Virulence of Marburg Virus Angola Compared to Mt. Elgon (Musoke) in Macaques: A Pooled Survival Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Aggregation of Data
2.2. Ethics Statement
2.3. Analysis
3. Results
3.1. Baseline Differences
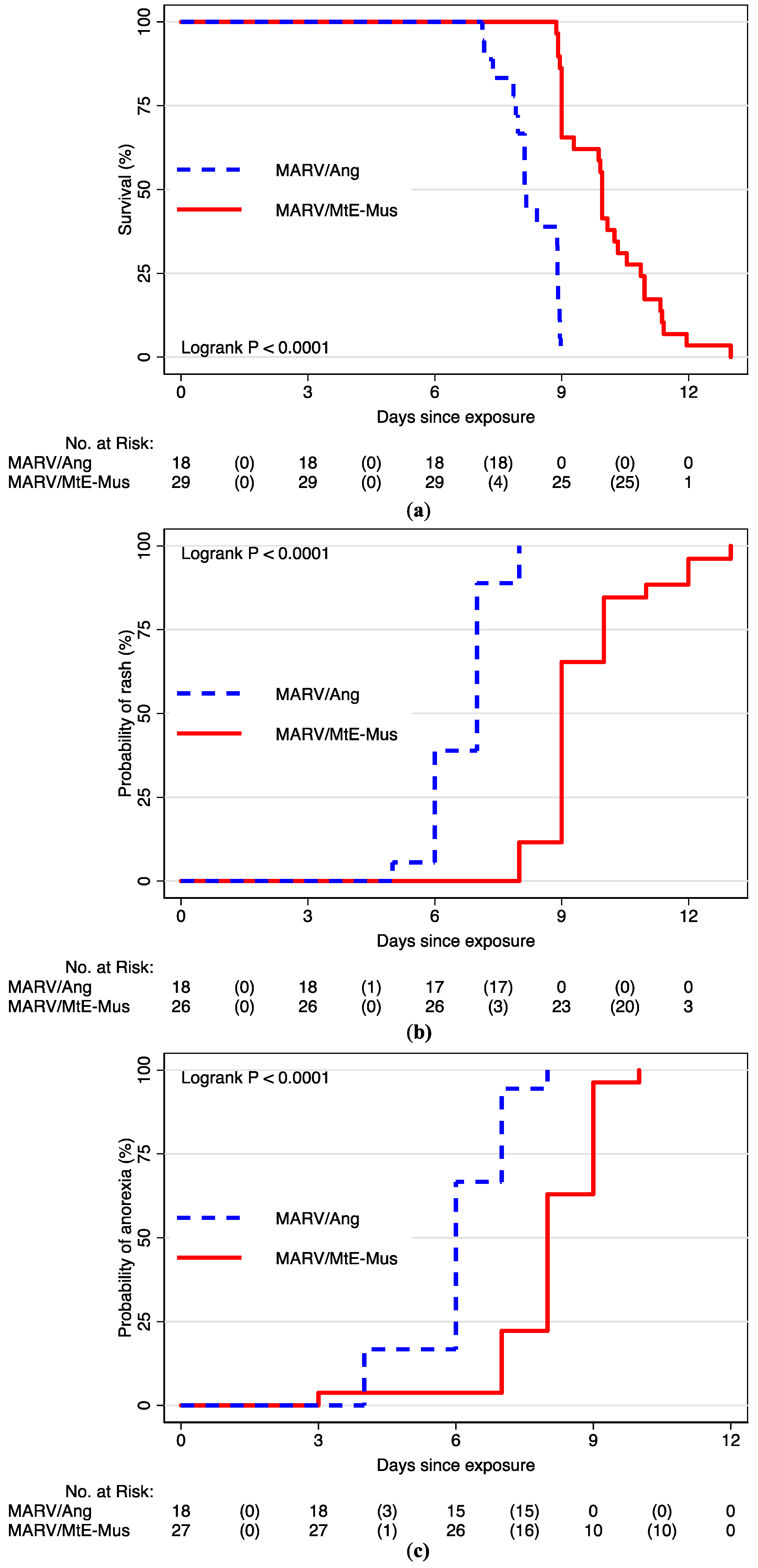
3.2. Time to Event Analyses
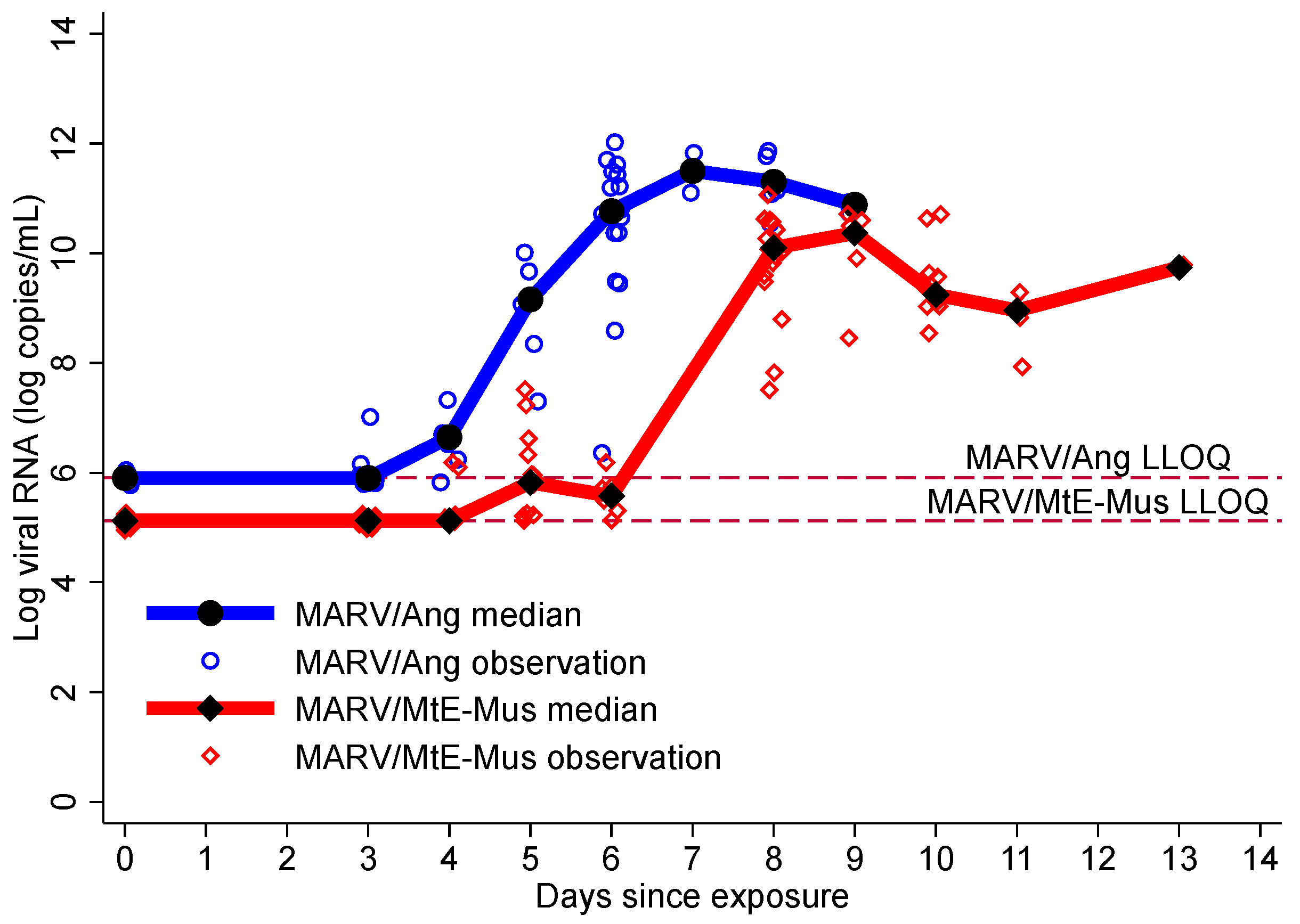
3.3. Viremia
3.4. Clinical Laboratory Parameters
3.4.1. Immune System Activation and Anemia
3.4.2. Moderate-to-Severe Organ Dysfunction
3.4.3. Coagulopathy
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Geisbert, T.W.; Hensley, L.E.; Jahrling, P.B.; Larsen, T.; Geisbert, J.B.; Paragas, J.; Young, H.A.; Fredeking, T.M.; Rote, W.E.; Vlasuk, G.P. Treatment of Ebola virus infection with a recombinant inhibitor of factor VIIa/tissue factor: A study in rhesus monkeys. Lancet 2003, 362, 1953–1958. [Google Scholar] [CrossRef]
- Sweileh, W.M. Global research trends of World Health Organization’s top eight emerging pathogens. Global Health 2017, 13, 9. [Google Scholar] [CrossRef] [PubMed]
- Nyakarahuka, L.; Kankya, C.; Krontveit, R.; Mayer, B.; Mwiine, F.N.; Lutwama, J.; Skjerve, E. How severe and prevalent are Ebola and Marburg viruses? A systematic review and meta-analysis of the case fatality rates and seroprevalence. BMC Infect. Dis. 2016, 16, 708. [Google Scholar] [CrossRef] [PubMed]
- Martini, G.A. Marburg agent disease: in man. Trans. R. Soc. Trop. Med. Hyg. 1969, 63, 295–302. [Google Scholar] [CrossRef]
- Brauburger, K.; Hume, A.J.; Mühlberger, E.; Olejnik, J. Forty-five years of Marburg virus research. Viruses 2012, 4, 1878–1927. [Google Scholar] [CrossRef] [PubMed]
- Chronology of Marburg Hemorrhagic Fever Outbreaks. Available online: https://www.cdc.gov/vhf/marburg/resources/outbreak-table.html (accessed on 12 November 2017).
- Roddy, P.; Weatherill, D.; Jeffs, B.; Abaakouk, Z.; Dorion, C.; Rodriguez-Martinez, J.; Palma, P.P.; de la Rosa, O.; Villa, L.; Grovas, I. The Medecins Sans Frontieres intervention in the Marburg hemorrhagic fever epidemic, Uige, Angola, 2005. II. Lessons learned in the community. J. Infect. Dis. 2007, 196, S162–S167. [Google Scholar] [CrossRef] [PubMed]
- van Paassen, J.; Bauer, M.P.; Arbous, M S.; Visser, L.G.; Schmidt-Chanasit, J.; Schilling, S.; Ölschlager, S.; Rieger, T.; Emmerich, P.; Schmetz, C.; et al. Acute liver failure, multiorgan failure, cerebral oedema, and activation of proangiogenic and antiangiogenic factors in a case of Marburg haemorrhagic fever. Lancet Infect. Dis. 2012, 12, 635–642. [Google Scholar] [CrossRef]
- Imported case of Marburg hemorrhagic fever―Colorado, 2008. MMWR Morb Mortal Wkly. Rep. 2009, 58, 1377–1381.
- Wasswa, H. Uganda grapples with new Marburg disease outbreak. BMJ 2017, 359, j5252. [Google Scholar] [CrossRef] [PubMed]
- Towner, J.S.; Khristova, M.L.; Sealy, T.K.; Vincent, M.J.; Erickson, B.R.; Bawiec, D.A.; Hartman, A.L.; Comer, J.A.; Zaki, S.R.; Ströher, U.; et al. Marburgvirus genomics and association with a large hemorrhagic fever outbreak in Angola. J. Virol. 2006, 80, 6497–6516. [Google Scholar] [CrossRef] [PubMed]
- Alonso, J.A.; Patterson, J.L. Sequence variability in viral genome non-coding regions likely contribute to observed differences in viral replication amongst MARV strains. Virology 2013, 440, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.H.; Johnson, B.K.; Isaacson, M.; Swanapoel, R.; Johnson, K.M.; Killey, M.; Bagshawe, A.; Siongok, T.; Keruga, W.K. Marburg-virus disease in Kenya. Lancet 1982, 1, 816–820. [Google Scholar] [CrossRef]
- Kuhn, J.H.; Dodd, L.E.; Wahl-Jensen, V.; Radoshitzky, S.R.; Bavari, S.; Jahrling, P.B. Evaluation of perceived threat differences posed by filovirus variants. Biosecur. Bioterror. 2011, 9, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Geisbert, T.W.; Strong, J.E.; Feldmann, H. Considerations in the use of nonhuman primate models of Ebola virus and Marburg virus infection. J. Infect. Dis. 2015, 212, S91–S97. [Google Scholar] [CrossRef] [PubMed]
- Daddario-DiCaprio, K.M.; Geisbert, T.W.; Geisbert, J.B.; Ströher, U.; Hensley, L.E.; Grolla, A.; Fritz, E.A.; Feldmann, F.; Feldmann, H.; Jones, S.M. Cross-protection against Marburg virus strains by using a live, attenuated recombinant vaccine. J. Virol 2006, 80, 9659–9666. [Google Scholar] [CrossRef] [PubMed]
- Fernando, L.; Qiu, X.; Melito, P.L.; Williams, K.J.; Feldmann, F.; Feldmann, H.; Jones, S.M.; Alimonti, J.B. Immune Response to Marburg Virus Angola Infection in Nonhuman Primates. J. Infect. Dis 2015, 212, S234–S241. [Google Scholar] [CrossRef] [PubMed]
- Kilgore, N.; Nuzum, E.O. An interagency collaboration to facilitate development of filovirus medical countermeasures. Viruses 2012, 4, 2312–2316. [Google Scholar] [CrossRef] [PubMed]
- Nuzum, E. Biodefense Vaccines, the Animal Rule and Collaboration, International Conference on Emerging Infectious Disease 2012, Atlanta, Georgia, 13 March 2012; Centers for Disease Control and Prevention: Atlanta, Georgia, 2012. [Google Scholar]
- Sullivan, N.J.; Martin, J.E.; Graham, B.S.; Nabel, G.J. Correlates of protective immunity for Ebola vaccines: Implications for regulatory approval by the animal rule. Nat. Rev. Microbiol. 2009, 7, 393–400. [Google Scholar] [CrossRef] [PubMed]
- FDA. Product Development under the Animal Rule: Guidance for Industry. 2016. [Google Scholar]
- Marburg haemorrhagic fever, Angola. Releve epidemiologique hebdomadaire 2005, 80, 158–159.
- Hevey, M.; Negley, D.; Pushko, P.; Smith, J.; Schmaljohn, A. Marburg virus vaccines based upon alphavirus replicons protect guinea pigs and nonhuman primates. Virology 1998, 251, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Johnston, S.C.; Lin, K.L.; Twenhafel, N.A.; Raymond, J.L.; Shamblin, J.D.; Wollen, S.E.; Wlazlowski, C.B.; Wilkinson, E.R.; Botto, M.A.; Goff, A.J. Dose Response of MARV/Angola Infection in Cynomolgus Macaques following IM or Aerosol Exposure. PloS ONE 2015, 10, e0138843. [Google Scholar] [CrossRef] [PubMed]
- Heald, A.E.; Charleston, J.S.; Iversen, P.L.; Warren, T.K.; Saoud, J.B.; Al-Ibrahim, M.; Wells, J.; Warfield, K.L.; Swenson, D.L.; Welch, L.S.; et al. AVI-7288 for Marburg Virus in Nonhuman Primates and Humans. N. Engl. J. Med. 2015, 373, 339–348. [Google Scholar] [CrossRef] [PubMed]
- Warren, T.K.; Wells, J.; Panchal, R.G.; Stuthman, K.S.; Garza, N.L.; Van Tongeren, S.A.; Dong, L.; Retterer, C.J.; Eaton, B.P.; Pegoraro, G.; et al. Protection against filovirus diseases by a novel broad-spectrum nucleoside analogue BCX4430. Nature 2014, 508, 402–405. [Google Scholar] [CrossRef] [PubMed]
- Warren, T.K.; Whitehouse, C.A.; Wells, J.; Welch, L.; Charleston, J.S.; Heald, A.; Nichols, D.K.; Mattix, M.E.; Palacios, G.; Kugleman, J.R.; et al. Delayed Time-to-Treatment of an Antisense Morpholino Oligomer Is Effective against Lethal Marburg Virus Infection in Cynomolgus Macaques. PLoS Negl. Trop. Dis. 2016, 10, e0004456. [Google Scholar] [CrossRef] [PubMed]
- Reisler, R.B.; Yu, C.; Donofrio, M.J.; Warren, T.K.; Wells, J.B.; Stuthman, K.S.; Garza, N.L.; Vantongeren, S.A.; Donnelly, G.C.; Kane, C.D.; et al. Clinical Laboratory Values as Early Indicators of Ebola Virus Infection in Nonhuman Primates. Emerg. Infect. Dis. 2017, 23, 1316–1324. [Google Scholar] [CrossRef] [PubMed]
- Council, N.R. Guide for the Care and Use of Laboratory Animals; National Academies Press: Washington, DC, USA, 2010. [Google Scholar]
- Ewers, E.C.; Pratt, W.D.; Twenhafel, N.A.; Shamblin, J.; Donnelly, G.; Esham, H.; Wlazlowski, C.; Johnson, J.C.; Botto, M.; Hensley, L.E.; Goff, A.J. Natural History of Aerosol Exposure with Marburg Virus in Rhesus Macaques. Viruses 2016, 8, 87. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.; Doane, C. APV Primate Formulary. 2003. Available online: http://agris.fao.org/agris-search/search.do?recordID=US201300105920 (accessed on 20 November 2018).
- Park, H.K.; Cho, J.W.; Lee, B.S.; Park, H.; Han, J.S.; Yang, M.J.; Im, W.J.; Park, D.Y.; Kim, W.J.; Han, S.C.; et al. Reference values of clinical pathology parameters in cynomolgus monkeys (Macaca fascicularis) used in preclinical studies. Laboratory Animal Res. 2016, 32, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Lofts, L.L.; Ibrahim, M.S.; Negley, D.L.; Hevey, M.C.; Schmaljohn, A.L. Genomic differences between guinea pig lethal and nonlethal Marburg virus variants. J. Infect. Dis. 2007, 196, S305–S312. [Google Scholar] [CrossRef] [PubMed]
- Roddy, P.; Thomas, S.L.; Jeffs, B.; Nascimento Folo, P.; Pablo Palma, P.; Moco Henrique, B.; Villa, L.; Damiao Machado, F.P.; Bernal, O.; Jones, S.M.; et al. Factors associated with Marburg hemorrhagic fever: Analysis of patient data from Uige, Angola. J. Infect. Dis. 2010, 201, 1909–1918. [Google Scholar] [CrossRef] [PubMed]
- Kortepeter, M.G.; Bausch, D.G.; Bray, M. Basic clinical and laboratory features of filoviral hemorrhagic fever. J. Infect. Dis 2011, S810–S816. [Google Scholar] [CrossRef] [PubMed]
- Timen, A.; Koopmans, M.P.; Vossen, A.C.; van Doornum, G.J.; Günther, S.; van den Berkmortel, F.; Verduin, K.M.; Dittrich, S.; Emmerich, P.; Osterhaus, A.D.; et al. Response to imported case of Marburg hemorrhagic fever, the Netherland. Emerg. Infect. Dis. 2009, 15, 1171–1175. [Google Scholar] [CrossRef] [PubMed]
- Shurtleff, A.C.; Bavari, S. Animal models for ebolavirus countermeasures discovery: what defines a useful model? Expert Opin. Drug Discov. 2015, 10, 685–702. [Google Scholar] [CrossRef] [PubMed]
- Speranza, E.; Bixler, S.L.; Altamura, L.A.; Arnold, C.E.; Pratt, W.D.; Taylor-Howell, C.; Burrows, C.; Aguilar, W.; Rossi, F.; Shamblin, J.D.; et al. A conserved transcriptional response to intranasal Ebola virus exposure in nonhuman primates prior to onset of fever. Sci. Transl. Med. 2018, 10, 434. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.P. Extrapolating from animals to humans. Sci. Transl. Med. 2012, 4, 151ps15. [Google Scholar] [CrossRef] [PubMed]
| Characteristic | Overall | MARV/Ang | MARV/MtE-Mus | p-value |
|---|---|---|---|---|
| (N = 47) | (N = 18; 38.3%) | (N = 29; 61.7%) | ||
| Sex: no. (%) | ||||
| Male | 25 (53.2) | 9 (50.0) | 16 (55.2) | 0.737 |
| Female | 22 (46.8) | 9 (50.0) | 13 (44.8) | |
| Age (years), mean (SD) | 5.63 (1.07) | 6.14 (1.12) | 5.31 (0.91) | 0.008 |
| Weight (kg), mean (SD) | 4.93 (1.40) | 5.12 (1.66) | 4.81 (1.22) | 0.456 |
| Outcome | Total | Overall | MARV/Ang | MARV/MtE-Mus |
|---|---|---|---|---|
| No. (%) | Median days (25th, 75th percentiles) | Median days (25th, 75th percentiles) | Median days (25th, 75th percentiles) | |
| Death | 47 (100) | 9.00 (9.00, 10.00) | 8.13 (8.00, 9.00) | 9.96 (9.00, 11.00) |
| Anorexia | 45 (96) | 7.00 (6.00, 8.00) | 6.00 (6.00, 7.00) | 8.00 (8.00, 9.00) |
| Rash | 44 (94) | 9.00 (7.00, 9.00) | 7.00 (6.00, 7.00) | 9.00 (9.00, 10.00) |
| Viremia (N = 41; ≥5.12 log copies/mL) | 41 (100) | 6.00 (4.00, 6.00) | 4.00 (3.00, 6.00) | 6.00 (5.00, 8.00) |
| High viremia (N = 41; ≥6.34 log copies/mL) | 39 (95.1) | 6.00 (5.00, 8.00) | 6.00 (4.00, 6.00) | 8.00 (8.00, 8.00) |
| Anemia * (N = 34) | 26 (76.5) | 6.00 (4.00, 10.00) | 7.00 (4.00, 8.00) | 5.00 (4.00, 10.00) |
| Immune activation | ||||
| 22 (84.6) | 8.00 (6.00, 9.00) | 4.00 (4.00, 4.00) | 8.00 (8.00, 9.00) |
| 19 (40.4) | 10.00 (9.00, 11.00) | 8.00 (8.00, 9.00) | 10.00 (10.00, ‡) |
| Organ dysfunction | ||||
| 39 (83.0) | 8.00 (6.00, 8.00) | 6.00 (6.00, 8.00) | 9.00 (8.00, 9.00) |
| 41 (87.2) | 6.00 (5.00, 8.00) | 5.00 (3.00, 6.00) | 8.00 (5.00, 8.00) |
| 18 (38.3) | 10.00 (9.00, 11.00) | 8.00 (8.00, 8.00) | 11.00 (10.00, 13.00) |
| Coagulopathy | ||||
| 10 (25.6) | 11.00 (10.00, 13.00) | 8.00 (8.00, 8.00) | 11.00 (10.00, 13.00) |
| 22 (61.1) | 8.00 (8.00, 10.00) | 6.00 (6.00, 8.00) | 9.00 (8.00, 10.00) |
| 22 (56.4) | 8.00 (8.00, 10.00) | 6.00 (6.00, 8.00) | 9.00 (8.00, 10.00) |
| 23 (56.1) | 8.00 (5.00, ‡) | 6.00 (5.00, ‡) | 8.00 (5.00, ‡) |
| Outcome | Crude HR (95% CI) | Adjusted HR (95% CI) |
|---|---|---|
| Death | 22.10 (7.08, 68.93) * | 25.08 (18.02, 34.91) * |
| Anorexia | 5.42 (2.55, 11.54) * | 6.19 (4.94, 7.76) * |
| Rash | 23.79 (6.49, 87.19) * | 21.48 (15.04, 30.70) * |
| Viremia (N = 41; ≥5.12 log copies/mL) | 1.70 (0.87, 3.32) | 1.51 (0.69, 3.30) |
| High viremia (N = 41) | 4.06 (1.86, 8.86) * | 3.75 (1.53, 9.18) |
| Anemia † (N = 34) | 1.20 (0.51, 2.81) | 1.51 (0.55, 4.18) |
| Immune activation | ||
| 23.94 (4.31, 133.04) * | 22.44 (3.34, 150.87) * |
| 8.98 (2.37, 33.95) * | 10.48 (2.39, 46.01) * |
| Organ dysfunction | ||
| 3.15 (1.53, 6.48) * | 3.41 (1.39, 8.37) |
| 2.62 (1.32, 5.23) * | 2.97 (1.27, 6.91) |
| 40.50 (5.05, 324.82) * | 43.40 (4.93, 382.04) * |
| Coagulopathy | ||
| 25.17 (2.86, 221.71) * | 20.30 (2.03, 203.02) |
| 5.32 (1.91, 14.79) * | 5.46 (1.73, 17.21) |
| 6.72 (2.35, 19.26) * | 7.37 (2.07, 26.28) * |
| 1.69 (0.72, 3.95) | 2.60 (0.95, 7.16) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blair, P.W.; Keshtkar-Jahromi, M.; Psoter, K.J.; Reisler, R.B.; Warren, T.K.; Johnston, S.C.; Goff, A.J.; Downey, L.G.; Bavari, S.; Cardile, A.P. Virulence of Marburg Virus Angola Compared to Mt. Elgon (Musoke) in Macaques: A Pooled Survival Analysis. Viruses 2018, 10, 658. https://doi.org/10.3390/v10110658
Blair PW, Keshtkar-Jahromi M, Psoter KJ, Reisler RB, Warren TK, Johnston SC, Goff AJ, Downey LG, Bavari S, Cardile AP. Virulence of Marburg Virus Angola Compared to Mt. Elgon (Musoke) in Macaques: A Pooled Survival Analysis. Viruses. 2018; 10(11):658. https://doi.org/10.3390/v10110658
Chicago/Turabian StyleBlair, Paul W., Maryam Keshtkar-Jahromi, Kevin J. Psoter, Ronald B. Reisler, Travis K. Warren, Sara C. Johnston, Arthur J. Goff, Lydia G. Downey, Sina Bavari, and Anthony P. Cardile. 2018. "Virulence of Marburg Virus Angola Compared to Mt. Elgon (Musoke) in Macaques: A Pooled Survival Analysis" Viruses 10, no. 11: 658. https://doi.org/10.3390/v10110658
APA StyleBlair, P. W., Keshtkar-Jahromi, M., Psoter, K. J., Reisler, R. B., Warren, T. K., Johnston, S. C., Goff, A. J., Downey, L. G., Bavari, S., & Cardile, A. P. (2018). Virulence of Marburg Virus Angola Compared to Mt. Elgon (Musoke) in Macaques: A Pooled Survival Analysis. Viruses, 10(11), 658. https://doi.org/10.3390/v10110658




