring Harb. Symp. Quant. Biol. 2001, 66, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Tapia, M.D.; Himmelbach, A.; Hohn, T. Molecular dissection of the Cauliflower mosaic virus translation transactivator. EMBO J. 1993, 12, 3305–3314. [Google Scholar] [CrossRef] [PubMed]
- Ryabova, L.A.; Hohn, T. Ribosome shunting in the cauliflower mosaic virus 35S RNA leader is a special case of reinitiation of translation functioning in plant and animal systems. Genes Dev. 2000, 14, 817–829. [Google Scholar]
- Park, H.S.; Himmelbach, A.; Browning, K.S.; Hohn, T.; Ryabova, L.A. A plant viral “Reinitiation” factor interacts with the host transcriptional machinery. Cell 2001, 106, 703–733. [Google Scholar] [CrossRef]
- Burks, E.A.; Bezerra, P.P.; Le, H.; Gallie, D.R.; Browning, K.S. Plant initiation factor 3 subunit composition resembles mammalian initiation factor 3 and has a novel subunit. J. Biol. Chem. 2001, 276, 2122–2131. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhang, H.P.; Feng, M.F.; Zuo, D.P.; Hu, Y.H.; Jiang, T. Transcriptome analysis of woodland strawberry (Fragaria vesca) response to the infection by Strawberry vein banding virus (SVBV). Virol. J. 2016, 13, 128. [Google Scholar] [CrossRef] [PubMed]
- Thomas, H.; Helen, R. Plant pararetroviruses: Replication and expression. Curr. Opin. Virol. 2013, 3, 621–628. [Google Scholar]
- Haas, M.; Bureau, M.; Geldreich, A.; Yot, P.; Keller, M. Cauliflower mosaic virus: Still in the news. Mol. Plant Pathol. 2002, 3, 19–429. [Google Scholar] [CrossRef]
- Thiebeauld, O.; Schepetilnikov, M.; Park, H.S.; Geldreich, A.; Kobayashi, K.; Keller, M.; Hohn, T.; Ryabova, L.A. A new plant protein interacts with eIF3 and 60S to enhance virus activated translation reinitiation. EMBO J. 2009, 28, 3171–3184. [Google Scholar] [CrossRef]
- Zhang, X.; Yuan, Y.R.; Pei, Y.; Lin, S.S.; Tuschl, T.; Patel, D.J.; Chua, N.H. Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis argonaute1 cleavage activity to counter plant defense. Genes Dev. 2006, 20, 3255–3268. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, Y.C.; Omarov, R.T.; Scholthof, H.B. Diverse and newly recognized effects associated with siRNA binding site modifications on the Tomato bushy stunt virus P19 silencing suppressor. J. Virol. 2009, 83, 2188–2200. [Google Scholar] [CrossRef] [PubMed]
- Brigneti, G.; Voinnet, O.; Li, W.X.; Ji, L.; Ding, S.; Baulcombe, D. Viral pathogenicity determinants are suppressorsof transgene silencing in Nicotiana benthamiana. EMBO J. 1998, 17, 6739–6746. [Google Scholar] [CrossRef] [PubMed]
- Park, H.S.; Browning, K.S.; Hohn, T.; Ryabova, L.A. Eucaryotic initiation factor 4B controls eIF3-mediated ribosomal entry of viral reinitiation factor. EMBO J. 2004, 23, 1381–1391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Hu, Y.; Jiang, L.; Rui, P.; Zhao, Q.; Feng, J.; Zuo, D.; Zhou, X.; Jiang, T. Strawberry Vein Banding Virus P6 Protein Is a Translation Trans-Activator and Its Activity Can be Suppressed by FveIF3g. Viruses 2018, 10, 717. https://doi.org/10.3390/v10120717
Li S, Hu Y, Jiang L, Rui P, Zhao Q, Feng J, Zuo D, Zhou X, Jiang T. Strawberry Vein Banding Virus P6 Protein Is a Translation Trans-Activator and Its Activity Can be Suppressed by FveIF3g. Viruses. 2018; 10(12):717. https://doi.org/10.3390/v10120717
Chicago/Turabian StyleLi, Shuai, Yahui Hu, Lei Jiang, Penghuan Rui, Qingqing Zhao, Jiying Feng, Dengpan Zuo, Xueping Zhou, and Tong Jiang. 2018. "Strawberry Vein Banding Virus P6 Protein Is a Translation Trans-Activator and Its Activity Can be Suppressed by FveIF3g" Viruses 10, no. 12: 717. https://doi.org/10.3390/v10120717
APA StyleLi, S., Hu, Y., Jiang, L., Rui, P., Zhao, Q., Feng, J., Zuo, D., Zhou, X., & Jiang, T. (2018). Strawberry Vein Banding Virus P6 Protein Is a Translation Trans-Activator and Its Activity Can be Suppressed by FveIF3g. Viruses, 10(12), 717. https://doi.org/10.3390/v10120717



